








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (3): 229-241.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0839
Previous Articles Next Articles
JIANG Lin-qi1( ), ZHAO Jia-ying1, ZHENG Fei-xiong1, YAO Xin-yi1, LI Xiao-xian1, YU Zhen-ming1,2(
), ZHAO Jia-ying1, ZHENG Fei-xiong1, YAO Xin-yi1, LI Xiao-xian1, YU Zhen-ming1,2( )
)
Received:2023-08-28
Online:2024-03-26
Published:2024-04-08
Contact:
YU Zhen-ming
E-mail:2978151322@qq.com;yuzhenming@zcmu.edu.cn
JIANG Lin-qi, ZHAO Jia-ying, ZHENG Fei-xiong, YAO Xin-yi, LI Xiao-xian, YU Zhen-ming. Identification and Expression Analysis of 14-3-3 Gene Family in Dendrobium officinale[J]. Biotechnology Bulletin, 2024, 40(3): 229-241.
| 引物名称Primer name | 上游引物Forward primer(5'-3') | 下游引物Reverse primer(5'-3') |
|---|---|---|
| EF-1α | GCTTGAGAAGGAGCCCAAGT | CCAACAGCCACAGTTTGTCG |
| DoGRF1 | GAAGCTGCTGAACAGTCATTG | CCCAGTCTAATTGGGTGAGTAG |
| DoGRF2 | AAAGAAGAGAGTCGTGGGAATG | GTAGCAGAAGGAATGAGATGGG |
| DoGRF3 | GCTGGAATCCTTCGTTTGTTG | TACCGATGGTAATCGCCTTTC |
| DoGRF4 | CGGTGGTATCCTCTCTTTGTTAG | AGCAAGATAGCGGTGGTAATC |
| DoGRF5 | AGCATCACGTGAAGAGGATTAG | CAGCAGACGACGAAGGAATAA |
| DoGRF6 | GAGGATCGTGTGGCGATTAT | GTAGCAGAGGGAATCAGATGAG |
| DoGRF7 | CCATAGGTACCTTGCTGAGTTT | AGTGCCAATCCCAACCTTATAG |
| DoGRF8 | CGGAGATGAAGGTATCCAACAG | TTGCAGTGTCCTCTACCAATC |
| DoGRF9 | GAGATGCTGCAATCGCTAATTC | CCGCTACACTAAACGCAAATG |
| DoGRF10 | ATCCGGCTCTTGAAGGAATG | GGAGCCAGAACAGACAAAGT |
| DoGRF11 | GGATGCGGAAGTCGCTAAT | ATGCTCGGCCAGACTAAAC |
| DoGRF12 | CCTCATTTCCGTCGCATACA | GCCTTCTTCCTTCTGCTCAA |
| DoGRF13 | AAAGTATATGGCGGAGGTGAAG | AATTGCTCTTCGTCCTCCTTAG |
| DoGRF14 | CATCGAGCAGAAGGAGGAAA | GGTGTGAGTCGAGTAACCTAAG |
| DoGRF15 | GGCTGAGAGGTATGAGGAAATG | AGGCTACCGAGAGGAGATTT |
| DoGRF16 | GCGGAGGATATTGCTCTTGT | GGCGAGCGTTCTCTGATAAA |
| DoGRF17 | TGAGAGAGAGCAGCAAGTTTAC | CTACTGTCAGCTCCACATCTAATC |
Table 1 Primer sequences used in this study
| 引物名称Primer name | 上游引物Forward primer(5'-3') | 下游引物Reverse primer(5'-3') |
|---|---|---|
| EF-1α | GCTTGAGAAGGAGCCCAAGT | CCAACAGCCACAGTTTGTCG |
