








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 338-348.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0340
Previous Articles Next Articles
YIN Yue1( ), QIN Xiao-ya1, MI Jia1, AN Wei1, HE Jun1, ZHANG Feng-feng2(
), QIN Xiao-ya1, MI Jia1, AN Wei1, HE Jun1, ZHANG Feng-feng2( )
)
Received:2025-03-31
Online:2026-03-26
Published:2026-04-23
Contact:
ZHANG Feng-feng
E-mail:yueyin0112@aliyun.com;zhfeng99998@163.com
YIN Yue, QIN Xiao-ya, MI Jia, AN Wei, HE Jun, ZHANG Feng-feng. Identification of FBN Gene Family and Its Relationship with Carotenoids Metabolism in Lyciumbarbarum[J]. Biotechnology Bulletin, 2026, 42(3): 338-348.
Fig. 1 Fruits of 'Ningqi 1' at different developmental stagesS1: Young fruit stage. S2: Green fruit stage. S3: Breaker stage. S4: Expanding stage. S5: Mature stage. The same below
基因名称 Gene name | 上游引物序列 Forward primer sequence (5′‒3′) | 下游引物序列 Reverse primer sequence (5′‒3′) | 用途 Purpose |
|---|---|---|---|
| LbaFBN1 | TCACAAGTGCGGTTCTGCGT | AGGAGCACGTTCAACAATCAGGT | 荧光定量 Quantitative real-time |
| LbaFBN2 | TTGAAGGGGAGTGGCAAATGA | CCATTTGCTTTGACAATCTGCTC | |
| LbaFBN3 | GTTCGAAGCCCCTCTAGAATACAG | TAATCTTTTGCCCAAACACATCC | |
| LbaFBN4 | CAAGAGCGCAAAGGTTCAAATAC | GCATAGGGTGTTGGAGTAAAGCA | |
| LbaFBN5 | CTCCTGGAAGAGCTCAAGGTGAG | GAAGGATCAACCATTTTCAAAACG | |
| LbaFBN6 | GTGTGATACCCATAAAAAGCCGT | GAATAAACAGGTTTCCTTGGTTGC | |
| LbaFBN8 | CTCCAGCAAGTTTTCTGTTGAGG | CAAAGCACCCTTTAGTTGCTCA | |
| LbaFBN9 | CCGGGAACTTATCACAGCTGC | CTCCCCTTGTTACACGTGTATCAG | |
| LbaFBN10 | GAACTCAAGGTAGGGGGCCTAGA | GCGTCTTTGAAACACAAACAAGG | |
| LbaFBN11 | CCTCCTCTCAAGGTTCCAATTCC | TCCTTAACGAGAACAAAAAGCCC | |
| LbaFBN12 | TGCAATAACACCAGACCAGTTG | CTTGTCATCTCTGCCTATCCTCG | |
| LbaFBN13 | ACCGCCAAAGCCAAAGGAAC | TTGGGTTCTTGGACTCAAGCTGA | |
| LbaActin | CAATCGGGTATTTCAAGGTCAAG | GAGCAGTGTTTCCCAGCATTG | |
| LbaFBN13-GFP | ACGAGCTGTACAAGGGTACCATGG CTTCCATCTCTTCTCTG | GGATCCTCTAGATGAACTAGTTAA GGCTTCAAGAGGGGAC | 亚细胞定位 Subcellular localization |
