








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 72-82.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1013
Previous Articles Next Articles
PANG Xin-li1,2( ), ZHANG Hong-bing1, LIU Xiao-qing2, WANG Yuan3, WU Ning-feng2, TIAN Jian3, GUAN Fei-fei2(
), ZHANG Hong-bing1, LIU Xiao-qing2, WANG Yuan3, WU Ning-feng2, TIAN Jian3, GUAN Fei-fei2( )
)
Received:2025-09-22
Online:2026-04-26
Published:2026-04-30
Contact:
GUAN Fei-fei
E-mail:pangxinli160703@icloud.com;guanfeifei@caas.cn
PANG Xin-li, ZHANG Hong-bing, LIU Xiao-qing, WANG Yuan, WU Ning-feng, TIAN Jian, GUAN Fei-fei. Research Advances in Enhancing the Soluble Expression of Recombinant Heterologous Proteins[J]. Biotechnology Bulletin, 2026, 42(4): 72-82.
宿主名称 Host name | 优点 Advantages | 缺点 Disadvantages | 适用蛋白类型 Applicable protein types | 常用优化策略举例 Example of common optimization strategies | 参考文献References |
|---|---|---|---|---|---|
| 大肠杆菌 E. coli | 生长快、成本低、遗传背景清晰、表达量高 | 缺乏复杂翻译后修饰、易形成包涵体 | 非糖基化蛋白、酶、抗体片段 | 使用工程化菌株(BL21,Rosetta)、优化诱导条件(Tunner,Lemo21)、使用融合标签 | [ |
| 枯草芽胞杆菌 B. subtilis | 非致病、分泌能力强、无内毒素 | 胞外蛋白酶多、遗传稳定性有时不足 | 工业酶(淀粉酶、蛋白酶)、抗原 | 蛋白酶缺陷株、信号肽优化、强启动子 | [ |
| 毕赤酵母 P. pastoris | 高密度发酵、分泌能力强、具备真核修饰能力 | 糖基化模式与人类不同、甲醇利用有安全隐患 | 分泌型蛋白、工业酶、部分糖蛋白 | AOX1启动子调控、温度控制、糖基化通路工程(ΔOCH1)、共底物喂养 | [ |
| 酿酒酵母 S.cerevisiae | 安全、遗传操作工具成熟 | 表达量相对较低、糖基化过度 | 疫苗抗原、食品酶、基础研究 | 糖基化通路改造(ΔOCH1,ΔALG3)、强启动子(GAL1等) | [ |
| 哺乳动物细胞(如CHO, HEK293) | 能完成复杂翻译后修饰、产物最接近天然 | 成本高、周期长、培养复杂 | 治疗性抗体、复杂糖蛋白、病毒样颗粒 | 细胞系工程(GS/KO系统)、培养基优化、过程控制 | [ |
| 丝状真菌(如木霉、曲霉) | 强大的蛋白分泌能力、能进行某些复杂修饰 | 遗传操作相对复杂、背景分泌蛋白多 | 工业水解酶(纤维素酶、淀粉酶) | 强启动子、蛋白酶缺陷株、分泌信号优化 | [ |
Table 1 Comparison of common expression host systems
宿主名称 Host name | 优点 Advantages | 缺点 Disadvantages | 适用蛋白类型 Applicable protein types | 常用优化策略举例 Example of common optimization strategies | 参考文献References |
|---|---|---|---|---|---|
| 大肠杆菌 E. coli | 生长快、成本低、遗传背景清晰、表达量高 | 缺乏复杂翻译后修饰、易形成包涵体 | 非糖基化蛋白、酶、抗体片段 | 使用工程化菌株(BL21,Rosetta)、优化诱导条件(Tunner,Lemo21)、使用融合标签 | [ |
| 枯草芽胞杆菌 B. subtilis | 非致病、分泌能力强、无内毒素 | 胞外蛋白酶多、遗传稳定性有时不足 | 工业酶(淀粉酶、蛋白酶)、抗原 | 蛋白酶缺陷株、信号肽优化、强启动子 | [ |
| 毕赤酵母 P. pastoris | 高密度发酵、分泌能力强、具备真核修饰能力 | 糖基化模式与人类不同、甲醇利用有安全隐患 | 分泌型蛋白、工业酶、部分糖蛋白 | AOX1启动子调控、温度控制、糖基化通路工程(ΔOCH1)、共底物喂养 | [ |
| 酿酒酵母 S.cerevisiae | 安全、遗传操作工具成熟 | 表达量相对较低、糖基化过度 | 疫苗抗原、食品酶、基础研究 | 糖基化通路改造(ΔOCH1,ΔALG3)、强启动子(GAL1等) | [ |
| 哺乳动物细胞(如CHO, HEK293) | 能完成复杂翻译后修饰、产物最接近天然 | 成本高、周期长、培养复杂 | 治疗性抗体、复杂糖蛋白、病毒样颗粒 | 细胞系工程(GS/KO系统)、培养基优化、过程控制 | [ |
| 丝状真菌(如木霉、曲霉) | 强大的蛋白分泌能力、能进行某些复杂修饰 | 遗传操作相对复杂、背景分泌蛋白多 | 工业水解酶(纤维素酶、淀粉酶) | 强启动子、蛋白酶缺陷株、分泌信号优化 | [ |
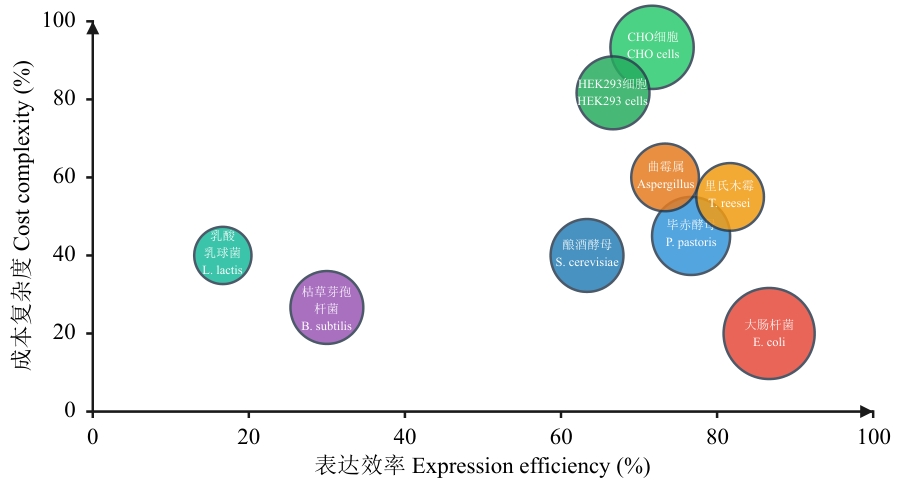
Fig. 2 Comparison of host systemsThe size of the bubbles indicates the frequency of use of the host system: The larger the bubble, the higher the frequency of use, and the smaller the bubble, the lower the frequency of use
