








生物技术通报 ›› 2026, Vol. 42 ›› Issue (1): 13-30.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0643
费思恬1,2( ), 侯鹰翔1,2, 李兰3, 张超1,2(
), 侯鹰翔1,2, 李兰3, 张超1,2( )
)
收稿日期:2025-06-19
出版日期:2026-01-26
发布日期:2026-02-04
通讯作者:
张超,男,博士,讲师,研究方向 :水稻分子生物学;E-mail: ricezhangchao@xnu.edu.cn作者简介:费思恬,女,硕士,助理实验师,研究方向 :水稻分子生物学;E-mail: feisitian@xnu.edu.cn
基金资助:
FEI Si-tian1,2( ), HOU Ying-xiang1,2, LI Lan3, ZHANG Chao1,2(
), HOU Ying-xiang1,2, LI Lan3, ZHANG Chao1,2( )
)
Received:2025-06-19
Published:2026-01-26
Online:2026-02-04
摘要:
赤霉素(GAs)是植物中一类重要的调控激素,广泛参与植物生长发育与逆境响应等多种生命过程。GA的合成及信号通路调控促成了作物育种的第一次绿色革命。SLR1是水稻中唯一的DELLA蛋白,是GA信号转导途径中关键的负调控因子,阻遏GA下游信号转导。SLR1还参与了脱落酸(ABA)、茉莉酸(JA)、油菜素内酯(BR)、独脚金内酯(SL)等激素途径,充当植物激素互作的“分子桥梁”。然而,SLR1蛋白不具备典型的DNA结合域,目前仍无证据表明SLR1能直接结合DNA序列,其主要通过与其他转录因子互作抑制或激活下游基因的表达来发挥调控功能。此外,SLR1自身的表达与功能也受到多种调控。在转录水平,SLR1基因受到OsYABBY4、OsWRKY36等转录因子的负调控;在蛋白水平,SLR1受到泛素化、糖基化、SUMO化、磷酸化等修饰,以及与其他蛋白互作对自身稳定性与活性的调控。SLR1不仅广泛调控水稻多种生长发育过程,还参与水稻响应多种生物与非生物胁迫,在水稻的整个生长周期中扮演了重要的角色。最新研究揭示SLR1在育种中的巨大潜力,通过调控SLR1在水稻中的蛋白丰度并使其维持中等含量,对提高水稻耐碱‒热性具有重要意义。本文综述了水稻SLR1分子结构和作用机制、在各植物激素途径中的串联作用、SLR1自身的蛋白修饰与调控以及具体的生物学功能,以期进一步发掘SLR1蛋白关联组分和调控网络,为水稻分子设计育种提供参考。
费思恬, 侯鹰翔, 李兰, 张超. 水稻赤霉素信号负调控因子SLR1的生物学功能及其调控网络[J]. 生物技术通报, 2026, 42(1): 13-30.
FEI Si-tian, HOU Ying-xiang, LI Lan, ZHANG Chao. Biological Functions and Regulatory Network of SLR1, a Negative Regulator of Gibberellin Signaling in Rice[J]. Biotechnology Bulletin, 2026, 42(1): 13-30.
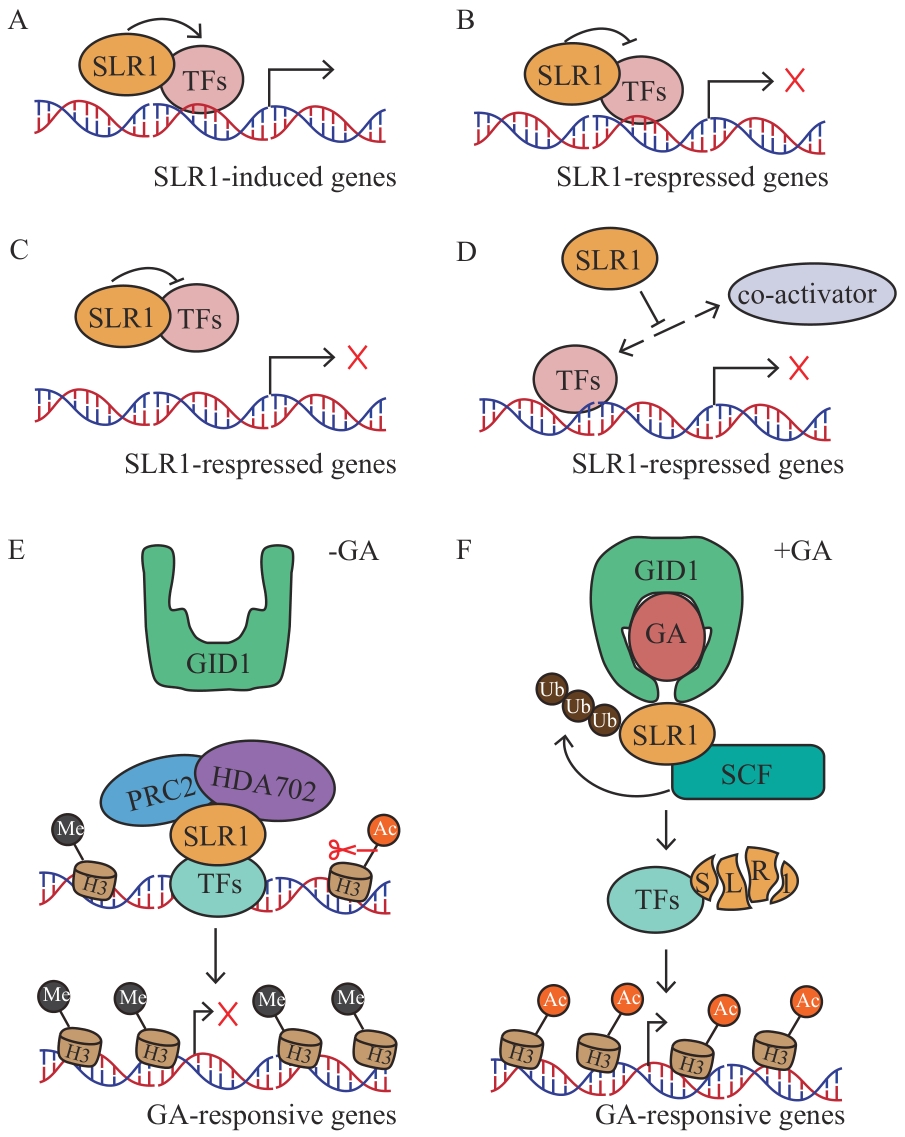
图2 SLR1介导下游基因表达的调控机制示意图A:SLR1结合TFs并激活后者的转录活性,正调控下游基因表达;B:SLR1结合TFs并抑制后者的转录活性,负调控下游基因表达;C:SLR1与TFs结合形成阻遏复合物,阻止TFs与下游基因顺式作用元件的结合;D:SLR1与转录共激活因子结合并阻止TFs与后者结合,抑制靶基因的表达;E:在缺乏GA的情况下,SLR1与PRC2和HDA702形成三元复合物,在GA响应基因上维持高H3K27me3水平和低H3K9ac水平并抑制其表达;F:在GA充足的情况下,SLR1被泛素化降解,不能与PRC2和HDA702等形成复合物,解除对GA响应基因的抑制作用
Fig. 2 Schematic diagram of the regulatory mechanism of downstream gene expression mediated by SLR1A: SLR1 binds to TFs and activates their transcriptional activity, thereby positively regulating the expressions of downstream genes. B: SLR1 binds to TFs and inhibits their transcriptional activity, thus negatively regulating the expressions of downstream genes. C: SLR1 binds to TFs to form a repressor complex, preventing TFs from binding to the cis-acting elements of downstream genes. D: SLR1 binds to transcriptional co-activators and blocks the binding between TFs and the latter, inhibiting the expressions of target genes. E: In the absence of GA, SLR1 forms a ternary complex with PRC2 and HDA702, maintaining high H3K27me3 levels and low H3K9ac levels on GA-responsive genes and inhibiting their expression. F: In the presence of sufficient GA, SLR1 is ubiquitinated and degraded, unable to form complexes with PRC2 and HDA702, thus relieving the inhibition of GA-responsive genes
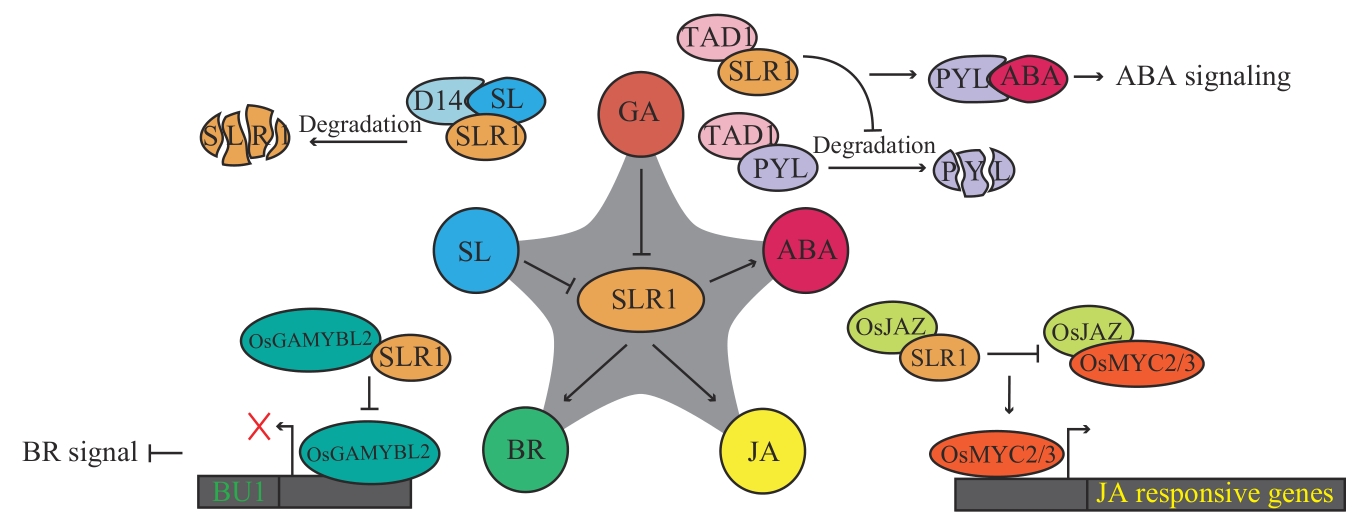
图3 SLR1参与其他植物激素信号途径圆形代表植物激素,椭圆形代表激素途径中的调节蛋白,分裂椭圆代表蛋白降解,→代表促进,┫或×代表抑制
Fig. 3 SLR1 involved in other plant hormone signaling pathwaysCircles refer to plant hormones, ellipses refers to regulatory proteins in hormone pathways, split ellipses refer to protein degradation, and “→” indicates promotion, while “┫” or “×” indicates inhibition
相关基因或蛋白 Related genes or proteins | 相关通路 Related pathways | 生物学功能 Biological function | 参考文献 Reference |
|---|---|---|---|
| GID1, GID2 | GA | 介导SLR1的降解,调控GA信号途径 Mediate the degradation of SLR1 and regulate the GA signaling pathway | [ |
| D14 | GA, SL | 促进SLR1的降解,调控水稻分蘖 Promote the degradation of SLR1 and regulate rice tillering | [ |
| D53 | SL, 氮素信号 | 高氮条件下,抑制D14介导SLR1的降解,调控氮素利用 Under high-nitrogen conditions, inhibit D14-mediated degradation of SLR1 to regulate nitrogen utilization | [ |
| PRC2, HDA702 | GA | 介导染色质沉默 Mediate chromatin silencing | [ |
| EL1 | GA | 促进SLR1磷酸化,调控GA信号途径 Promote the phosphorylation of SLR1, regulate the GA signaling pathway | [ |
| OsSPY | GA | 激活SLR1的抑制活性 Activate the inhibitory activity of SLR1 | [ |
| OsYABBY4 | GA | 调控小穗发育 Regulate spikelet development | [ |
| OsWRKY36 | GA | 抑制GA信号和调控植株 Inhibit GA signaling and regulate plant height | [ |
| OsMYB91 | GA, ABA | 平衡水稻生长和非生物胁迫抗性 Balance rice growth and abiotic stress resistance | [ |
