








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 121-135.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0826
Previous Articles Next Articles
JIANG Yi1( ), GAO Ming-yan1, ZHANG Di1, FU Li-xia2, CHENG Xu1, CHEN Xiang3, GONG Jian-sen1, ZHANG Ying1, YANG Qin1, YU Yan1(
), GAO Ming-yan1, ZHANG Di1, FU Li-xia2, CHENG Xu1, CHEN Xiang3, GONG Jian-sen1, ZHANG Ying1, YANG Qin1, YU Yan1( )
)
Received:2025-07-31
Online:2026-02-26
Published:2026-03-17
Contact:
YU Yan
E-mail:yijiang620@126.com;yuyan_1020@126.com
JIANG Yi, GAO Ming-yan, ZHANG Di, FU Li-xia, CHENG Xu, CHEN Xiang, GONG Jian-sen, ZHANG Ying, YANG Qin, YU Yan. Host Proteases and Coronavirus Invasion: From S Protein Cleavage Mechanism to Development and Applications of Protease Inhibitor[J]. Biotechnology Bulletin, 2026, 42(2): 121-135.
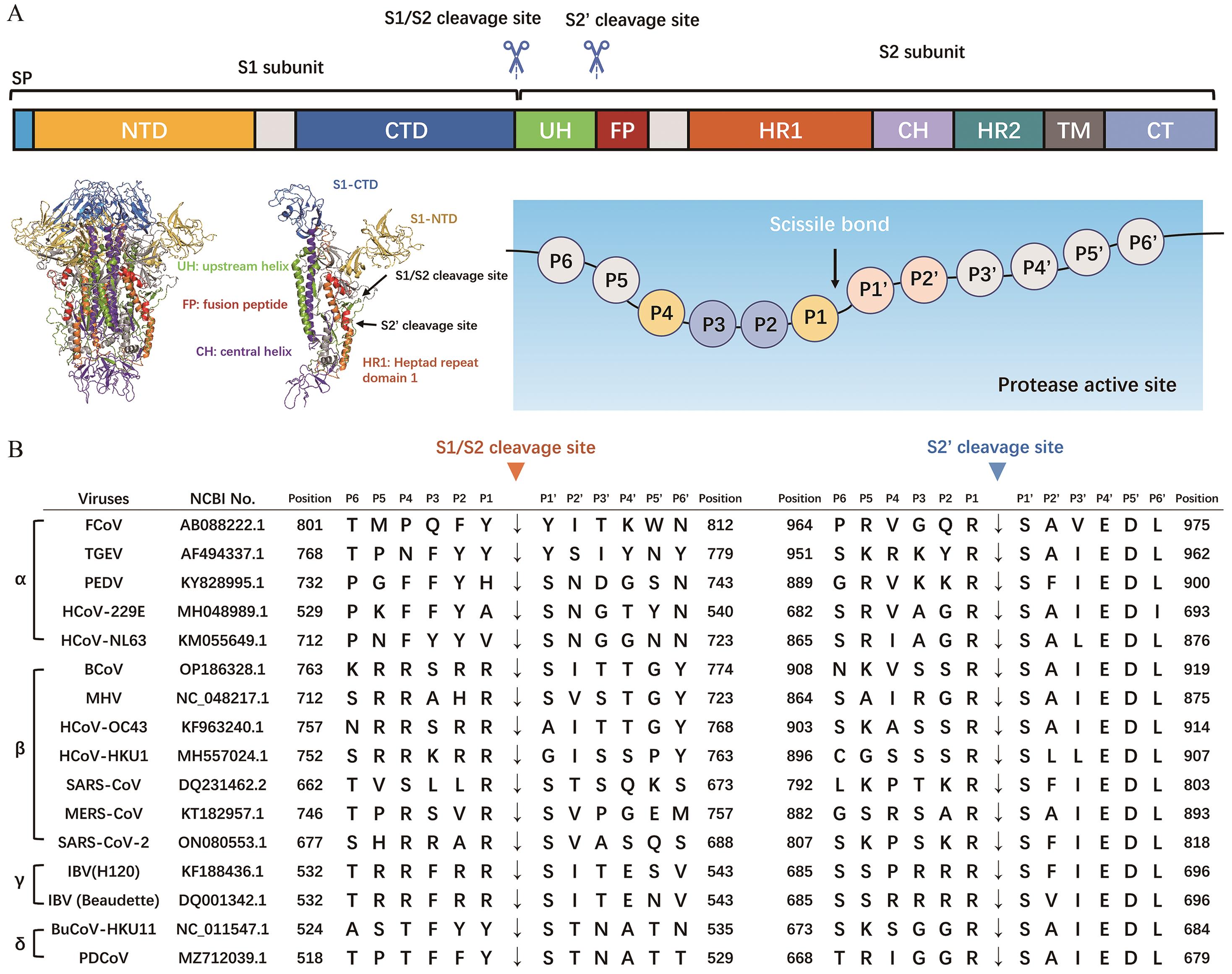
Fig. 2 Cleavage site motifs of proteases among different coronavirusesA: Schematic of protease activation site. B: S1/S2 or S2′ cleavage site motifs of different coronaviruses
蛋白酶 Protease | 细胞定位 Cellular localization | 作用途径 Pathway | 识别基序 Recognized motif P2-P1↓P1′-P2′ | 是否依赖酸性环境 Acid dependence | 参考文献 References |
|---|---|---|---|---|---|
| Furin | TGN | 生物合成 | RXK/RR↓S | 不依赖 | [ |
| TMPRSS2 | 跨膜 | 质膜途径 | K/R↓S | 不依赖 | [ |
| CTSL/B | 内质体/溶酶体 | 内体途径 | R/K/G/A/S/T/Q/Y↓ | 依赖,pH 3.0-6.5 | [ |
| HAT | 跨膜 | 质膜途径 | K/R↓ | 不依赖 | [ |
| 胰蛋白酶 | 细胞外 | 质膜途径 | 选择性不高,主要K/R↓S | 中性pH值,最适pH 8.0 | [ |
| 弹性蛋白酶 | 细胞外 | 质膜途径 | A/S/G/V/W/Y/F/L/C/M↓ | 不依赖 | [ |
| 纤溶酶 | 细胞外 | 质膜途径 | K/R↓ | 不依赖 | [ |
Table 1 Distribution and cleavage properties of proteases in different hosts
蛋白酶 Protease | 细胞定位 Cellular localization | 作用途径 Pathway | 识别基序 Recognized motif P2-P1↓P1′-P2′ | 是否依赖酸性环境 Acid dependence | 参考文献 References |
|---|---|---|---|---|---|
| Furin | TGN | 生物合成 | RXK/RR↓S | 不依赖 | [ |
| TMPRSS2 | 跨膜 | 质膜途径 | K/R↓S | 不依赖 | [ |
| CTSL/B | 内质体/溶酶体 | 内体途径 | R/K/G/A/S/T/Q/Y↓ | 依赖,pH 3.0-6.5 | [ |
| HAT | 跨膜 | 质膜途径 | K/R↓ | 不依赖 | [ |
| 胰蛋白酶 | 细胞外 | 质膜途径 | 选择性不高,主要K/R↓S | 中性pH值,最适pH 8.0 | [ |
| 弹性蛋白酶 | 细胞外 | 质膜途径 | A/S/G/V/W/Y/F/L/C/M↓ | 不依赖 | [ |
| 纤溶酶 | 细胞外 | 质膜途径 | K/R↓ | 不依赖 | [ |
| [72] | Regan AD, Shraybman R, Cohen RD, et al. Differential role for low pH and cathepsin-mediated cleavage of the viral spike protein during entry of serotype II feline coronaviruses [J]. Vet Microbiol, 2008, 132(3/4): 235-248. |
| [73] | Xiao WW, Xiong YX, Wang YC, et al. Cathepsin L and transmembrane serine protease 11E mediate trypsin-independent entry of porcine deltacoronavirus into Huh7 cells [J]. J Virol, 2025, 99(9): e0105525. |
| [74] | Bertram S, Glowacka I, Müller MA, et al. Cleavage and activation of the severe acute respiratory syndrome coronavirus spike protein by human airway trypsin-like protease [J]. J Virol, 2011, 85(24): 13363-13372. |
| [75] | Ren WY, Hong WQ, Yang JY, et al. SARS-CoV-2 Delta and Omicron variants resist spike cleavage by human airway trypsin-like protease [J]. J Clin Investig, 2024, 134(18): e174304. |
| [76] | Kishimoto M, Uemura K, Sanaki T, et al. TMPRSS11D and TMPRSS13 activate the SARS-CoV-2 spike protein [J]. Viruses, 2021, 13(3): 384. |
| [77] | Wicht O, Li WT, Willems L, et al. Proteolytic activation of the porcine epidemic diarrhea coronavirus spike fusion protein by trypsin in cell culture [J]. J Virol, 2014, 88(14): 7952-7961. |
| [78] | Costello DA, Millet JK, Hsia CY, et al. Single particle assay of coronavirus membrane fusion with proteinaceous receptor-embedded supported bilayers [J]. Biomaterials, 2013, 34(32): 7895-7904. |
