








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (7): 108-116.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0174
Previous Articles Next Articles
WU Shu-ning1,2( ), SU Yong-ping1,2, LI Dong-xue3, BAI Ying-guo3, LIU Bo2(
), SU Yong-ping1,2, LI Dong-xue3, BAI Ying-guo3, LIU Bo2( ), ZHANG Zhi-wei1(
), ZHANG Zhi-wei1( )
)
Received:2024-02-21
Online:2024-07-26
Published:2024-07-30
Contact:
LIU Bo, ZHANG Zhi-wei
E-mail:wsn06012023@163.com;lfb2500@163.com;zhiweizhang2012@163.com
WU Shu-ning, SU Yong-ping, LI Dong-xue, BAI Ying-guo, LIU Bo, ZHANG Zhi-wei. Design and Application of a Cumate-inducible Promoter for Corynebacterium glutamicum[J]. Biotechnology Bulletin, 2024, 40(7): 108-116.
| 引物名称 Primer name | 引物序列 Primer sequence(5' -3') | 引物用途 Primer usage |
|---|---|---|
| cymR-F | CCAGGGTGGTTTTTCTTTCTATCGCTTGAACTTAGCGTAGC | 扩增CymR |
| cymR-R | AGAGTCAATTCAGGGTGGTGAATATGAGCCCGAAGCGCCGCAC | |
| LacIq-F | ATTCACCACCCTGAATTGACTCT | 扩增PlacIq |
| LacIq-R | GGATCAGCTTGCAATTCGCGCGCGAAGGCGAAGCGGCATTTACG | |
| Mazf-F | CACGGCGAAAGGATACTCATGGTAAGCCGATACGTACCCG | 扩增mazf |
| Mazf-R | GTCGACCTGCAGGCATGCCTACCCAATCAGTACGTTAATTTTGG | |
| gfp-F gfp-R | GCATGCCTGCAGGTCGACATGGTGAGCAAGGGCGAGGA ACCCGGGGATCCTCTAGATTACTTGTACAGCTCGTCCATG | 扩增gfp |
| cymR/H10CuO-F cymR/H10CuO-R | ATGGGTCTTGTTGTTGGCAGACCGTATCCAAAGCATCCG AGGAAGAGTGGTTTTGTGCTCATGAGTATCCTTTCGCCGTGCT | 扩增cymR/H10CuO |
| recET-F | ATGAGCACAAAACCACTCTTCCT | 扩增recET |
| recET-R | CTTCCTGGCATCTTCCAGCAAAAAAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAATTATTCCTCTGAATTATCGATTACACTG | |
| H10CuO/cpf1-F | GTTCGCGGAATCATGACCGCTCAACCCTTACCGGTCG | PH10-CuO扩增 |
| H10CuO/cpf1-R | TCACAAACTCTTGGTAGATGGACATGAGTATCCTTTCGCCGTGCTC | |
