








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (1): 1-12.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0953
GAN Chen-lu1( ), YOU Yu-ting1, XIE Han-dan1,2, ZENG Zi-xian1,2, ZHU Bo1,2(
), YOU Yu-ting1, XIE Han-dan1,2, ZENG Zi-xian1,2, ZHU Bo1,2( )
)
Received:2025-09-05
Online:2026-01-26
Published:2026-02-04
Contact:
ZHU Bo
E-mail:gancl34@163.com;bozhu@sicnu.edu.cn
GAN Chen-lu, YOU Yu-ting, XIE Han-dan, ZENG Zi-xian, ZHU Bo. Research Progress in Flavin Monooxygenases in Plants[J]. Biotechnology Bulletin, 2026, 42(1): 1-12.
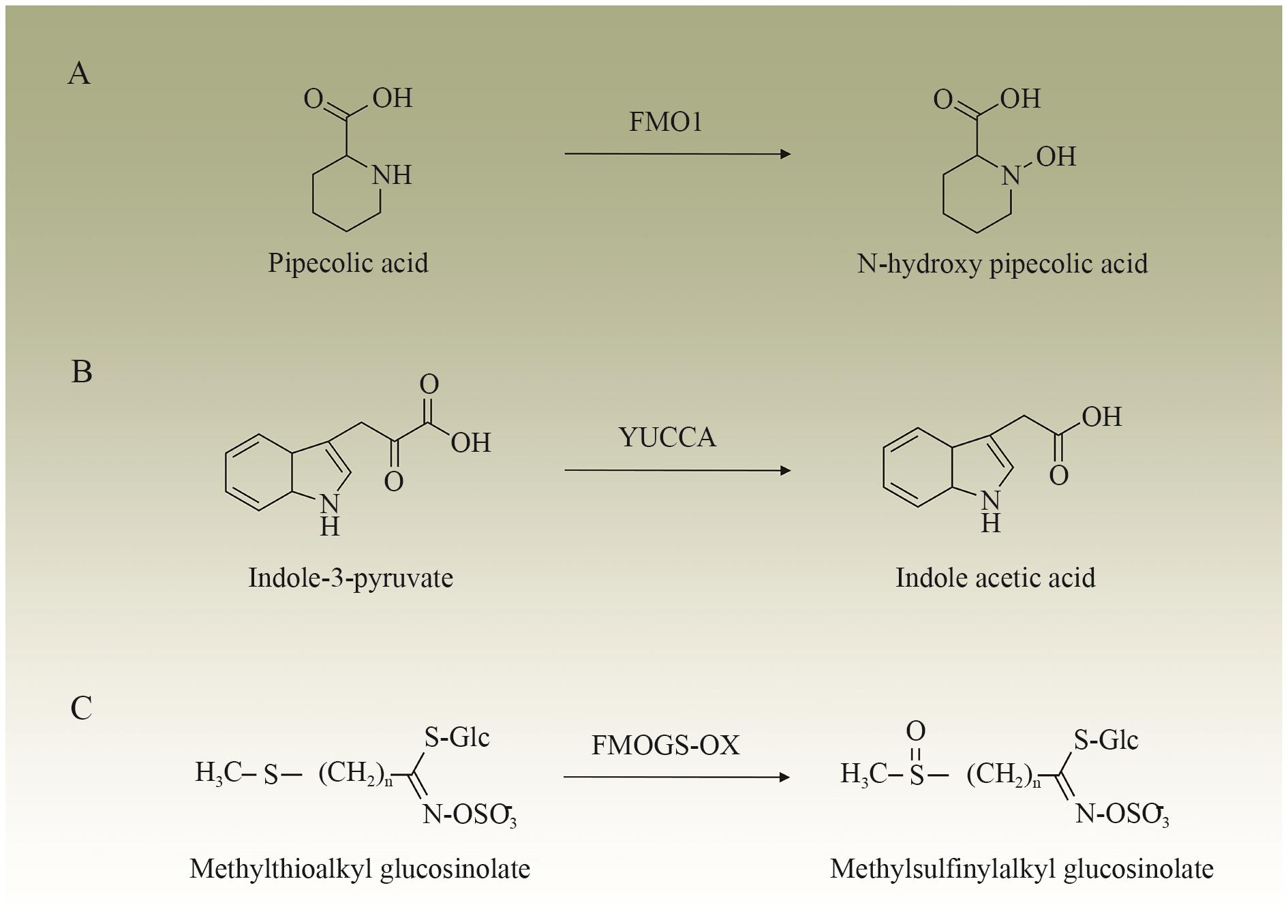
Fig. 2 Catalytic reaction of flavin monooxygenases in plantsA: FMO1 catalyzes the transformation of pipecolic acid to N-hydroxy piperidine pipecolic acid. B: YUCCAs catalyze the conversion of indole-3-pyruvate to indole acetic acid. C: FMOGS-OXs catalyze the conversion of methylthioalkyl glucosinolates to methylsulfinylalkyl glucosinolates
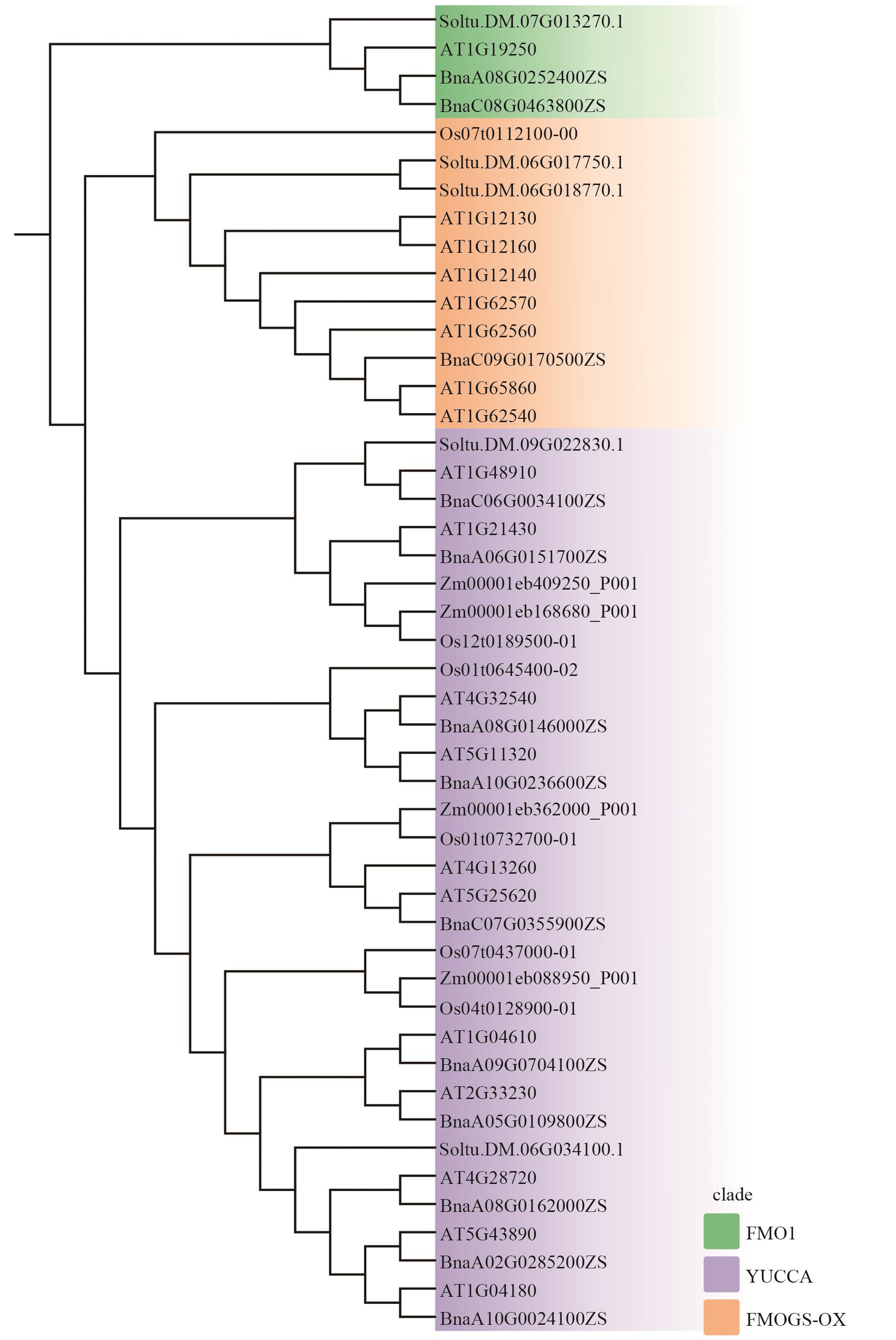
Fig. 3 Phyleigenic trees of flavin monooxygenases in Arabidopsis thaliana, Brassica napus, Oryza sativa, Zea mays, and Solanum tuberosumGreen: FMO1. Purple: YUCCAs. Orange: FMOGS-OXs. At (Arabidopsis thaliana); Bna (Brassica napus); Os (Oryza sativa); Zm (Zea mays); St (Solanum tuberosum)
生理过程 Physiological process | 蛋白类别 Protein category | 植物种类 Plant species | 基因名称 Gene name | 功能描述 Function | 参考文献 References |
|---|---|---|---|---|---|
生长发育 Growth and development | FMO1 | 拟南芥 | FMO1 | 根的发育 | [ |
| 其他 | TaFMO1-5B | 根的发育 | [ | ||
| YUCCAs | 拟南芥 | YUC2、YUC5、YUC8、YUC9 | 茎、叶、花序发育 | [ | |
| 水稻 | OsYUC1、OsYUC2、OsYUC11 | 冠根、籽粒、胚乳发育 | [ | ||
| 玉米 | spi1 | 腋生分生组织的形成 | [ | ||
| 其他 | SiYUC2、SiYUC6、SiYUC8、SiYUC11、Turnera YUC6、FvYUC6、EbYUC2、PpYUC11、CsYUC10、CmYUC6、CmYUC11 | 雄配子体、花粉、穗子、果实、叶片发育及胚胎形成 | [ | ||
| FMO1 | 拟南芥 | FMO1 | 光胁迫 | [ | |
| 其他 | FMO1 | 干旱胁迫 | [ | ||
非生物胁迫 Abiotic stress | YUCCAs | 拟南芥 | YUC6、YUC8 | 干旱、高温、重金属胁迫 | [ |
| 其他 | CsYUC8/9、CsYUC10b | 高温、低温、盐胁迫 | [ | ||
| FMOGS-OXs | 拟南芥 | FMOGS-OX2-7 | 干旱、低温、盐、渗透胁迫及激素处理 | [ | |
| 其他 | TsFMO | 盐胁迫 | [ | ||
生物胁迫 Biological stress | FMO1 | 拟南芥 | FMO1 | 系统性获得抗性(SAR)、降解病原毒素 | [ |
| 其他 | FMO | NHP合成、抑制黄酮合成并促进木质素积累 | [ | ||
| YUCCAs | 其他 | Bs3 | 触发超敏反应 | [ | |
| FMOGS-OXs | 拟南芥 | FMOGS-OXs | GSL生物合成 | [ | |
次生代谢 Secondary metabolism | FMOGS-OXs | 拟南芥 | FMOGS-OXs | GSL生物合成 | [ |
| 其他 | FMOGS-OX2、BrrFMOGS-OX2、BrrFMOGS-OX5.1、BrrFMOGS-OX5.2、BrrFMOGS-OX6.1、BrrFMOGS-OX6.2、FMOGS-OXs、AsFMO1 | GSL生物合成、催化蒜氨酸S-氧化 | [ |
Table 1 Physiological functions of three types of flavin monooxygenases in different species
生理过程 Physiological process | 蛋白类别 Protein category | 植物种类 Plant species | 基因名称 Gene name | 功能描述 Function | 参考文献 References |
|---|---|---|---|---|---|
生长发育 Growth and development | FMO1 | 拟南芥 | FMO1 | 根的发育 | [ |
| 其他 | TaFMO1-5B | 根的发育 | [ | ||
| YUCCAs | 拟南芥 | YUC2、YUC5、YUC8、YUC9 | 茎、叶、花序发育 | [ | |
| 水稻 | OsYUC1、OsYUC2、OsYUC11 | 冠根、籽粒、胚乳发育 | [ | ||
| 玉米 | spi1 | 腋生分生组织的形成 | [ | ||