| [79] | Matsuyama S, Ujike M, Morikawa S, et al. Protease-mediated enhancement of severe acute respiratory syndrome coronavirus infection [J]. Proc Natl Acad Sci U S A, 2005, 102(35): 12543-12547. |
| [80] | Jaimes JA, Millet JK, Whittaker GR. Proteolytic cleavage of the SARS-CoV-2 spike protein and the role of the novel S1/S2 site [J]. iScience, 2020, 23(6): 101212. |
| [81] | Kim Y, Jang G, Lee DR, et al. Trypsin enhances SARS-CoV-2 infection by facilitating viral entry [J]. Arch Virol, 2022, 167(2): 441-458. |
| [82] | Hofmann M, Wyler R. Propagation of the virus of porcine epidemic diarrhea in cell culture [J]. J Clin Microbiol, 1988, 26(11): 2235-2239. |
| [83] | Shirato K, Matsuyama S, Ujike M, et al. Role of proteases in the release of porcine epidemic diarrhea virus from infected cells [J]. J Virol, 2011, 85(15): 7872-7880. |
| [84] | Xiao WW, Li Z, Chen CQ, et al. Revisiting the roles of trypsin in the productive infection of porcine deltacoronavirus in porcine-derived cells [J]. Virology, 2025, 604: 110453. |
| [85] | Korkmaz B, Gauthier F. Chapter 587 - Elastase-2/Leukocyte Elastase [M]//Rawlings ND, Salvesen G. Handbook of Proteolytic Enzymes (Third Edition). Academic Press. 2013: 2653-2661. |
| [86] | Heald-Sargent T, Gallagher T. Ready, set, fuse! the coronavirus spike protein and acquisition of fusion competence [J]. Viruses, 2012, 4(4): 557-580. |
| [87] | Kawabata K, Hagio T, Matsuoka S. The role of neutrophil elastase in acute lung injury [J]. Eur J Pharmacol, 2002, 451(1): 1-10. |
| [88] | Szturmowicz M, Demkow U. Neutrophil extracellular traps (NETs) in severe SARS-CoV-2 lung disease [J]. Int J Mol Sci, 2021, 22(16): 8854. |
| [89] | Kim Y, Oh C, Shivanna V, et al. Trypsin-independent porcine epidemic diarrhea virus US strain with altered virus entry mechanism [J]. BMC Vet Res, 2017, 13(1): 356. |
| [90] | Ji HL, Zhao RZ, Matalon S, et al. Elevated plasmin(ogen) as a common risk factor for COVID-19 susceptibility [J]. Physiol Rev, 2020, 100(3): 1065-1075. |
| [91] | Kam YW, Okumura Y, Kido H, et al. Cleavage of the SARS coronavirus spike glycoprotein by airway proteases enhances virus entry into human bronchial epithelial cells in vitro [J]. PLoS One, 2009, 4(11): e7870. |
| [92] | Kaur U, Chakrabarti SS, Ojha B, et al. Targeting host cell proteases to prevent SARS-CoV-2 invasion [J]. Curr Drug Targets, 2021, 22(2): 192-201. |
| [93] | Glowacka I, Bertram S, Müller MA, et al. Evidence that TMPRSS2 activates the severe acute respiratory syndrome coronavirus spike protein for membrane fusion and reduces viral control by the humoral immune response [J]. J Virol, 2011, 85(9): 4122-4134. |
| [94] | Han YT, Ma YL, Wang ZQ, et al. TMPRSS13 promotes the cell entry of swine acute diarrhea syndrome coronavirus [J]. J Med Virol, 2024, 96(6): e29712. |
| [95] | McKee DL, Sternberg A, Stange U, et al. Candidate drugs against SARS-CoV-2 and COVID-19 [J]. Pharmacol Res, 2020, 157: 104859. |
| [96] | Hernández-Mitre MP, Morpeth SC, Venkatesh B, et al. TMPRSS2 inhibitors for the treatment of COVID-19 in adults: a systematic review and meta-analysis of randomized clinical trials of nafamostat and camostat mesylate [J]. Clin Microbiol Infect, 2024, 30(6): 743-754. |
| [97] | Hoffmann M, Schroeder S, Kleine-Weber H, et al. Nafamostat mesylate blocks activation of SARS-CoV-2: new treatment option for COVID-19 [J]. Antimicrob Agents Chemother, 2020, 64(6): e00754-20. |