Table 1 Primer sequence information used in this study
基因名称 Gene name | 上游引物序列 Forward primer sequence (5′‒3′) | 下游引物序列 Reverse primer sequence (5′‒3′) | 用途 Purpose |
|---|---|---|---|
| LbaFBN1 | TCACAAGTGCGGTTCTGCGT | AGGAGCACGTTCAACAATCAGGT | 荧光定量 Quantitative real-time |
| LbaFBN2 | TTGAAGGGGAGTGGCAAATGA | CCATTTGCTTTGACAATCTGCTC | |
| LbaFBN3 | GTTCGAAGCCCCTCTAGAATACAG | TAATCTTTTGCCCAAACACATCC | |
| LbaFBN4 | CAAGAGCGCAAAGGTTCAAATAC | GCATAGGGTGTTGGAGTAAAGCA | |
| LbaFBN5 | CTCCTGGAAGAGCTCAAGGTGAG | GAAGGATCAACCATTTTCAAAACG | |
| LbaFBN6 | GTGTGATACCCATAAAAAGCCGT | GAATAAACAGGTTTCCTTGGTTGC | |
| LbaFBN8 | CTCCAGCAAGTTTTCTGTTGAGG | CAAAGCACCCTTTAGTTGCTCA | |
| LbaFBN9 | CCGGGAACTTATCACAGCTGC | CTCCCCTTGTTACACGTGTATCAG | |
| LbaFBN10 | GAACTCAAGGTAGGGGGCCTAGA | GCGTCTTTGAAACACAAACAAGG | |
| LbaFBN11 | CCTCCTCTCAAGGTTCCAATTCC | TCCTTAACGAGAACAAAAAGCCC | |
| LbaFBN12 | TGCAATAACACCAGACCAGTTG | CTTGTCATCTCTGCCTATCCTCG | |
| LbaFBN13 | ACCGCCAAAGCCAAAGGAAC | TTGGGTTCTTGGACTCAAGCTGA | |
| LbaActin | CAATCGGGTATTTCAAGGTCAAG | GAGCAGTGTTTCCCAGCATTG | |
| LbaFBN13-GFP | ACGAGCTGTACAAGGGTACCATGG CTTCCATCTCTTCTCTG | GGATCCTCTAGATGAACTAGTTAA GGCTTCAAGAGGGGAC | 亚细胞定位 Subcellular localization |
基因名称 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids | 相对分子量 Molecular weight (kD) | 理论等电点 Theoretical pI | 总平均疏水性指数 Grand average of hydropathicity | 亚细胞定位 Subcellular location |
|---|---|---|---|---|---|---|
| LbaFBN1 | Lba01g00762 | 216 | 24.61 | 7.78 | -0.345 | 叶绿体 Chloroplast |
| LbaFBN2 | Lba02g00336 | 410 | 45.99 | 9.15 | -0.208 | 细胞胞质 Cytoplasmic |
| LbaFBN3 | Lba04g00802 | 332 | 36.45 | 4.73 | -0.361 | 叶绿体 Chloroplast |
| LbaFBN4 | Lba04g01207 | 699 | 78.47 | 8.09 | -0.257 | 细胞核 Nuclear |
| LbaFBN5 | Lba06g00949 | 335 | 38.38 | 9.31 | -0.289 | 叶绿体 Chloroplast |
| LbaFBN6 | Lba06g02827 | 246 | 27.63 | 9.82 | -0.299 | 叶绿体 Chloroplast |
| LbaFBN7 | Lba08g00868 | 198 | 22.34 | 7.54 | -0.215 | 细胞核 Nuclear |
