








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (3): 202-214.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0982
Previous Articles Next Articles
TIAN Chun-yan( ), LI Xu-juan, LI Chun-jia, MAO Jun, LIU Xin-long(
), LI Xu-juan, LI Chun-jia, MAO Jun, LIU Xin-long( )
)
Received:2023-10-19
Online:2024-03-26
Published:2024-04-08
Contact:
LIU Xin-long
E-mail:tianchy89@126.com;lxlgood868@163.com
TIAN Chun-yan, LI Xu-juan, LI Chun-jia, MAO Jun, LIU Xin-long. Genome-wide Analysis of Codon Usage Bias in Saccharum Species and Its Phylogenetically Related Species Erianthus fulvus[J]. Biotechnology Bulletin, 2024, 40(3): 202-214.
| 核苷酸组成和ENC Nucleotide composition and ENC | 材料 Materials | 平均值 Mean | |||
|---|---|---|---|---|---|
| LA-purple | NP-X | AP85-441 | Yunnan2009-3 | ||
| A3s/% | 15.61±8.47 | 15.65±8.14 | 14.91±8.01 | 16.41±8.38 | 15.65 |
| T3s/% | 20.19±11.13 | 19.95±10.52 | 19.00±10.41 | 21.14±10.95 | 20.07 |
| G3s/% | 31.24±8.23 | 31.11±7.78 | 31.95±7.95 | 30.42±7.84 | 31.18 |
| C3s/% | 32.97±12.91 | 33.28±12.31 | 34.14±12.13 | 32.04±12.66 | 33.11 |
| GC/% | 56.21±0.10 | 56.42±0.09 | 57.53±0.09 | 55.02±0.09 | 56.30 |
| GC1/% | 58.39±7.29 | 58.53±6.76 | 59.29±6.80 | 57.67±6.86 | 58.47 |
| GC2/% | 45.73±7.79 | 46.05±7.34 | 46.87±7.57 | 44.67±7.06 | 45.83 |
| GC3/% | 64.21±18.91 | 64.39±18.02 | 66.09±17.78 | 62.46±18.68 | 64.29 |
| GC3s/% | 62.89±0.20 | 63.10±0.19 | 64.91±0.18 | 61.06±0.19 | 62.99 |
| ENC | 47.97±9.11 | 48.74±8.93 | 48.19±9.26 | 48.89±8.82 | 48.45 |
Table 1 Nucleotide composition and ENC of CDS sequences in Saccharum species and its phylogenetically related species E. fulvus
| 核苷酸组成和ENC Nucleotide composition and ENC | 材料 Materials | 平均值 Mean | |||
|---|---|---|---|---|---|
| LA-purple | NP-X | AP85-441 | Yunnan2009-3 | ||
| A3s/% | 15.61±8.47 | 15.65±8.14 | 14.91±8.01 | 16.41±8.38 | 15.65 |
| T3s/% | 20.19±11.13 | 19.95±10.52 | 19.00±10.41 | 21.14±10.95 | 20.07 |
| G3s/% | 31.24±8.23 | 31.11±7.78 | 31.95±7.95 | 30.42±7.84 | 31.18 |
| C3s/% | 32.97±12.91 | 33.28±12.31 | 34.14±12.13 | 32.04±12.66 | 33.11 |
| GC/% | 56.21±0.10 | 56.42±0.09 | 57.53±0.09 | 55.02±0.09 | 56.30 |
| GC1/% | 58.39±7.29 | 58.53±6.76 | 59.29±6.80 | 57.67±6.86 | 58.47 |
| GC2/% | 45.73±7.79 | 46.05±7.34 | 46.87±7.57 | 44.67±7.06 | 45.83 |
| GC3/% | 64.21±18.91 | 64.39±18.02 | 66.09±17.78 | 62.46±18.68 | 64.29 |
| GC3s/% | 62.89±0.20 | 63.10±0.19 | 64.91±0.18 | 61.06±0.19 | 62.99 |
| ENC | 47.97±9.11 | 48.74±8.93 | 48.19±9.26 | 48.89±8.82 | 48.45 |
| ENC值 ENC value | LA-purple | NP-X | AP85-441 | Yunnan2009-3 | ||||
|---|---|---|---|---|---|---|---|---|
| 数目 | 百分比/% | 数目 | 百分比/% | 数目 | 百分比/% | 数目 | 百分比/% | |
| ENC≤35 | 34 381 | 14.15 | 15 880 | 12.10 | 8 642 | 14.23 | 4 290 | 12.05 |
| 35<ENC≤45 | 46 920 | 19.32 | 24 450 | 18.64 | 11 548 | 19.07 | 5 955 | 16.72 |
| ENC˃45 | 161 603 | 66.53 | 90 849 | 69.26 | 40 414 | 66.70 | 25 371 | 71.23 |
Table 2 Frequency distribution and percentage of ENC
| ENC值 ENC value | LA-purple | NP-X | AP85-441 | Yunnan2009-3 | ||||
|---|---|---|---|---|---|---|---|---|
| 数目 | 百分比/% | 数目 | 百分比/% | 数目 | 百分比/% | 数目 | 百分比/% | |
| ENC≤35 | 34 381 | 14.15 | 15 880 | 12.10 | 8 642 | 14.23 | 4 290 | 12.05 |
| 35<ENC≤45 | 46 920 | 19.32 | 24 450 | 18.64 | 11 548 | 19.07 | 5 955 | 16.72 |
| ENC˃45 | 161 603 | 66.53 | 90 849 | 69.26 | 40 414 | 66.70 | 25 371 | 71.23 |
| 氨基酸 Amino acid | 密码子 Codon | LA-purple | NP-X | AP85-441 | Yunnan2009-3 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GW | HEB | LEB | ∆RSCU | GW | HEB | LEB | ∆RSCU | GW | HEB | LEB | ∆RSCU | GW | HEB | LEB | ∆RSCU | |||||
| Phe | UUU | 0.76 | 0.05 | 0.95 | -0.90 | 0.73 | 0.06 | 0.95 | -0.89 | 0.69 | 0.19 | 0.93 | -0.74 | 0.78 | 0.06 | 0.96 | -0.90 | |||
| UUC** | 1.24 | 1.95 | 1.05 | 0.90 | 1.27 | 1.94 | 1.05 | 0.89 | 1.31 | 1.81 | 1.07 | 0.74 | 1.22 | 1.94 | 1.04 | 0.90 | ||||