| 其他 | SiYUC2、SiYUC6、SiYUC8、SiYUC11、Turnera YUC6、FvYUC6、EbYUC2、PpYUC11、CsYUC10、CmYUC6、CmYUC11 | 雄配子体、花粉、穗子、果实、叶片发育及胚胎形成 | [ | ||
| FMO1 | 拟南芥 | FMO1 | 光胁迫 | [ | |
| 其他 | FMO1 | 干旱胁迫 | [ | ||
非生物胁迫 Abiotic stress | YUCCAs | 拟南芥 | YUC6、YUC8 | 干旱、高温、重金属胁迫 | [ |
| 其他 | CsYUC8/9、CsYUC10b | 高温、低温、盐胁迫 | [ | ||
| FMOGS-OXs | 拟南芥 | FMOGS-OX2-7 | 干旱、低温、盐、渗透胁迫及激素处理 | [ | |
| 其他 | TsFMO | 盐胁迫 | [ | ||
生物胁迫 Biological stress | FMO1 | 拟南芥 | FMO1 | 系统性获得抗性(SAR)、降解病原毒素 | [ |
| 其他 | FMO | NHP合成、抑制黄酮合成并促进木质素积累 | [ | ||
| YUCCAs | 其他 | Bs3 | 触发超敏反应 | [ | |
| FMOGS-OXs | 拟南芥 | FMOGS-OXs | GSL生物合成 | [ | |
次生代谢 Secondary metabolism | FMOGS-OXs | 拟南芥 | FMOGS-OXs | GSL生物合成 | [ |
| 其他 | FMOGS-OX2、BrrFMOGS-OX2、BrrFMOGS-OX5.1、BrrFMOGS-OX5.2、BrrFMOGS-OX6.1、BrrFMOGS-OX6.2、FMOGS-OXs、AsFMO1 | GSL生物合成、催化蒜氨酸S-氧化 | [ |
| [1] | Mitchell AJ, Weng J-K. Unleashing the synthetic power of plant oxygenases: from mechanism to application [J]. Plant Physiol, 2019, 179(3): 813-829. |
| [2] | Thodberg S, Jakobsen Neilson EH. The “green” FMOs: diversity, functionality and application of plant flavoproteins [J]. Catalysts, 2020, 10(3): 329. |
| [3] | Schlaich NL. Flavin-containing monooxygenases in plants: looking beyond detox [J]. Trends Plant Sci, 2007, 12(9): 412-418. |
| [4] | Ge CN, Gao CJ, Chen QG, et al. ESCRT-dependent vacuolar sorting and degradation of the auxin biosynthetic enzyme YUC1 flavin monooxygenase [J]. J Integr Plant Biol, 2019, 61(9): 968-973. |
| [5] | Kriechbaumer V, Wang PW, Hawes C, et al. Alternative splicing of the auxin biosynthesis gene YUCCA4 determines its subcellular compartmentation [J]. Plant J, 2012, 70(2): 292-302. |
| [6] | Kim JI, Sharkhuu A, Jin JB, et al. yucca6, a dominant mutation in Arabidopsis, affects auxin accumulation and auxin-related phenotypes [J]. Plant Physiol, 2007, 145(3): 722-735. |
| [7] | Zhang YY, Mao QS, Ma RJ, et al. Genome-wide identification and expression analysis of the PpYUCCA gene family in weeping peach trees (Prunus persica ‘pendula’) [J]. Horticulturae, 2022, 8(10): 878. |
| [8] | Zhang K, Zhang JF, Cui C, et al. Genome-wide identification and expression profiling of the YUCCA gene family in Brassica napus [J]. Oil Crop Sci, 2022, 7(3): 103-111. |
| [9] | Li J, Kristiansen KA, Hansen BG, et al. Cellular and subcellular localization of flavin-monooxygenases involved in glucosinolate biosynthesis [J]. J Exp Bot, 2011, 62(3): 1337-1346. |
| [10] | Huijbers MME, Montersino S, Westphal AH, et al. Flavin dependent monooxygenases [J]. Arch Biochem Biophys, 2014, 544: 2-17. |
| [11] | Dai XH, Mashiguchi K, Chen QG, et al. The biochemical mechanism of auxin biosynthesis by an Arabidopsis yucca flavin -containing-containing monooxygenase [J]. J Biol Chem, 2013, 288(3): 1448-1457. |
| [12] | Yildiz I, Mantz M, Hartmann M, et al. The mobile SAR signal N-hydroxypipecolic acid induces NPR1-dependent transcriptional reprogramming and immune priming [J]. Plant Physiol, 2021, 186(3): 1679-1705. |