| LbaFBN8 | Lba08g01931 | 302 | 33.68 | 6.78 | -0.088 | 叶绿体 Chloroplast |
| LbaFBN9 | Lba09g00405 | 279 | 30.17 | 6.61 | -0.247 | 叶绿体 Chloroplast |
| LbaFBN10 | Lba09g02050 | 289 | 32.20 | 9.04 | -0.271 | 质膜 Plasma membrane |
| LbaFBN11 | Lba09g02224 | 378 | 41.83 | 4.54 | -0.352 | 叶绿体 Chloroplast |
| LbaFBN12 | Lba10g00821 | 752 | 84.03 | 8.33 | 0.002 | 质膜 Plasma membrane |
| LbaFBN13 | Lba12g00894 | 321 | 35.37 | 5.28 | -0.342 | 叶绿体 Chloroplast |
Table 2 Basic information of LbaFBN gene family in L. barbarum No.1
基因名称 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids | 相对分子量 Molecular weight (kD) | 理论等电点 Theoretical pI | 总平均疏水性指数 Grand average of hydropathicity | 亚细胞定位 Subcellular location |
|---|---|---|---|---|---|---|
| LbaFBN1 | Lba01g00762 | 216 | 24.61 | 7.78 | -0.345 | 叶绿体 Chloroplast |
| LbaFBN2 | Lba02g00336 | 410 | 45.99 | 9.15 | -0.208 | 细胞胞质 Cytoplasmic |
| LbaFBN3 | Lba04g00802 | 332 | 36.45 | 4.73 | -0.361 | 叶绿体 Chloroplast |
| LbaFBN4 | Lba04g01207 | 699 | 78.47 | 8.09 | -0.257 | 细胞核 Nuclear |
| LbaFBN5 | Lba06g00949 | 335 | 38.38 | 9.31 | -0.289 | 叶绿体 Chloroplast |
| LbaFBN6 | Lba06g02827 | 246 | 27.63 | 9.82 | -0.299 | 叶绿体 Chloroplast |
| LbaFBN7 | Lba08g00868 | 198 | 22.34 | 7.54 | -0.215 | 细胞核 Nuclear |
| LbaFBN8 | Lba08g01931 | 302 | 33.68 | 6.78 | -0.088 | 叶绿体 Chloroplast |
| LbaFBN9 | Lba09g00405 | 279 | 30.17 | 6.61 | -0.247 | 叶绿体 Chloroplast |
| LbaFBN10 | Lba09g02050 | 289 | 32.20 | 9.04 | -0.271 | 质膜 Plasma membrane |
| LbaFBN11 | Lba09g02224 | 378 | 41.83 | 4.54 | -0.352 | 叶绿体 Chloroplast |
| LbaFBN12 | Lba10g00821 | 752 | 84.03 | 8.33 | 0.002 | 质膜 Plasma membrane |
| LbaFBN13 | Lba12g00894 | 321 | 35.37 | 5.28 | -0.342 | 叶绿体 Chloroplast |
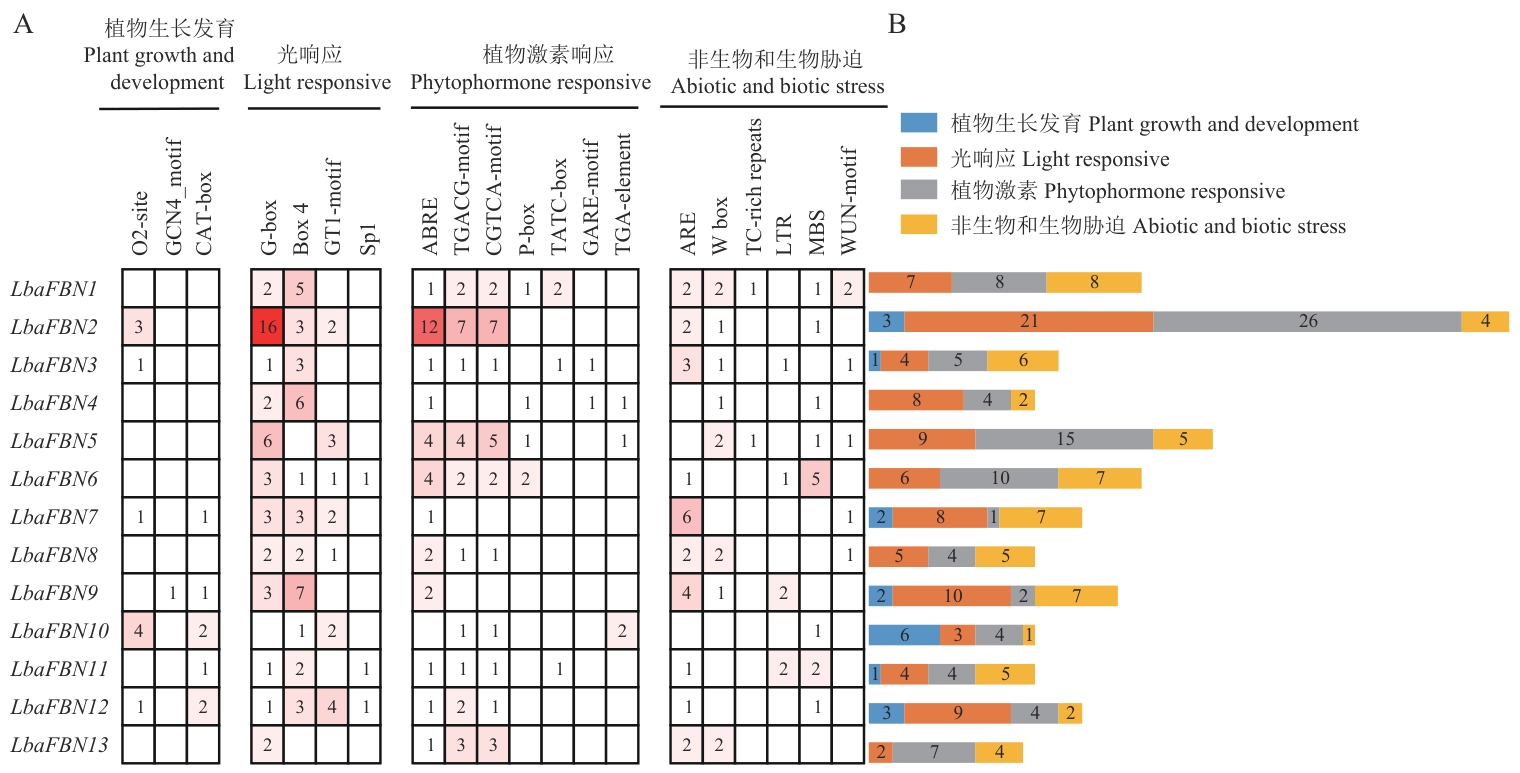
Fig. 5 Analysis of cis-acting elements of LbaFBN genes promotersIn Fig. A, the different colors and numbers of the grids indicate the numbers of different promoter elements in these LbaFBN genes. In Fig. B, the different colored histogram indicates the sum of the cis-acting elements in each category