| Leu | UUA | 0.39 | 0.01 | 0.53 | -0.52 | 0.37 | 0.01 | 0.52 | -0.51 | 0.33 | 0.04 | 0.51 | -0.47 | 0.41 | 0.01 | 0.53 | -0.52 | |||
| UUG | 0.94 | 0.14 | 1.11 | -0.97 | 0.92 | 0.16 | 1.11 | -0.95 | 0.86 | 0.08 | 1.10 | -1.02 | 0.97 | 0.15 | 1.12 | -0.97 | ||||
| CUU | 1.04 | 0.13 | 1.27 | -1.14 | 1.03 | 0.15 | 1.24 | -1.09 | 1.00 | 1.30 | 1.26 | 0.04 | 1.08 | 0.14 | 1.26 | -1.12 | ||||
| CUC** | 1.59 | 2.99 | 1.20 | 1.79 | 1.60 | 2.95 | 1.21 | 1.74 | 1.72 | 2.09 | 1.23 | 0.86 | 1.51 | 2.94 | 1.19 | 1.75 | ||||
| CUA | 0.50 | 0.07 | 0.66 | -0.59 | 0.50 | 0.08 | 0.67 | -0.59 | 0.46 | 0.88 | 0.65 | 0.23 | 0.54 | 0.08 | 0.67 | -0.59 | ||||
| CUG** | 1.54 | 2.65 | 1.24 | 1.41 | 1.58 | 2.65 | 1.24 | 1.41 | 1.64 | 1.61 | 1.26 | 0.35 | 1.49 | 2.67 | 1.22 | 1.45 | ||||
| Ile | AUU | 1.04 | 0.10 | 1.17 | -1.07 | 1.01 | 0.11 | 1.16 | -1.05 | 0.97 | 0.31 | 1.17 | -0.86 | 1.05 | 0.12 | 1.17 | -1.05 | |||
| AUC** | 1.34 | 2.81 | 1.04 | 1.77 | 1.35 | 2.78 | 1.04 | 1.74 | 1.43 | 2.48 | 1.05 | 1.43 | 1.29 | 2.78 | 1.03 | 1.75 | ||||
| AUA | 0.63 | 0.09 | 0.78 | -0.69 | 0.64 | 0.10 | 0.79 | -0.69 | 0.60 | 0.21 | 0.78 | -0.57 | 0.66 | 0.10 | 0.80 | -0.70 | ||||
| Val | GUU | 0.95 | 0.09 | 1.23 | -1.14 | 0.94 | 0.1 | 1.21 | -1.11 | 0.90 | 0.78 | 1.22 | -0.44 | 1.00 | 0.09 | 1.23 | -1.14 | |||
| GUC | 1.15 | 1.69 | 0.97 | 0.72 | 1.17 | 1.69 | 0.99 | 0.70 | 1.22 | 1.48 | 0.99 | 0.49 | 1.12 | 1.67 | 0.97 | 0.70 | ||||
| GUA | 0.41 | 0.04 | 0.58 | -0.54 | 0.41 | 0.05 | 0.58 | -0.53 | 0.38 | 0.53 | 0.57 | -0.04 | 0.44 | 0.05 | 0.59 | -0.54 | ||||
| GUG** | 1.48 | 2.18 | 1.22 | 0.96 | 1.48 | 2.17 | 1.22 | 0.95 | 1.50 | 1.21 | 1.01 | 0.20 | 1.44 | 2.19 | 1.21 | 0.98 | ||||
| Ser | UCU | 1.03 | 0.15 | 1.17 | -1.02 | 1.00 | 0.16 | 1.16 | -1.00 | 0.96 | 0.58 | 1.14 | -0.56 | 1.06 | 0.16 | 1.17 | -1.01 | |||
| UCC** | 1.25 | 2.24 | 1.06 | 1.18 | 1.24 | 2.18 | 1.07 | 1.11 | 1.32 | 1.85 | 1.08 | 0.77 | 1.19 | 2.17 | 1.03 | 1.14 | ||||
| UCA | 0.99 | 0.13 | 1.18 | -1.05 | 0.99 | 0.14 | 1.17 | -1.03 | 0.94 | 0.33 | 1.16 | -0.83 | 1.04 | 0.14 | 1.20 | -1.06 | ||||
| UCG | 0.80 | 1.65 | 0.68 | 0.97 | 0.81 | 1.64 | 0.69 | 0.95 | 0.87 | 1.21 | 0.70 | 0.51 | 0.76 | 1.66 | 0.68 | 0.98 | ||||
| AGU | 0.72 | 0.07 | 0.85 | -0.78 | 0.70 | 0.08 | 0.85 | -0.77 | 0.64 | 0.30 | 0.84 | -0.54 | 0.75 | 0.08 | 0.87 | -0.79 | ||||
| AGC** | 1.22 | 1.76 | 1.05 | 0.71 | 1.25 | 1.80 | 1.07 | 0.73 | 1.27 | 1.73 | 1.07 | 0.66 | 1.20 | 1.79 | 1.05 | 0.74 | ||||
| Pro | CCU | 0.98 | 0.20 | 1.15 | -0.95 | 0.96 | 0.21 | 1.13 | -0.92 | 0.92 | 1.20 | 1.14 | 0.06 | 1.02 | 0.21 | 1.17 | -0.96 | |||
| CCC | 0.88 | 1.38 | 0.80 | 0.58 | 0.88 | 1.36 | 0.81 | 0.55 | 0.92 | 0.95 | 0.82 | 0.13 | 0.85 | 1.36 | 0.79 | 0.57 | ||||
| CCA | 1.03 | 0.20 | 1.20 | -1.00 | 1.03 | 0.22 | 1.20 | -0.98 | 0.96 | 0.70 | 1.18 | -0.48 | 1.08 | 0.21 | 1.22 | -1.01 | ||||
| CCG | 1.11 | 2.22 | 0.85 | 1.37 | 1.12 | 2.21 | 0.85 | 1.36 | 1.20 | 1.14 | 0.86 | 0.28 | 1.05 | 2.22 | 0.82 | 1.40 | ||||
| Thr | ACU | 0.95 | 0.10 | 1.13 | -1.03 | 0.91 | 0.11 | 1.12 | -1.01 | 0.87 | 0.38 | 1.12 | -0.74 | 0.97 | 0.11 | 1.14 | -1.03 | |||
| ACC | 1.20 | 1.91 | 1.00 | 0.91 | 1.20 | 1.87 | 1.00 | 0.87 | 1.26 | 1.72 | 1.01 | 0.71 | 1.17 | 1.87 | 0.98 | 0.89 | ||||
| ACA | 1.00 | 0.11 | 1.22 | -1.11 | 1.01 | 0.12 | 1.21 | -1.09 | 0.94 | 0.33 | 1.21 | -0.88 | 1.06 | 0.12 | 1.24 | -1.12 | ||||
| ACG | 0.85 | 1.88 | 0.65 | 1.23 | 0.88 | 1.89 | 0.67 | 1.22 | 0.93 | 1.58 | 0.67 | 0.91 | 0.81 | 1.90 | 0.65 | 1.25 | ||||
| Ala | GCU | 0.92 | 0.15 | 1.16 | -1.01 | 0.90 | 0.16 | 1.14 | -0.98 | 0.85 | 1.21 | 1.14 | 0.07 | 0.95 | 0.16 | 1.17 | -1.01 | |||
| GCC** | 1.27 | 1.91 | 1.02 | 0.89 | 1.28 | 1.90 | 1.03 | 0.87 | 1.33 | 1.14 | 1.03 | 0.11 | 1.24 | 1.88 | 1.02 | 0.86 | ||||
| GCA | 0.81 | 0.14 | 1.06 | -0.92 | 0.81 | 0.15 | 1.05 | -0.90 | 0.75 | 0.66 | 1.04 | -0.38 | 0.86 | 0.14 | 1.06 | -0.92 | ||||