| [98] | Li K, Meyerholz DK, Bartlett JA, et al. The TMPRSS2 inhibitor nafamostat reduces SARS-CoV-2 pulmonary infection in mouse models of COVID-19 [J]. mBio, 2021, 12(4): e00970-21. |
| [99] | Lucas JM, Heinlein C, Kim T, et al. The androgen-regulated protease TMPRSS2 activates a proteolytic cascade involving components of the tumor microenvironment and promotes prostate cancer metastasis [J]. Cancer Discov, 2014, 4(11): 1310-1325. |
| [100] | Wettstein L, Weil T, Conzelmann C, et al. Alpha-1 antitrypsin inhibits TMPRSS2 protease activity and SARS-CoV-2 infection [J]. Nat Commun, 2021, 12: 1726. |
| [101] | Walker B, Lynas JF. Strategies for the inhibition of serine proteases [J]. Cell Mol Life Sci CMLS, 2001, 58(4): 596-624. |
| [102] | Singh S, O’Reilly S, Gewaid H, et al. Reactive centre loop mutagenesis of SerpinB3 to target TMPRSS2 and furin: inhibition of SARS-CoV-2 cell entry and replication [J]. Int J Mol Sci, 2022, 23(20): 12522. |
| [103] | Keller C, Böttcher-Friebertshäuser E, Lohoff M. TMPRSS2, a novel host-directed drug target against SARS-CoV-2 [J]. Signal Transduct Target Ther, 2022, 7: 251. |
| [104] | Mahoney M, Damalanka VC, Tartell MA, et al. A novel class of TMPRSS2 inhibitors potently block SARS-CoV-2 and MERS-CoV viral entry and protect human epithelial lung cells [J]. Proc Natl Acad Sci U S A, 2021, 118(43): e2108728118. |
| [105] | Boon ACM, Bricker TL, Fritch EJ, et al. Efficacy of host cell serine protease inhibitor MM3122 against SARS-CoV-2 for treatment and prevention of COVID-19 [J]. J Virol, 2024, 98(5): e01903-23. |
| [106] | Cheng YW, Chao TL, Li CL, et al. Furin inhibitors block SARS-CoV-2 spike protein cleavage to suppress virus production and cytopathic effects [J]. Cell Rep, 2020, 33(2): 108254. |
| [107] | Tanikawa T, Hayashi T, Suzuki R, et al. Inhibitory effect of honokiol on furin-like activity and SARS-CoV-2 infection [J]. J Tradit Complementary Med, 2022, 12(1): 69-72. |
| [108] | Jean F, Stella K, Thomas L, et al. α1-Antitrypsin Portland, a bioengineered serpin highly selective for furin: Application as an antipathogenic agent [J]. Proc Natl Acad Sci U S A, 1998, 95(13): 7293-7298. |
| [1] | Woo PCY, Lau SKP, Huang Y, et al. Coronavirus diversity, phylogeny and interspecies jumping [J]. Exp Biol Med (Maywood), 2009, 234(10): 1117-1127. |
| [2] | Drosten C, Günther S, Preiser W, et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome [J]. N Engl J Med, 2003, 348(20): 1967-1976. |
| [3] | van Boheemen S, de Graaf M, Lauber C, et al. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans [J]. mBio, 2012, 3(6): e00473-12. |
| [4] | Enjuanes L, Almazán F, Sola I, et al. Biochemical aspects of coronavirus replication and virus-host interaction [J]. Annu Rev Microbiol, 2006, 60: 211-230. |
| [5] | Guillén J, Pérez-Berná AJ, Moreno MR, et al. Identification of the membrane-active regions of the severe acute respiratory syndrome coronavirus spike membrane glycoprotein using a 16/18-mer peptide scan: implications for the viral fusion mechanism [J]. J Virol, 2005, 79(3): 1743-1752. |
| [6] | Xing LX, Liu ZM, Wang XL, et al. Early fusion intermediate of ACE2-using coronavirus spike acting as an antiviral target [J]. Cell, 2025, 188(5): 1297-1314.e24. |
| [7] | Belouzard S, Chu VC, Whittaker GR. Activation of the SARS coronavirus spike protein via sequential proteolytic cleavage at two distinct sites [J]. Proc Natl Acad Sci U S A, 2009, 106(14): 5871-5876. |