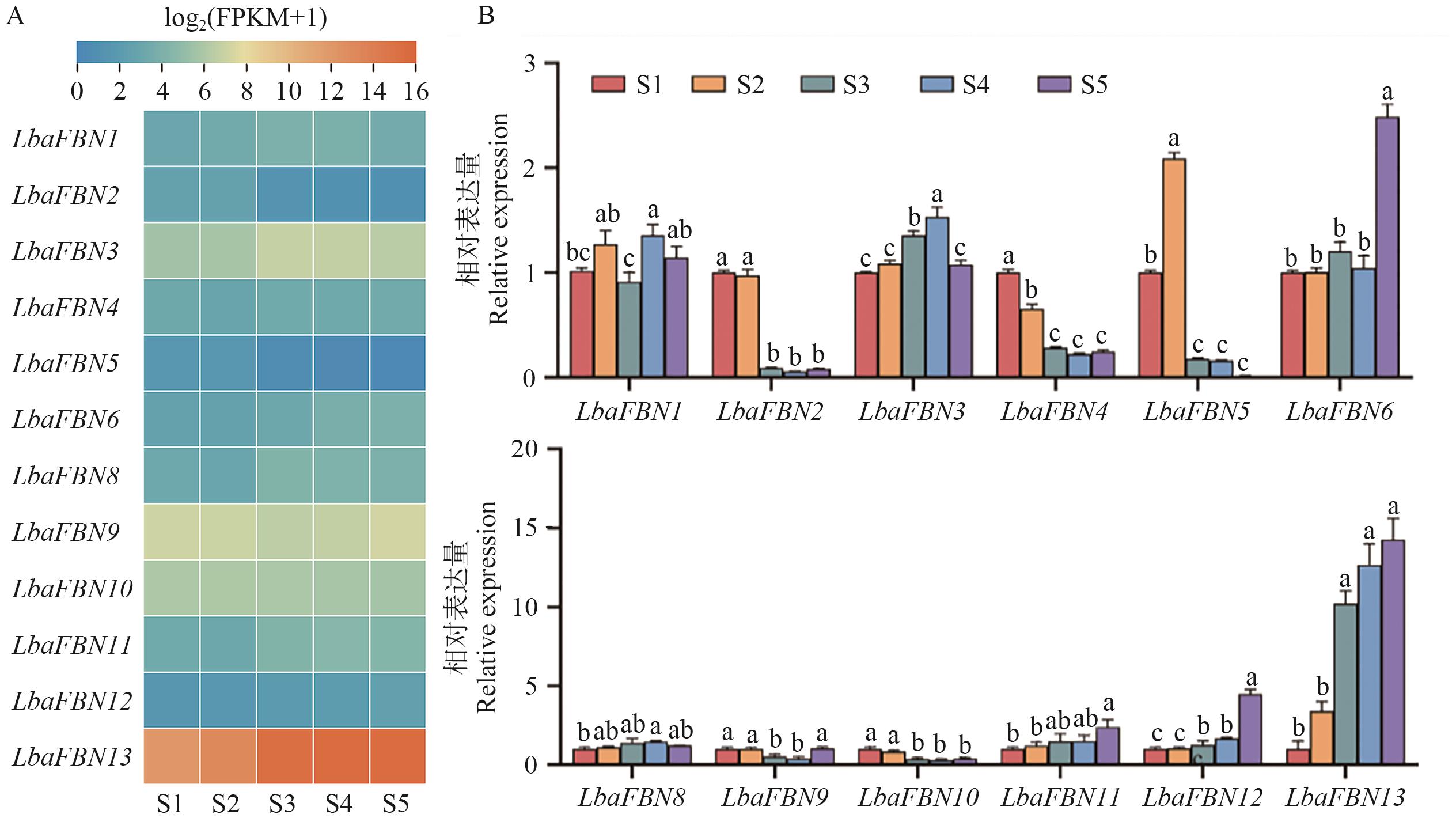
Fig. 7 Expression profile of LbaFBNs genes in 'Ningqi 1' during five fruit developmental stagesA: Transcript profiling of LbaFBN genes based on RNA-seq analysis. B: Gene expression patterns of LbaFBN genes using RT-qPCR validation. The different letters indicate significant differences at the 0.05 level. The same below
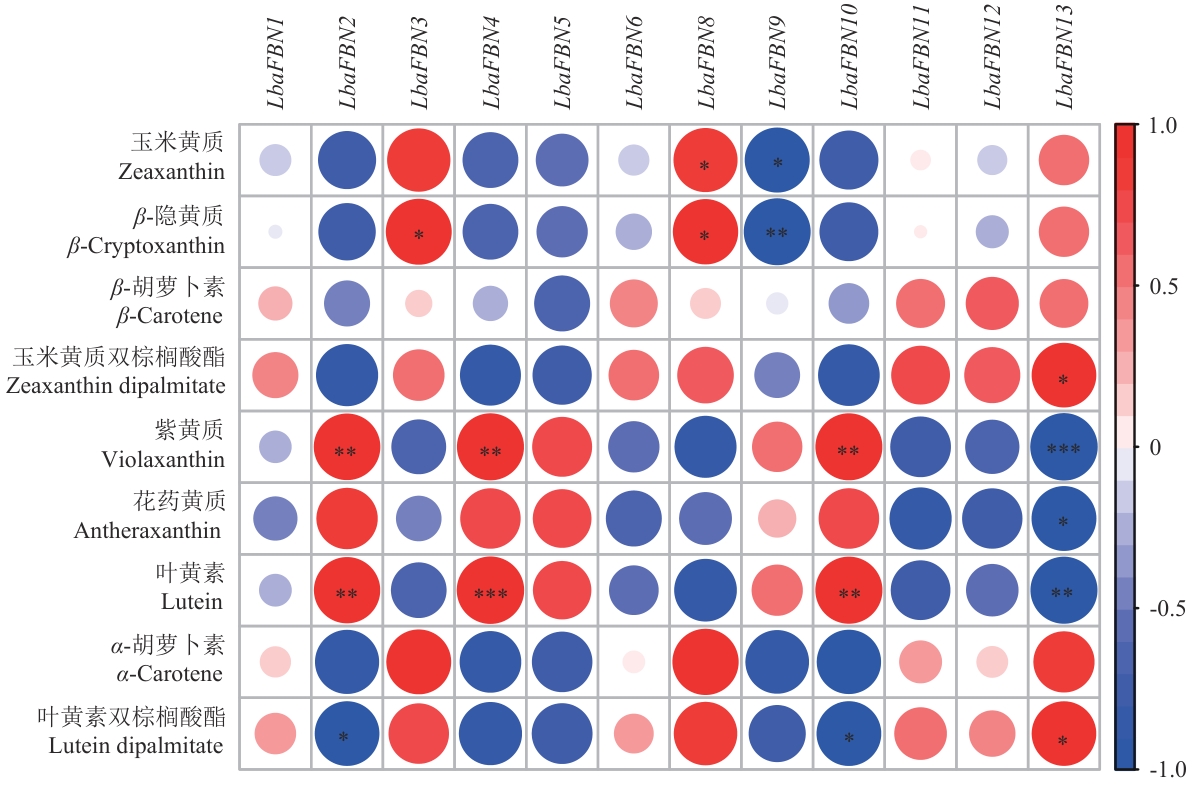
Fig. 9 Correlation analysis between LbaFBNs gene expression and carotenoids content*, **, and *** indicates significant levels at P<0.5, P<0.01, and P<0.001, respectively
| [1] | Cunningham FX Jr, Tice AB, Pham C, et al. Inactivation of genes encoding plastoglobuli-like proteins in Synechocystis sp. PCC 6803 leads to a light-sensitive phenotype [J]. J Bacteriol, 2010, 192(6): 1700-1709. |
| [2] | Simkin AJ, Gaffé J, Alcaraz JP, et al. Fibrillin influence on plastid ultrastructure and pigment content in tomato fruit [J]. Phytochemistry, 2007, 68(11): 1545-1556. |
| [3] | Deruère J, Bouvier F, Steppuhn J, et al. Structure and expression of two plant genes encoding chromoplast-specific proteins: occurrence of partially spliced transcripts [J]. Biochem Biophys Res Commun, 1994, 199(3): 1144-1150. |
| [4] | 周慧梅, 谢德颖, 邹娟子, 等. 植物质体Fibrillin蛋白家族的起源、结构和功能 [J]. 中国农业科技导报, 2014, 16(4): 41-49. |
| Zhou HM, Xie DY, Zou JZ, et al. Origin, structure and function of plastid-associated fibrillin protein family in plants [J]. J Agric Sci Technol, 2014, 16(4): 41-49. | |
| [5] | Sun TH, Rao S, Zhou XS, et al. Plant carotenoids: recent advances and future perspectives [J]. Mol Hortic, 2022, 2(1): 3. |
| [6] | Kim I, Kim HU. The mysterious role of fibrillin in plastid metabolism: current advances in understanding [J]. J Exp Bot, 2022, 73(9): 2751-2764. |
| [7] | Zhou XS, Sun TH, Owens L, et al. Carotenoid sequestration protein FIBRILLIN participates in CmOR-regulated β-carotene accumulation in melon [J]. Plant Physiol, 2023, 193(1): 643-660. |