| GCG | 1.00 | 1.80 | 0.76 | 1.04 | 1.01 | 1.79 | 0.78 | 1.01 | 1.07 | 1.00 | 0.79 | 0.21 | 0.95 | 1.81 | 0.75 | 1.06 | ||||
| Tyr | UAU | 0.81 | 0.05 | 1.00 | -0.95 | 0.77 | 0.05 | 1.01 | -0.96 | 0.72 | 0.2 | 0.99 | -0.79 | 0.83 | 0.05 | 1.02 | -0.97 | |||
| UAC** | 1.19 | 1.95 | 1.00 | 0.95 | 1.23 | 1.95 | 0.99 | 0.96 | 1.28 | 1.80 | 1.01 | 0.79 | 1.17 | 1.95 | 0.98 | 0.97 | ||||
| His | CAU | 0.93 | 0.12 | 1.09 | -0.97 | 0.90 | 0.13 | 1.08 | -0.95 | 0.84 | 0.95 | 1.07 | -0.12 | 0.96 | 0.13 | 1.10 | -0.97 | |||
| CAC | 1.07 | 1.88 | 0.91 | 0.97 | 1.10 | 1.87 | 0.92 | 0.95 | 1.16 | 1.05 | 0.93 | 0.12 | 1.04 | 1.87 | 0.90 | 0.97 | ||||
| Gln | CAA | 0.75 | 0.10 | 0.88 | -0.78 | 0.73 | 0.11 | 0.88 | -0.77 | 0.67 | 1.19 | 0.86 | 0.33 | 0.79 | 0.11 | 0.90 | -0.79 | |||
| CAG* | 1.25 | 1.90 | 1.12 | 0.78 | 1.27 | 1.89 | 1.12 | 0.77 | 1.33 | 0.81 | 1.14 | -0.33 | 1.21 | 1.89 | 1.10 | 0.79 | ||||
| Asn | AAU | 0.92 | 0.09 | 1.06 | -0.97 | 0.90 | 0.10 | 1.06 | -0.96 | 0.86 | 0.61 | 1.04 | -0.43 | 0.95 | 0.11 | 1.06 | -0.95 | |||
| AAC | 1.08 | 1.91 | 0.94 | 0.97 | 1.10 | 1.90 | 0.94 | 0.96 | 1.14 | 1.39 | 0.96 | 0.43 | 1.05 | 1.89 | 0.94 | 0.95 | ||||
| Lys | AAA | 0.65 | 0.09 | 0.79 | -0.70 | 0.66 | 0.09 | 0.80 | -0.71 | 0.62 | 0.25 | 0.79 | -0.54 | 0.68 | 0.09 | 0.80 | -0.71 | |||
| AAG** | 1.35 | 1.91 | 1.21 | 0.70 | 1.34 | 1.91 | 1.2 | 0.71 | 1.38 | 1.75 | 1.21 | 0.54 | 1.32 | 1.91 | 1.20 | 0.71 | ||||
| Asp | GAU | 0.96 | 0.12 | 1.15 | -1.03 | 0.94 | 0.12 | 1.14 | -1.02 | 0.89 | 0.77 | 1.14 | -0.37 | 0.99 | 0.12 | 1.15 | -1.03 | |||
| GAC | 1.04 | 1.88 | 0.85 | 1.03 | 1.06 | 1.88 | 0.86 | 1.02 | 1.11 | 1.23 | 0.86 | 0.37 | 1.01 | 1.88 | 0.85 | 1.03 | ||||
| Glu | GAA | 0.73 | 0.10 | 0.87 | -0.77 | 0.73 | 0.12 | 0.88 | -0.76 | 0.68 | 0.90 | 0.87 | 0.03 | 0.77 | 0.11 | 0.88 | -0.77 | |||
| GAG* | 1.27 | 1.90 | 1.13 | 0.77 | 1.27 | 1.88 | 1.12 | 0.76 | 1.32 | 1.10 | 1.13 | -0.03 | 1.23 | 1.89 | 1.12 | 0.77 | ||||
| Cys | UGU | 0.70 | 0.06 | 0.87 | -0.81 | 0.67 | 0.06 | 0.86 | -0.80 | 0.62 | 0.42 | 0.84 | -0.42 | 0.72 | 0.06 | 0.87 | -0.81 | |||
| UGC** | 1.30 | 1.94 | 1.13 | 0.81 | 1.33 | 1.94 | 1.14 | 0.80 | 1.38 | 1.58 | 1.16 | 0.42 | 1.28 | 1.94 | 1.13 | 0.81 | ||||
| Arg | CGU | 0.63 | 0.17 | 0.77 | -0.60 | 0.62 | 0.18 | 0.77 | -0.59 | 0.59 | 1.15 | 0.76 | 0.39 | 0.62 | 0.18 | 0.77 | -0.59 | |||
| CGC** | 1.42 | 2.86 | 1.10 | 1.76 | 1.43 | 2.79 | 1.11 | 1.68 | 1.53 | 1.92 | 1.11 | 0.81 | 1.35 | 2.78 | 1.10 | 1.68 | ||||
| CGA | 0.45 | 0.11 | 0.63 | -0.52 | 0.46 | 0.12 | 0.64 | -0.52 | 0.45 | 1.28 | 0.64 | 0.64 | 0.46 | 0.12 | 0.63 | -0.51 | ||||
| CGG | 1.06 | 1.82 | 0.94 | 0.88 | 1.09 | 1.82 | 0.96 | 0.86 | 1.17 | 1.44 | 0.98 | 0.46 | 1.02 | 1.84 | 0.94 | 0.90 | ||||
| AGA | 1.00 | 0.09 | 1.19 | -1.10 | 0.96 | 0.18 | 0.77 | -0.59 | 0.87 | 0.06 | 1.15 | -1.09 | 1.07 | 0.10 | 1.20 | -1.10 | ||||
| AGG | 1.44 | 0.95 | 1.38 | -0.43 | 1.45 | 2.79 | 1.11 | 1.68 | 1.39 | 0.15 | 1.35 | -1.20 | 1.48 | 0.98 | 1.37 | -0.39 | ||||
| Gly | GGU | 0.83 | 0.17 | 1.02 | -0.85 | 0.81 | 0.17 | 1.00 | -0.83 | 0.77 | 0.67 | 0.98 | -0.31 | 0.86 | 0.16 | 1.02 | -0.86 | |||
| GGC** | 1.52 | 2.67 | 1.10 | 1.57 | 1.53 | 2.65 | 1.13 | 1.52 | 1.59 | 1.57 | 1.13 | 0.44 | 1.47 | 2.65 | 1.10 | 1.55 | ||||
| GGA | 0.79 | 0.19 | 1.01 | -0.82 | 0.79 | 0.20 | 1.02 | -0.82 | 0.75 | 1.03 | 1.01 | 0.02 | 0.82 | 0.19 | 1.02 | -0.83 | ||||
| GGG | 0.86 | 0.98 | 0.87 | 0.11 | 0.87 | 0.98 | 0.86 | 0.12 | 0.89 | 0.73 | 0.88 | -0.15 | 0.86 | 0.99 | 0.86 | 0.13 | ||||
| Trp | UGG | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | |||
| Met | AUG | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | |||
| Ter | UAA | 0.93 | 0.38 | 0.74 | -0.36 | 0.66 | 0.42 | 0.73 | -0.31 | 0.66 | 0.54 | 0.73 | -0.19 | 0.67 | 0.38 | 0.73 | -0.35 | |||