| [8] | Millet JK, Whittaker GR. Host cell proteases: Critical determinants of coronavirus tropism and pathogenesis [J]. Virus Res, 2015, 202: 120-134. |
| [9] | de Haan CAM, Stadler K, Godeke GJ, et al. Cleavage inhibition of the murine coronavirus spike protein by a furin-like enzyme affects cell-cell but not virus-cell fusion [J]. J Virol, 2004, 78(11): 6048-6054. |
| [10] | Park JE, Li K, Barlan A, et al. Proteolytic processing of Middle East respiratory syndrome coronavirus spikes expands virus tropism [J]. Proc Natl Acad Sci U S A, 2016, 113(43): 12262-12267. |
| [11] | Millet JK, Whittaker GR. Host cell entry of Middle East respiratory syndrome coronavirus after two-step, furin-mediated activation of the spike protein [J]. Proc Natl Acad Sci U S A, 2014, 111(42): 15214-15219. |
| [12] | Yamada Y, Liu DX. Proteolytic activation of the spike protein at a novel RRRR/S motif is implicated in furin-dependent entry, syncytium formation, and infectivity of coronavirus infectious bronchitis virus in cultured cells [J]. J Virol, 2009, 83(17): 8744-8758. |
| [13] | Hansen GH, Delmas B, Besnardeau L, et al. The coronavirus transmissible gastroenteritis virus causes infection after receptor-mediated endocytosis and acid-dependent fusion with an intracellular compartment [J]. J Virol, 1998, 72(1): 527-534. |
| [14] | Shen LW, Mao HJ, Wu YL, et al. TMPRSS2: a potential target for treatment of influenza virus and coronavirus infections [J]. Biochimie, 2017, 142: 1-10. |
| [15] | Seidah NG, Prat A. The biology and therapeutic targeting of the proprotein convertases [J]. Nat Rev Drug Discov, 2012, 11(5): 367-383. |
| [16] | de Haan CAM, Haijema BJ, Schellen P, et al. Cleavage of group 1 coronavirus spike proteins: how furin cleavage is traded off against heparan sulfate binding upon cell culture adaptation [J]. J Virol, 2008, 82(12): 6078-6083. |
| [17] | de Haan CAM, Rottier PJM. Molecular interactions in the assembly of coronaviruses [J]. Adv Virus Res, 2005, 64: 165-230. |
| [18] | Choe Y, Leonetti F, Greenbaum DC, et al. Substrate profiling of cysteine proteases using a combinatorial peptide library identifies functionally unique specificities [J]. J Biol Chem, 2006, 281(18): 12824-12832. |
| [19] | Kawase M, Shirato K, van der Hoek L, et al. Simultaneous treatment of human bronchial epithelial cells with serine and cysteine protease inhibitors prevents severe acute respiratory syndrome coronavirus entry [J]. J Virol, 2012, 86(12): 6537-6545. |
| [20] | Henrich S, Cameron A, Bourenkov GP, et al. The crystal structure of the proprotein processing proteinase furin explains its stringent specificity [J]. Nat Struct Mol Biol, 2003, 10(7): 520-526. |
| [21] | Nakayama K. Furin: a mammalian subtilisin/Kex2p-like endoprotease involved in processing of a wide variety of precursor proteins [J]. Biochem J, 1997, 327 ( Pt 3)(Pt 3): 625-635. |
| [22] | Khatri R, Lohiya B, Kaur G, et al. Understanding the role of conserved proline and serine residues in the SARS-CoV-2 spike cleavage sites in the virus entry, fusion, and infectivity [J]. 3 Biotech, 2023, 13(10): 323. |
| [23] | Kirschke H. Chapter 410 - Cathepsin L [M]//Rawlings ND, Salvesen G. Handbook of Proteolytic Enzymes (Third Edition). Academic Press: Elsevier, 2013: 1808-1817. |
| [24] | Zhao MM, Zhu Y, Zhang L, et al. Novel cleavage sites identified in SARS-CoV-2 spike protein reveal mechanism for cathepsin L-facilitated viral infection and treatment strategies [J]. Cell Discov, 2022, 8: 53. |