| RTD1 | GA | 负调控SLR1蛋白的转录和蛋白水平 Negatively regulate the transcription and protein level of SLR1 protein | [ |
| OsNPC6 | GA | 调控中胚轴伸长 Regulate mesocotyl elongation | [ |
| CIPK31 | GA | 抑制SLR1蛋白降解 Inhibit the degradation of SLR1 protein | [ |
| OsGAMYBL2 | GA, BR | 调控GA合成和BR信号 Regulate GA biosynthesis and BR signaling | [ |
| OsJAZ8, OsJAZ9 | JA | 调控水稻生长发育与抗逆 Regulate rice growth and development as well as stress resistance | [ |
| MYC2/3 | JA | 调控JA响应基因的表达 Regulate the expression of JA-responsive genes | [ |
| SP8, P2, M | GA, JA | 介导广谱抗病毒防御反应 Mediate broad-spectrum antiviral defense response | [ |
| TAD1 | ABA | 抑制ABA信号 Inhibit ABA signaling | [ |
| MOC1 | GA | 调控株高和分蘖 Regulate plant height and tillering | [ |
| NGR5 | GA | 提高氮素利用、促进分蘖 Enhance nitrogen utilization and promote tillering | [ |
| OsMADS23 | GA, SL | 抑制D14基因转录,促进分蘖 Suppress the transcription of the D14 gene and promote tillering | [ |
| OsIDD2 | GA | 调控细胞增殖 Regulate cell proliferation | [ |
| OsNAC29/31 | GA | 调控纤维素合成 Regulate cellulose synthesis | [ |
| OsKNAT7 | GA | 调控次生细胞壁的合成 Regulate the synthesis of secondary cell walls | [ |
| OsMYB103L | GA | 调控纤维素生物和次生细胞壁合成 Regulate cellulose and secondary cell wall biosynthesis | [ |
| OsNAC055 | 木质素合成途径 | 调控木质素合成 Regulate lignin biosynthesis | [ |
| GAMYB | GA | 调控水稻育性 Regulate rice fertility | [ |
| UDT1, TDR | GA | 提高绒毡层发育相关基因的表达,调控水稻的育性 Enhance the expressions of tapetum development-related genes and regulate rice fertility | [ |
| OsMS188 | GA | 调控孢粉素生物合成,促进花粉壁的形成 Regulate sporopollenin biosynthesis and promote pollen wall formation | [ |
| OSH1 | GA | 调控次生细胞壁合成 Regulate secondary cell wall biosynthesis | [ |
| RID1 | GA | 调控营养生长向生殖生长的转变 Regulate the transition from vegetative growth to reproductive growth | [ |
| GHD7 | GA | 调控抽穗期和植株形态 Regulate rice heading date and plant morphology | [ |
| qSH1, OSH15, SNB | GA | 调控木质素含量和落粒性 Regulate lignin biosynthesis and seed shattering | [ |
| OsNAC120 | GA, ABA | 调控干旱胁迫响应 Regulate the response to drought stress | [ |
| OsBURP3, OsSUS1 | GA | 调控干旱胁迫响应 Regulate the response to drought stress | [ |
| OsPIL13/14 | GA, 光信号 | 调控黑暗或盐胁迫下的幼苗生长 Regulate seedling growth under darkness or salt stress | [ |
| OsNF-YA3 | GA, ABA | 调控生长发育与渗透胁迫耐受性 Regulate growth and development as well as osmotic stress tolerance | [ |
| IDD10, bZIP23 | GA, 氮素信号 | 调控氮素吸收和盐碱耐受性 Regulate nitrogen absorption and saline-alkali tolerance | [ |
| OsGRF6 | GA | 调控水稻生长和耐寒性 Regulate rice growth and cold tolerance | [ |
| OsNuCYP20-2 | GA | 促进SLR1的降解,促进细胞伸长 Promote the degradation of SLR1 and cell elongation | [ |
| Sub1A | GA, ET, ABA | 促进SLR1的积累,抑制GA信号 Promote the accumulation of SLR1 and inhibit GA signaling | [ |
| GRF4 | GA, BR | 调节根系代谢和氮素利用 Regulate root metabolism and nitrogen utilization | [ |
| OsWRKY71 | GA | 调控逆境下的根系发育 Regulate root development under stress conditions | [ |
| OsSPL7, OsSPL14 | GA | 调控Xoo的抗性 Regulate resistance to Xanthomonasoryzae pv. oryzae (Xoo) | [ |
| LPA1 | GA | 对水稻纹枯病rice sheath blight(ShB)的抗性 Resistance to rice sheath blight (ShB) | [ |
表1 与SLR1相关的基因或蛋白的生物学功能
Table 1 Biological functions of genes or proteins associated with SLR1
相关基因或蛋白 Related genes or proteins | 相关通路 Related pathways | 生物学功能 Biological function | 参考文献 Reference |
|---|---|---|---|
| GID1, GID2 | GA | 介导SLR1的降解,调控GA信号途径 Mediate the degradation of SLR1 and regulate the GA signaling pathway | [ |
| D14 | GA, SL | 促进SLR1的降解,调控水稻分蘖 Promote the degradation of SLR1 and regulate rice tillering | [ |
| D53 | SL, 氮素信号 | 高氮条件下,抑制D14介导SLR1的降解,调控氮素利用 Under high-nitrogen conditions, inhibit D14-mediated degradation of SLR1 to regulate nitrogen utilization | [ |
| PRC2, HDA702 | GA | 介导染色质沉默 Mediate chromatin silencing | [ |
| EL1 | GA | 促进SLR1磷酸化,调控GA信号途径 Promote the phosphorylation of SLR1, regulate the GA signaling pathway | [ |
| OsSPY | GA | 激活SLR1的抑制活性 Activate the inhibitory activity of SLR1 | [ |
| OsYABBY4 | GA | 调控小穗发育 Regulate spikelet development | [ |
| OsWRKY36 | GA | 抑制GA信号和调控植株 Inhibit GA signaling and regulate plant height | [ |
| OsMYB91 | GA, ABA | 平衡水稻生长和非生物胁迫抗性 Balance rice growth and abiotic stress resistance | [ |
| RTD1 | GA | 负调控SLR1蛋白的转录和蛋白水平 Negatively regulate the transcription and protein level of SLR1 protein | [ |