| [13] | Chen YC, Holmes EC, Rajniak J, et al. N-hydroxy-pipecolic acid is a mobile metabolite that induces systemic disease resistance in Arabidopsis [J]. Proc Natl Acad Sci U S A, 2018, 115(21): 4920-4929. |
| [14] | Hou XH, Liu SN, Pierri F, et al. Allelic analyses of the Arabidopsis YUC1 locus reveal residues and domains essential for the functions of YUC family of flavin monooxygenases [J]. J Integr Plant Biol, 2011, 53(1): 54-62. |
| [15] | Ahn G, Jeong SY, Khan HA, et al. FAD and NADPH binding sites of YUCCA6 are essential for chaperone activity and oxidative stress tolerance in Arabidopsis thaliana [J]. Plant Physiol Biochem, 2025, 218: 109335. |
| [16] | Jing Li BGH. Subclade of flavin-monooxygenases involved in aliphatic glucosinolate biosynthesis [J]. Plant Physiol, 2008, 148(3): 1721-1733. |
| [17] | Sun S, Bakkeren G. A bird’s-eye view: exploration of the flavin-containing monooxygenase superfamily in common wheat [J]. Front Plant Sci, 2024, 15: 1369299. |
| [18] | Gaba Y, Bhowal B, Pareek A, et al. Genomic survey of flavin monooxygenases in wild and cultivated rice provides insight into evolution and functional diversities [J]. Int J Mol Sci, 2023, 24(4): 4190. |
| [19] | Turnaev II, Gunbin KV, Suslov VV, et al. The phylogeny of class B flavoprotein monooxygenases and the origin of the YUCCA protein family [J]. Plants, 2020, 9(9): 1092. |
| [20] | Wang CY, Liu Y, Li SS, et al. Origin of plant auxin biosynthesis in charophyte algae [J]. Trends Plant Sci, 2014, 19(12): 741-743. |
| [21] | Matthes MS, Best NB, Robil JM, et al. Auxin EvoDevo: conservation and diversification of genes regulating auxin biosynthesis, transport, and signaling [J]. Mol Plant, 2019, 12(3): 298-320. |
| [22] | Hofberger JA, Lyons E, Edger PP, et al. Whole genome and tandem duplicate retention facilitated glucosinolate pathway diversification in the mustard family [J]. Genome Biol Evol, 2013, 5(11): 2155-2173. |
| [23] | Cang W, Sheng YX, Evivie ER, et al. Lineage-specific evolution of flavin-containing monooxygenases involved in aliphatic glucosinolate side-chain modification [J]. J Syst Evol, 2018, 56(2): 92-104. |
| [24] | Xie JM, Chen YR, Cai GJ, et al. Tree Visualization By One Table (tvBOT): a web application for visualizing, modifying and annotating phylogenetic trees [J]. Nucleic Acids Res, 2023, 51(W1): W587-W592. |
| [25] | Zhang T, Li RN, Xing JL, et al. The YUCCA-auxin-WOX11 module controls crown root development in rice [J]. Front Plant Sci, 2018, 9: 523. |
| [26] | Jiang HJ, Zhai KE, Ye XF, et al. The endosperm-specific expression of YUCCA Genes enhances rice grain filling [J]. Phyton, 2022, 91(12): 2633-2648. |
| [27] | Abu-Zaitoon YM, Bennett K, Normanly J, et al. A large increase in IAA during development of rice grains correlates with the expression of tryptophan aminotransferase OsTAR1 and a grain-specific YUCCA [J]. Physiol Plant, 2012, 146(4): 487-499. |