| [25] | Evnin Luke B. Substrate specificity of trypsin investigated by using a genetic selection [J]. Proc Natl Acad Sci U S A, 1990, 87(17): 6659-6663. |
| [26] | Baird JTT, Craik CS. Chapter 575 - Trypsin [M]//Rawlings ND, Salvesen G. Handbook of Proteolytic Enzymes (Third Edition). Academic Press: Elsevier, 2013: 2594-2600. |
| [27] | Belouzard S, Madu I, Whittaker GR. Elastase-mediated activation of the severe acute respiratory syndrome coronavirus spike protein at discrete sites within the S2 domain [J]. J Biol Chem, 2010, 285(30): 22758-22763. |
| [28] | Bieth, 2004. Leukocyte elastase. In: Barrett, Rawlings, Woessner (Eds.), Handbook of Proteolytic Enzymes [M]. vol. 2, second ed, pp. 1517-1523. |
| [29] | Backes BJ, Harris JL, Leonetti F, et al. Synthesis of positional-scanning libraries of fluorogenic peptide substrates to define the extended substrate specificity of plasmin and thrombin [J]. Nat Biotechnol, 2000, 18(2): 187-193. |
| [30] | Kleine-Weber H, Elzayat MT, Hoffmann M, et al. Functional analysis of potential cleavage sites in the MERS-coronavirus spike protein [J]. Sci Rep, 2018, 8: 16597. |
| [31] | Reinke LM, Spiegel M, Plegge T, et al. Different residues in the SARS-CoV spike protein determine cleavage and activation by the host cell protease TMPRSS2 [J]. PLoS One, 2017, 12(6): e0179177. |
| [32] | Jackson CB, Farzan M, Chen B, et al. Mechanisms of SARS-CoV-2 entry into cells [J]. Nat Rev Mol Cell Biol, 2022, 23(1): 3-20. |
| [33] | Rajah MM, Hubert M, Bishop E, et al. SARS-CoV-2 Alpha, Beta, and Delta variants display enhanced Spike-mediated syncytia formation [J]. EMBO J, 2021, 40(24): e108944. |
| [34] | Escalera A, Gonzalez-Reiche AS, Aslam S, et al. Mutations in SARS-CoV-2 variants of concern link to increased spike cleavage and virus transmission [J]. Cell Host Microbe, 2022, 30(3): 373-387.e7. |
| [35] | Liu Y, Liu JY, Johnson BA, et al. Delta spike P681R mutation enhances SARS-CoV-2 fitness over Alpha variant [J]. Cell Rep, 2022, 39(7): 110829. |
| [36] | Saito A, Irie T, Suzuki R, et al. Enhanced fusogenicity and pathogenicity of SARS-CoV-2 Delta P681R mutation [J]. Nature, 2022, 602(7896): 300-306. |
| [109] | Varughese KI, Ahmed FR, Carey PR, et al. Crystal structure of a papain-E-64 complex [J]. Biochemistry, 1989, 28(3): 1330-1332. |
| [110] | Hu J, Gao QZ, He CL, et al. Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2 [J]. Genes Dis, 2020, 7(4): 551-557. |
| [111] | Mellott DM, Tseng CT, Drelich A, et al. A clinical-stage cysteine protease inhibitor blocks SARS-CoV-2 infection of human and monkey cells [J]. ACS Chem Biol, 2021, 16(4): 642-650. |
| [112] | Zhou YC, Vedantham P, Lu K, et al. Protease inhibitors targeting coronavirus and filovirus entry [J]. Antivir Res, 2015, 116: 76-84. |
| [113] | 周雯雯, 尤宝庆, 郑怡凡, 等. 组织蛋白酶L小分子抑制剂的抗SARS-CoV-2活性研究 [J]. 药学学报, 2024, 59(3): 600-607. |
| Zhou WW, You BQ, Zheng YF, et al. Anti-SARS-CoV-2 activity of small molecule inhibitors of cathepsin L [J]. Acta Pharm Sin, 2024, 59(3): 600-607. | |
| [114] | Zhou N, Pan T, Zhang JS, et al. Glycopeptide antibiotics potently inhibit cathepsin L in the late endosome/lysosome and block the entry of Ebola virus, middle east respiratory syndrome coronavirus (MERS-CoV), and severe acute respiratory syndrome coronavirus (SARS-CoV) [J]. J Biol Chem, 2016, 291(17): 9218-9232. |
| [115] | Mao BL, Le-Trilling VTK, Tang HH, et al. Diphyllin elicits a doubled-pronged attack on the entry of SARS-CoV-2 by inhibiting cathepsin L and furin [J]. Virus Res, 2024, 350: 199485. |
