








Biotechnology Bulletin ›› 2023, Vol. 39 ›› Issue (6): 102-108.doi: 10.13560/j.cnki.biotech.bull.1985.2022-1058
Previous Articles Next Articles
CHEN Yong1, LI Ya-xin2, WANG Ya-xuan2, LIANG Lu-jie2, FENG Si-yuan2, Tian Guo-bao2,3( )
)
Received:2022-08-25
Online:2023-06-26
Published:2023-07-07
Contact:
Tian Guo-bao
E-mail:tiangb@mail.sysu.edu.cn
CHEN Yong, LI Ya-xin, WANG Ya-xuan, LIANG Lu-jie, FENG Si-yuan, Tian Guo-bao. Research Progress in the Molecular Mechanism of MCR-1 Mediated Polymyxin Resistance[J]. Biotechnology Bulletin, 2023, 39(6): 102-108.
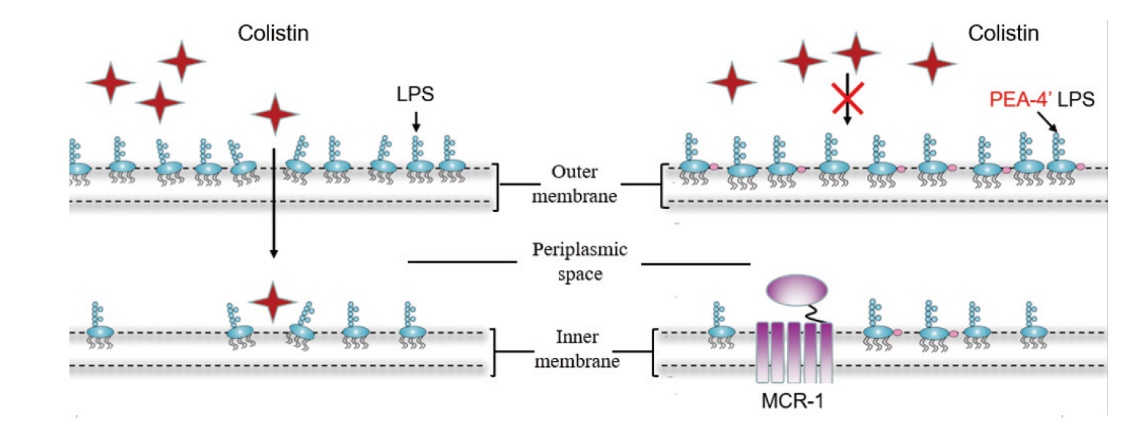
Fig. 1 Resistance mechanism of MCR-1-mediated bacterium producing polymyxin MCR-1 transfers the positively charged phosphoethanolamine(PEA)group on the phosphatidylethanolamine(PE)molecule to the phospholipid A on the bacterial cell outer membrane to generate PEA-4' Lipid A, which increases the positive charge on the bacterial cell membrane, thus reducing the affinity of polymyxin and lipopolysaccharide, and ultimately leads to the generation of low-level drug resistance phenotype
| [1] |
Talbot GH, Bradley J, Edwards JE Jr, et al. Bad bugs need drugs: an update on the development pipeline from the antimicrobial availability task force of the infectious diseases society of America[J]. Clin Infect Dis, 2006, 42(5): 657-668.
doi: 10.1086/499819 pmid: 16447111 |
| [2] |
Kaye KS, Pogue JM, Tran TB, et al. Agents of last resort: polymyxin resistance[J]. Infect Dis Clin North Am, 2016, 30(2): 391-414.
doi: 10.1016/j.idc.2016.02.005 URL |
| [3] |
Falagas ME, Kasiakou SK. Colistin: the revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections[J]. Clin Infect Dis, 2005, 40(9): 1333-1341.
doi: 10.1086/429323 pmid: 15825037 |
| [4] |
Falagas ME, Rafailidis PI, Matthaiou DK. Resistance to polymyxins: mechanisms, frequency and treatment options[J]. Drug Resist Updat, 2010, 13(4/5): 132-138.
doi: 10.1016/j.drup.2010.05.002 URL |
| [5] |
Son SJ, Huang RJ, Squire CJ, et al. MCR-1: a promising target for structure-based design of inhibitors to tackle polymyxin resistance[J]. Drug Discov Today, 2019, 24(1): 206-216.
doi: S1359-6446(18)30025-4 pmid: 30036574 |
| [6] |
Li J, Nation RL, Milne RW, et al. Evaluation of colistin as an agent against multi-resistant Gram-negative bacteria[J]. Int J Antimicrob Agents, 2005, 25(1): 11-25.