| [28] | Meng Q, Zhang RL, Wang YN, et al. Genome-wide characterization and haplotypic variation analysis of the YUC gene family in foxtail millet (Setaria italica) [J]. Int J Mol Sci, 2023, 24(21): 15637. |
| [29] | Gallavotti A, Barazesh S, Malcomber S, et al. sparse inflorescence1encodes a monocot-specificYUCCA-like gene required for vegetative and reproductive development in maize [J]. Proc Natl Acad Sci U S A, 2008, 105(39): 15196-15201. |
| [30] | Henning PM, Shore JS, McCubbin AG. The S-gene YUC6 pleiotropically determines male mating type and pollen size in heterostylous Turnera (Passifloraceae): a novel neofunctionalization of the YUCCA gene family [J]. Plants, 2022, 11(19): 2640. |
| [31] | Zhu Q, Lu YC, Xiong JL, et al. Development of a stable genetic transformation system in Erigeron breviscapus: a case study with EbYUC2 in relation to leaf number and flowering time [J]. Planta, 2024, 259(5): 98. |
| [32] | Liu H, Xie WF, Zhang L, et al. Auxin biosynthesis by the YUCCA6 flavin monooxygenase gene in woodland strawberry [J]. J Integr Plant Biol, 2014, 56(4): 350-363. |
| [33] | Tatsuki M, Soeno K, Shimada Y, et al. Insertion of a transposon-like sequence in the 5'-flanking region of the YUCCA gene causes the stony hard phenotype [J]. Plant J, 2018, 96(4): 815-827. |
| [34] | Li SN, Wang CH, Zhou XY, et al. The curvature of cucumber fruits is associated with spatial variation in auxin accumulation and expression of a YUCCA biosynthesis gene [J]. Hortic Res, 2020, 7: 135. |
| [35] | Zheng L, Zhang L, Duan K, et al. YUCCA type auxin biosynthesis genes encoding flavin monooxygenases in melon: Genome-wide identification and developmental expression analysis [J]. S Afr N J Bot, 2016, 102: 142-152. |
| [36] | Müller-Moulé P, Nozue K, Pytlak ML, et al. YUCCA auxin biosynthetic genes are required for Arabidopsis shade avoidance [J]. PeerJ, 2016, 4: e2574. |
| [37] | Cheng YF, Dai XH, Zhao YD. Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation inArabidopsis [J]. Plant Cell, 2007, 19(8): 2430-2439. |
| [38] | Woodward C, Bemis SM, Hill EJ, et al. Interaction of auxin and ERECTA in elaborating Arabidopsis inflorescence architecture revealed by the activation tagging of a new member of the YUCCA family putative flavin monooxygenases [J]. Plant Physiol, 2005, 139(1): 192-203. |
| [39] | Chen PY, Umeda M. DNA double-strand breaks induce the expression of flavin-containing monooxygenase and reduce root meristem size in Arabidopsis thaliana [J]. Genes Cells, 2015, 20(8): 636-646. |
| [40] | Zhao P, Ma XY, Zhang RZ, et al. Integration of genome-wide association study, linkage analysis, and population transcriptome analysis to reveal the TaFMO1-5B modulating seminal root growth in bread wheat [J]. Plant J, 2023, 116(5): 1385-1400. |