| [8] | Iglesias-Sanchez A, Navarro-Carcelen J, Morelli L, et al. Arabidopsis FIBRILLIN6 influences carotenoid biosynthesis by directly promoting phytoene synthase activity [J]. Plant Physiol, 2024, 194(3): 1662-1673. |
| [9] | Wang Y, Tian C, Na QT, et al. The role of SlCHRC in carotenoid biosynthesis and plastid development in tomato fruit [J]. Int J Biol Macromol, 2024, 281: 136354. |
| [10] | El-Sappah AH, Li J, Yan K, et al. Fibrillin gene family and its role in plant growth, development, and abiotic stress [J]. Front Plant Sci, 2024, 15: 1453974. |
| [11] | Chen CJ, Wu Y, Li JW, et al. TBtools-II: a "one for all, all for one" bioinformatics platform for biological big-data mining [J]. Mol Plant, 2023, 16(11): 1733-1742. |
| [12] | Liu JM, Zhang Y, Shen Q, et al. Identification of the FBN gene family in tomato and functional analysis of SlFBN11 in the electron transport under low night temperature [J]. Int J Biol Macromol, 2024, 283: 137181. |
| [13] | Singh DK, McNellis TW. Fibrillin protein function: the tip of the iceberg [J]. Trends Plant Sci, 2011, 16(8): 432-441. |
| [14] | Li JJ, Li XK, Khatab AA, et al. Phylogeny, structural diversity and genome-wide expression analysis of fibrillin family genes in rice [J]. Phytochemistry, 2020, 175: 112377. |
| [15] | Sun HR, Ren M, Zhang JN. Genome-wide identification and expression analysis of fibrillin (FBN) gene family in tomato (Solanum lycopersicum L.) [J]. PeerJ, 2022, 10: e13414. |
| [16] | Jiang YY, Hu HC, Ma YH, et al. Genome-wide identification and characterization of the fibrillin gene family in Triticum aestivum [J]. PeerJ, 2020, 8: e9225. |
| [17] | Zafer MZ, Tahir MHN, Khan Z, et al. Genome-wide characterization and sequence polymorphism analyses of Glycine max fibrillin (FBN) revealed its role in response to drought condition [J]. Genes, 2023, 14(6): 1188. |
| [18] | 曾钦羽. 芥菜型油菜BjFBNs基因功能初步研究 [D]. 武汉: 华中农业大学, 2023. |