doi: 10.1016/j.ijantimicag.2004.10.001 URL |
| [7] |
Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study[J]. Lancet Infect Dis, 2016, 16(2): 161-168.
doi: 10.1016/S1473-3099(15)00424-7 URL |
| [8] |
Shen ZQ, Wang Y, Shen YB, et al. Early emergence of mcr-1 in Escherichia coli from food-producing animals[J]. Lancet Infect Dis, 2016, 16(3): 293.
doi: 10.1016/S1473-3099(16)00061-X URL |
| [9] |
Haenni M, Poirel L, Kieffer N, et al. Co-occurrence of extended spectrum β lactamase and MCR-1 encoding genes on plasmids[J]. Lancet Infect Dis, 2016, 16(3): 281-282.
doi: 10.1016/S1473-3099(16)00007-4 pmid: 26774244 |
| [10] |
Pham Thanh D, Thanh Tuyen H, Nguyen Thi Nguyen TT, et al. Inducible colistin resistance via a disrupted plasmid-borne mcr-1 gene in a 2008 Vietnamese Shigella sonnei isolate[J]. J Antimicrob Chemother, 2016, 71(8): 2314-2317.
doi: 10.1093/jac/dkw173 pmid: 27246235 |
| [11] |
Jorgensen SB, Soraas A, Arnesen LS, et al. First environmental sample containing plasmid-mediated colistin-resistant ESBL-producing Escherichia coli detected in Norway[J]. APMIS, 2017, 125(9): 822-825.
doi: 10.1111/apm.2017.125.issue-9 URL |
| [12] |
Ling ZR, Yin WJ, Shen ZQ, et al. Epidemiology of mobile colistin resistance genes mcr-1 to mcr-9[J]. J Antimicrob Chemother, 2020, 75(11): 3087-3095.
doi: 10.1093/jac/dkaa205 pmid: 32514524 |
| [13] |
Nang SC, Li J, Velkov T. The rise and spread of mcr plasmid-mediated polymyxin resistance[J]. Crit Rev Microbiol, 2019, 45(2): 131-161.
doi: 10.1080/1040841X.2018.1492902 pmid: 31122100 |
| [14] |
Li RC, Xie MM, Zhang JF, et al. Genetic characterization of mcr-1-bearing plasmids to depict molecular mechanisms underlying dissemination of the colistin resistance determinant[J]. J Antimicrob Chemother, 2017, 72(2): 393-401.
doi: 10.1093/jac/dkw411 pmid: 28073961 |
| [15] |
He YZ, Li XP, Miao YY, et al. The ISApl12 dimer circular intermediate participates in mcr-1 transposition[J]. Front Microbiol, 2019, 10: 15.
doi: 10.3389/fmicb.2019.00015 URL |
| [16] |
Sun J, Fang LX, Wu ZW, et al. Genetic analysis of the IncX4 plasmids: implications for a unique pattern in the mcr-1 acquisition[J]. Sci Rep, 2017, 7(1): 424.
doi: 10.1038/s41598-017-00095-x |
| [17] |
Li RC, Zhang P, Yang XR, et al. Identification of a novel hybrid plasmid coproducing MCR-1 and MCR-3 variant from an Escherichia coli strain[J]. J Antimicrob Chemother, 2019, 74(6): 1517-1520.
doi: 10.1093/jac/dkz058 URL |
| [18] |
Matamoros S, van Hattem JM, Arcilla MS, et al. Global phylogenetic analysis of Escherichia coli and plasmids carrying the mcr-1 gene indicates bacterial diversity but plasmid restriction[J]. Sci Rep, 2017, 7(1): 15364.
doi: 10.1038/s41598-017-15539-7 |
| [19] |
Wu RJ, Yi LX, Yu LF, et al. Fitness advantage of mcr-1-bearing IncI2 and IncX4 plasmids in vitro[J]. Front Microbiol, 2018, 9: 331.
doi: 10.3389/fmicb.2018.00331 URL |
| [20] | Singh S, Pathak A, Kumar A, et al. Emergence of chromosome-borne colistin resistance gene mcr-1 in clinical isolates of Klebsiella pneumoniae from India[J]. Antimicrob Agents Chemother, 2018, 62(2): e01885-e01817. |
| [21] |
Lu XY, Xiao X, Liu Y, et al. Chromosome-mediated mcr-1 in Escherichia coli strain L73 from a goose[J]. Int J Antimicrob Agents, 2019, 54(1): 99-101.