| [41] | Catalá R, López-Cobollo R, Berbís MÁ, et al. Trimethylamine N-oxide is a new plant molecule that promotes abiotic stress tolerance [J]. Sci Adv, 2021, 7(21): eabd9296. |
| [42] | Cao HY, Liu RN, Zhang JH, et al. Improving sulforaphane content in transgenic broccoli plants by overexpressing MAM1, FMOGS-OX2, and Myrosinase [J]. Plant Cell Tissue Organ Cult, 2021, 146(3): 461-471. |
| [43] | Wang LL, Zhou YL, Ding Y, et al. Novel flavin-containing monooxygenase protein FMO1 interacts with CAT2 to negatively regulate drought tolerance through ROS homeostasis and ABA signaling pathway in tomato [J]. Hortic Res, 2023, 10(4): uhad037. |
| [44] | Cha JY, Kim WY, Kang SB, et al. A novel thiol-reductase activity of Arabidopsis YUC6 confers drought tolerance independently of auxin biosynthesis [J]. Nat Commun, 2015, 6: 8041. |
| [45] | Kim JI, Baek D, Park HC, et al. Overexpression of Arabidopsis YUCCA6 in potato results in high-auxin developmental phenotypes and enhanced resistance to water deficit [J]. Mol Plant, 2013, 6(2): 337-349. |
| [46] | Ke QB, Wang Z, Ji CY, et al. Transgenic poplar expressing Arabidopsis YUCCA6 exhibits auxin-overproduction phenotypes and increased tolerance to abiotic stress [J]. Plant Physiol Biochem, 2015, 94: 19-27. |
| [47] | Nicolas-Espinosa J, Garcia-Ibañez P, Lopez-Zaplana A, et al. Confronting secondary metabolites with water uptake and transport in plants under abiotic stress [J]. Int J Mol Sci, 2023, 24(3): 2826. |
| [48] | Zhang TY, Liu R, Zheng JY, et al. Insights into glucosinolate accumulation and metabolic pathways in Isatis indigotica Fort [J]. BMC Plant Biol, 2022, 22(1): 78. |
| [49] | Sun JQ, Qi LL, Li YN, et al. PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating Arabidopsis hypocotyl growth [J]. PLoS Genet, 2012, 8(3): e1002594. |
| [50] | Yan SS, Che G, Ding L, et al. Different cucumber CsYUC genes regulate response to abiotic stresses and flower development [J]. Sci Rep, 2016, 6: 20760. |
| [51] | Ye X, Kang BG, Osburn LD, et al. Identification of the flavin-dependent monooxygenase-encoding YUCCA gene family in Populustrichocarpa and their expression in vegetative tissues and in response to hormone and environmental stresses [J]. Plant Cell Tissue Organ Cult, 2009, 97(3): 271-283. |
| [52] | Gao Q, Gao F, Wang RC, et al. Molecular cloning, expression, and polyclonal antibody production of a novel flavin-containing monooxygenase from Thellungiella Halophila [J]. Plant Mol Biol Report, 2009, 27(1): 94-101. |
| [53] | Zhao HY, Li D, Liu YQ, et al. Flavin-containing monooxygenases FMOGS-OXs integrate flowering transition and salt tolerance in Arabidopsis thaliana [J]. Physiol Plant, 2024, 176(2): e14287. |
| [54] | Czarnocka W, Fichman Y, Bernacki M, et al. FMO1 is involved in excess light stress-induced signal transduction and cell death signaling [J]. Cells, 2020, 9(10): 2163. |