| LysE-LF | TCAGTGGAACGAAAACTCACTGCGCGAGCAAGGAGAGTAC | 扩增lysE左同源臂 |
| LysE-LR | CGCCATGACAAACAAGGTGG | |
| LysE-RF | GTGGCACCGAGTCGGTGCTTTTTTTGAGGTAAGCGATGCCACCCCA | 扩增lysE右同源臂 |
| LysE-RR | TCTCATCCGCCAAAACAGCC GTTATGGTTTGCCGTCATGGCAGCATCTACAACAGTAGAAATTCGGATCCAT- TATACCTAGGACTGAGCTAGCTGTCAACCAAGAAGCTACCTCGTTGAACA | |
| asd/H10CuO-F | CCACCTTGTTTGTCATGGCGGCTCAACCCTTACCGGTCGGCTCTAAGCCGGCGGCGTATGGTAAGCTCTGTTATGTATAGTCCGAGCACGGCGAAAGGATACTC ATGACCACCATCGCAGTTGTT | 扩增asd |
| asd/H10CuO-R | AAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAATTACTTAACCAGCAGCTCAGCG |
Table 1 Primers used in this study
| 引物名称 Primer name | 引物序列 Primer sequence(5' -3') | 引物用途 Primer usage |
|---|---|---|
| cymR-F | CCAGGGTGGTTTTTCTTTCTATCGCTTGAACTTAGCGTAGC | 扩增CymR |
| cymR-R | AGAGTCAATTCAGGGTGGTGAATATGAGCCCGAAGCGCCGCAC | |
| LacIq-F | ATTCACCACCCTGAATTGACTCT | 扩增PlacIq |
| LacIq-R | GGATCAGCTTGCAATTCGCGCGCGAAGGCGAAGCGGCATTTACG | |
| Mazf-F | CACGGCGAAAGGATACTCATGGTAAGCCGATACGTACCCG | 扩增mazf |
| Mazf-R | GTCGACCTGCAGGCATGCCTACCCAATCAGTACGTTAATTTTGG | |
| gfp-F gfp-R | GCATGCCTGCAGGTCGACATGGTGAGCAAGGGCGAGGA ACCCGGGGATCCTCTAGATTACTTGTACAGCTCGTCCATG | 扩增gfp |
| cymR/H10CuO-F cymR/H10CuO-R | ATGGGTCTTGTTGTTGGCAGACCGTATCCAAAGCATCCG AGGAAGAGTGGTTTTGTGCTCATGAGTATCCTTTCGCCGTGCT | 扩增cymR/H10CuO |
| recET-F | ATGAGCACAAAACCACTCTTCCT | 扩增recET |
| recET-R | CTTCCTGGCATCTTCCAGCAAAAAAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAATTATTCCTCTGAATTATCGATTACACTG | |
| H10CuO/cpf1-F | GTTCGCGGAATCATGACCGCTCAACCCTTACCGGTCG | PH10-CuO扩增 |
| H10CuO/cpf1-R | TCACAAACTCTTGGTAGATGGACATGAGTATCCTTTCGCCGTGCTC | |
| LysE-LF | TCAGTGGAACGAAAACTCACTGCGCGAGCAAGGAGAGTAC | 扩增lysE左同源臂 |