| Zeng QY. Preliminary study of BjFBNs gene function in Brassica juncea [D]. Wuhan: Huazhong Agricultural University, 2023. | |
| [19] | Pandey A, Sharma P, Mishra D, et al. Genome-wide identification of the fibrillin gene family in chickpea (Cicer arietinum L.) and its response to drought stress [J]. Int J Biol Macromol, 2023, 234: 123757. |
| [20] | 肖佳, 高昊, 周正群, 等. 枸杞属中枸杞红素类成分研究进展 [J]. 科学通报, 2017, 62(16): 1691-1698. |
| Xiao J, Gao H, Zhou ZQ, et al. Recent progress in the study of Zeaxanthin dipalmitate [J]. Chin Sci Bull, 2017, 62(16): 1691-1698. | |
| [21] | Juan C, Montesano D, Manes J, et al. Carotenoids present in goji berries Lycium barbarum L. are suitable to protect against mycotoxins effects: an in vitro study of bioavailability [J]. J Funct Foods, 2022, 92: 105049. |
| [22] | 尹跃, 秦小雅, 赵建华, 等. 枸杞SWEET基因家族鉴定及其与可溶性糖含量关系 [J]. 西北植物学报, 2024, 44(3): 396-407. |
| Yin Y, Qin XY, Zhao JH, et al. Identification of the SWEET gene family and its relationship with soluble sugar content in wolfberry [J]. Acta Bot Boreali Occidentalia Sin, 2024, 44(3): 396-407. | |
| [23] | Liu YL, Zeng SH, Sun W, et al. Comparative analysis of carotenoid accumulation in two goji (Lycium barbarum L. and L. ruthenicum Murr.) fruits [J]. BMC Plant Biol, 2014, 14: 269. |
| [24] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2 (-Delta Delta C(T)) Method [J]. Methods, 2001, 25(4): 402-408. |
| [25] | 邓昌哲, 安飞飞, 李开绵, 等. 外源ABA及其抑制剂钨酸钠对木薯块根类胡萝卜素相关基因和蛋白的影响 [J]. 生物技术通报, 2017, 33(11): 76-83. |
| Deng CZ, An FF, Li KM, et al. Effects of ABA and its synthesis inhibitor sodium tungstate on carotenoid associated genes and enzymes of cassava tuber root [J]. Biotechnol Bull, 2017, 33(11): 76-83. | |
| [26] | Gámez-Arjona FM, Raynaud S, Ragel P, et al. Starch synthase 4 is located in the thylakoid membrane and interacts with plastoglobule-associated proteins in Arabidopsis [J]. Plant J, 2014, 80(2): 305-316. |
| [27] | Torres-Romero D, Gómez-Zambrano Á, Serrato AJ, et al. Arabidopsis fibrillin 1-2 subfamily members exert their functions via specific protein-protein interactions [J]. J Exp Bot, 2022, 73(3): 903-914. |