| [116] | Li YH, Wang K, Sun HM, et al. Omicsynin B4 potently blocks coronavirus infection by inhibiting host proteases cathepsin L and TMPRSS2 [J]. Antivir Res, 2023, 214: 105606. |
| [117] | Essalmani R, Jain J, Susan-Resiga D, et al. Distinctive roles of furin and TMPRSS2 in SARS-CoV-2 infectivity [J]. J Virol, 2022, 96(8): e00128-22. |
| [37] | Wicht O, Burkard C, de Haan CAM, et al. Identification and characterization of a proteolytically primed form of the murine coronavirus spike proteins after fusion with the target cell [J]. J Virol, 2014, 88(9): 4943-4952. |
| [38] | Choi A, Kots ED, Singleton DT, et al. Analysis of the molecular determinants for furin cleavage of the spike protein S1/S2 site in defined strains of the prototype coronavirus murine hepatitis virus (MHV) [J]. Virus Res, 2024, 340: 199283. |
| [39] | Hoffmann M, Kleine-Weber H, Pöhlmann S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells [J]. Mol Cell, 2020, 78(4): 779-784.e5. |
| [40] | Escalera A, Laporte M, Turner S, et al. The impact of S2 mutations on Omicron SARS-CoV-2 cell surface expression and fusogenicity [J]. Emerg Microbes Infect, 2024, 13: 2297553. |
| [41] | Park SB, Khan M, Chiliveri SC, et al. SARS-CoV-2 Omicron variants harbor spike protein mutations responsible for their attenuated fusogenic phenotype [J]. Commun Biol, 2023, 6: 556. |
| [42] | Johnson BA, Xie XP, Bailey AL, et al. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis [J]. Nature, 2021, 591(7849): 293-299. |
| [43] | Hoffmann M, Kleine-Weber H, Graichen L, et al. Acquisition of a multibasic cleavage site does not increase MERS-CoV entry into Calu-3 human lung cells [J]. J Virol, 2024, 98(11): e01305-24. |
| [44] | Cheng JL, Zhao Y, Hu YX, et al. The furin-S2’ site in avian coronavirus plays a key role in central nervous system damage progression [J]. J Virol, 2021, 95(11): JVI.02447-JVI.02420. |
| [45] | Li WT, Wicht O, van Kuppeveld FJM, et al. A single point mutation creating a furin cleavage site in the spike protein renders porcine epidemic diarrhea coronavirus trypsin independent for cell entry and fusion [J]. J Virol, 2015, 89(15): 8077-8081. |
| [46] | Bickerton E, Maier HJ, Stevenson-Leggett P, et al. The S2 subunit of infectious bronchitis virus beaudette is a determinant of cellular tropism [J]. J Virol, 2018, 92(19): e01044-18. |
| [47] | Jiang Y, Gao MY, Cheng X, et al. The V617I substitution in avian coronavirus IBV spike protein plays a crucial role in adaptation to primary chicken kidney cells [J]. Front Microbiol, 2020, 11: 604335. |
| [48] | 姜逸, 周生, 俞燕, 等. 表达不同致病型S基因重组鸡传染性支气管炎病毒的细胞嗜性研究 [J]. 病毒学报, 2017, 33(5): 738-744. |
| Jiang Y, Zhou S, Yu Y, et al. Cell tropism of the recombinant chicken infectious bronchitis virus expressing the S gene of different pathogenic strains [J]. Chin J Virol, 2017, 33(5): 738-744. | |
| [49] | Jiang Y, Xue M, Tang MJ, et al. Adaptation of the infectious bronchitis virus H120 vaccine strain to Vero cell lines [J]. Vet Microbiol, 2023, 280: 109709. |
| [50] | Jiang Y, Cheng X, Gao MY, et al. Two mutations on S2 subunit were critical for Vero cell tropism expansion of infectious bronchitis virus HV80 [J]. Vet Microbiol, 2024, 294: 110134. |
| [51] | Liu DX, Liang JQ, Fung TS. Human coronavirus-229E, -OC43, -NL63, and-HKU1 (Coronaviridae) [M]//Encyclopedia of Virology. Amsterdam: Elsevier, 2021: 428-440. |
| [52] | Belouzard S, Millet JK, Licitra BN, et al. Mechanisms of coronavirus cell entry mediated by the viral spike protein [J]. Viruses, 2012, 4(6): 1011-1033. |