| OsNPC6 | GA | 调控中胚轴伸长 Regulate mesocotyl elongation | [ |
| CIPK31 | GA | 抑制SLR1蛋白降解 Inhibit the degradation of SLR1 protein | [ |
| OsGAMYBL2 | GA, BR | 调控GA合成和BR信号 Regulate GA biosynthesis and BR signaling | [ |
| OsJAZ8, OsJAZ9 | JA | 调控水稻生长发育与抗逆 Regulate rice growth and development as well as stress resistance | [ |
| MYC2/3 | JA | 调控JA响应基因的表达 Regulate the expression of JA-responsive genes | [ |
| SP8, P2, M | GA, JA | 介导广谱抗病毒防御反应 Mediate broad-spectrum antiviral defense response | [ |
| TAD1 | ABA | 抑制ABA信号 Inhibit ABA signaling | [ |
| MOC1 | GA | 调控株高和分蘖 Regulate plant height and tillering | [ |
| NGR5 | GA | 提高氮素利用、促进分蘖 Enhance nitrogen utilization and promote tillering | [ |
| OsMADS23 | GA, SL | 抑制D14基因转录,促进分蘖 Suppress the transcription of the D14 gene and promote tillering | [ |
| OsIDD2 | GA | 调控细胞增殖 Regulate cell proliferation | [ |
| OsNAC29/31 | GA | 调控纤维素合成 Regulate cellulose synthesis | [ |
| OsKNAT7 | GA | 调控次生细胞壁的合成 Regulate the synthesis of secondary cell walls | [ |
| OsMYB103L | GA | 调控纤维素生物和次生细胞壁合成 Regulate cellulose and secondary cell wall biosynthesis | [ |
| OsNAC055 | 木质素合成途径 | 调控木质素合成 Regulate lignin biosynthesis | [ |
| GAMYB | GA | 调控水稻育性 Regulate rice fertility | [ |
| UDT1, TDR | GA | 提高绒毡层发育相关基因的表达,调控水稻的育性 Enhance the expressions of tapetum development-related genes and regulate rice fertility | [ |
| OsMS188 | GA | 调控孢粉素生物合成,促进花粉壁的形成 Regulate sporopollenin biosynthesis and promote pollen wall formation | [ |
| OSH1 | GA | 调控次生细胞壁合成 Regulate secondary cell wall biosynthesis | [ |
| RID1 | GA | 调控营养生长向生殖生长的转变 Regulate the transition from vegetative growth to reproductive growth | [ |
| GHD7 | GA | 调控抽穗期和植株形态 Regulate rice heading date and plant morphology | [ |
| qSH1, OSH15, SNB | GA | 调控木质素含量和落粒性 Regulate lignin biosynthesis and seed shattering | [ |
| OsNAC120 | GA, ABA | 调控干旱胁迫响应 Regulate the response to drought stress | [ |
| OsBURP3, OsSUS1 | GA | 调控干旱胁迫响应 Regulate the response to drought stress | [ |
| OsPIL13/14 | GA, 光信号 | 调控黑暗或盐胁迫下的幼苗生长 Regulate seedling growth under darkness or salt stress | [ |
| OsNF-YA3 | GA, ABA | 调控生长发育与渗透胁迫耐受性 Regulate growth and development as well as osmotic stress tolerance | [ |
| IDD10, bZIP23 | GA, 氮素信号 | 调控氮素吸收和盐碱耐受性 Regulate nitrogen absorption and saline-alkali tolerance | [ |
| OsGRF6 | GA | 调控水稻生长和耐寒性 Regulate rice growth and cold tolerance | [ |
| OsNuCYP20-2 | GA | 促进SLR1的降解,促进细胞伸长 Promote the degradation of SLR1 and cell elongation | [ |
| Sub1A | GA, ET, ABA | 促进SLR1的积累,抑制GA信号 Promote the accumulation of SLR1 and inhibit GA signaling | [ |
| GRF4 | GA, BR | 调节根系代谢和氮素利用 Regulate root metabolism and nitrogen utilization | [ |
| OsWRKY71 | GA | 调控逆境下的根系发育 Regulate root development under stress conditions | [ |
| OsSPL7, OsSPL14 | GA | 调控Xoo的抗性 Regulate resistance to Xanthomonasoryzae pv. oryzae (Xoo) | [ |
| LPA1 | GA | 对水稻纹枯病rice sheath blight(ShB)的抗性 Resistance to rice sheath blight (ShB) | [ |
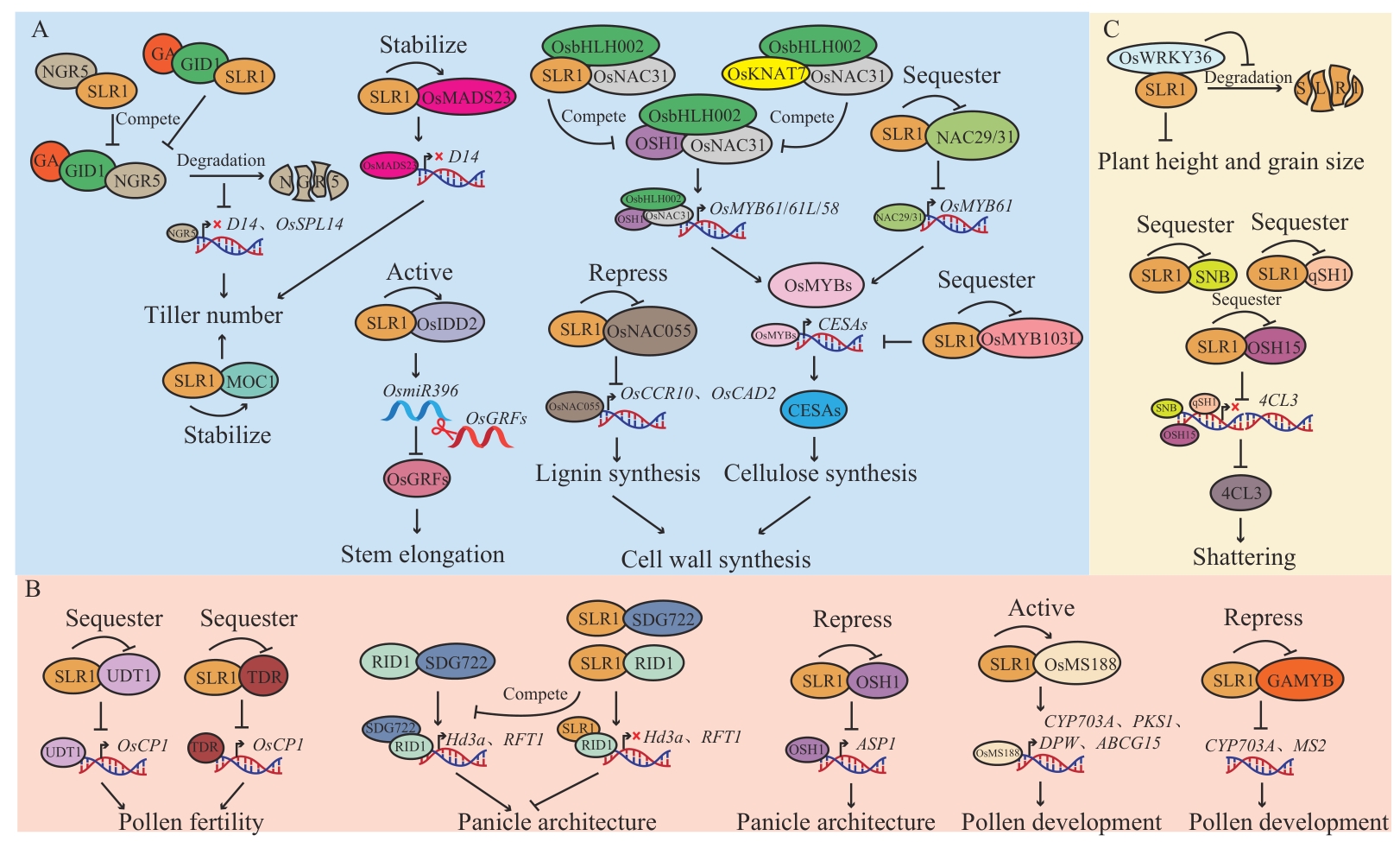
图4 SLR1调控水稻生长发育A:SLR1参与调控水稻营养生长;B:SLR1参与调控水稻花发育;C:SLR1参与调控水稻种子发育;椭圆形代表水稻生长发育途径中的调节蛋白,分裂椭圆代表蛋白降解,→代表促进,┫或×代表抑制
Fig. 4 SLR1 regulates rice growth and developmentA: SLR1 is involved in regulating the vegetative growth of rice; B: SLR1 is involved in regulating rice flower development; C: SLR1 is involved in regulating rice seed development. Ellipses rindicate regulatory proteins in rice growth and development pathways, split ellipses indicate protein degradation, “→” indicates promotion, and “┫” or “×” indicates inhibition
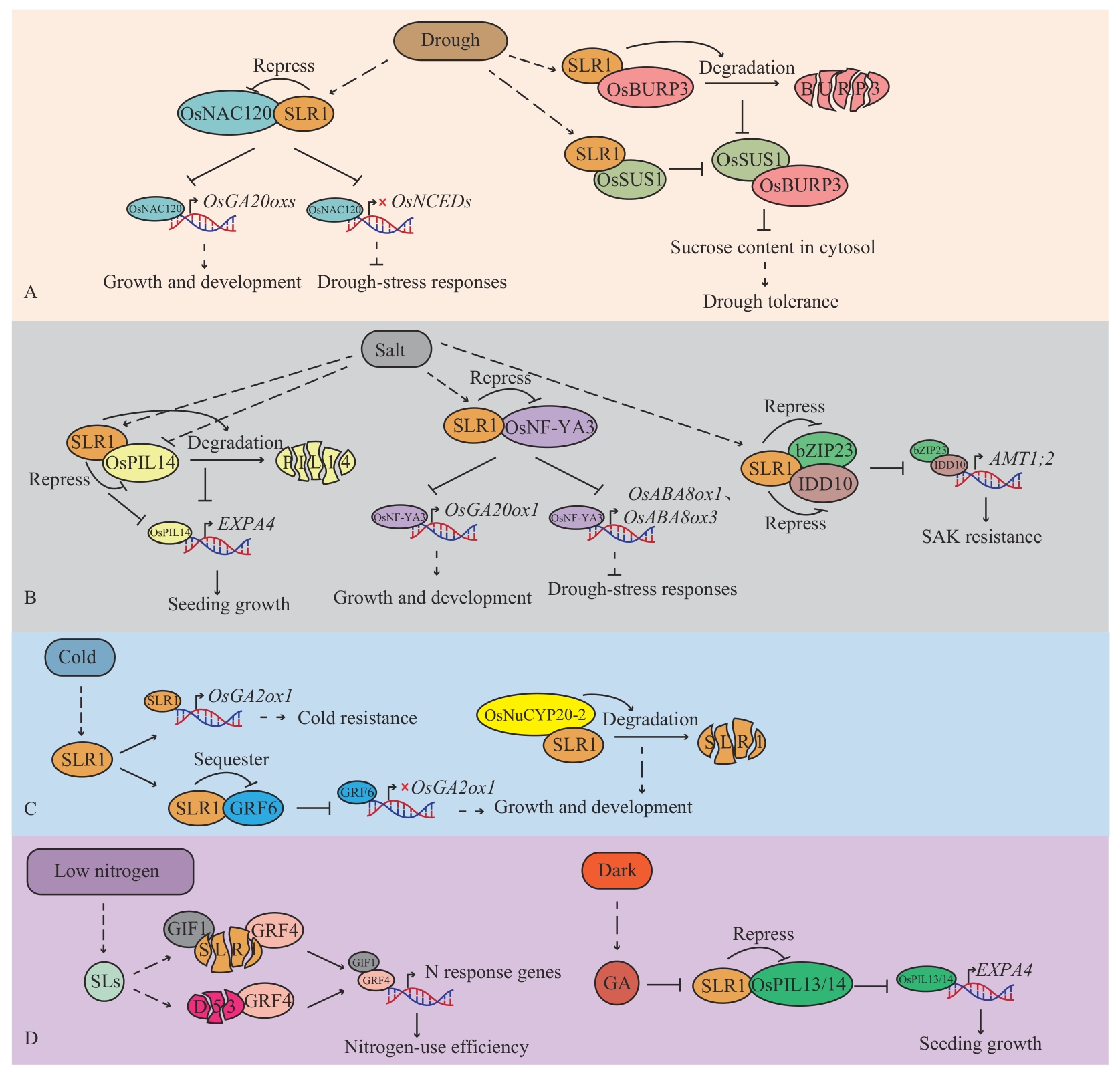
图5 SLR1调控非生物胁迫A:SLR1调控干旱胁迫;B:SLR1调控盐胁迫;C:SLR1调控冷胁迫;D:SLR1调控低氮胁迫。圆矩形代表非生物胁迫,椭圆形代表水稻响应非生物胁迫途径中的调节蛋白,分裂椭圆代表蛋白降解,→代表促进,┫或×代表抑制,虚线箭头代表间接促进
Fig. 5 SLR1 regulates abiotic stressA: SLR1 regulates drought stress; B: SLR1 regulates salt stress; C: SLR1 regulates cold stress; D: SLR1 regulates low-nitrogen stress. Rounded rectangles indicate abiotic stresses, Ellipses indicate regulatory proteins in the rice abiotic stress response pathway, split ellipses indicate protein degradation, “→” indicates promotion, “┫” or “×” indicates inhibition, and dashed arrows indicate indirect promotion