| [1] | CHEN Deng-ke, LAN Gang, XIA Zhi, HOU Bao-guo, YANG Liu-liu, CAO Cai-rong, LI Peng-bo, WU Cui-cui. Identification of ZF-HD Gene Family in Arachis hypogaea and Analysis in Response to Abiotic Stress [J]. Biotechnology Bulletin, 2026, 42(4): 114-128. |
| [2] | ZENG Ting, ZHANG Lan, LUO Rui. Functional Analysis of the Transcription Factor MpR2R3-MYB17 in Regulating Gemma Development in Marchantia polymorpha L. [J]. Biotechnology Bulletin, 2026, 42(1): 208-217. |
| [3] | REN Yun-er, WU Guo-qiang, CHENG Bin, WEI Ming. Genome-wide Identification of the BvATGs Genes Family in Sugar Beet (Beta vulgaris L.) and Analysis of Their Expression Pattern under Salt Stress [J]. Biotechnology Bulletin, 2026, 42(1): 184-197. |
| [4] | YANG Juan, FENG Hui, JI Nai-zhe, SUN Li-ping, WANG Yun, ZHANG Jia-nan, ZHAO Shi-wei. Cloning and Functional Analysis of AP2/ERF Transcription Factors RcERF4 and RcRAP2-12 in Rose [J]. Biotechnology Bulletin, 2026, 42(1): 150-160. |
| [5] | XU Xiao-ping, YANG Cheng-long, HE Xing, GUO Wen-jie, WU Jian, FANG Shao-zhong. Cloning of the LoAPS1 and Its Function Analysis during the Process of Dormancy Release in Lilium [J]. Biotechnology Bulletin, 2025, 41(9): 195-206. |
| [6] | DONG Xiang-xiang, MIAO Bai-ling, XU He-juan, CHEN Juan-juan, LI Liang-jie, GONG Shou-fu, ZHU Qing-song. Bioinformatics Analysis and Flowering Regulation Function of FveBBX20 Gene in Woodland Strawberry [J]. Biotechnology Bulletin, 2025, 41(9): 115-123. |
| [7] | LAI Shi-yu, LIANG Qiao-lan, WEI Lie-xin, NIU Er-bo, CHEN Ying-e, ZHOU Xin, YANG Si-zheng, WANG Bo. The Role of NbJAZ3 in the Infection of Nicotiana benthamiana by Alfalfa Mosaic Virus [J]. Biotechnology Bulletin, 2025, 41(8): 186-196. |
| [8] | LI Xin-ni, LI Jun-yi, MA Xue-hua, HE Wei, LI Jia-li, YU Jia, CAO Xiao-ning, QIAO Zhi-jun, LIU Si-chen. Identification of the PMEI Gene Family of Pectin Methylesterase Inhibitor in Foxtail Millet and Analysis of Its Response to Abiotic Stress [J]. Biotechnology Bulletin, 2025, 41(7): 150-163. |