| UAG | 0.93 | 0.90 | 0.90 | 0.00 | 0.93 | 0.92 | 0.91 | 0.01 | 0.92 | 0.50 | 0.89 | -0.39 | 0.94 | 0.91 | 0.91 | 0.00 | ||||
| UGA | 1.42 | 1.00 | 1.00 | 0.00 | 1.41 | 1.00 | 1.00 | 0.00 | 1.43 | 1.96 | 1.38 | 0.58 | 1.39 | 1.70 | 1.35 | 0.35 | ||||
Table 3 Optimal codons analysis of Saccharum species and its phylogenetically related species E. fulvus
| 氨基酸 Amino acid | 密码子 Codon | LA-purple | NP-X | AP85-441 | Yunnan2009-3 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GW | HEB | LEB | ∆RSCU | GW | HEB | LEB | ∆RSCU | GW | HEB | LEB | ∆RSCU | GW | HEB | LEB | ∆RSCU | |||||
| Phe | UUU | 0.76 | 0.05 | 0.95 | -0.90 | 0.73 | 0.06 | 0.95 | -0.89 | 0.69 | 0.19 | 0.93 | -0.74 | 0.78 | 0.06 | 0.96 | -0.90 | |||
| UUC** | 1.24 | 1.95 | 1.05 | 0.90 | 1.27 | 1.94 | 1.05 | 0.89 | 1.31 | 1.81 | 1.07 | 0.74 | 1.22 | 1.94 | 1.04 | 0.90 | ||||
| Leu | UUA | 0.39 | 0.01 | 0.53 | -0.52 | 0.37 | 0.01 | 0.52 | -0.51 | 0.33 | 0.04 | 0.51 | -0.47 | 0.41 | 0.01 | 0.53 | -0.52 | |||
| UUG | 0.94 | 0.14 | 1.11 | -0.97 | 0.92 | 0.16 | 1.11 | -0.95 | 0.86 | 0.08 | 1.10 | -1.02 | 0.97 | 0.15 | 1.12 | -0.97 | ||||
| CUU | 1.04 | 0.13 | 1.27 | -1.14 | 1.03 | 0.15 | 1.24 | -1.09 | 1.00 | 1.30 | 1.26 | 0.04 | 1.08 | 0.14 | 1.26 | -1.12 | ||||
| CUC** | 1.59 | 2.99 | 1.20 | 1.79 | 1.60 | 2.95 | 1.21 | 1.74 | 1.72 | 2.09 | 1.23 | 0.86 | 1.51 | 2.94 | 1.19 | 1.75 | ||||
| CUA | 0.50 | 0.07 | 0.66 | -0.59 | 0.50 | 0.08 | 0.67 | -0.59 | 0.46 | 0.88 | 0.65 | 0.23 | 0.54 | 0.08 | 0.67 | -0.59 | ||||
| CUG** | 1.54 | 2.65 | 1.24 | 1.41 | 1.58 | 2.65 | 1.24 | 1.41 | 1.64 | 1.61 | 1.26 | 0.35 | 1.49 | 2.67 | 1.22 | 1.45 | ||||
| Ile | AUU | 1.04 | 0.10 | 1.17 | -1.07 | 1.01 | 0.11 | 1.16 | -1.05 | 0.97 | 0.31 | 1.17 | -0.86 | 1.05 | 0.12 | 1.17 | -1.05 | |||
| AUC** | 1.34 | 2.81 | 1.04 | 1.77 | 1.35 | 2.78 | 1.04 | 1.74 | 1.43 | 2.48 | 1.05 | 1.43 | 1.29 | 2.78 | 1.03 | 1.75 | ||||
| AUA | 0.63 | 0.09 | 0.78 | -0.69 | 0.64 | 0.10 | 0.79 | -0.69 | 0.60 | 0.21 | 0.78 | -0.57 | 0.66 | 0.10 | 0.80 | -0.70 | ||||
| Val | GUU | 0.95 | 0.09 | 1.23 | -1.14 | 0.94 | 0.1 | 1.21 | -1.11 | 0.90 | 0.78 | 1.22 | -0.44 | 1.00 | 0.09 | 1.23 | -1.14 | |||
| GUC | 1.15 | 1.69 | 0.97 | 0.72 | 1.17 | 1.69 | 0.99 | 0.70 | 1.22 | 1.48 | 0.99 | 0.49 | 1.12 | 1.67 | 0.97 | 0.70 | ||||
| GUA | 0.41 | 0.04 | 0.58 | -0.54 | 0.41 | 0.05 | 0.58 | -0.53 | 0.38 | 0.53 | 0.57 | -0.04 | 0.44 | 0.05 | 0.59 | -0.54 | ||||
| GUG** | 1.48 | 2.18 | 1.22 | 0.96 | 1.48 | 2.17 | 1.22 | 0.95 | 1.50 | 1.21 | 1.01 | 0.20 | 1.44 | 2.19 | 1.21 | 0.98 | ||||
| Ser | UCU | 1.03 | 0.15 | 1.17 | -1.02 | 1.00 | 0.16 | 1.16 | -1.00 | 0.96 | 0.58 | 1.14 | -0.56 | 1.06 | 0.16 | 1.17 | -1.01 | |||
| UCC** | 1.25 | 2.24 | 1.06 | 1.18 | 1.24 | 2.18 | 1.07 | 1.11 | 1.32 | 1.85 | 1.08 | 0.77 | 1.19 | 2.17 | 1.03 | 1.14 | ||||
| UCA | 0.99 | 0.13 | 1.18 | -1.05 | 0.99 | 0.14 | 1.17 | -1.03 | 0.94 | 0.33 | 1.16 | -0.83 | 1.04 | 0.14 | 1.20 | -1.06 | ||||
| UCG | 0.80 | 1.65 | 0.68 | 0.97 | 0.81 | 1.64 | 0.69 | 0.95 | 0.87 | 1.21 | 0.70 | 0.51 | 0.76 | 1.66 | 0.68 | 0.98 | ||||
| AGU | 0.72 | 0.07 | 0.85 | -0.78 | 0.70 | 0.08 | 0.85 | -0.77 | 0.64 | 0.30 | 0.84 | -0.54 | 0.75 | 0.08 | 0.87 | -0.79 | ||||
| AGC** | 1.22 | 1.76 | 1.05 | 0.71 | 1.25 | 1.80 | 1.07 | 0.73 | 1.27 | 1.73 | 1.07 | 0.66 | 1.20 | 1.79 | 1.05 | 0.74 | ||||
| Pro | CCU | 0.98 | 0.20 | 1.15 | -0.95 | 0.96 | 0.21 | 1.13 | -0.92 | 0.92 | 1.20 | 1.14 | 0.06 | 1.02 | 0.21 | 1.17 | -0.96 | |||
| CCC | 0.88 | 1.38 | 0.80 | 0.58 | 0.88 | 1.36 | 0.81 | 0.55 | 0.92 | 0.95 | 0.82 | 0.13 | 0.85 | 1.36 | 0.79 | 0.57 | ||||
| CCA | 1.03 | 0.20 | 1.20 | -1.00 | 1.03 | 0.22 | 1.20 | -0.98 | 0.96 | 0.70 | 1.18 | -0.48 | 1.08 | 0.21 | 1.22 | -1.01 | ||||
| CCG | 1.11 | 2.22 | 0.85 | 1.37 | 1.12 | 2.21 | 0.85 | 1.36 | 1.20 | 1.14 | 0.86 | 0.28 | 1.05 | 2.22 | 0.82 | 1.40 | ||||