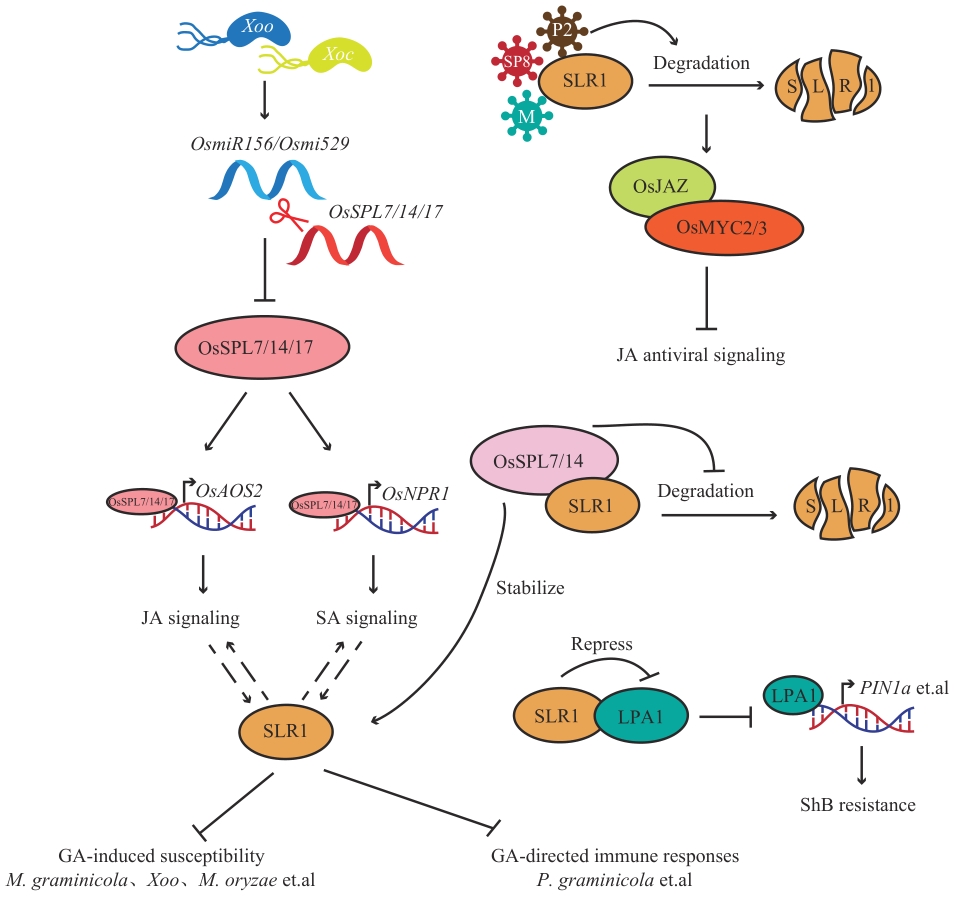
图6 SLR1调控生物胁迫椭圆形代表水稻响应生物胁迫途径中的调节蛋白,分裂椭圆代表蛋白降解,→代表促进,┫代表抑制,虚线箭头代表间接促进
Fig. 6 SLR1 regulates biological stressEllipses indicate regulatory proteins in the rice biotic stress response pathway, split ellipses indicate protein degradation, “→” indicates promotion, “┫” indicates inhibition, and dashed arrows indicate indirect promotion
| [1] | Hedden P, Proebsting WM. Genetic analysis of gibberellin biosynthesis [J]. Plant Physiol, 1999, 119(2): 365-370. |
| [2] | Olszewski N, Sun TP, Gubler F. Gibberellin signaling: biosynthesis, catabolism, and response pathways [J]. Plant Cell, 2002, 14(): S61-S80. |
| [3] | Richards DE, King KE, Ait-Ali T, et al. How gibberellin regulates plant growth and development: a molecular genetic analysis of gibberellin signaling [J]. Annu Rev Plant Physiol Plant Mol Biol, 2001, 52: 67-88. |
| [4] | Khush GS. Green revolution: preparing for the 21st century [J]. Genome, 1999, 42(4): 646-655. |
| [5] | Pingali PL. Green revolution: impacts, limits, and the path ahead [J]. Proc Natl Acad Sci USA, 2012, 109(31): 12302-12308. |
| [6] | Monna L, Kitazawa N, Yoshino R, et al. Positional cloning of rice semidwarfing gene, sd-1: rice "green revolution gene" encodes a mutant enzyme involved in gibberellin synthesis [J]. DNA Res, 2002, 9(1): 11-17. |
| [7] | Spielmeyer W, Ellis MH, Chandler PM. Semidwarf (sd-1), "green revolution" rice, contains a defective gibberellin 20-oxidase gene [J]. Proc Natl Acad Sci USA, 2002, 99(13): 9043-9048. |
| [8] | Hedden P. The genes of the green revolution [J]. Trends Genet, 2003, 19(1): 5-9. |
| [9] | Sun TP, Gubler F. Molecular mechanism of gibberellin signaling in plants [J]. Annu Rev Plant Biol, 2004, 55: 197-223. |
| [10] | Pysh LD, Wysocka-Diller JW, Camilleri C, et al. The GRAS gene family in Arabidopsis: sequence characterization and basic expression analysis of the SCARECROW-LIKE genes [J]. Plant J, 1999, 18(1): 111-119. |
| [11] | Ogawa M, Kusano T, Katsumi M, et al. Rice gibberellin-insensitive gene homolog, OsGAI, encodes a nuclear-localized protein capable of gene activation at transcriptional level [J]. Gene, 2000, 245(1): 21-29. |
| [12] | Fu XD, Richards DE, Ait-Ali T, et al. Gibberellin-mediated proteasome-dependent degradation of the barley DELLA protein SLN1 repressor [J]. Plant Cell, 2002, 14(12): 3191-3200. |
| [13] | Jiang CF, Fu XD. GA action: turning on de-DELLA repressing signaling [J]. Curr Opin Plant Biol, 2007, 10(5): 461-465. |
| [14] | Ikeda A, Ueguchi-Tanaka M, Sonoda Y, et al. Slender rice, a constitutive gibberellin response mutant, is caused by a null mutation of the SLR1 gene, an ortholog of the height-regulating gene GAI/RGA/RHT/D8 [J]. Plant Cell, 2001, 13(5): 999-1010. |
| [15] | Willige BC, Ghosh S, Nill C, et al. The DELLA domain of GA INSENSITIVE mediates the interaction with the GA INSENSITIVE DWARF1A gibberellin receptor of Arabidopsis [J]. Plant Cell, 2007, 19(4): 1209-1220. |
| [16] | Ueguchi-Tanaka M, Nakajima M, Katoh E, et al. Molecular interactions of a soluble gibberellin receptor, GID1, with a rice DELLA protein, SLR1, and gibberellin [J]. Plant Cell, 2007, 19(7): 2140-2155. |
| [17] | Van De Velde K, Ruelens P, Geuten K, et al. Exploiting DELLA signaling in cereals [J]. Trends Plant Sci, 2017, 22(10): 880-893. |
| [18] | Li MZ, An FY, Li WY, et al. DELLA proteins interact with FLC to repress flowering transition [J]. J Integr Plant Biol, 2016, 58(7): 642-655. |
| [19] | Gomi K, Sasaki A, Itoh H, et al. GID2, an F-box subunit of the SCF E3 complex, specifically interacts with phosphorylated SLR1 protein and regulates the gibberellin-dependent degradation of SLR1 in rice [J]. Plant J, 2004, 37(4): 626-634. |
| [20] | Itoh H, Ueguchi-Tanaka M, Sato Y, et al. The gibberellin signaling pathway is regulated by the appearance and disappearance of SLENDER RICE1 in nuclei [J]. Plant Cell, 2002, 14(1): 57-70. |
| [21] | 白云赫, 朱旭东, 樊秀彩, 等. 植物DELLA蛋白及其应答赤霉素信号调控植物生长发育的研究进展 [J]. 分子植物育种, 2019, 17(8): 2509-2516. |
| Bai YH, Zhu XD, Fan XC, et al. Reseach progress of plant DELLA proteins and its response to gibberellin signal regulating plant growth and development [J]. Mol Plant Breed, 2019, 17(8): 2509-2516. | |