doi: 10.1016/j.ijantimicag.2019.03.003 URL |
| [22] | Zhou HW, Zhang T, Ma JH, et al. Occurrence of plasmid- and chromosome-carried mcr-1 in waterborne Enterobacteriaceae in China[J]. Antimicrob Agents Chemother, 2017, 61(8): e00017-e00017. |
| [23] | Yamaguchi T, Kawahara R, Hamamoto K, et al. High prevalence of colistin-resistant Escherichia coli with chromosomally carried mcr-1 in healthy residents in Vietnam[J]. mSphere, 2020, 5(2): e00117-e00120. |
| [24] |
Wang Y, Xu CY, Zhang R, et al. Changes in colistin resistance and mcr-1 abundance in Escherichia coli of animal and human origins following the ban of colistin-positive additives in China: an epidemiological comparative study[J]. Lancet Infect Dis, 2020, 20(10): 1161-1171.
doi: 10.1016/S1473-3099(20)30149-3 URL |
| [25] |
Liu YY, Zhou QL, He WY, et al. Mcr-1 and plasmid prevalence in Escherichia coli from livestock[J]. Lancet Infect Dis, 2020, 20(10): 1126.
doi: 10.1016/S1473-3099(20)30697-6 URL |
| [26] |
Shen C, Zhong LL, Yang YQ, et al. Dynamics of mcr-1 prevalence and mcr-1-positive Escherichia coli after the cessation of colistin use as a feed additive for animals in China: a prospective cross-sectional and whole genome sequencing-based molecular epidemiological study[J]. Lancet Microbe, 2020, 1(1): e34-e43.
doi: 10.1016/S2666-5247(20)30005-7 URL |
| [27] |
Shen C, Zhong LL, Zhong ZJ, et al. Prevalence of mcr-1 in colonized inpatients, China, 2011-2019[J]. Emerg Infect Dis, 2021, 27(9): 2502-2504.
doi: 10.3201/eid2709.203642 URL |
| [28] |
Usui M, Nozawa Y, Fukuda A, et al. Decreased colistin resistance and mcr-1 prevalence in pig-derived Escherichia coli in Japan after banning colistin as a feed additive[J]. J Glob Antimicrob Resist, 2021, 24: 383-386.
doi: 10.1016/j.jgar.2021.01.016 URL |
| [29] |
Fournier C, Aires-de-Sousa M, Nordmann P, et al. Occurrence of CTX-M-15- and MCR-1-producing Enterobacterales in pigs in Portugal: evidence of direct links with antibiotic selective pressure[J]. Int J Antimicrob Agents, 2020, 55(2): 105802.
doi: 10.1016/j.ijantimicag.2019.09.006 URL |
| [30] |
Sabnis A, Hagart KL, Klöckner A, et al. Colistin kills bacteria by targeting lipopolysaccharide in the cytoplasmic membrane[J]. eLife, 2021, 10: e65836.
doi: 10.7554/eLife.65836 URL |
| [31] |
MacNair CR, Stokes JM, Carfrae LA, et al. Overcoming mcr-1 mediated colistin resistance with colistin in combination with other antibiotics[J]. Nat Commun, 2018, 9(1): 458.
doi: 10.1038/s41467-018-02875-z pmid: 29386620 |
| [32] |
Bialvaei AZ, Samadi Kafil H. Colistin, mechanisms and prevalence of resistance[J]. Curr Med Res Opin, 2015, 31(4): 707-721.
doi: 10.1185/03007995.2015.1018989 pmid: 25697677 |
| [33] |
Velkov T, Roberts KD, Nation RL, et al. Pharmacology of polymyxins: new insights into an ‘old’ class of antibiotics[J]. Future Microbiol, 2013, 8(6): 711-724.
doi: 10.2217/fmb.13.39 pmid: 23701329 |
| [34] |
Chatzidimitriou M, Kavvada A, Kavvadas D, et al. Mcr genes conferring colistin resistance in enterobacterales; a five year overview[J]. Acta Med Acad, 2021, 50(3): 365-371.
doi: 10.5644/ama2006-124.355 pmid: 35164512 |
| [35] |
Hinchliffe P, Yang QE, Portal E, et al. Insights into the mechanistic basis of plasmid-mediated colistin resistance from crystal structures of the catalytic domain of MCR-1[J]. Sci Rep, 2017, 7(1): 39392.
doi: 10.1038/srep39392 |
| [36] |
Coates K, Walsh TR, Spencer J, et al. 1.12 Å resolution crystal structure of the catalytic domain of the plasmid-mediated colistin resistance determinant MCR-2[J]. Acta Crystallogr Sect F, 2017, 73(8): 443-449.