| LysE-LR | CGCCATGACAAACAAGGTGG | |
| LysE-RF | GTGGCACCGAGTCGGTGCTTTTTTTGAGGTAAGCGATGCCACCCCA | 扩增lysE右同源臂 |
| LysE-RR | TCTCATCCGCCAAAACAGCC GTTATGGTTTGCCGTCATGGCAGCATCTACAACAGTAGAAATTCGGATCCAT- TATACCTAGGACTGAGCTAGCTGTCAACCAAGAAGCTACCTCGTTGAACA | |
| asd/H10CuO-F | CCACCTTGTTTGTCATGGCGGCTCAACCCTTACCGGTCGGCTCTAAGCCGGCGGCGTATGGTAAGCTCTGTTATGTATAGTCCGAGCACGGCGAAAGGATACTC ATGACCACCATCGCAGTTGTT | 扩增asd |
| asd/H10CuO-R | AAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAATTACTTAACCAGCAGCTCAGCG |
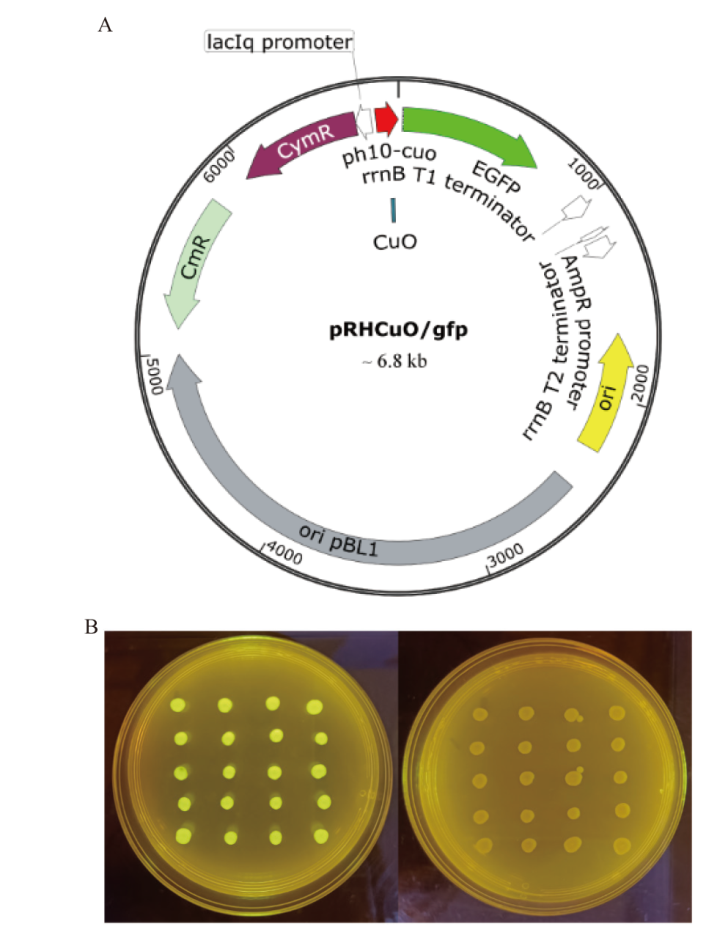
Fig. 3 Characterization of PH10-CuO A: Plasmid pRHCuO/gfp. B: Profiles of positive transformants harboring pRHCuO/gfp on different media (Left: chloramphenicol-contained LB medium with 4-isopropylbenzoic acid; right: chloramphenicol-contained LB medium)
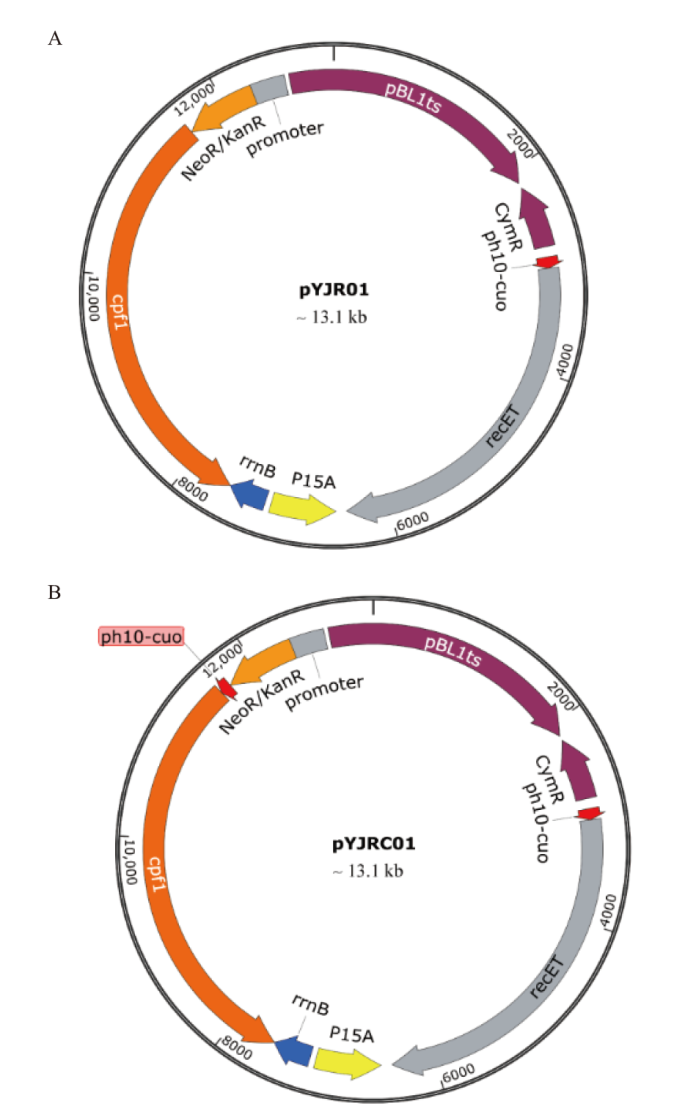
Fig. 5 Plasmids via gene editing A: PH10-CuO regulating the plasmid pYJR01 expressed with recET. B: PH10-CuO regulating the plasmid pYJR01 expressed with recET and cas12a
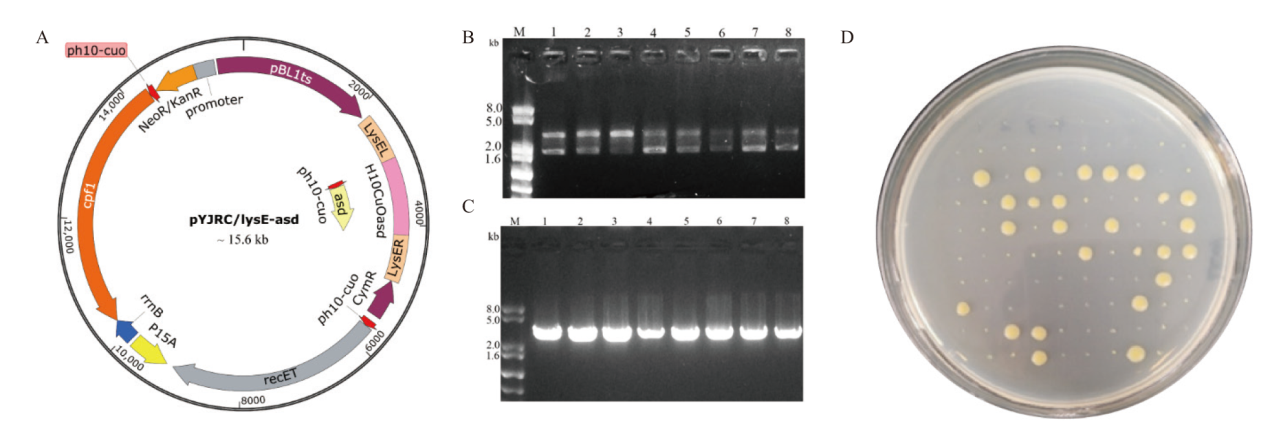
Fig. 6 Gene editing and plasmid elimination A: Plasmid pYJRC/lysE-asd for deleting lysE and overexpressing asd. B: Transformants verification(Lane M: DNA marker; lane 1-8: PCR products with the chromosomal DNA of transformants on solid medium BHISG containing 25 μg/mL kanamycin and 12.5 μg/mL 4-isopropylbenzoic acid as templates. C: Transformants verification(Lane M: DNA marker; lane 1-8: PCR products with the chromosomal DNA of transformants cultured in liquid medium BHISG containing 25 μg/mL kanamycin and 12.5 μg/mL 4-isopropylbenzoic acid for 24 h as templates). D: Screen of transformants without plasmid pYJRC/lysE-asd