| [22] | Hirano K, Asano K, Tsuji H, et al. Characterization of the molecular mechanism underlying gibberellin perception complex formation in rice [J]. Plant Cell, 2010, 22(8): 2680-2696. |
| [23] | Hellens RP, Allan AC, Friel EN, et al. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants [J]. Plant Methods, 2005, 1: 13. |
| [24] | Hauvermale AL, Ariizumi T, Steber CM. Gibberellin signaling: a theme and variations on DELLA repression [J]. Plant Physiol, 2012, 160(1): 83-92. |
| [25] | Ueguchi-Tanaka M. Gibberellin metabolism and signaling [J]. Biosci Biotechnol Biochem, 2023, 87(10): 1093-1101. |
| [26] | Phokas A, Coates JC. Evolution of DELLA function and signaling in land plants [J]. Evol Dev, 2021, 23(3): 137-154. |
| [27] | Yano K, Aya K, Hirano K, et al. Comprehensive gene expression analysis of rice aleurone cells: probing the existence of an alternative gibberellin receptor [J]. Plant Physiol, 2015, 167(2): 531-544. |
| [28] | Sun HW, Guo XL, Zhu XL, et al. Strigolactone and gibberellin signaling coordinately regulate metabolic adaptations to changes in nitrogen availability in rice [J]. Mol Plant, 2023, 16(3): 588-598. |
| [29] | Zhao Y, Zhou DX. Epigenomic modification and epigenetic regulation in rice [J]. J Genet Genom, 2012, 39(7): 307-315. |
| [30] | Chen XS, Zhou DX. Rice epigenomics and epigenetics: challenges and opportunities [J]. Curr Opin Plant Biol, 2013, 16(2): 164-169. |
| [31] | Li TT, Chen XS, Zhong XC, et al. Jumonji C domain protein JMJ705-mediated removal of histone H3 lysine 27 trimethylation is involved in defense-related gene activation in rice [J]. Plant Cell, 2013, 25(11): 4725-4736. |
| [32] | Li JJ, Li Q, Wang WT, et al. DELLA-mediated gene repression is maintained by chromatin modification in rice [J]. EMBO J, 2023, 42(21): e114220. |
| [33] | Itoh H, Matsuoka M, Steber CM. A role for the ubiquitin-26S-proteasome pathway in gibberellin signaling [J]. Trends Plant Sci, 2003, 8(10): 492-497. |
| [34] | Mashita O, Koishihara H, Fukui K, et al. Discovery and identification of 2-methoxy-1-naphthaldehyde as a novel strigolactone-signaling inhibitor [J]. J Pestic Sci, 2016, 41(3): 71-78. |
| [35] | Han SN, Liu YX, Bao A, et al. OsCSN1 regulates the growth of rice seedlings through the GA signaling pathway in blue light [J]. J Plant Physiol, 2023, 280: 153904. |
| [36] | Fernandes T, Gonçalves NM, Matiolli CC, et al. SUMOylation of rice DELLA SLR1 modulates transcriptional responses and improves yield under salt stress [J]. Planta, 2024, 260(6): 136. |
| [37] | Itoh H, Sasaki A, Ueguchi-Tanaka M, et al. Dissection of the phosphorylation of rice DELLA protein, SLENDER RICE1 [J]. Plant Cell Physiol, 2005, 46(8): 1392-1399. |
| [38] | Dai C, Xue HW. Rice early flowering1, a CKI, phosphorylates DELLA protein SLR1 to negatively regulate gibberellin signalling [J]. EMBO J, 2010, 29(11): 1916-1927. |
| [39] | Shimada A, Ueguchi-Tanaka M, Sakamoto T, et al. The rice SPINDLY gene functions as a negative regulator of gibberellin signaling by controlling the suppressive function of the DELLA protein, SLR1 and modulating brassinosteroid synthesis [J]. Plant J, 2006, 48(3): 390-402. |
| [40] | Yano K, Morinaka Y, Wang FM, et al. GWAS with principal component analysis identifies a gene comprehensively controlling rice architecture [J]. Proc Natl Acad Sci USA, 2019, 116(42): 21262-21267. |
| [41] | Yang C, Ma YM, Li JX. The rice YABBY4 gene regulates plant growth and development through modulating the gibberellin pathway [J]. J Exp Bot, 2016, 67(18): 5545-5556. |
| [42] | Lan J, Lin QB, Zhou CL, et al. Small grain and semi-dwarf 3, a WRKY transcription factor, negatively regulates plant height and grain size by stabilizing SLR1 expression in rice [J]. Plant Mol Biol, 2020, 104(4/5): 429-450. |
| [43] | Zhu N, Cheng SF, Liu XY, et al. The R2R3-type MYB gene OsMYB91 has a function in coordinating plant growth and salt stress tolerance in rice [J]. Plant Sci, 2015, 236: 146-156. |