| Thr | ACU | 0.95 | 0.10 | 1.13 | -1.03 | 0.91 | 0.11 | 1.12 | -1.01 | 0.87 | 0.38 | 1.12 | -0.74 | 0.97 | 0.11 | 1.14 | -1.03 | |||
| ACC | 1.20 | 1.91 | 1.00 | 0.91 | 1.20 | 1.87 | 1.00 | 0.87 | 1.26 | 1.72 | 1.01 | 0.71 | 1.17 | 1.87 | 0.98 | 0.89 | ||||
| ACA | 1.00 | 0.11 | 1.22 | -1.11 | 1.01 | 0.12 | 1.21 | -1.09 | 0.94 | 0.33 | 1.21 | -0.88 | 1.06 | 0.12 | 1.24 | -1.12 | ||||
| ACG | 0.85 | 1.88 | 0.65 | 1.23 | 0.88 | 1.89 | 0.67 | 1.22 | 0.93 | 1.58 | 0.67 | 0.91 | 0.81 | 1.90 | 0.65 | 1.25 | ||||
| Ala | GCU | 0.92 | 0.15 | 1.16 | -1.01 | 0.90 | 0.16 | 1.14 | -0.98 | 0.85 | 1.21 | 1.14 | 0.07 | 0.95 | 0.16 | 1.17 | -1.01 | |||
| GCC** | 1.27 | 1.91 | 1.02 | 0.89 | 1.28 | 1.90 | 1.03 | 0.87 | 1.33 | 1.14 | 1.03 | 0.11 | 1.24 | 1.88 | 1.02 | 0.86 | ||||
| GCA | 0.81 | 0.14 | 1.06 | -0.92 | 0.81 | 0.15 | 1.05 | -0.90 | 0.75 | 0.66 | 1.04 | -0.38 | 0.86 | 0.14 | 1.06 | -0.92 | ||||
| GCG | 1.00 | 1.80 | 0.76 | 1.04 | 1.01 | 1.79 | 0.78 | 1.01 | 1.07 | 1.00 | 0.79 | 0.21 | 0.95 | 1.81 | 0.75 | 1.06 | ||||
| Tyr | UAU | 0.81 | 0.05 | 1.00 | -0.95 | 0.77 | 0.05 | 1.01 | -0.96 | 0.72 | 0.2 | 0.99 | -0.79 | 0.83 | 0.05 | 1.02 | -0.97 | |||
| UAC** | 1.19 | 1.95 | 1.00 | 0.95 | 1.23 | 1.95 | 0.99 | 0.96 | 1.28 | 1.80 | 1.01 | 0.79 | 1.17 | 1.95 | 0.98 | 0.97 | ||||
| His | CAU | 0.93 | 0.12 | 1.09 | -0.97 | 0.90 | 0.13 | 1.08 | -0.95 | 0.84 | 0.95 | 1.07 | -0.12 | 0.96 | 0.13 | 1.10 | -0.97 | |||
| CAC | 1.07 | 1.88 | 0.91 | 0.97 | 1.10 | 1.87 | 0.92 | 0.95 | 1.16 | 1.05 | 0.93 | 0.12 | 1.04 | 1.87 | 0.90 | 0.97 | ||||
| Gln | CAA | 0.75 | 0.10 | 0.88 | -0.78 | 0.73 | 0.11 | 0.88 | -0.77 | 0.67 | 1.19 | 0.86 | 0.33 | 0.79 | 0.11 | 0.90 | -0.79 | |||
| CAG* | 1.25 | 1.90 | 1.12 | 0.78 | 1.27 | 1.89 | 1.12 | 0.77 | 1.33 | 0.81 | 1.14 | -0.33 | 1.21 | 1.89 | 1.10 | 0.79 | ||||
| Asn | AAU | 0.92 | 0.09 | 1.06 | -0.97 | 0.90 | 0.10 | 1.06 | -0.96 | 0.86 | 0.61 | 1.04 | -0.43 | 0.95 | 0.11 | 1.06 | -0.95 | |||
| AAC | 1.08 | 1.91 | 0.94 | 0.97 | 1.10 | 1.90 | 0.94 | 0.96 | 1.14 | 1.39 | 0.96 | 0.43 | 1.05 | 1.89 | 0.94 | 0.95 | ||||
| Lys | AAA | 0.65 | 0.09 | 0.79 | -0.70 | 0.66 | 0.09 | 0.80 | -0.71 | 0.62 | 0.25 | 0.79 | -0.54 | 0.68 | 0.09 | 0.80 | -0.71 | |||
| AAG** | 1.35 | 1.91 | 1.21 | 0.70 | 1.34 | 1.91 | 1.2 | 0.71 | 1.38 | 1.75 | 1.21 | 0.54 | 1.32 | 1.91 | 1.20 | 0.71 | ||||
| Asp | GAU | 0.96 | 0.12 | 1.15 | -1.03 | 0.94 | 0.12 | 1.14 | -1.02 | 0.89 | 0.77 | 1.14 | -0.37 | 0.99 | 0.12 | 1.15 | -1.03 | |||
| GAC | 1.04 | 1.88 | 0.85 | 1.03 | 1.06 | 1.88 | 0.86 | 1.02 | 1.11 | 1.23 | 0.86 | 0.37 | 1.01 | 1.88 | 0.85 | 1.03 | ||||
| Glu | GAA | 0.73 | 0.10 | 0.87 | -0.77 | 0.73 | 0.12 | 0.88 | -0.76 | 0.68 | 0.90 | 0.87 | 0.03 | 0.77 | 0.11 | 0.88 | -0.77 | |||
| GAG* | 1.27 | 1.90 | 1.13 | 0.77 | 1.27 | 1.88 | 1.12 | 0.76 | 1.32 | 1.10 | 1.13 | -0.03 | 1.23 | 1.89 | 1.12 | 0.77 | ||||
| Cys | UGU | 0.70 | 0.06 | 0.87 | -0.81 | 0.67 | 0.06 | 0.86 | -0.80 | 0.62 | 0.42 | 0.84 | -0.42 | 0.72 | 0.06 | 0.87 | -0.81 | |||
| UGC** | 1.30 | 1.94 | 1.13 | 0.81 | 1.33 | 1.94 | 1.14 | 0.80 | 1.38 | 1.58 | 1.16 | 0.42 | 1.28 | 1.94 | 1.13 | 0.81 | ||||
| Arg | CGU | 0.63 | 0.17 | 0.77 | -0.60 | 0.62 | 0.18 | 0.77 | -0.59 | 0.59 | 1.15 | 0.76 | 0.39 | 0.62 | 0.18 | 0.77 | -0.59 | |||
| CGC** | 1.42 | 2.86 | 1.10 | 1.76 | 1.43 | 2.79 | 1.11 | 1.68 | 1.53 | 1.92 | 1.11 | 0.81 | 1.35 | 2.78 | 1.10 | 1.68 | ||||
| CGA | 0.45 | 0.11 | 0.63 | -0.52 | 0.46 | 0.12 | 0.64 | -0.52 | 0.45 | 1.28 | 0.64 | 0.64 | 0.46 | 0.12 | 0.63 | -0.51 | ||||
| CGG | 1.06 | 1.82 | 0.94 | 0.88 | 1.09 | 1.82 | 0.96 | 0.86 | 1.17 | 1.44 | 0.98 | 0.46 | 1.02 | 1.84 | 0.94 | 0.90 | ||||
| AGA | 1.00 | 0.09 | 1.19 | -1.10 | 0.96 | 0.18 | 0.77 | -0.59 | 0.87 | 0.06 | 1.15 | -1.09 | 1.07 | 0.10 | 1.20 | -1.10 | ||||
| AGG | 1.44 | 0.95 | 1.38 | -0.43 | 1.45 | 2.79 | 1.11 | 1.68 | 1.39 | 0.15 | 1.35 | -1.20 | 1.48 | 0.98 | 1.37 | -0.39 | ||||