| [1] | Becker J, Rohles CM, Wittmann C. Metabolically engineered Corynebacterium glutamicum for bio-based production of chemicals, fuels, materials, and healthcare products[J]. Metab Eng, 2018, 50: 122-141. |
| [2] | Zhao NN, Qian L, Luo GJ, et al. Synthetic biology approaches to access renewable carbon source utilization in Corynebacterium glutamicum[J]. Appl Microbiol Biotechnol, 2018, 102(22): 9517-9529. |
| [3] |
Kalinowski J, Bathe B, Bartels D, et al. The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of L-aspartate-derived amino acids and vitamins[J]. J Biotechnol, 2003, 104(1-3): 5-25.
doi: 10.1016/s0168-1656(03)00154-8 pmid: 12948626 |
| [4] |
Wendisch VF, Bott M, Eikmanns BJ. Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for biotechnological production of organic acids and amino acids[J]. Curr Opin Microbiol, 2006, 9(3): 268-274.
pmid: 16617034 |
| [5] |
Blazeck J, Alper HS. Promoter engineering: recent advances in controlling transcription at the most fundamental level[J]. Biotechnol J, 2013, 8(1): 46-58.
doi: 10.1002/biot.201200120 pmid: 22890821 |
| [6] | Xu N, Wei L, Liu J. Recent advances in the applications of promoter engineering for the optimization of metabolite biosynthesis[J]. World J Microbiol Biotechnol, 2019, 35(2): 33. |
| [7] | 刘莫识, 刘娇, 孙冠男, 等. 谷氨酸棒杆菌人工合成启动子文库的构建及应用[J]. 生物工程学报, 2022, 38(2): 831-842. |
| Liu MS, Liu J, Sun GN, et al. Construction and application of a synthetic promoter library for Corynebacterium glutamicum[J]. Chin J Biotechnol, 2022, 38(2): 831-842. | |
| [8] |
Eaton RW. P-Cymene catabolic pathway in Pseudomonas putida F1: cloning and characterization of DNA encoding conversion of p-cymene to p-cumate[J]. J Bacteriol, 1997, 179(10): 3171-3180.