| DoGRF1 | GAAGCTGCTGAACAGTCATTG | CCCAGTCTAATTGGGTGAGTAG |
| DoGRF2 | AAAGAAGAGAGTCGTGGGAATG | GTAGCAGAAGGAATGAGATGGG |
| DoGRF3 | GCTGGAATCCTTCGTTTGTTG | TACCGATGGTAATCGCCTTTC |
| DoGRF4 | CGGTGGTATCCTCTCTTTGTTAG | AGCAAGATAGCGGTGGTAATC |
| DoGRF5 | AGCATCACGTGAAGAGGATTAG | CAGCAGACGACGAAGGAATAA |
| DoGRF6 | GAGGATCGTGTGGCGATTAT | GTAGCAGAGGGAATCAGATGAG |
| DoGRF7 | CCATAGGTACCTTGCTGAGTTT | AGTGCCAATCCCAACCTTATAG |
| DoGRF8 | CGGAGATGAAGGTATCCAACAG | TTGCAGTGTCCTCTACCAATC |
| DoGRF9 | GAGATGCTGCAATCGCTAATTC | CCGCTACACTAAACGCAAATG |
| DoGRF10 | ATCCGGCTCTTGAAGGAATG | GGAGCCAGAACAGACAAAGT |
| DoGRF11 | GGATGCGGAAGTCGCTAAT | ATGCTCGGCCAGACTAAAC |
| DoGRF12 | CCTCATTTCCGTCGCATACA | GCCTTCTTCCTTCTGCTCAA |
| DoGRF13 | AAAGTATATGGCGGAGGTGAAG | AATTGCTCTTCGTCCTCCTTAG |
| DoGRF14 | CATCGAGCAGAAGGAGGAAA | GGTGTGAGTCGAGTAACCTAAG |
| DoGRF15 | GGCTGAGAGGTATGAGGAAATG | AGGCTACCGAGAGGAGATTT |
| DoGRF16 | GCGGAGGATATTGCTCTTGT | GGCGAGCGTTCTCTGATAAA |
| DoGRF17 | TGAGAGAGAGCAGCAAGTTTAC | CTACTGTCAGCTCCACATCTAATC |
| 蛋白名称 Protein name | 氨基酸数Number of amino acids/aa | 分子量Molecular weight/kD | 等电点Isoelectric point | 不稳定系数 Instability index | 脂肪系数 Aliphatic index | 总平均亲水系数 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| DoGRF1 | 248 | 28.24 | 4.85 | 43.69 | 86.98 | -0.444 | 细胞核Nucleus |
| DoGRF2 | 299 | 34.17 | 5.18 | 42.37 | 91.10 | -0.341 | 细胞核Nucleus |
| DoGRF3 | 265 | 29.77 | 4.96 | 41.61 | 91.40 | -0.283 | 细胞核Nucleus |
| DoGRF4 | 266 | 29.93 | 4.96 | 43.50 | 93.31 | -0.293 | 细胞核Nucleus |
| DoGRF5 | 267 | 30.00 | 4.83 | 39.17 | 81.24 | -0.537 | 细胞核Nucleus |
| DoGRF6 | 261 | 29.48 | 6.40 | 42.95 | 87.13 | -0.357 | 细胞核Nucleus |
| DoGRF7 | 250 | 28.25 | 4.99 | 43.68 | 86.04 | -0.358 | 细胞核Nucleus |
| DoGRF8 | 267 | 30.31 | 4.87 | 41.45 | 80.41 | -0.552 | 细胞核Nucleus |
| DoGRF9 | 260 | 29.48 | 4.74 | 47.93 | 91.23 | -0.483 | 细胞核Nucleus |
| DoGRF10 | 257 | 28.94 | 4.73 | 48.82 | 83.23 | -0.472 | 细胞核Nucleus |
| DoGRF11 | 257 | 29.10 | 4.82 | 49.74 | 86.65 | -0.503 | 细胞核Nucleus |