| [44] | Zhang YH, Liu K, Zhu XM, et al. Rice tocopherol deficiency 1 encodes a homogentisate phytyltransferase essential for tocopherol biosynthesis and plant development in rice [J]. Plant Cell Rep, 2018, 37(5): 775-787. |
| [45] | Yang D, Liu X, Yin XM, et al. Rice non-specific phospholipase C6 is involved in mesocotyl elongation [J]. Plant Cell Physiol, 2021, 62(6): 985-1000. |
| [46] | Chen JS, Wang ST, Jiang SQ, et al. Overexpression of Calcineurin B-like interacting protein kinase 31 promotes lodging and sheath blight resistance in rice [J]. Plants, 2024, 13(10): 1306. |
| [47] | Davière JM, Achard P. A pivotal role of DELLAs in regulating multiple hormone signals [J]. Mol Plant, 2016, 9(1): 10-20. |
| [48] | 赵春丽, 王晓, 陈家兰, 等. 植物DELLA蛋白家族研究进展 [J]. 应用与环境生物学报, 2020, 26(5): 1299-1308. |
| Zhao CL, Wang X, Chen JL, et al. Progress in research on plant DELLA family proteins [J]. Chin J Appl Environ Biol, 2020, 26(5): 1299-1308. | |
| [49] | Eckardt NA. GA perception and signal transduction: molecular interactions of the GA receptor GID1 with GA and the DELLA protein SLR1 in rice [J]. Plant Cell, 2007, 19(7): 2095-2097. |
| [50] | Asano K, Hirano K, Ueguchi-Tanaka M, et al. Isolation and characterization of dominant dwarf mutants, Slr1-d, in rice [J]. Mol Genet Genomics, 2009, 281(2): 223-231. |
| [51] | Jung YJ, Kim JH, Lee HJ, et al. Generation and transcriptome profiling of Slr1-d7 and Slr1-d8 mutant lines with a new semi-dominant dwarf allele of SLR1 using the CRISPR/Cas9 system in rice [J]. Int J Mol Sci, 2020, 21(15): 5492. |
| [52] | Zentella R, Zhang ZL, Park M, et al. Global analysis of DELLA direct targets in early gibberellin signaling in Arabidopsis [J]. Plant Cell, 2007, 19(10): 3037-3057. |
| [53] | Tsuji H, Aya K, Ueguchi-Tanaka M, et al. GAMYB controls different sets of genes and is differentially regulated by microRNA in aleurone cells and anthers [J]. Plant J, 2006, 47(3): 427-444. |
| [54] | Gao J, Chen H, Yang HF, et al. A brassinosteroid responsive miRNA-target module regulates gibberellin biosynthesis and plant development [J]. New Phytol, 2018, 220(2): 488-501. |
| [55] | Hou XL, Lee LYC, Xia KF, et al. DELLAs modulate jasmonate signaling via competitive binding to JAZs [J]. Dev Cell, 2010, 19(6): 884-894. |
| [56] | Um TY, Lee HY, Lee S, et al. Jasmonate zim-domain protein 9 interacts with slender rice 1 to mediate the antagonistic interaction between jasmonic and gibberellic acid signals in rice [J]. Front Plant Sci, 2018, 9: 1866. |
| [57] | Li LL, Zhang HH, Yang ZH, et al. Independently evolved viral effectors convergently suppress DELLA protein SLR1-mediated broad-spectrum antiviral immunity in rice [J]. Nat Commun, 2022, 13(1): 6920. |
| [58] | Nakamura H, Xue YL, Miyakawa T, et al. Molecular mechanism of strigolactone perception by DWARF14 [J]. Nat Commun, 2013, 4: 2613. |
| [59] | Liao ZG, Zhang YC, Yu Q, et al. Coordination of growth and drought responses by GA-ABA signaling in rice [J]. New Phytol, 2023, 240(3): 1149-1161. |
| [60] | Liao ZG, Yu H, Duan JB, et al. SLR1 inhibits MOC1 degradation to coordinate tiller number and plant height in rice [J]. Nat Commun, 2019, 10(1): 2738. |
| [61] | Wu K, Wang SS, Song WZ, et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice [J]. Science, 2020, 367(6478): eaaz2046. |
| [62] | Li XX, Xie ZZ, Qin T, et al. The SLR1-OsMADS23-D14 module mediates the crosstalk between strigolactone and gibberellin signaling to control rice tillering [J]. New Phytol, 2025, 246(5): 2137-2154. |
| [63] | Zhuang H, Li YF. Strigolactone and gibberellin crosstalk: the role of the SLR1-OsMADS23-D14 module in regulating rice tiller development [J]. New Phytol, 2025, 246(5): 1893-1895. |
| [64] | Lu YZ, Feng Z, Meng YL, et al. SLENDER RICE1 and Oryza sativa INDETERMINATE DOMAIN2 regulating OsmiR396 are involved in stem elongation [J]. Plant Physiol, 2020, 182(4): 2213-2227. |
| [65] | Huang DB, Wang SG, Zhang BC, et al. A gibberellin-mediated DELLA-NAC signaling cascade regulates cellulose synthesis in rice [J]. Plant Cell, 2015, 27(6): 1681-1696. |
| [66] | Chen Y, Qi HY, Yang LJ, et al. The OsbHLH002/OsICE1-OSH1 module orchestrates secondary cell wall formation in rice [J]. Cell Rep, 2023, 42(7): 112702. |