名称 Model name | 类型 Type | 应用层面/目标Application level/Objective | 核心优势功能/原理 Key feature/Mechanism | 优势 Advantages | 局限性 Limitations | 参考文献References |
|---|---|---|---|---|---|---|
| CNN启动子模型 | 预测 | 宿主/转录调控 | 使用卷积神经网络分析启动子序列,预测其强度 | 实现对启动子活性的高精度(R²=0.79-0.84)预测和精细调控 | 依赖大量标注数据 | [ |
| EMOPEC | 预测 | 宿主/翻译起始 | 使用随机森林算法,定量揭示SD序列与蛋白表达水平的关系 | 可理性设计RBS以提升异源蛋白表达量 | 对真核系统适用性有限 | [ |
| UTR-LM | 预测 | 宿主/翻译效率 | 基于Transformer架构,通过5'UTR序列预测平均核糖体负载量(MRL) | 仅凭序列即可准确预测,实验验证可将表达量提升32.5% | 模型可解释性弱 | [ |
| MPEPE | 预测 | 密码子/协同效应 | 深度学习预测高表达倾向的氨基酸替换,再经保守性分析筛选位点 | 综合考虑氨基酸替换和密码子使用,揭示协同效应 | 优化流程相对复杂 | [ |
| MLD-NCS | 预测 | 密码子/翻译起始 | 结合LSTM与注意力机制,优化mRNA 5'端前30个密码子 | 有效避免翻译停滞,在枯草芽胞杆菌中表达量提升5.41倍 | 主要优化翻译起始阶段 | [ |
| MPB-EXP/MUT | 预测 | 氨基酸/可溶性设计 | 基于蛋白质语言模型,从序列学习高可溶性特征并设计突变 | 能有效改善多种蛋白的可溶性,实现从几乎不表达到可溶的跃迁 | 生成序列的可靠性需验证 | [ |
| C-terminal composition 分析 | 预测 | 氨基酸/表达丰度 | 统计分析C末端氨基酸组成(带电荷残基)与蛋白表达水平的关联 | 规则简单明确,易于应用(如在C端引入Lys/Arg) | 普适性有待考察 | [ |
| GMMA | 预测 | 氨基酸/突变组合 | 从多突变体数据中推断单点突变效应,并组合有利突变 | 能够发现突变间的叠加效应,实现表达量的数十至数百倍提升 | 依赖于大量的实验数据 | [ |
| ZymCTRL | 生成 | 宿主/全序列设计 | 条件语言模型,根据酶学性质定向生成酶序列及配套调控元件 | 实现“按需定制”的全序列人工设计,整合性强 | 长序列功能验证挑战大 | [ |
| DeepCodon | 生成 | 密码子/翻译优化 | 深度学习模型优化密码子使用频率,提高翻译效率 | 在多种宿主中均实现2-10倍的表达量提升,通用性强 | 对mRNA高级结构考虑不足 | [ |
| RFdiffusion | 生成 | 氨基酸/结构设计 | 通过迭代去噪过程,从随机噪声中生成具有目标结构的蛋白质骨架 | 能够从头设计复杂蛋白结构(如对称寡聚体),突破性强 | 计算资源需求极高 | [ |
| ProteinMPNN | 生成 | 氨基酸/结构设计 | 基于给定的蛋白质骨架结构,逆折叠生成兼容的氨基酸序列 | 序列回收率高,设计的序列可溶性和表达量显著提升(最高20倍) | 严重依赖输入结构的准确性 | [ |
Table 2 Summary of artificial intelligence models for optimizing soluble protein expression
名称 Model name | 类型 Type | 应用层面/目标Application level/Objective | 核心优势功能/原理 Key feature/Mechanism | 优势 Advantages | 局限性 Limitations | 参考文献References |
|---|---|---|---|---|---|---|
| CNN启动子模型 | 预测 | 宿主/转录调控 | 使用卷积神经网络分析启动子序列,预测其强度 | 实现对启动子活性的高精度(R²=0.79-0.84)预测和精细调控 | 依赖大量标注数据 | [ |
| EMOPEC | 预测 | 宿主/翻译起始 | 使用随机森林算法,定量揭示SD序列与蛋白表达水平的关系 | 可理性设计RBS以提升异源蛋白表达量 | 对真核系统适用性有限 | [ |
| UTR-LM | 预测 | 宿主/翻译效率 | 基于Transformer架构,通过5'UTR序列预测平均核糖体负载量(MRL) | 仅凭序列即可准确预测,实验验证可将表达量提升32.5% | 模型可解释性弱 | [ |
| MPEPE | 预测 | 密码子/协同效应 | 深度学习预测高表达倾向的氨基酸替换,再经保守性分析筛选位点 | 综合考虑氨基酸替换和密码子使用,揭示协同效应 | 优化流程相对复杂 | [ |
| MLD-NCS | 预测 | 密码子/翻译起始 | 结合LSTM与注意力机制,优化mRNA 5'端前30个密码子 | 有效避免翻译停滞,在枯草芽胞杆菌中表达量提升5.41倍 | 主要优化翻译起始阶段 | [ |
| MPB-EXP/MUT | 预测 | 氨基酸/可溶性设计 | 基于蛋白质语言模型,从序列学习高可溶性特征并设计突变 | 能有效改善多种蛋白的可溶性,实现从几乎不表达到可溶的跃迁 | 生成序列的可靠性需验证 | [ |
| C-terminal composition 分析 | 预测 | 氨基酸/表达丰度 | 统计分析C末端氨基酸组成(带电荷残基)与蛋白表达水平的关联 | 规则简单明确,易于应用(如在C端引入Lys/Arg) | 普适性有待考察 | [ |
| GMMA | 预测 | 氨基酸/突变组合 | 从多突变体数据中推断单点突变效应,并组合有利突变 | 能够发现突变间的叠加效应,实现表达量的数十至数百倍提升 | 依赖于大量的实验数据 | [ |