doi: 10.1107/S2053230X17009669 URL |
| [37] | Sun J, Xu YC, Gao RS, et al. Deciphering MCR-2 colistin resistance[J]. mBio, 2017, 8(3): e00625-e00617. |
| [38] |
Xu YC, Lin JX, Cui T, et al. Mechanistic insights into transferable polymyxin resistance among gut bacteria[J]. J Biol Chem, 2018, 293(12): 4350-4365.
doi: 10.1074/jbc.RA117.000924 pmid: 29462787 |
| [39] |
Ma GX, Zhu YF, Yu ZC, et al. High resolution crystal structure of the catalytic domain of MCR-1[J]. Sci Rep, 2016, 6: 39540.
doi: 10.1038/srep39540 pmid: 28000749 |
| [40] |
Gao RS, Hu YF, Li ZC, et al. Dissemination and mechanism for the MCR-1 colistin resistance[J]. PLoS Pathog, 2016, 12(11): e1005957.
doi: 10.1371/journal.ppat.1005957 URL |
| [41] |
Zhang XF, Doi Y, Huang X, et al. Possible transmission of mcr-1-harboring Escherichia coli between companion animals and human[J]. Emerg Infect Dis, 2016, 22(9): 1679-1681.
doi: 10.3201/eid2209.160464 URL |
| [42] | Xu YC, Wei WH, Lei S, et al. An evolutionarily conserved mechanism for intrinsic and transferable polymyxin resistance[J]. mBio, 2018, 9(2): e02317-e02317. |
| [43] |
Zhang HM, Srinivas S, Xu YC, et al. Genetic and biochemical mechanisms for bacterial lipid A modifiers associated with polymyxin resistance[J]. Trends Biochem Sci, 2019, 44(11): 973-988.
doi: S0968-0004(19)30135-5 pmid: 31279652 |
| [44] |
Sun J, Zhang HM, Liu YH, et al. Towards understanding MCR-like colistin resistance[J]. Trends Microbiol, 2018, 26(9): 794-808.
doi: S0966-842X(18)30042-8 pmid: 29525421 |
| [45] |
Xu YC, Chen HY, Zhang HM, et al. The MCR-3 inside linker appears as a facilitator of colistin resistance[J]. Cell Rep, 2021, 35(7): 109135.
doi: 10.1016/j.celrep.2021.109135 URL |
| [46] | Powers MJ, Trent MS. Phospholipid retention in the absence of asymmetry strengthens the outer membrane permeability barrier to last-resort antibiotics[J]. Proc Natl Acad Sci USA, 2018, 115(36): E8518-E8527. |
| [47] |
Wang XY, Quinn PJ, Yan AX. Kdo2-lipid A: structural diversity and impact on immunopharmacology[J]. Biol Rev, 2015, 90(2): 408-427.
doi: 10.1111/brv.2015.90.issue-2 URL |
| [48] |
Mandela E, Stubenrauch CJ, Ryoo D, et al. Adaptation of the periplasm to maintain spatial constraints essential for cell envelope processes and cell viability[J]. eLife, 2022, 11: e73516.
doi: 10.7554/eLife.73516 URL |
| [49] | Sutterlin HA, Shi HD, May KL, et al. Disruption of lipid homeostasis in the Gram-negative cell envelope activates a novel cell death pathway[J]. Proc Natl Acad Sci USA, 2016, 113(11): E1565-E1574. |
| [50] |
Yang J, Wang HH, Lu YY, et al. A ProQ/FinO family protein involved in plasmid copy number control favours fitness of bacteria carrying mcr-1-bearing IncI2 plasmids[J]. Nucleic Acids Res, 2021, 49(7): 3981-3996.
doi: 10.1093/nar/gkab149 pmid: 33721023 |
| [51] |
Yang QE, Li M, Spiller OB, et al. Balancing mcr-1 expression and bacterial survival is a delicate equilibrium between essential cellular defence mechanisms[J]. Nat Commun, 2017, 8(1): 2054.
doi: 10.1038/s41467-017-02149-0 |
| [52] |
Yi LX, Durand R, Grenier F, et al. PixR, a novel activator of conjugative transfer of IncX4 resistance plasmids, mitigates the fitness cost of mcr-1 carriage in Escherichia coli[J]. mBio, 2022, 13(1): e0320921.
doi: 10.1128/mbio.03209-21 URL |
| [53] |
Feng SY, Liang WF, Li JC, et al. MCR-1-dependent lipid remodelling compromises the viability of Gram-negative bacteria[J]. Emerg Microbes Infect, 2022, 11(1): 1236-1249.