| [55] | Cha JY, Jeong SY, Ahn G, et al. The thiol-reductase activity of YUCCA6 enhances nickel heavy metal stress tolerance in Arabidopsis [J]. Front Plant Sci, 2022, 13: 1007542. |
| [56] | Bray EA. Genes commonly regulated by water-deficit stress in Arabidopsis thaliana [J]. J Exp Bot, 2004, 55(407): 2331-2341. |
| [57] | Li YM, Li R, Sawada Y, et al. Abscisic acid-mediated induction of FLAVIN-CONTAINING MONOOXYGENASE 2 leads to reduced accumulation of methylthioalkyl glucosinolates in Arabidopsis thaliana [J]. Plant Sci, 2021, 303: 110764. |
| [58] | Kong WW, Li J, Yu QY, et al. Two novel flavin-containing monooxygenases involved in biosynthesis of aliphatic glucosinolates [J]. Front Plant Sci, 2016, 7: 1292. |
| [59] | Tatiana E Mishina JZ. The Arabidopsis flavin -dependent-dependent monooxygenase FMO1 is an essential component of biologically induced systemic acquired resistance [J]. Plant Physiol, 2006, 141(4): 1666-1675. |
| [60] | Hartmann M, Zeier T, Bernsdorff F, et al. Flavin monooxygenase-generated N-hydroxypipecolic acid is a critical element of plant systemic immunity [J]. Cell, 2018, 173(2): 456-469.e16. |
| [61] | Koch M, Vorwerk S, Masur C, et al. A role for a flavin-containing mono-oxygenase in resistance against microbial pathogens in Arabidopsis [J]. Plant J, 2006, 47(4): 629-639. |
| [62] | Bartsch M, Gobbato E, Bednarek P, et al. Salicylic acid-independent ENHANCED DISEASE SUSCEPTIBILITY1 signaling in Arabidopsis immunity and cell death is regulated by the monooxygenase FMO1 and the nudix hydrolase NUDT7 [J]. Plant Cell, 2006, 18(4): 1038-1051. |
| [63] | Ling YM, Xiong XP, Yang WL, et al. Comparative analysis of transcriptomics and metabolomics reveals defense mechanisms in melon cultivars against pseudoperonospora cubensis infection [J]. Int J Mol Sci, 2023, 24(24): 17552. |
| [64] | Christina Krönauer JK. Cell death triggered by the YUCCA-like Bs3 protein coincides with accumulation of salicylic acid and pipecolic acid but not of indole-3-acetic acid [J]. Plant Physiol, 2019, 180(3): 1647-1659. |
| [65] | Kliebenstein DJ, Kroymann J, Mitchell-Olds T. The glucosinolate-myrosinase system in an ecological and evolutionary context [J]. Curr Opin Plant Biol, 2005, 8(3): 264-271. |
| [66] | Daniel J Kliebenstein JK. Genetic control of natural variation in Arabidopsis glucosinolate accumulation [J]. Plant Physiol, 2001, 126(2): 811-825. |
| [67] | Geiselhardt S, Yoneya K, Blenn B, et al. Egg laying of cabbage white butterfly (Pieris brassicae) on Arabidopsis thaliana affects subsequent performance of the larvae [J]. PLoS One, 2013, 8(3): e59661. |
| [68] | Hopkins RJ, van Dam NM, van Loon JJA. Role of glucosinolates in insect-plant relationships and multitrophic interactions [J]. Annu Rev Entomol, 2009, 54: 57-83. |
| [69] | Calmes B, N’Guyen G, Dumur J, et al. Glucosinolate-derived isothiocyanates impact mitochondrial function in fungal cells and elicit an oxidative stress response necessary for growth recovery [J]. Front Plant Sci, 2015, 6: 414. |