pmid: 9150211 |
| [9] | Choi YJ, Morel L, Le François T, et al. Novel, versatile, and tightly regulated expression system for Escherichia coli strains[J]. Appl Environ Microbiol, 2010, 76(15): 5058-5066. |
| [10] | Seo SO, Schmidt-Dannert C. Development of a synthetic cumate-inducible gene expression system for Bacillus[J]. Appl Microbiol Biotechnol, 2019, 103(1): 303-313. |
| [11] | Bai CX, van Wezel GP. CUBIC: a versatile cumate-based inducible CRISPRi system in Streptomyces[J]. ACS Synth Biol, 2023, 12(10): 3143-3147. |
| [12] | Pöschel L, Gehr E, Jordan P, et al. Expression of toxic genes in Methylorubrum extorquens with a tightly repressed, cumate-inducible promoter[J]. Antonie Van Leeuwenhoek, 2023, 116(12): 1285-1294. |
| [13] | Pan HQ, Shim A, Lubin MB, et al. Hopanoid lipids promote soybean- Bradyrhizobium symbiosis[J]. bioRxiv, 2023: 2023.09.04.556284. |
| [14] |
Mullick A, Xu Y, Warren R, et al. The cumate gene-switch: a system for regulated expression in mammalian cells[J]. BMC Biotechnol, 2006, 6: 43.
pmid: 17083727 |
| [15] | 张灵. 谷氨酸棒杆菌严格调控的启动子系统的建立[D]. 广州: 华南理工大学, 2021. |
| Zhang L. Establishment of tightly regulated promoter system in Corynebacterium glutamicum[D]. Guangzhou: South China University of Technology, 2021. | |
| [16] | Wei L, Xu N, Wang YR, et al. Promoter library-based module combination(PLMC)technology for optimization of threonine biosynthesis in Corynebacterium glutamicum[J]. Appl Microbiol Biotechnol, 2018, 102(9): 4117-4130. |
| [17] | Li Y, Ai YQ, Zhang JZ, et al. A novel expression vector for Corynebacterium glutamicum with an auxotrophy complementation system[J]. Plasmid, 2020, 107: 102476. |
| [18] | Zhang J, Yang FY, Yang YP, et al. Optimizing a CRISPR-Cpf1-based genome engineering system for Corynebacterium glutamicum[J]. Microb Cell Fact, 2019, 18(1): 60. |
| [19] | Su R, Wang T, Bo TD, et al. Enhanced production of D-pantothenic acid in Corynebacterium glutamicum using an efficient CRISPR-Cpf1 genome editing method[J]. Microb Cell Fact, 2023, 22(1):3. |
| [20] |
Kallscheuer N, Vogt M, Stenzel A, et al. Construction of a Corynebacterium glutamicum platform strain for the production of stilbenes and(2S)-flavanones[J]. Metab Eng, 2016, 38: 47-55.
doi: S1096-7176(16)30047-7 pmid: 27288926 |
| [21] |
上官玲玲, 卢慧芳, 夏会丽, 等. 谷氨酸棒杆菌细胞工厂构建与应用的研究进展[J]. 食品与发酵工业, 2022, 48(17): 313-320.
doi: 10.13995/j.cnki.11-1802/ts.030792 |
| Shangguan LL, Lu HF, Xia HL, et al. Research progress on construction and application of Corynebacterium glutamicum cell factory[J]. Food Ferment Ind, 2022, 48(17): 313-320. | |
| [22] | 龙梦飞, 徐美娟, 张显, 等. 合成生物学与代谢工程在谷氨酸棒杆菌产氨基酸中的应用[J]. 中国科学: 生命科学, 2019, 49(5): 541-552. |
| Long MF, Xu MJ, Zhang X, et al. Synthetic biology and metabolic engineering for amino acid production in Corynebacterium glutamicum[J]. Sci Sin Vitae, 2019, 49(5): 541-552. | |
| [23] | Wang B, Hu QT, Zhang Y, et al. A RecET-assisted CRISPR-Cas9 genome editing in Corynebacterium glutamicum[J]. Microb Cell Fact, 2018, 17(1): 63. |
| [24] | Deng C, Wu YK, Lv XQ, et al. Refactoring transcription factors for metabolic engineering[J]. Biotechnol Adv, 2022, 57: 107935. |
| [25] | Lu XY, Zhang MM, Li G, et al. Applications and research advances in the delivery of CRISPR/Cas9 systems for the treatment of inherited diseases[J]. Int J Mol Sci, 2023, 24(17): 13202. |
| [26] |
Liu X, Wu SR, Xu J, et al. Application of CRISPR/Cas9 in plant biology[J]. Acta Pharm Sin B, 2017, 7(3): 292-302.
doi: 10.1016/j.apsb.2017.01.002 pmid: 28589077 |
| [27] |
Tian PF, Wang J, Shen XL, et al. Fundamental CRISPR-Cas9 tools and current applications in microbial systems[J]. Synth Syst Biotechnol, 2017, 2(3): 219-225.