| [9] | XU Hui-zhen, SHANTWANA Ghimire, RAJU Kharel, YUE Yun, SI Huai-jun, TANG Xun. Analysis of the Potato SUMO E3 Ligase Gene Family and Cloning and Expression Pattern of StSIZ1 [J]. Biotechnology Bulletin, 2025, 41(6): 119-129. |
| [10] | CHENG Shan, WANG Hui-wei, CHEN Chen, ZHU Ya-jing, LI Chun-xin, BIE Hai, WANG Shu-feng, CHEN Xian-gong, ZHANG Xiang-ge. Cloning of MYB Transcription Factor Gene CeMYB154 and Analysis of Salt Tolerance Function in Cyperus esculentus [J]. Biotechnology Bulletin, 2025, 41(6): 218-228. |
| [11] | YANG Chun, WANG Xiao-qian, WANG Hong-jun, CHAO Yue-hui. Cloning, Subcellular Localization and Expression Analysis of MtZHD4 Gene from Medicago truncatula [J]. Biotechnology Bulletin, 2025, 41(5): 244-254. |
| [12] | HUANG Jin-heng, HUANG Xi, ZHANG Jia-yan, ZHOU Xin-yu, LIAO Pei-ran, YANG Quan. Identification and Expression Analysis of the C3H Gene Family in Grona styracifolia across Different Varieties [J]. Biotechnology Bulletin, 2025, 41(4): 243-255. |
| [13] | BAN Qiu-yan, ZHAO Xin-yue, CHI Wen-jing, LI Jun-sheng, WANG Qiong, XIA Yao, LIANG Li-yun, HE Wei, LI Ye-yun, ZHAO Guang-shan. Cloning of Phytochrome Interaction Factor CsPIF3a and Its Response to Light and Temperature Stress in Camellia sinensis [J]. Biotechnology Bulletin, 2025, 41(4): 256-265. |
| [14] | LU Yong-jie, XIA Hai-qian, LI Yong-ling, ZHANG Wen-jian, YU Jing, ZHAO Hui-na, WANG Bing, XU Ben-bo, LEI Bo. Cloning and Expression Analysis of AP2/ERF Transcription Factor NtESR2 in Nicotiana tabacum [J]. Biotechnology Bulletin, 2025, 41(4): 266-277. |
| [15] | LIU Fang, DU Qian-qian, HE Hao, XIAO Gang, YAN Zhong-yuan, HAO Xiao-hua. Mechanism of miR172b/c-BnMSH7.A1 Module Responding to Cu 2+ Stress in Brassica napus [J]. Biotechnology Bulletin, 2025, 41(2): 139-149. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||