| Gly | GGU | 0.83 | 0.17 | 1.02 | -0.85 | 0.81 | 0.17 | 1.00 | -0.83 | 0.77 | 0.67 | 0.98 | -0.31 | 0.86 | 0.16 | 1.02 | -0.86 | |||
| GGC** | 1.52 | 2.67 | 1.10 | 1.57 | 1.53 | 2.65 | 1.13 | 1.52 | 1.59 | 1.57 | 1.13 | 0.44 | 1.47 | 2.65 | 1.10 | 1.55 | ||||
| GGA | 0.79 | 0.19 | 1.01 | -0.82 | 0.79 | 0.20 | 1.02 | -0.82 | 0.75 | 1.03 | 1.01 | 0.02 | 0.82 | 0.19 | 1.02 | -0.83 | ||||
| GGG | 0.86 | 0.98 | 0.87 | 0.11 | 0.87 | 0.98 | 0.86 | 0.12 | 0.89 | 0.73 | 0.88 | -0.15 | 0.86 | 0.99 | 0.86 | 0.13 | ||||
| Trp | UGG | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | |||
| Met | AUG | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 1.00 | 0.00 | |||
| Ter | UAA | 0.93 | 0.38 | 0.74 | -0.36 | 0.66 | 0.42 | 0.73 | -0.31 | 0.66 | 0.54 | 0.73 | -0.19 | 0.67 | 0.38 | 0.73 | -0.35 | |||
| UAG | 0.93 | 0.90 | 0.90 | 0.00 | 0.93 | 0.92 | 0.91 | 0.01 | 0.92 | 0.50 | 0.89 | -0.39 | 0.94 | 0.91 | 0.91 | 0.00 | ||||
| UGA | 1.42 | 1.00 | 1.00 | 0.00 | 1.41 | 1.00 | 1.00 | 0.00 | 1.43 | 1.96 | 1.38 | 0.58 | 1.39 | 1.70 | 1.35 | 0.35 | ||||
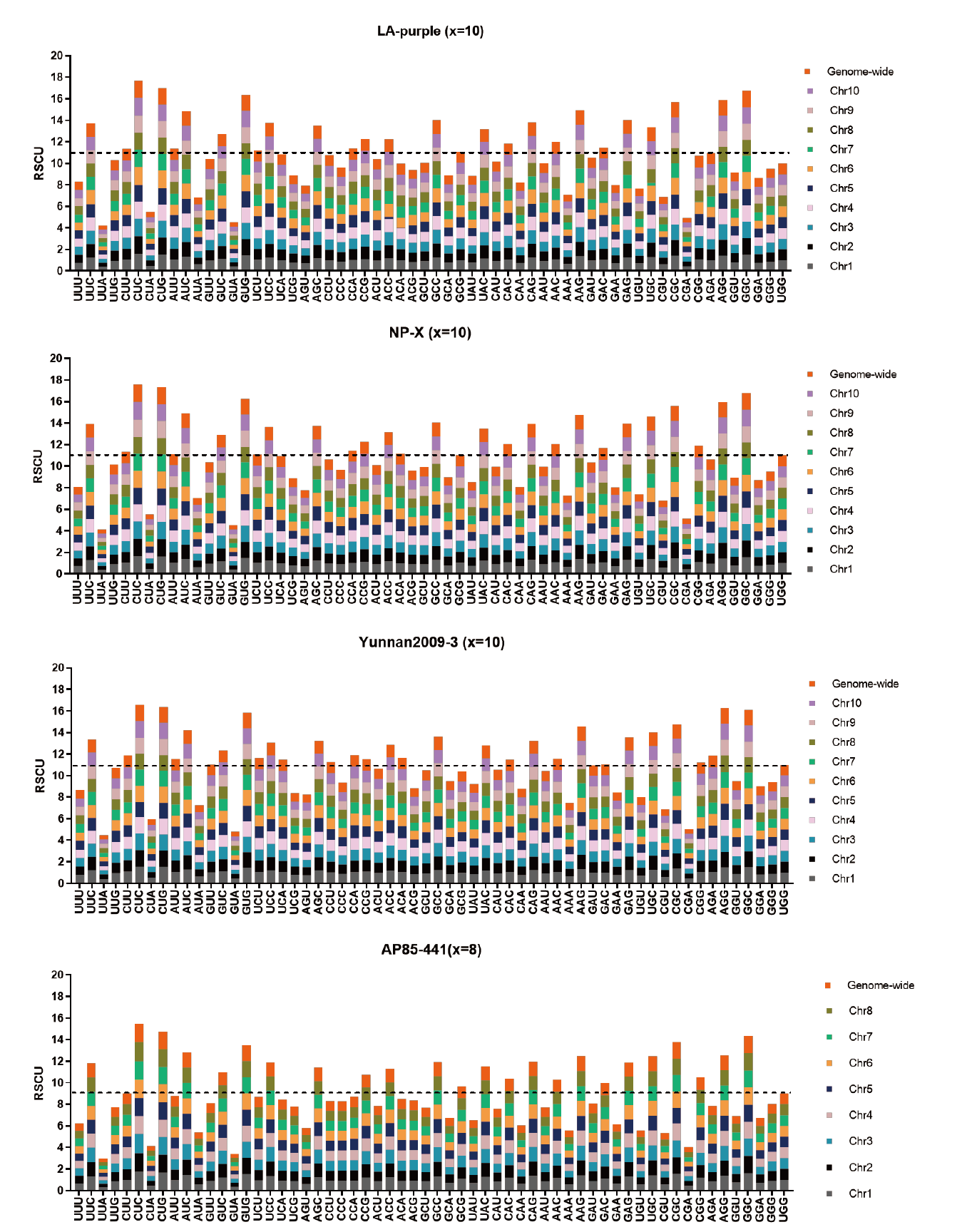
Fig. 5 Analysis of codon RSCU on genome-wide and chromosome set level in four materials The dotted lines in the figure refers to the RSCU value when RSCU is 1 on both genome-wide level and every chromosome sets. Codons with RSCU above the dotted line in the figures can be regarded as preferred synonymous codons for this amino acid

Fig. 7 Comparison analysis of codon usage bias between sugarcane and other seven organisms based on RSCU value Ss: Saccharum spp.(including Saccharum species and E. fulvus); Zm: Z. mays; Sb: S. bicolor; Os: O. ativa; At: A. thaliana; Nt: N. tabacum; Ec: E. coli; Sc: S. cerevisiae
| [1] |
Ikemura T. Codon usage and tRNA content in unicellular and multicellular organisms[J]. Mol Biol Evol, 1985, 2(1): 13-34.
doi: 10.1093/oxfordjournals.molbev.a040335 pmid: 3916708 |
| [2] |
Bulmer M. The selection-mutation-drift theory of synonymous codon usage[J]. Genetics, 1991, 129(3): 897-907.
doi: 10.1093/genetics/129.3.897 pmid: 1752426 |
| [3] |
Hershberg R, Petrov DA. Selection on codon bias[J]. Annu Rev Genet, 2008, 42: 287-299.
doi: 10.1146/annurev.genet.42.110807.091442 pmid: 18983258 |
| [4] | 张乐, 金龙国, 罗玲, 等. 大豆基因组和转录组的核基因密码子使用偏好性分析[J]. 作物学报, 2011, 37(6): 965-974. |
| Zhang L, Jin LG, Luo L, et al. Analysis of nuclear gene codon bias on soybean genome and transcriptome[J]. Acta Agron Sin, 2011, 37(6): 965-974. | |
| [5] | 李亚麒, 黄家雄, 娄予强, 等. 小粒咖啡铁皮卡叶绿体基因组密码子偏好性分析[J]. 西北林学院学报, 2023, 38(2): 92-99. |
| Li YQ, Huang JX, Lou YQ, et al. Codon usage bias of the chloroplast genome in Coffea arabica‘Typica’[J]. J Northwest For Univ, 2023, 38(2): 92-99. | |
| [6] |
Tao P, Dai L, Luo MC, et al. Analysis of synonymous codon usage in classical swine fever virus[J]. Virus Genes, 2009, 38(1): 104-112.
doi: 10.1007/s11262-008-0296-z pmid: 18958611 |
| [7] |
Das S, Paul S, Dutta C. Synonymous codon usage in adenoviruses: influence of mutation, selection and protein hydropathy[J]. Virus Res, 2006, 117(2): 227-236.
pmid: 16307819 |
| [8] |
Yadav MK, Swati D. Comparative genome analysis of six malarial parasites using codon usage bias based tools[J]. Bioinformation, 2012, 8(24): 1230-1239.
doi: 10.6026/97320630081230 pmid: 23275725 |
| [9] | 赖瑞联, 冯新, 陈瑾, 等. 橄榄查尔酮异构酶基因CHI的密码子偏好模式[J]. 应用与环境生物学报, 2017, 23(5): 945-951. |
| Lai RL, Feng X, Chen J, et al. Codon usage pattern of chalcone isomerase gene(CHI)in Canarium album(Lour.) Raeusch[J]. Chin J Appl Environ Biol, 2017, 23(5): 945-951. | |
| [10] |
寇莹莹, 宋英今, 杨少辉, 等. 植酸酶phyA基因的密码子优化及其在大豆中的表达[J]. 作物学报, 2016, 42(12): 1798-1804.
doi: 10.3724/SP.J.1006.2016.01798 |
|
Kou YY, Song YJ, Yang SH, et al. Codon optimization and expression of phyA gene in soybean(Glycine max merr.)[J]. Acta Agron Sin, 2016, 42(12): 1798-1804.