| DoGRF12 | 257 | 28.95 | 4.85 | 48.62 | 81.36 | -0.508 | 细胞核Nucleus |
| DoGRF13 | 254 | 28.57 | 5.94 | 51.58 | 90.39 | -0.259 | 细胞核Nucleus |
| DoGRF14 | 146 | 16.29 | 6.83 | 50.72 | 79.66 | -0.125 | 叶绿体Chloroplast |
| DoGRF15 | 278 | 30.57 | 9.05 | 45.76 | 94.21 | -0.042 | 细胞核Nucleus |
| DoGRF16 | 266 | 29.51 | 8.24 | 40.65 | 95.86 | -0.021 | 细胞核Nucleus |
| DoGRF17 | 246 | 27.61 | 9.03 | 39.78 | 94.47 | -0.128 | 叶绿体Chloroplast |
Table 2 Physicochemical properties of DoGRF proteins in D. officinale
| 蛋白名称 Protein name | 氨基酸数Number of amino acids/aa | 分子量Molecular weight/kD | 等电点Isoelectric point | 不稳定系数 Instability index | 脂肪系数 Aliphatic index | 总平均亲水系数 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| DoGRF1 | 248 | 28.24 | 4.85 | 43.69 | 86.98 | -0.444 | 细胞核Nucleus |
| DoGRF2 | 299 | 34.17 | 5.18 | 42.37 | 91.10 | -0.341 | 细胞核Nucleus |
| DoGRF3 | 265 | 29.77 | 4.96 | 41.61 | 91.40 | -0.283 | 细胞核Nucleus |
| DoGRF4 | 266 | 29.93 | 4.96 | 43.50 | 93.31 | -0.293 | 细胞核Nucleus |
| DoGRF5 | 267 | 30.00 | 4.83 | 39.17 | 81.24 | -0.537 | 细胞核Nucleus |
| DoGRF6 | 261 | 29.48 | 6.40 | 42.95 | 87.13 | -0.357 | 细胞核Nucleus |
| DoGRF7 | 250 | 28.25 | 4.99 | 43.68 | 86.04 | -0.358 | 细胞核Nucleus |
| DoGRF8 | 267 | 30.31 | 4.87 | 41.45 | 80.41 | -0.552 | 细胞核Nucleus |
| DoGRF9 | 260 | 29.48 | 4.74 | 47.93 | 91.23 | -0.483 | 细胞核Nucleus |
| DoGRF10 | 257 | 28.94 | 4.73 | 48.82 | 83.23 | -0.472 | 细胞核Nucleus |
| DoGRF11 | 257 | 29.10 | 4.82 | 49.74 | 86.65 | -0.503 | 细胞核Nucleus |
| DoGRF12 | 257 | 28.95 | 4.85 | 48.62 | 81.36 | -0.508 | 细胞核Nucleus |
| DoGRF13 | 254 | 28.57 | 5.94 | 51.58 | 90.39 | -0.259 | 细胞核Nucleus |
| DoGRF14 | 146 | 16.29 | 6.83 | 50.72 | 79.66 | -0.125 | 叶绿体Chloroplast |
| DoGRF15 | 278 | 30.57 | 9.05 | 45.76 | 94.21 | -0.042 | 细胞核Nucleus |
| DoGRF16 | 266 | 29.51 | 8.24 | 40.65 | 95.86 | -0.021 | 细胞核Nucleus |
| DoGRF17 | 246 | 27.61 | 9.03 | 39.78 | 94.47 | -0.128 | 叶绿体Chloroplast |
| 蛋白名称Protein name | α-螺旋 Alpha helix | 延伸链 Extended strand | β-折叠 Beta turn | 随机卷曲 Random coil |