| ZymCTRL | 生成 | 宿主/全序列设计 | 条件语言模型,根据酶学性质定向生成酶序列及配套调控元件 | 实现“按需定制”的全序列人工设计,整合性强 | 长序列功能验证挑战大 | [ |
| DeepCodon | 生成 | 密码子/翻译优化 | 深度学习模型优化密码子使用频率,提高翻译效率 | 在多种宿主中均实现2-10倍的表达量提升,通用性强 | 对mRNA高级结构考虑不足 | [ |
| RFdiffusion | 生成 | 氨基酸/结构设计 | 通过迭代去噪过程,从随机噪声中生成具有目标结构的蛋白质骨架 | 能够从头设计复杂蛋白结构(如对称寡聚体),突破性强 | 计算资源需求极高 | [ |
| ProteinMPNN | 生成 | 氨基酸/结构设计 | 基于给定的蛋白质骨架结构,逆折叠生成兼容的氨基酸序列 | 序列回收率高,设计的序列可溶性和表达量显著提升(最高20倍) | 严重依赖输入结构的准确性 | [ |
| [1] | Buck CB, Trus BL. The papillomavirus virion: a machine built to hide molecular Achilles’ heels [M]//Viral Molecular Machines. Boston, MA: Springer US, 2011: 403-422. |
| [2] | Corbett KS, Edwards DK, Leist SR, et al. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness [J]. Nature, 2020, 586(7830): 567-571. |
| [3] | Kanaya S, Yamada Y, Kinouchi M, et al. codon usage and tRNA genes in eukaryotes: correlation of codon usage diversity with translation efficiency and with CG-dinucleotide usage as assessed by multivariate analysis [J]. J Mol Evol, 2001, 53(4): 290-298. |
| [4] | Musto H, Romero H, Zavala A. Translational selection is operative for synonymous codon usage in Clostridium perfringens and Clostridium acetobutylicum [J]. Microbiology, 2003, 149(4): 855-863. |
| [5] | Duret L, Galtier N. Biased gene conversion and the evolution of mammalian genomic landscapes [J]. Annu Rev Genom Hum Genet, 2009, 10: 285-311. |
| [6] | Lajoie MJ, Rovner AJ, Goodman DB, et al. Genomically recoded organisms expand biological functions [J]. Science, 2013, 342(6156): 357-360. |
| [7] | Martin RW, Des Soye BJ, Kwon YC, et al. Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids [J]. Nat Commun, 2018, 9: 1203. |
| [8] | Ding ZD, Guan FF, Xu GS, et al. MPEPE, a predictive approach to improve protein expression in E. coli based on deep learning [J]. Comput Struct Biotechnol J, 2022, 20: 1142-1153. |
| [9] | Yan ZL, Chu WR, Sheng YH, et al. Integrating deep learning and synthetic biology: a co-design approach for enhancing gene expression via N-terminal coding sequences [J]. ACS Synth Biol, 2024, 13(9): 2960-2968. |
| [10] | Watson JL, Juergens D, Bennett NR, et al. De novo design of protein structure and function with RFdiffusion [J]. Nature, 2023, 620(7976): 1089-1100. |
| [11] | Fu HG, Liang YB, Zhong XQ, et al. Codon optimization with deep learning to enhance protein expression [J]. Sci Rep, 2020, 10: 17617. |
| [12] | Cabrita LD, Gilis D, Robertson AL, et al. Enhancing the stability and solubility of TEV protease using in silico design [J]. Protein Sci, 2007, 16(11): 2360-2367. |
| [13] | Jiang SM, Li CH, Zhang WW, et al. Directed evolution and structural analysis of N-carbamoyl-D-amino acid amidohydrolase provide insights into recombinant protein solubility in Escherichia coli [J]. Biochem J, 2007, 402(3): 429-437. |