| [67] | Ye YF, Liu BM, Zhao M, et al. CEF1/OsMYB103L is involved in GA-mediated regulation of secondary wall biosynthesis in rice [J]. Plant Mol Biol, 2015, 89(4/5): 385-401. |
| [68] | Liu YF, Wu Q, Qin ZL, et al. Transcription factor OsNAC055 regulates GA-mediated lignin biosynthesis in rice straw [J]. Plant Sci, 2022, 325: 111455. |
| [69] | Aya K, Ueguchi-Tanaka M, Kondo M, et al. Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB [J]. Plant Cell, 2009, 21(5): 1453-1472. |
| [70] | 宋欣玥. SLR1调控水稻雄蕊发育的功能研究 [D]. 上海: 上海师范大学, 2023. |
| Song XY. Study on the function of SLR1 in regulating stamen development in rice [D]. Shanghai: Shanghai Normal University, 2023. | |
| [71] | Tang JQ, Tian XJ, Mei EY, et al. WRKY53 negatively regulates rice cold tolerance at the booting stage by fine-tuning anther gibberellin levels [J]. Plant Cell, 2022, 34(11): 4495-4515. |
| [72] | Jin Y, Song XY, Chang HZ, et al. The GA-DELLA-OsMS188 module controls male reproductive development in rice [J]. New Phytol, 2022, 233(6): 2629-2642. |
| [73] | Su S, Hong J, Chen XF, et al. Gibberellins orchestrate panicle architecture mediated by DELLA-KNOX signalling in rice [J]. Plant Biotechnol J, 2021, 19(11): 2304-2318. |
| [74] | Zhang S, Deng L, Cheng R, et al. RID1 sets rice heading date by balancing its binding with SLR1 and SDG722 [J]. J Integr Plant Biol, 2022, 64(1): 149-165. |
| [75] | Liu RJ, Feng QF, Li PB, et al. GLW7.1, a strong functional allele of Ghd7, enhances grain size in rice [J]. Int J Mol Sci, 2022, 23(15): 8715. |
| [76] | Wu H, He Q, He B, et al. Gibberellin signaling regulates lignin biosynthesis to modulate rice seed shattering [J]. Plant Cell, 2023, 35(12): 4383-4404. |
| [77] | Xie ZZ, Jin L, Sun Y, et al. OsNAC120 balances plant growth and drought tolerance by integrating GA and ABA signaling in rice [J]. Plant Commun, 2024, 5(3): 100782. |
| [78] | Huang JS, Xie B, Xian FJ, et al. Gibberellin signalling mediates nucleocytoplasmic trafficking of Sucrose Synthase 1 to regulate the drought tolerance in rice [J]. Plant Biotechnol J, 2025, 23(6): 1909-1926. |
| [79] | Lyu YS, Dong XL, Niu SP, et al. An orchestrated ethylene-gibberellin signaling cascade contributes to mesocotyl elongation and emergence of rice direct seeding [J]. J Integr Plant Biol, 2024, 66(7): 1427-1439. |
| [80] | Mo WP, Tang WJ, Du YX, et al. PHYTOCHROME-INTERACTING FACTOR-LIKE14 and SLENDER RICE1 interaction controls seedling growth under salt stress [J]. Plant Physiol, 2020, 184(1): 506-517. |
| [81] | Jin XK, Zhang YF, Li XX, et al. OsNF-YA3 regulates plant growth and osmotic stress tolerance by interacting with SLR1 and SAPK9 in rice [J]. Plant J, 2023, 114(4): 914-933. |
| [82] | Li Z, Chen H, Guan QJ, et al. Gibberellic acid signaling promotes resistance to saline-alkaline stress by increasing the uptake of ammonium in rice [J]. Plant Physiol Biochem, 2024, 207: 108424. |
| [83] | Li ZT, Wang B, Zhang ZY, et al. OsGRF6 interacts with SLR1 to regulate OsGA2ox1 expression for coordinating chilling tolerance and growth in rice [J]. J Plant Physiol, 2021, 260: 153406. |
| [84] | Ge Q, Zhang YY, Xu YY, et al. Cyclophilin OsCYP20-2 with a novel variant integrates defense and cell elongation for chilling response in rice [J]. New Phytol, 2020, 225(6): 2453-2467. |
| [85] | Mittal L, Tayyeba S, Sinha AK. Finding a breather for Oryza sativa: Understanding hormone signalling pathways involved in rice plants to submergence stress [J]. Plant Cell Environ, 2022, 45(2): 279-295. |
| [86] | Fukao T, Bailey-Serres J. Submergence tolerance conferred by Sub1A is mediated by SLR1 and SLRL1 restriction of gibberellin responses in rice [J]. Proc Natl Acad Sci USA, 2008, 105(43): 16814-16819. |
| [87] | Schmitz AJ, Folsom JJ, Jikamaru Y, et al. SUB1A-mediated submergence tolerance response in rice involves differential regulation of the brassinosteroid pathway [J]. New Phytol, 2013, 198(4): 1060-1070. |
| [88] | Li S, Tian YH, Wu K, et al. Modulating plant growth-metabolism coordination for sustainable agriculture [J]. Nature, 2018, 560(7720): 595-600. |