|---|---|---|---|---|
| DoGRF1 | 52.42 | 12.54 | 5.41 | 29.63 |
| DoGRF2 | 62.54 | 11.04 | 3.34 | 23.08 |
| DoGRF3 | 49.75 | 14.39 | 5.30 | 30.56 |
| DoGRF4 | 52.31 | 14.58 | 6.25 | 26.85 |
| DoGRF5 | 59.20 | 11.66 | 3.37 | 25.77 |
| DoGRF6 | 67.43 | 6.51 | 3.07 | 22.99 |
| DoGRF7 | 52.96 | 12.81 | 5.17 | 29.06 |
| DoGRF8 | 50.84 | 10.60 | 5.06 | 33.49 |
| DoGRF9 | 51.75 | 11.05 | 4.85 | 32.35 |
| DoGRF10 | 52.48 | 15.45 | 2.04 | 30.03 |
| DoGRF11 | 52.06 | 11.60 | 4.38 | 31.96 |
| DoGRF12 | 49.77 | 10.42 | 5.09 | 34.72 |
| DoGRF13 | 44.19 | 16.28 | 5.04 | 34.50 |
| DoGRF14 | 34.91 | 15.98 | 10.65 | 38.46 |
| DoGRF15 | 46.38 | 13.62 | 8.09 | 31.91 |
| DoGRF16 | 45.36 | 12.500 | 5.85 | 36.29 |
| DoGRF17 | 43.03 | 13.75 | 6.97 | 36.25 |
Table 3 Analysis of secondary structure of DoGRF proteins in D. officinale
| 蛋白名称Protein name | α-螺旋 Alpha helix | 延伸链 Extended strand | β-折叠 Beta turn | 随机卷曲 Random coil |
|---|---|---|---|---|
| DoGRF1 | 52.42 | 12.54 | 5.41 | 29.63 |
| DoGRF2 | 62.54 | 11.04 | 3.34 | 23.08 |
| DoGRF3 | 49.75 | 14.39 | 5.30 | 30.56 |
| DoGRF4 | 52.31 | 14.58 | 6.25 | 26.85 |
| DoGRF5 | 59.20 | 11.66 | 3.37 | 25.77 |
| DoGRF6 | 67.43 | 6.51 | 3.07 | 22.99 |
| DoGRF7 | 52.96 | 12.81 | 5.17 | 29.06 |
| DoGRF8 | 50.84 | 10.60 | 5.06 | 33.49 |
| DoGRF9 | 51.75 | 11.05 | 4.85 | 32.35 |
| DoGRF10 | 52.48 | 15.45 | 2.04 | 30.03 |
| DoGRF11 | 52.06 | 11.60 | 4.38 | 31.96 |
| DoGRF12 | 49.77 | 10.42 | 5.09 | 34.72 |
| DoGRF13 | 44.19 | 16.28 | 5.04 | 34.50 |
| DoGRF14 | 34.91 | 15.98 | 10.65 | 38.46 |
| DoGRF15 | 46.38 | 13.62 | 8.09 | 31.91 |
| DoGRF16 | 45.36 | 12.500 | 5.85 | 36.29 |
| DoGRF17 | 43.03 | 13.75 | 6.97 | 36.25 |
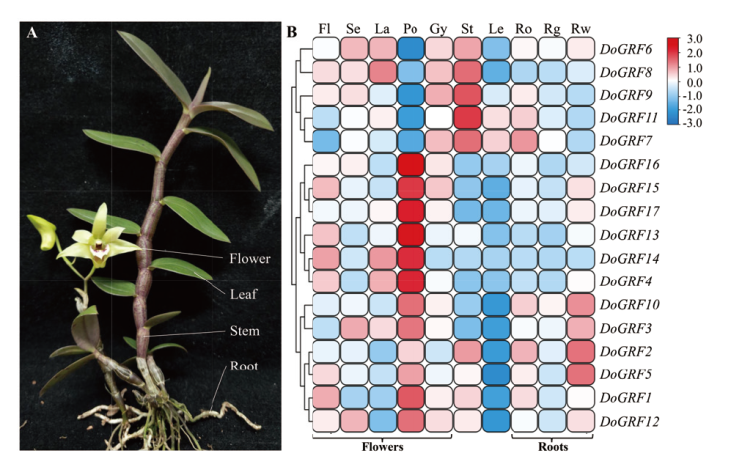
Fig. 6 Tissue specific expression analysis of DoGRF family genes in D. officinale Fl:Flower bud;Se:sepal;La:labellum;Po:pollinium;Gy:gynostemium;St:stem;Le:leaf;Ro:root;Rg:green root tip;Rw:white part root
| [1] |
Zhang HM, Zhu JH, Gong ZZ, et al. Abiotic stress responses in plants[J]. Nat Rev Genet, 2022, 23(2): 104-119.