| [14] | Jung S, Park S. Improving the expression yield of Candida antarctica lipase B in Escherichia coli by mutagenesis [J]. Biotechnol Lett, 2008, 30(4): 717-722. |
| [15] | Jonet MA, Mahadi NM, Murad AMA, et al. Optimization of a heterologous signal peptide by site-directed mutagenesis for improved secretion of recombinant proteins in Escherichia coli [J]. Microb Physiol, 2012, 22(1): 48-58. |
| [16] | Ito Y, Ishigami M, Hashiba N, et al. Avoiding entry into intracellular protein degradation pathways by signal mutations increases protein secretion in Pichia pastoris [J]. Microb Biotechnol, 2022, 15(9): 2364-2378. |
| [17] | Skoczinski P, Volkenborn K, Fulton A, et al. Contribution of single amino acid and codon substitutions to the production and secretion of a lipase by Bacillus subtilis [J]. Microb Cell Fact, 2017, 16(1): 160. |
| [18] | Liu TY, Zhang YY, Li YJ, et al. Effective gene expression prediction and optimization from protein sequences [J]. Adv Sci, 2025, 12(8): 2407664. |
| [19] | Weber M, Burgos R, Yus E, et al. Impact of C‐terminal amino acid composition on protein expression in bacteria [J]. Mol Syst Biol, 2020, 16(5): MSB199208. |
| [20] | Norrild RK, Johansson KE, O'Shea C, et al. Increasing protein stability by inferring substitution effects from high-throughput experiments [J]. Cell Rep Meth, 2022, 2(11): 100333. |
| [21] | Sumida KH, Núñez-Franco R, Kalvet I, et al. Improving protein expression, stability, and function with ProteinMPNN [J]. J Am Chem Soc, 2024, 146(3): 2054-2061. |
| [22] | Studier FW. Use of bacteriophage T7 lysozyme to improve an inducible T7 expression system [J]. J Mol Biol, 1991, 219(1): 37-44. |
| [23] | İncir İ, Kaplan Ö. Escherichia coli as a versatile cell factory: Advances and challenges in recombinant protein production [J]. Protein Expr Purif, 2024, 219: 106463. |
| [24] | Makino T, Skretas G, Georgiou G. Strain engineering for improved expression of recombinant proteins in bacteria [J]. Microb Cell Fact, 2011, 10(1): 32. |
| [25] | Tegel H, Tourle S, Ottosson J, et al. Increased levels of recombinant human proteins with the Escherichia coli strain Rosetta(DE3) [J]. Protein Expr Purif, 2010, 69(2): 159-167. |
| [26] | Phillips TA, VanBogelen RA, Neidhardt FC. Lon gene product of Escherichia coli is a heat-shock protein [J]. J Bacteriol, 1984, 159(1): 283-287. |
| [27] | van Wijk DJS. Tuning Escherichia coli for membrane protein overexpression [J]. Proc Natl Acad Sci U S A, 2008, 105(38): 14371-14376. |
| [28] | Schlegel S, Rujas E, Ytterberg AJ, et al. Optimizing heterologous protein production in the periplasm of E. coli by regulating gene expression levels [J]. Microb Cell Fact, 2013, 12(1): 24. |