doi: 10.3724/SP.J.1006.2016.01798 URL |
|
| [11] | 周宗梁, 林智敏, 耿丽丽, 等. 水稻中cry1Ah1基因密码子优化方案的比较[J]. 生物工程学报, 2012, 28(10): 1184-1194. |
| Zhou ZL, Lin ZM, Geng LL, et al. Comparison of codon optimizations of cry1Ah1 gene in rice[J]. Chin J Biotechnol, 2012, 28(10): 1184-1194. | |
| [12] | 陈惠, 赵海霞, 王红宁, 等. 植酸酶基因中稀有密码子的改造提高其在毕赤酵母中的表达量[J]. 中国生物化学与分子生物学报, 2005, 21(2): 170-174. |
| Chen H, Zhao HX, Wang HN, et al. Increasing expression level of phytase gene(phyA)in Pichia pastoris by changing rare codons[J]. Chin J Biochem Mol Biol, 2005, 21(2): 170-174. | |
| [13] |
Menzella HG. Comparison of two codon optimization strategies to enhance recombinant protein production in Escherichia coli[J]. Microb Cell Fact, 2011, 10: 15.
doi: 10.1186/1475-2859-10-15 pmid: 21371320 |
| [14] |
Jabeen R, Khan MS, Zafar Y, et al. Codon optimization of cry1Ab gene for hyper expression in plant organelles[J]. Mol Biol Rep, 2010, 37(2): 1011-1017.
doi: 10.1007/s11033-009-9802-1 pmid: 19757171 |
| [15] |
Hirai H, Kashima Y, Hayashi K, et al. Efficient expression of laccase gene from white-rot fungus Schizophyllum commune in a transgenic tobacco plant[J]. FEMS Microbiol Lett, 2008, 286(1): 130-135.
doi: 10.1111/fml.2008.286.issue-1 URL |
| [16] |
D'Hont A. Unraveling the genome structure of polyploids using FISH and GISH; examples of sugarcane and banana[J]. Cytogenet Genome Res, 2005, 109(1-3): 27-33.
doi: 10.1159/000082378 pmid: 15753555 |
| [17] |
Zhang JS, Zhang XT, Tang HB, et al. Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L[J]. Nat Genet, 2018, 50(11): 1565-1573.
doi: 10.1038/s41588-018-0237-2 |
| [18] |
Zhang Q, Qi YY, Pan HR, et al. Genomic insights into the recent chromosome reduction of autopolyploid sugarcane Saccharum spontaneum[J]. Nat Genet, 2022, 54(6): 885-896.
doi: 10.1038/s41588-022-01084-1 pmid: 35654976 |
| [19] |
Wang TY, Wang BY, Hua XT, et al. A complete gap-free diploid genome in Saccharum complex and the genomic footprints of evolution in the highly polyploid Saccharum genus[J]. Nat Plants, 2023, 9(4): 554-571.
doi: 10.1038/s41477-023-01378-0 |
| [20] |
Zhang WJ, Zhou J, Li ZF, et al. Comparative analysis of codon usage patterns among mitochondrion, chloroplast and nuclear genes in Triticum aestivum L[J]. J Integr Plant Biol, 2007, 49(2): 246-254.
doi: 10.1111/jipb.2007.49.issue-2 URL |
| [21] | 冯展, 江媛, 郑燕, 等. 肉苁蓉属植物叶绿体基因组密码子偏好性分析[J]. 中草药, 2023, 54(5): 1540-1550. |
| Feng Z, Jiang Y, Zheng Y, et al. Codon use bias analysis of chloroplast genome of Cistanche[J]. Chin Tradit Herb Drugs, 2023, 54(5): 1540-1550. | |
| [22] |
Wright F. The ‘effective number of codons’ used in a gene[J]. Gene, 1990, 87(1): 23-29.
doi: 10.1016/0378-1119(90)90491-9 pmid: 2110097 |
| [23] |
Jiang Y, Deng F, Wang HL, et al. An extensive analysis on the global codon usage pattern of baculoviruses[J]. Arch Virol, 2008, 153(12): 2273-2282.
doi: 10.1007/s00705-008-0260-1 pmid: 19030954 |
| [24] |
张以忠, 曾文艺, 邓琳琼, 等. 甘蓝S-位点基因SRK、SLG和SP11/SCR密码子偏好性分析[J]. 作物学报, 2022, 48(5): 1152-1168.
doi: 10.3724/SP.J.1006.2022.14003 |
|
Zhang YZ, Zeng WY, Deng LQ, et al. Codon usage bias analysis of S-locus genes SRK, SLG, and SP11/SCR in Brassica oleracea[J]. Acta Agron Sin, 2022, 48(5): 1152-1168.
doi: 10.3724/SP.J.1006.2022.14003 URL |
|
| [25] | 金刚, 覃旭, 龙凌云, 等. 剑麻叶绿体基因组编码序列密码子的使用特征[J]. 福建农林大学学报: 自然科学版, 2018, 47(6): 705-710. |
| Jin G, Qin X, Long LY, et al. Characteristics of codon usage in the chloroplast protein-coding genes of Agave hybrid No.11648[J]. J Fujian Agric For Univ Nat Sci Ed, 2018, 47(6): 705-710. | |
| [26] | 雷佳欣, 张丽娟, 高丹丹, 等. 文冠果基因组密码子偏好性分析[J]. 西北林学院学报, 2023, 38(4): 104-110. |
| Lei JX, Zhang LJ, Gao DD, et al. Codon usage bias of Xanthoceras sorbifolium genome[J]. J Northwest For Univ, 2023, 38(4): 104-110. | |
| [27] |
赖瑞联, 林玉玲, 钟春水, 等. 龙眼生长素受体基因TIR1密码子偏好性分析[J]. 园艺学报, 2016, 43(4): 771-780.
doi: 10.16420/j.issn.0513-353x.2015-0662 |
| Lai RL, Lin YL, Zhong CS, et al. Analysis of codon bias of auxin receptor gene TIR1 in Dimocarpus longan[J]. Acta Hortic Sin, 2016, 43(4): 771-780. | |
| [28] |
Liu QP, Feng Y, Zhao XA, et al. Synonymous codon usage bias in Oryza sativa[J]. Plant Sci, 2004, 167(1): 101-105.
doi: 10.1016/j.plantsci.2004.03.003 URL |
| [29] | 刘汉梅, 何瑞, 张怀渝, 等. 玉米同义密码子偏爱性分析[J]. 农业生物技术学报, 2010, 18(3): 456-461. |
| Liu HM, He R, Zhang HY, et al. Analysis of synonymous codon bias in maize[J]. J Agric Biotechnol, 2010, 18(3): 456-461. | |
| [30] | 张晓峰, 薛庆中. 水稻和拟南芥NBS-LRR基因家族同义密码子使用偏好的比较[J]. 作物学报, 2005, 31(5): 596-602. |
| Zhang XF, Xue QZ. Synonymous codon bias of NBS-LRR gene family in rice and Arabidopsis[J]. Acta Agron Sin, 2005, 31(5): 596-602. | |
| [31] |
Kawabe A, Miyashita NT. Patterns of codon usage bias in three dicot and four monocot plant species[J]. Genes Genet Syst, 2003, 78(5): 343-352.
doi: 10.1266/ggs.78.343 pmid: 14676425 |
| [32] |
Comeron JM, Aguadé M. An evaluation of measures of synonymous codon usage bias[J]. J Mol Evol, 1998, 47(3): 268-274.
doi: 10.1007/pl00006384 pmid: 9732453 |
| [33] |
唐玉娟, 赵英, 黄国弟, 等. 芒果叶绿体基因组密码子使用偏好性分析[J]. 热带作物学报, 2021, 42(8): 2143-2150.
doi: 10.3969/j.issn.1000-2561.2021.08.004 |
| Tang YJ, Zhao Y, Huang GD, et al. Analysis on codon usage bias of chloroplast genes from mango[J]. Chin J Trop Crops, 2021, 42(8): 2143-2150. | |
| [34] |
杨祥燕, 蔡元保, 谭秦亮, 等. 菠萝叶绿体基因组密码子偏好性分析[J]. 热带作物学报, 2022, 43(3): 439-446.