| [53] | Shirato K, Kawase M, Matsuyama S. Middle east respiratory syndrome coronavirus infection mediated by the transmembrane serine protease TMPRSS2 [J]. J Virol, 2013, 87(23): 12552-12561. |
| [54] | Bestle D, Heindl MR, Limburg H, et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells [J]. Life Sci Alliance, 2020, 3(9): e202000786. |
| [55] | Shi W, Fan WL, Bai J, et al. TMPRSS2 and MSPL facilitate trypsin-independent porcine epidemic diarrhea virus replication in vero cells [J]. Viruses, 2017, 9(5): 114. |
| [56] | Lukassen S, Chua RL, Trefzer T, et al. SARS-CoV-2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells [J]. EMBO J, 2020: e105114. |
| [57] | Kakizaki M, Hashimoto R, Nagata N, et al. The respective roles of TMPRSS2 and cathepsins for SARS-CoV-2 infection in human respiratory organoids [J]. J Virol, 2025, 99: e01853-24. |
| [58] | Yang Y, Liu C, Du LY, et al. Two mutations were critical for bat-to-human transmission of middle east respiratory syndrome coronavirus [J]. J Virol, 2015, 89(17): 9119-9123. |
| [59] | Turk V, Stoka V, Vasiljeva O, et al. Cysteine cathepsins: From structure, function and regulation to new frontiers [J]. Biochim Biophys Acta Proteins Proteom, 2012, 1824(1): 68-88. |
| [60] | Biniossek ML, Nägler DK, Becker-Pauly C, et al. Proteomic identification of protease cleavage sites characterizes prime and non-prime specificity of cysteine cathepsins B, L, and S [J]. J Proteome Res, 2011, 10(12): 5363-5373. |
| [61] | Bosch BJ, Bartelink W, Rottier PJM. Cathepsin L functionally cleaves the severe acute respiratory syndrome coronavirus class I fusion protein upstream of rather than adjacent to the fusion peptide [J]. J Virol, 2008, 82(17): 8887-8890. |
| [62] | Willett BJ, Grove J, MacLean OA, et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway [J]. Nat Microbiol, 2022, 7(8): 1161-1179. |
| [63] | Simmons G, Gosalia DN, Rennekamp AJ, et al. Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry [J]. Proc Natl Acad Sci U S A, 2005, 102(33): 11876-11881. |
| [64] | Gierer S, Bertram S, Kaup F, et al. The spike protein of the emerging betacoronavirus EMC uses a novel coronavirus receptor for entry, can be activated by TMPRSS2, and is targeted by neutralizing antibodies [J]. J Virol, 2013, 87(10): 5502-5511. |
| [65] | Ou TL, Mou HH, Zhang LZ, et al. Hydroxychloroquine-mediated inhibition of SARS-CoV-2 entry is attenuated by TMPRSS2 [J]. PLoS Pathog, 2021, 17(1): e1009212. |
| [66] | Ou XY, Liu Y, Lei XB, et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV [J]. Nat Commun, 2020, 11: 1620. |
| [67] | Daniloski Z, Jordan TX, Wessels HH, et al. Identification of required host factors for SARS-CoV-2 infection in human cells [J]. Cell, 2021, 184(1): 92-105.e16. |
| [68] | Bolsinger MM, Drobny A, Wilfling S, et al. SARS-CoV-2 spike protein induces time-dependent CTSL upregulation in HeLa cells and alveolarspheres [J]. J Cell Biochem, 2024, 125(9): e30627. |
| [69] | Kawase M, Shirato K, Matsuyama S, et al. Protease-mediated entry via the endosome of human coronavirus 229E [J]. J Virol, 2009, 83(2): 712-721. |
| [70] | Qiu ZZ, Hingley ST, Simmons G, et al. Endosomal proteolysis by cathepsins is necessary for murine coronavirus mouse hepatitis virus type 2 spike-mediated entry [J]. J Virol, 2006, 80(12): 5768-5776. |
| [71] | Park JE, Cruz DJM, Shin HJ. Clathrin- and serine proteases-dependent uptake of porcine epidemic diarrhea virus into Vero cells [J]. Virus Res, 2014, 191: 21-29. |