doi: 10.1038/s41576-021-00413-0 |
| [2] |
严静, 蔡易熹, 陈燕兰, 等. 铁皮石斛茎、叶、花的活性成分及综合利用研究进展[J]. 食品与发酵工业, 2021, 47(17): 299-306.
doi: 10.13995/j.cnki.11-1802/ts.026336 |
| Yan J, Cai YX, Chen YL, et al. Research progress in active components and comprehensive utilization of stems, leaves and flowers of Dendrobium officinale[J]. Food Ferment Ind, 2021, 47(17): 299-306. | |
| [3] |
Yu ZM, Yang ZY, Teixeira da Silva JA, et al. Influence of low temperature on physiology and bioactivity of postharvest Dendrobium officinale stems[J]. Postharvest Biol Technol, 2019, 148: 97-106.
doi: 10.1016/j.postharvbio.2018.10.014 URL |
| [4] |
Niu ZT, Zhu F, Fan YJ, et al. The chromosome-level reference genome assembly for Dendrobium officinale and its utility of functional genomics research and molecular breeding study[J]. Acta Pharm Sin B, 2021, 11(7): 2080-2092.
doi: 10.1016/j.apsb.2021.01.019 URL |
| [5] |
Zhao X, Li F, Li K. The 14-3-3 proteins: regulators of plant metabolism and stress responses[J]. Plant Biol, 2021, 23(4): 531-539.
doi: 10.1111/plb.v23.4 URL |
| [6] |
Ren JX, Zhang P, Dai YB, et al. Evolution of the 14-3-3 gene family in monocotyledons and dicotyledons and validation of MdGRF13 function in transgenic Arabidopsis thaliana[J]. Plant Cell Rep, 2023, 42(8): 1345-1364.
doi: 10.1007/s00299-023-03035-4 |
| [7] |
Huang Y, Wang WS, Yu H, et al. The role of 14-3-3 proteins in plant growth and response to abiotic stress[J]. Plant Cell Rep, 2022, 41(4): 833-852.
doi: 10.1007/s00299-021-02803-4 |
| [8] | 李芳, 滕建晒, 陈宣钦. 14-3-3蛋白参与植物应答非生物胁迫的研究进展[J]. 植物科学学报, 2018, 36(3): 459-469. |
| Li F, Teng JS, Chen XQ. Research progress on the 14-3-3 protein involved in plant responses to abiotic stress[J]. Plant Sci J, 2018, 36(3): 459-469. | |
| [9] | 桑婷, 颜秀娟, 裴红霞, 等. 辣椒14-3-3蛋白基因家族全基因组鉴定与表达特征分析[J]. 分子植物育种, 2021, 19(16): 5268-5278. |
| Sang T, Yan XJ, Pei HX, et al. Genome-wide identification and expression characteristics analysis of 14-3-3 gene family in pepper(Capsicum annuum)[J]. Mol Plant Breed, 2021, 19(16): 5268-5278. | |
| [10] |
He FY, Duan SG, Jian YQ, et al. Genome-wide identification and gene expression analysis of the 14-3-3 gene family in potato(Solanum tuberosum L.)[J]. BMC Genomics, 2022, 23(1): 811.
doi: 10.1186/s12864-022-09037-y |
| [11] |
Xu MY, Hu ZY, Lai W, et al. Comprehensive analysis of 14-3-3 family genes and their responses to cold and drought stress in cucumber[J]. Funct Plant Biol, 2021, 48(12): 1264-1276.
doi: 10.1071/FP21022 pmid: 34635203 |
| [12] |
Chen F, Li Q, Sun LX, et al. The rice 14-3-3 gene family and its involvement in responses to biotic and abiotic stress[J]. DNA Res, 2006, 13(2): 53-63.
doi: 10.1093/dnares/dsl001 URL |
| [13] |
Pan RR, Wang YJ, An FF, et al. Genome-wide identification and characterization of 14-3-3 gene family related to negative regulation of starch accumulation in storage root of Manihot esculenta[J]. Front Plant Sci, 2023, 14: 1184903.