doi: 10.3969/j.issn.1000-2561.2022.03.001 |
| Yang XY, Cai YB, Tan QL, et al. Analysis of codon usage bias in the chloroplast genome of Ananas comosus[J]. Chin J Trop Crops, 2022, 43(3): 439-446. | |
| [35] |
Wen Y, Zou ZL, Li HS, et al. Analysis of codon usage patterns in Morus notabilis based on genome and transcriptome data[J]. Genome, 2017, 60(6): 473-484.
doi: 10.1139/gen-2016-0129 URL |
| [36] | 王占军, 丁亮, 蔡倩文, 等. 3种木薯全基因组的密码子偏好性模式与变异来源比较[J]. 应用与环境生物学报, 2021, 27(4): 1013-1021. |
| Wang ZJ, Ding L, Cai QW, et al. Comparison of codon preference patterns and variation sources in Manihot esculenta Crantz genomes[J]. Chin J Appl Environ Biol, 2021, 27(4): 1013-1021. | |
| [37] | 毛立彦, 黄秋伟, 龙凌云, 等. 7种睡莲属植物叶绿体基因组密码子偏好性分析[J]. 西北林学院学报, 2022, 37(2): 98-107. |
| Mao LY, Huang QW, Long LY, et al. Comparative analysis of codon usage bias in chloroplast genomes of seven Nymphaea species[J]. J Northwest For Univ, 2022, 37(2): 98-107. | |
| [38] | 冯瑞云, 梅超, 王慧杰, 等. 籽粒苋叶绿体基因组密码子偏好性分析[J]. 中国草地学报, 2019, 41(4): 8-15. |
| Feng RY, Mei C, Wang HJ, et al. Analysis of codon usage in the chloroplast genome of grain amaranth(Amaranthus hypochondriacus L.)[J]. Chin J Grassland, 2019, 41(4): 8-15. | |
| [39] | 原晓龙, 李云琴, 张劲峰, 等. 乳油木叶绿体基因组密码子偏好性分析[J]. 分子植物育种, 2020, 18(17): 5658-5664. |
| Yuan XL, Li YQ, Zhang JF, et al. Codon usage bias analysis of chloroplast genome in Vitellaria paradoxa[J]. Mol Plant Breed, 2020, 18(17): 5658-5664. | |
| [40] |
Tang DF, Wei F, Cai ZQ, et al. Analysis of codon usage bias and evolution in the chloroplast genome of Mesona chinensis Benth[J]. Dev Genes Evol, 2021, 231(1-2): 1-9.
doi: 10.1007/s00427-020-00670-9 |
| [1] | YANG Wei-cheng, SUN Yan, YANG Qian, WANG Zhuang-lin, MA Ju-hua, XUE Jin-ai, LI Run-zhi. Genome-wide Identification of the FAX family in Gossypium hirsutum and Functional Analysis of GhFAX1 [J]. Biotechnology Bulletin, 2024, 40(3): 155-169. |
| [2] | GONG Li-li, YU Hua, YANG Jie, CHEN Tian-chi, ZHAO Shuang-ying, WU Yue-yan. Identification and Analysis of Grape(Vitis vinifera L.)CYP707A Gene Family and Functional Verification to Fruit Ripening [J]. Biotechnology Bulletin, 2024, 40(2): 160-171. |
| [3] | LU Yu-dan, LIU Xiao-chi, FENG Xin, CHEN Gui-xin, CHEN Yi-ting. Identification of the Kiwifruit BBX Gene Family and Analysis of Their Transcriptional Characteristics [J]. Biotechnology Bulletin, 2024, 40(2): 172-182. |
| [4] | YANG Yu-qing, TAN Juan, WANG Fang, PENG Shun-li, CHEN Jie, TAN Ming-yan, LYU Mei-yan, ZHOU Fu-yu, LIU Sheng-chuan. Research and Application Progress in Chloroplast Genome of Tea Plant(Camellia sinensis) [J]. Biotechnology Bulletin, 2024, 40(2): 20-30. |
| [5] | WANG Teng-hui, GE Wen-dong, LUO Ya-fang, FAN Zhen-yu, WANG Yu-shu. Gene Mapping of Kale White Leaves Based on Whole Genome Re-sequencing of Extreme Mixed Pool(BSA) [J]. Biotechnology Bulletin, 2023, 39(9): 176-182. |
| [6] | LI Xue-qi, ZHANG Su-jie, YU Man, HUANG Jin-guang, ZHOU Huan-bin. Establishment of CRISPR/CasX-based Genome Editing Technology in Rice [J]. Biotechnology Bulletin, 2023, 39(9): 40-48. |
| [7] | FANG Lan, LI Yan-yan, JIANG Jian-wei, CHENG Sheng, SUN Zheng-xiang, ZHOU Yi. Isolation, Identification and Growth-promoting Characteristics of Endohyphal Bacterium 7-2H from Endophytic Fungi of Spiranthes sinensis [J]. Biotechnology Bulletin, 2023, 39(8): 272-282. |
| [8] | RAO Zi-huan, XIE Zhi-xiong. Isolation and Identification of a Cellulose-degrading Strain of Olivibacter jilunii and Analysis of Its Degradability [J]. Biotechnology Bulletin, 2023, 39(8): 283-290. |
| [9] | GUO Shao-hua, MAO Hui-li, LIU Zheng-quan, FU Mei-yuan, ZHAO Ping-yuan, MA Wen-bo, LI Xu-dong, GUAN Jian-yi. Whole Genome Sequencing and Comparative Genome Analysis of a Fish-derived Pathogenic Aeromonas Hydrophila Strain XDMG [J]. Biotechnology Bulletin, 2023, 39(8): 291-306. |
| [10] | ZHANG Dao-lei, GAN Yu-jun, LE Liang, PU Li. Epigenetic Regulation of Yield-related Traits in Maize and Epibreeding [J]. Biotechnology Bulletin, 2023, 39(8): 31-42. |
| [11] | SHI Jia-xin, LIU Kai, ZHU Jin-jie, QI Xian-tao, XIE Chuan-xiao, LIU Chang-lin. Gene Editing Reshaping Maize Plant Type for Increasing Hybrid Yield [J]. Biotechnology Bulletin, 2023, 39(8): 62-69. |
| [12] | DU Dong-dong, QIAN Jing, LI Si-qi, LIU Wen-fei, WEI Xiang-li, LIU Chang-yong, LUO Rui-feng, KANG Li-chao. Whole Genome Sequencing and Analysis of Listeria monocytogenes Strain LMXJ15 [J]. Biotechnology Bulletin, 2023, 39(7): 298-306. |
| [13] | LI Yu-zhen, MEI Tian-xiu, LI Zhi-wen, WANG Qi, LI Jun, ZOU Yue, ZHAO Xin-qing. Advances in Genomic Studies and Metabolic Engineering of Red Yeasts [J]. Biotechnology Bulletin, 2023, 39(7): 67-79. |
| [14] | YIN Ming-hua, YU Huan-yuan, XIAO Xin-yi, WANG Yu-ting. Chloroplast Genomic Characterization and Phylogenetic Analysis of Colocasia esculenta L. Schoot var. cormosus cv. ‘Hongyayu’ from Jiangxi Yanshan [J]. Biotechnology Bulletin, 2023, 39(6): 233-247. |
| [15] | ZHANG Lu-yang, HAN Wen-long, XU Xiao-wen, YAO Jian, LI Fang-fang, TIAN Xiao-yuan, ZHANG Zhi-qiang. Identification and Expression Analysis of the Tobacco TCP Gene Family [J]. Biotechnology Bulletin, 2023, 39(6): 248-258. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||