| [1] | LIU Yu, LIU Xiao-xiao, ZHENG Jian-hao, LI Bin, HE Hong-hong, MAO Li. Preparation and Identification of Monoclonal Antibodies against the Bovine Coronavirus [J]. Biotechnology Bulletin, 2026, 42(1): 67-75. |
| [2] | TAN Yu-rong, CHEN Dong-liang, YANG Shou-zhen, LAI Zhen-guang, TANG Xiang-min, SUN Zu-dong, ZENG Wei-ying. Functioal Analysis on GmKTI1-like Gene of Soybean Resistance to Bean Pyralid (Lamprosema indicata) [J]. Biotechnology Bulletin, 2025, 41(6): 99-108. |
| [3] | RUZHA Yelizhati, YANG Yu. Strategies for Increasing Heterologous Protein Expression in Pichia pastoris [J]. Biotechnology Bulletin, 2024, 40(3): 118-134. |
| [4] | WANG Xiang-kun, SONG Xue-hong, LIU Jin-long, GUO Pei-hong, ZHUANG Xiao-feng, WEI Liang-meng, ZHOU Fan, ZHANG Shu-yu, GAO Pan-pan, WEI Kai. Novel Coronavirus Subunit Vaccine and Screening of Its Efficient Immune Enhancer [J]. Biotechnology Bulletin, 2023, 39(1): 305-314. |
| [5] | HU Xiu-wen, LIU Hua, WANG Yu, TANG Xue-ming, WANG Jin-bin, ZENG Hai-juan, JIANG Wei, LI Hong. Application of CRISPR-Cas System in Nucleic Acid Detection [J]. Biotechnology Bulletin, 2021, 37(9): 266-273. |
| [6] | QU Huan, LI Cheng, CHEN Rui, LIAO Yi-jie, CAO San-jie, WEN Yi-ping, YAN Qi-gui, HUANG Xiao-bo. Truncated Expression of the S1-CTD Fragment of Porcine Deltacoronavirus and Establishment of an Indirect ELISA for Detecting Its Antibody [J]. Biotechnology Bulletin, 2021, 37(5): 273-280. |
| [7] | CHEN Rui, FU Jia-yu, LIU Hao-yu, LI Cheng, ZHAO Yu-jia, HU Jing-fei, QU Huan, CAO San-jie, WEN Xin-tian, WEN Yi-ping, ZHAO Qin, WU Rui, HUANG Xiao-bo. Prokaryotic Expression and Polyclonal Antibody Preparation of Porcine Deltacoronavirus(PDCoV)N Protein [J]. Biotechnology Bulletin, 2020, 36(8): 104-110. |
| [8] | LIANG Li-qin, YAN Jing, ZHANG Xin, HAO Ze-ting, DUAN Jiang-yan. Research Progresses on the Development and Application of CRISPR Technology [J]. Biotechnology Bulletin, 2018, 34(5): 9-16. |
| [9] | WAN Qing-qing, LI Jie, ZENG Yu, ZHANG Ming-ming, ZHAO Xin-qing, BAI Feng-wu. Enhanced Production of Cellobiohydrolase in the Recombinant Saccharomyces cerevisiae by MRP8 Overexpression [J]. Biotechnology Bulletin, 2018, 34(5): 94-100. |
| [10] | ZHANG Qian-qian, LIU Kang-tai, HAN Zong-sheng, LI Rong-gui. Antiserum Levels in Mice Inoculated with Spike Protein S1 Subunit and Its Coding DNA from Recombinant Middle East Respiratory Syndrome Virus [J]. Biotechnology Bulletin, 2018, 34(10): 150-156. |
| [11] | WANG Yu, WANG Jian-hua, LIU Yun-jun. Digestive Stability of AM79 EPSPS Protein in Simulative Gastric and Intestinal Fluid [J]. Biotechnology Bulletin, 2017, 33(6): 176-181. |
| [12] | SU Peng, GONG Guo-li. Research Progress on Optimizing the Expression of Exogenous Proteins in Escherichia coli [J]. Biotechnology Bulletin, 2017, 33(2): 16-23. |
| [13] | CONG Hua-jian, WU Shuan-hu, TIAN Jian, CHU Xiao-yu, WU Ning-feng. Prediction of Optimal Growth Temperature of Bacterium Based on the Homologous Proteins [J]. Biotechnology Bulletin, 2016, 32(3): 155-165. |
| [14] | Liu Wen, Jia Yihua, Fang Li, Liu Changai, Chen Hao, Lu Kaimin, Hu Wei. Expression, Self-assembly and Antioxidation of Escherichia coli DPS [J]. Biotechnology Bulletin, 2015, 31(11): 202-206. |
| [15] | Zhao Xiayun, Xian Dengyu, Song Ming, Tang Qinglin. Research Progress of MIKC-type MADS-box Protein Regulation on Flowering [J]. Biotechnology Bulletin, 2014, 30(7): 8-15. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||