| [29] | Turner P, Holst O, Karlsson EN. Optimized expression of soluble cyclomaltodextrinase of thermophilic origin in Escherichia coli by using a soluble fusion-tag and by tuning of inducer concentration [J]. Protein Expr Purif, 2005, 39(1): 54-60. |
| [30] | Singh A, Upadhyay V, Upadhyay AK, et al. Protein recovery from inclusion bodies of Escherichia coli using mild solubilization process [J]. Microb Cell Fact, 2015, 14(1): 41. |
| [31] | Gu Y, Xu XH, Wu YK, et al. Advances and prospects of Bacillus subtilis cellular factories: From rational design to industrial applications [J]. Metab Eng, 2018, 50: 109-121. |
| [32] | Chen YZ, Li MM, Yan MC, et al. Bacillus subtilis: current and future modification strategies as a protein secreting factory [J]. World J Microbiol Biotechnol, 2024, 40(6): 195. |
| [33] | Rojas Contreras JA, Pedraza-Reyes M, Ordoñez LG, et al. Replicative and integrative plasmids for production of human interferon gamma in Bacillus subtilis [J]. Plasmid, 2010, 64(3): 170-176. |
| [34] | De S, Van D. Control of prokaryotic translational initiation by mRNA secondary structure [J]. Prog Nucleic Acid Res Mol Biol, 1990, 38: 1-35. |
| [35] | Looser V, Bruhlmann B, Bumbak F, et al. Cultivation strategies to enhance productivity of Pichia pastoris: a review [J]. Biotechnol Adv, 2015, 33(6): 1177-1193. |
| [36] | Mayson BE, Kilburn DG, Zamost BL, et al. Effects of methanol concentration on expression levels of recombinant protein in fed-batch cultures of Pichia methanolica [J]. Biotechnol Bioeng, 2003, 81(3): 291-298. |
| [37] | Çelik E, Çalık P, Oliver SG. Metabolic flux analysis for recombinant protein production by Pichia pastoris using dual carbon sources: Effects of methanol feeding rate [J]. Biotechnol Bioeng, 2010, 105(2): 317-329. |
| [38] | Arias CAD, de Araujo Viana Marques D, Malpiedi LP, et al. Cultivation of Pichia pastoris carrying the scFv anti LDL (-) antibody fragment. Effect of preculture carbon source [J]. Braz J Microbiol, 2017, 48(3): 419-426. |
| [39] | Azadi S, Mahboubi A, Naghdi N, et al. Evaluation of sorbitol-methanol co-feeding strategy on production of recombinant human growth hormone in Pichia pastoris [J]. Iran J Pharm Res, 2017, 16(4): 1555-1564. |
| [40] | Jahic M, Wallberg F, Bollok M, et al. Temperature limited fed-batch technique for control of proteolysis in Pichia pastoris bioreactor cultures [J]. Microb Cell Fact, 2003, 2(1): 6. |
| [41] | Tang HT, Wang SH, Wang JJ, et al. N-hypermannose glycosylation disruption enhances recombinant protein production by regulating secretory pathway and cell wall integrity in Saccharomyces cerevisiae [J]. Sci Rep, 2016, 6: 25654. |