doi: 10.1080/22221751.2022.2065934 URL |
| [1] | LI Hai-li, LANG Li-min, ZHANG Qing-xian, YOU Yi, ZHU Wen-hao, WANG Zhi-fang, ZHANG Li-xian, WANG Ke-ling. Identification and Drug Resistance of Escherichia coli Simultaneously Producing Carbapenemase NDM-1 and NDM-5 [J]. Biotechnology Bulletin, 2022, 38(9): 106-115. |
| [2] | LIU Cheng-cheng, HU Xiao-fang, FENG You-jun. Antimicrobial Resistance:Biochemical Mechanisms and Countermeasures [J]. Biotechnology Bulletin, 2022, 38(9): 4-16. |
| [3] | ZHAO Yan-kun, LIU Hui-min, MENG Lu, WANG Cheng, WANG Jia-qi, ZHENG Nan. Research Progress in Heteroresistance of Escherichia coli [J]. Biotechnology Bulletin, 2022, 38(9): 59-71. |
| [4] | LI Liu, MU Ying-chun, LIU Lu, ZHANG Hong-yu, XU Jin-hua, YANG Zhen, QIAO Lu, SONG Jin-long. Research Progress on Contamination Control of Fluoroquinolone Antibiotics and Drug Resistance Genes [J]. Biotechnology Bulletin, 2022, 38(9): 84-95. |
| [5] | LIU Xiao-li, TONG Zhen-yi, ZHAO Liang, YIN Li, LIU Chen-guang. Research Progress in Non-antibiotic Active Substances Against Helicobacter pylori [J]. Biotechnology Bulletin, 2022, 38(9): 96-105. |
| [6] | CHEN Fu-nuan, HUANG Yu, CAI Jia, WANG Zhong-liang, JIAN Ji-chang, WANG Bei. Structure of ABC Transporter and Research Progress of It in Bacterial Pathogenicity [J]. Biotechnology Bulletin, 2022, 38(6): 43-52. |
| [7] | ZHU Hao, ZHANG Yan-wei, LIU Run, LIANG Yan, YANG Yi, XU Tian-le, YANG Zhang-ping. Research Progress in Antibiotic Adjuvant and Antibiotics in Antibacterial Aspects [J]. Biotechnology Bulletin, 2022, 38(6): 66-73. |
| [8] | TIAN Lu, WU Mi, GOU Jing-xuan, GONG Guo-li. Research and Application Progress of Bacteriocin [J]. Biotechnology Bulletin, 2021, 37(4): 224-233. |
| [9] | MEI Fen, LI Rui-wei, ZHANG Juan-juan, ZUO Rong-xia, ZOU Yun-lian, SHEN Tao, SA Ya-lian. Construction of Plasmids for Knocking out ALOX5 Gene Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2020, 36(3): 110-114. |
| [10] | FENG Gao, ZHANG Yu-chen , GOU Min, CHEN Ya-ting. Response of Butyrate-oxidizing Microbial Community to the Co-effects of Antibiotics and Activated Carbon [J]. Biotechnology Bulletin, 2019, 35(8): 64-76. |
| [11] | LI Di-yin, HE Yong-xing, HAN Jian-ting, LI Kun, WANG Zhi-ping, LI Miao-hui. Functional Identification of RstA in Pseudomonas fluorescens Strain 2P24 [J]. Biotechnology Bulletin, 2019, 35(6): 107-113. |
| [12] | DOU Peng-peng, WANG Li, ZHANG Hua, ZHENG Yao. Molecular Identification and Drug Resistance Analysis of Plesiomonas shigellode Isolated from Fish [J]. Biotechnology Bulletin, 2019, 35(11): 118-123. |
| [13] | ZHAO Yan-kun, LIU Hui-min, WANG Shuai, CAI Jian-xing, WANG Cheng, CHEN He. Research Progress on Drug Resistance of Staphylococcus aureus in Bovine Mastitis [J]. Biotechnology Bulletin, 2018, 34(10): 18-25. |
| [14] | Li Wenbin, Li Zengbo, Liu Xianjun. Research on Biological Activity of Antibiotic Analogues in SC-04 Culture [J]. Biotechnology Bulletin, 2013, 0(5): 190-193. |
| [15] | Zhang Yuxia, Liang Qiong, Yong Guoxin. Research of Reversal Effect on K562/A02 Cell Line with a Novel Marine Bioactive Substances [J]. Biotechnology Bulletin, 2013, 0(2): 177-183. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||