| [89] | Mirza Z, Gupta M. Iron reprogrammes the root system architecture by regulating OsWRKY71 in arsenic-stressed rice (Oryza sativa L.) [J]. Plant Mol Biol, 2024, 114(1): 11. |
| [90] | De Vleesschauwer D, Seifi HS, Filipe O, et al. The DELLA protein SLR1 integrates and amplifies salicylic acid- and jasmonic acid-dependent innate immunity in rice [J]. Plant Physiol, 2016, 170(3): 1831-1847. |
| [91] | Liu MM, Shi ZY, Zhang XH, et al. Inducible overexpression of Ideal Plant Architecture1 improves both yield and disease resistance in rice [J]. Nat Plants, 2019, 5(4): 389-400. |
| [92] | Sun Q, Li TY, Li DD, et al. Overexpression of Loose Plant Architecture 1 increases planting density and resistance to sheath blight disease via activation of PIN-FORMED 1a in rice [J]. Plant Biotechnol J, 2019, 17(5): 855-857. |
| [93] | Sakata T, Oda S, Tsunaga Y, et al. Reduction of gibberellin by low temperature disrupts pollen development in rice [J]. Plant Physiol, 2014, 164(4): 2011-2019. |
| [94] | Guo SQ, Chen YX, Ju YL, et al. Fine-tuning gibberellin improves rice alkali-thermal tolerance and yield [J]. Nature, 2025, 639(8053): 162-171. |
| [95] | Lu L, Chen XY, Tan QY, et al. Gibberellin-mediated sensitivity of rice roots to aluminum stress [J]. Plants, 2024, 13(4): 543. |
| [96] | Hui SG, Ke YG, Chen D, et al. Rice microRNA156/529- SQUAMOSA PROMOTER BINDING PROTEIN-LIKE7/14/17 modules regulate defenses against bacteria [J]. Plant Physiol, 2023, 192(3): 2537-2553. |
| [97] | Zhu HY, Chen H, Kantharaj V, et al. SLR1-LPA1 signal regulates sheath blight resistance and lamina joint angle in rice [J]. Plant Physiol Biochem, 2025, 222: 109689. |
| [98] | De Vleesschauwer D, Van Buyten E, Satoh K, et al. Brassinosteroids antagonize gibberellin- and salicylate-mediated root immunity in rice [J]. Plant Physiol, 2012, 158(4): 1833-1846. |
| [99] | Zhang J, Luo T, Wang WW, et al. Silencing OsSLR1 enhances the resistance of rice to the brown planthopper Nilaparvata lugens [J]. Plant Cell Environ, 2017, 40(10): 2147-2159. |
| [100] | Yimer HZ, Nahar K, Kyndt T, et al. Gibberellin antagonizes jasmonate-induced defense against Meloidogyne graminicola in rice [J]. New Phytol, 2018, 218(2): 646-660. |
| [101] | Itoh H, Shimada A, Ueguchi-Tanaka M, et al. Overexpression of a GRAS protein lacking the DELLA domain confers altered gibberellin responses in rice [J]. Plant J, 2005, 44(4): 669-679. |
| [102] | Liu T, Gu JY, Xu CJ, et al. Overproduction of OsSLRL2 alters the development of transgenic Arabidopsis plants [J]. Biochem Biophys Res Commun, 2007, 358(4): 983-989. |
| [103] | Wang JD, Wang J, Huang LC, et al. ABA-mediated regulation of rice grain quality and seed dormancy via the NF-YB1-SLRL2-bHLH144 Module [J]. Nat Commun, 2024, 15(1): 4493. |
| [1] | 杨跃琴, 邢英, 仲子荷, 田维军, 杨雪清, 王建旭. 甲基汞胁迫下水稻OsMATE34的表达及功能分析[J]. 生物技术通报, 2026, 42(1): 86-94. |
| [2] | 王芳, 邵会茹, 吕林龙, 赵点, 胡振, 吕建珍, 姜亮. 植物和细菌TurboID邻近蛋白标记方法的建立[J]. 生物技术通报, 2025, 41(9): 44-53. |
| [3] | 邓美壁, 严浪, 詹志田, 朱敏, 和玉兵. RUBY辅助的水稻高效CRISPR基因编辑[J]. 生物技术通报, 2025, 41(8): 65-73. |
| [4] | 侯鹰翔, 费思恬, 黎妮, 李兰, 宋松泉, 王伟平, 张超. 水稻miRNAs响应生物胁迫研究进展[J]. 生物技术通报, 2025, 41(7): 69-80. |
| [5] | 吴浩, 董伟峰, 贺子天, 李艳肖, 谢辉, 孙明哲, 沈阳, 孙晓丽. 水稻BXL基因家族的全基因组鉴定及表达分析[J]. 生物技术通报, 2025, 41(6): 87-98. |
| [6] | 杜量衡, 唐黄磊, 张治国. 控制水稻光响应基因ELM1的图位克隆[J]. 生物技术通报, 2025, 41(5): 82-89. |
| [7] | 刘园园, 陈析丰, 钱前, 高振宇. 水稻穗发育调控的分子机制研究进展[J]. 生物技术通报, 2025, 41(5): 1-13. |
| [8] | 陈晓军, 惠建, 马洪文, 白海波, 钟楠, 李稼润, 樊云芳. 利用单碱基基因编辑技术创制OsALS抗除草剂水稻种质资源[J]. 生物技术通报, 2025, 41(4): 106-114. |
| [9] | 覃悦, 杨妍, 张磊, 卢丽丽, 李先平, 蒋伟. 二倍体和四倍体马铃薯StGAox基因鉴定与比较分析[J]. 生物技术通报, 2025, 41(3): 146-160. |
| [10] | 李欣芃, 张武汉, 张莉, 舒服, 何强, 郭杨, 邓华凤, 王悦, 孙平勇. γ射线诱变创制水稻突变体及其分子鉴定[J]. 生物技术通报, 2025, 41(3): 35-43. |
| [11] | 方慧敏, 顾艺枢, 张晶, 张龙. 水稻叶片淀粉的分离与理化性质分析[J]. 生物技术通报, 2025, 41(2): 51-57. |
| [12] | 段若昕, 陈盈盈, 林金星, 李瑞丽. 种子休眠及其调控机制研究进展[J]. 生物技术通报, 2025, 41(12): 16-26. |
| [13] | 张晓丹, 尹铮, 刘清晨, 李雪梅, 刘晓华, 梁美霞. 基于代谢组和转录组联合解析山桃响应冻害机制[J]. 生物技术通报, 2025, 41(12): 225-239. |
| [14] | 王若若, 曲鹏坤, 张欣, 王珞, 朱英, 田双一, 赵德刚. 杜仲EuGIF1基因鉴定及其表达和蛋白互作分析[J]. 生物技术通报, 2025, 41(12): 267-279. |
| [15] | 范艳飞, 叶露幻, 李雨桐, 王钏跞, 张瑞, 罗建华, 王鹏. 运用小麦杂交株系挖掘环式光合电子传递调控基因并应用于作物高光效改造[J]. 生物技术通报, 2025, 41(10): 72-86. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||