| [42] | 贺铁凡, 徐沙, 张阁元, 等. 重构酿酒酵母N-糖基化途径生产人源化糖蛋白 [J]. 微生物学报, 2014, 54(5): 509-516. |
| He TF, Xu S, Zhang GY, et al. Reconstruction of N-glycosylation pathway for producing human glycoproteins in Saccharomyces cerevisiae [J]. Acta Microbiol Sin, 2014, 54(5): 509-516. | |
| [43] | Liu DJ, Garrigues S, de Vries RP. Heterologous protein production in filamentous fungi [J]. Appl Microbiol Biotechnol, 2023, 107(16): 5019-5033. |
| [44] | Ha TK, Kim D, Kim CL, et al. Factors affecting the quality of therapeutic proteins in recombinant Chinese Hamster ovary cell culture [J]. Biotechnol Adv, 2022, 54: 107831. |
| [45] | Fu YS, Han ZM, Cheng WT, et al. Improvement strategies for transient gene expression in mammalian cells [J]. Appl Microbiol Biotechnol, 2024, 108(1): 480. |
| [46] | Kotopka BJ, Smolke CD. Model-driven generation of artificial yeast promoters [J]. Nat Commun, 2020, 11: 2113. |
| [47] | Bonde MT, Pedersen M, Klausen MS, et al. Predictable tuning of protein expression in bacteria [J]. Nat Meth, 2016, 13(3): 233-236. |
| [48] | Chu YY, Yu D, Li YP, et al. A 5' UTR language model for decoding untranslated regions of mRNA and function predictions [J]. Nat Mach Intell, 2024, 6(4): 449-460. |
| [49] | Zrimec J, Fu XZ, Muhammad AS, et al. Controlling gene expression with deep generative design of regulatory DNA [J]. Nat Commun, 2022, 13: 5099. |
| [50] | Munsamy G, Lindner S, Lorenz P, et al. ZymCTRL: a conditional language model for the controllable generation of artificial enzymes [J]. NeurIPS, 2022. |
| [51] | Yang KK, Wu Z, Arnold FH. Machine-learning-guided directed evolution for protein engineering [J]. Nat Meth, 2019, 16(8): 687-694. |
| [52] | Quax TEF, Claassens NJ, Söll D, et al. Codon bias as a means to fine-tune gene expression [J]. Mol Cell, 2015, 59(2): 149-161. |
| [53] | Rocklin GJ, Chidyausiku TM, Goreshnik I, et al. Global analysis of protein folding using massively parallel design, synthesis, and testing [J]. Science, 2017, 357(6347): 168-175. |
| [1] | TIAN Chun-yan, LI Xu-juan, LI Chun-jia, MAO Jun, LIU Xin-long. Genome-wide Analysis of Codon Usage Bias in Saccharum Species and Its Phylogenetically Related Species Erianthus fulvus [J]. Biotechnology Bulletin, 2024, 40(3): 202-214. |
| [2] | YI Hua-Wei, TANG Xiao-Feng. Research Progress on the Prediction of Protein Stability Based on Amino Acid Sequence and Simulated Structure [J]. Biotechnology Bulletin, 2017, 33(4): 83-89. |
| [3] | Wang Shishan, Chen Yanke, Yang Jun. The Gene Sequence Optimization of Membrane Protein in Prokaryotic Expression System [J]. Biotechnology Bulletin, 2015, 31(12): 50-55. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||