doi: 10.1016/j.synbio.2017.08.006 pmid: 29318202 |
| [28] |
Joung J, Konermann S, Gootenberg JS, et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening[J]. Nat Protoc, 2017, 12(4): 828-863.
doi: 10.1038/nprot.2017.016 pmid: 28333914 |
| [29] | Liu J, Wang Y, Lu YJ, et al. Development of a CRISPR/Cas9 genome editing toolbox for Corynebacterium glutamicum[J]. Microb Cell Fact, 2017, 16(1): 205. |
| [30] | 王钰, 郑平, 孙际宾. 谷氨酸棒杆菌的代谢工程使能技术研究进展[J]. 生物工程学报, 2021, 37(5): 1603-1618. |
| Wang Y, Zheng P, Sun JB. Recent advances in developing enabling technologies for Corynebacterium glutamicum metabolic engineering[J]. Chin J Biotechnol, 2021, 37(5): 1603-1618. | |
| [31] |
Jiang Y, Qian FH, Yang JJ, et al. CRISPR-Cpf1 assisted genome editing of Corynebacterium glutamicum[J]. Nat Commun, 2017, 8: 15179.
doi: 10.1038/ncomms15179 pmid: 28469274 |
| [32] | 杨帆, 李寅. 新一代基因组编辑系统CRISPR/Cpf1[J]. 生物工程学报, 2017, 33(3): 361-371. |
| Yang F, Li Y. The new generation tool for CRISPR genome editing: CRISPR/Cpf1[J]. Chin J Biotechnol, 2017, 33(3): 361-371. | |
| [33] | 包心茹, 陈卯森, 钟洁, 等. CRISPR/Cas12a基因组编辑技术及应用[J]. 中国生物工程杂志, 2023, 43(10): 32-42. |
| Bao XR, Chen MS, Zhong J, et al. Characteristics and application of CRISPR/Cas12a genome editing technology[J]. China Biotechnol, 2023, 43(10): 32-42. | |
| [34] | Li N, Wang M, Yu SQ, et al. Optimization of CRISPR-Cas9 through promoter replacement and efficient production of L-homoserine in Corynebacterium glutamicum[J]. Biotechnol J, 2021, 16(8): e2100093. |
| [1] | XUE Ning, WANG Jin, LI Shi-xin, LIU Ye, CHENG Hai-jiao, ZHANG Yue, MAO Yu-feng, WANG Meng. Construction of L-phenylalanine High-producing Corynebacterium glutamicum Engineered Strains via Multi-gene Simultaneous Regulation Combined with High-throughput Screening [J]. Biotechnology Bulletin, 2023, 39(9): 268-280. |
| [2] | LIU Jia-hui, LIU Ye, HUA Er-bing, WANG Meng. PAM Extension of Cytosine Base Editing Tool in Corynebacterium glutamicum [J]. Biotechnology Bulletin, 2023, 39(9): 49-57. |
| [3] | CHEN Guang, LI Jia, DU Rui-ying, WANG Xu. pOsHAK1:OsFLN2 Expression Enhances the Drought Tolerance by Altering Sugar Metabolism in Rice [J]. Biotechnology Bulletin, 2022, 38(8): 92-100. |
| [4] | NIE Zhi-hua, ZHU Lei-lei. Effects of Biotin on L-Glutamate Efflux Mediated by MscCG in Fermentation Process [J]. Biotechnology Bulletin, 2020, 36(10): 150-155. |
| [5] | GAO Ran-ran, LIU Qian, SUN Wen-liang, SUN Zhi-yong, LIU Hao, TIAN Chao-guang. Construction of a Recombinant Protein Expression System in Neurospora crassa [J]. Biotechnology Bulletin, 2016, 32(7): 160-169. |
| [6] | Li Tian, Sun Jingkuan, Liu Jingtao. Research Advances on Plant Promoter [J]. Biotechnology Bulletin, 2015, 31(2): 18-25. |
| [7] | Yu Xiaoxia, Tian Jian, Liu Xiaoqing, Wu Ningfeng. Research Progress of Bacillus subtilis Expression System and Its Promoter Regulatory Elements [J]. Biotechnology Bulletin, 2015, 31(2): 35-44. |
| [8] | Li Zhuoxue, Chen Xinbo. Research Advances on Plant Inducible Promoters and Related Cis-acting Elements [J]. Biotechnology Bulletin, 2015, 31(10): 8-15. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||