| [70] | Michael Dalgaard Mikkelsen BLP. Modulation of CYP79 genes and glucosinolate profiles in Arabidopsis by defense signaling pathways [J]. Plant Physiol, 2003, 131(1): 298-308. |
| [71] | Hansen BG, Kliebenstein DJ, Halkier BA. Identification of a flavin-monooxygenase as the S-oxygenating enzyme in aliphatic glucosinolate biosynthesis in Arabidopsis [J]. Plant J, 2007, 50(5): 902-910. |
| [72] | Li ZS, Liu YM, Li LY, et al. Transcriptome reveals the gene expression patterns of sulforaphane metabolism in broccoli florets [J]. PLoS One, 2019, 14(3): e0213902. |
| [73] | Li ZS, Liu GM, He HJ, et al. Effects of nanocarbon solution treatment on the nutrients and glucosinolate metabolism in broccoli [J]. Food Chem, 2022, 15: 100429. |
| [74] | Lee YS, Ku K-M, Becker TM, et al. Chemopreventive glucosinolate accumulation in various broccoli and collard tissues: Microfluidic-based targeted transcriptomics for by-product valorization [J]. PLoS One, 2017, 12(9): e0185112. |
| [75] | Yang Y, Hu Y, Yue YL, et al. Expression profiles of glucosinolate biosynthetic genes in turnip (Brassica rapa var. Rapa) at different developmental stages and effect of transformed flavin-containing monooxygenase genes on hairy root glucosinolate content [J]. J Sci Food Agric, 2020, 100(3): 1064-1071. |
| [76] | Zhao YJ, Chen ZY, Chen JX, et al. Comparative transcriptomic analyses of glucosinolate metabolic genes during the formation of Chinese kale seeds [J]. BMC Plant Biol, 2021, 21(1): 394. |
| [77] | Yoshimoto N, Onuma M, Mizuno S, et al. Identification of a flavin-containing S-oxygenating monooxygenase involved in alliin biosynthesis in garlic [J]. Plant J, 2015, 83(6): 941-951. |
| [78] | Grubb CD, Abel S. Glucosinolate metabolism and its control [J]. Trends Plant Sci, 2006, 11(2): 89-100. |
| [1] | LONG Lin-xi, ZENG Yin-ping, WANG Qian, DENG Yu-ping, GE Min-qian, CHEN Yan-zhuo, LI Xin-juan, YANG Jun, ZOU Jian. Identification of Sunflower GH3 Gene Family and Analysis of Their Function in Flower Development [J]. Biotechnology Bulletin, 2026, 42(1): 125-138. |
| [2] | HE Qi-chen, YANG Yang, ALIYA Waili, TANG Xin-yue, LI Zhong-xi, CHEN Yong-kun, CHEN Ling-na. Investigation into the Family Characteristics of the Lavender Copper Amine Oxidase Gene and the Role of LaCuAO1 in Bioamine Degradation [J]. Biotechnology Bulletin, 2026, 42(1): 114-124. |
| [3] | YANG Dan, JIN Ya-rong, MAO Chun-li, WANG Bi-xian, ZHANG Ya-ning, YANG Zhi-yi, ZHOU Zhi-yao, YANG Rui-ming, FAN Heng-rui, HUANG Lin-kai, YAN Hai-dong. Identification and Expression Analysis of C2H2 Gene Family in Cenchrus purpureus [J]. Biotechnology Bulletin, 2026, 42(1): 251-261. |
| [4] | LI Jian-bin, HOU Jia-e, LI Lei-lin, AI Ming-tao, LIU Tian-tai, CUI Xiu-ming, YANG Qian. Genome-wide Identification of Panax notoginseng Lipoxygenases Coupled in Response to Methyl-jasmonate and Wounding [J]. Biotechnology Bulletin, 2026, 42(1): 218-229. |
| [5] | CHENG Ting-ting, LIU Jun, WANG Li-li, LIAN Cong-long, WEI Wen-jun, GUO Hui, WU Yao-lin, YANG Jing-fan, LAN Jin-xu, CHEN Sui-qing. Genome-wide Identification of the Chalcone Isomerase Gene Family in Eucommia ulmoides and Analysis of Their Expression Patterns [J]. Biotechnology Bulletin, 2025, 41(9): 242-255. |
| [6] | LI Shan, MA Deng-hui, MA Hong-yi, YAO Wen-kong, YIN Xiao. Identification and Expression Analysis of SKP1 Gene Family in Grapevine (Vitis vinifera L.) [J]. Biotechnology Bulletin, 2025, 41(9): 147-158. |
| [7] | HUANG Guo-dong, DENG Yu-xing, CHENG Hong-wei, DAN Yan-nan, ZHOU Hui-wen, WU Lan-hua. Genome-wide Identification and Expression Analysis of the ZIP Gene Family in Soybean [J]. Biotechnology Bulletin, 2025, 41(9): 71-81. |
| [8] | HUA Wen-ping, LIU Fei, HAO Jia-xin, CHEN Chen. Identification and Expression Patterns Analysis of ADH Gene Family in Salvia miltiorrhiza [J]. Biotechnology Bulletin, 2025, 41(8): 211-219. |
| [9] | LA Gui-xiao, ZHAO Yu-long, DAI Dan-dan, YU Yong-liang, GUO Hong-xia, SHI Gui-xia, JIA Hui, YANG Tie-gang. Identification of Plasma Membrane H+-ATPase Gene Family in Safflower and Expression Analysis in Response to Low Nitrogen and Low Phosphorus Stress [J]. Biotechnology Bulletin, 2025, 41(8): 220-233. |
| [10] | HUANG Shi-yu, TIAN Shan-shan, YANG Tian-wei, GAO Man-rong, ZHANG Shang-wen. Genome-wide Identification and Expression Pattern Analysis of WRI1 Gene Family in Erythropalum scandens [J]. Biotechnology Bulletin, 2025, 41(8): 242-254. |
| [11] | CHENG Xue, FU Ying, CHAI Xiao-jiao, WANG Hong-yan, DENG Xin. Identification of LHC Gene Family in Setaria italica and Expression Analysis under Abiotic Stresses [J]. Biotechnology Bulletin, 2025, 41(8): 102-114. |
| [12] | REN Rui-bin, SI Er-jing, WAN Guang-you, WANG Jun-cheng, YAO Li-rong, ZHANG Hong, MA Xiao-le, LI Bao-chun, WANG Hua-jun, MENG Ya-xiong. Identification and Expression Analysis of GH17 Gene Family of Pyrenophora graminea [J]. Biotechnology Bulletin, 2025, 41(8): 146-154. |
| [13] | ZHANG Ze, YANG Xiu-li, NING Dong-xian. Identification of 4CL Gene Family in Arachis hypogaea L. and Expression Analysis in Response to Drought and Salt Stress [J]. Biotechnology Bulletin, 2025, 41(7): 117-127. |
| [14] | LI Xin-ni, LI Jun-yi, MA Xue-hua, HE Wei, LI Jia-li, YU Jia, CAO Xiao-ning, QIAO Zhi-jun, LIU Si-chen. Identification of the PMEI Gene Family of Pectin Methylesterase Inhibitor in Foxtail Millet and Analysis of Its Response to Abiotic Stress [J]. Biotechnology Bulletin, 2025, 41(7): 150-163. |
| [15] | LI Kai-yue, DENG Xiao-xia, YIN Yuan, DU Ya-tong, XU Yuan-jing, WANG Jing-hong, YU Song, LIN Ji-xiang. Identification of LEA Gene Family and Analysis on Its Response to Aluminum Stress in Ricinus communis L. [J]. Biotechnology Bulletin, 2025, 41(7): 128-138. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||