doi: 10.3389/fpls.2023.1184903 URL |
| [14] |
Zhang ZB, Wang XK, Wang S, et al. Expansion and diversification of the 14-3-3 gene family in Camellia sinensis[J]. J Mol Evol, 2022, 90(3-4): 296-306.
doi: 10.1007/s00239-022-10060-6 |
| [15] |
Liu Q, Zhang SH, Liu B. 14-3-3 proteins: Macro-regulators with great potential for improving abiotic stress tolerance in plants[J]. Biochem Biophys Res Commun, 2016, 477(1): 9-13.
doi: 10.1016/j.bbrc.2016.05.120 URL |
| [16] |
Camoni L, Visconti S, Aducci P, et al. 14-3-3 proteins in plant hormone signaling: doing several things at once[J]. Front Plant Sci, 2018, 9: 297.
doi: 10.3389/fpls.2018.00297 pmid: 29593761 |
| [17] |
Yang ZJ, Wang CW, Xue Y, et al. Calcium-activated 14-3-3 proteins as a molecular switch in salt stress tolerance[J]. Nat Commun, 2019, 10(1): 1199.
doi: 10.1038/s41467-019-09181-2 pmid: 30867421 |
| [18] |
Yu ZM, Zhang GH, Teixeira da Silva JA, et al. Genome-wide identification and analysis of DNA methyltransferase and demethylase gene families in Dendrobium officinale reveal their potential functions in polysaccharide accumulation[J]. BMC Plant Biol, 2021, 21(1): 21.
doi: 10.1186/s12870-020-02811-8 |
| [19] |
Zhang MZ, Liu N, Teixeira da Silva JA, et al. Physiological and transcriptomic analysis uncovers salinity stress mechanisms in a facultative crassulacean acid metabolism plant Dendrobium officinale[J]. Front Plant Sci, 2022, 13: 1028245.
doi: 10.3389/fpls.2022.1028245 URL |
| [20] |
Lu SN, Wang JY, Chitsaz F, et al. CDD/SPARCLE: the conserved domain database in 2020[J]. Nucleic Acids Res, 2020, 48(D1): D265-D268.
doi: 10.1093/nar/gkz991 URL |
| [21] |
Potter SC, Luciani A, Eddy SR, et al. HMMER web server: 2018 update[J]. Nucleic Acids Res, 2018, 46(W1): W200-W204.
doi: 10.1093/nar/gky448 URL |
| [22] |
Chen CJ, Chen H, Zhang Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data[J]. Mol Plant, 2020, 13(8): 1194-1202.
doi: S1674-2052(20)30187-8 pmid: 32585190 |
| [23] |
Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms[J]. Mol Biol Evol, 2018, 35(6): 1547-1549.
doi: 10.1093/molbev/msy096 pmid: 29722887 |
| [24] |
Zhang GQ, Liu KW, Li Z, et al. The Apostasia genome and the evolution of orchids[J]. Nature, 2017, 549(7672): 379-383.
doi: 10.1038/nature23897 URL |
| [25] |
Pertea M, Kim D, Pertea GM, et al. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown[J]. Nat Protoc, 2016, 11(9): 1650-1667.
doi: 10.1038/nprot.2016.095 pmid: 27560171 |
| [26] | 俞振明, 赵聪慧, 张明泽, 等. 益智不同组织多糖含量及其生物合成途径分析[J]. 热带亚热带植物学报, 2021, 29(6): 669-677. |
| Yu ZM, Zhao CH, Zhang MZ, et al. Analysis of polysaccharide content and biosynthesis pathway in different tissues of Alpinia oxyphylla[J]. J Trop Subtrop Bot, 2021, 29(6): 669-677. | |
| [27] |
Yu ZM, Zhang GH, Teixeira da Silva JA, et al. The methyl jasmonate-responsive transcription factor DobHLH4 promotes DoTPS10, which is involved in linalool biosynthesis in Dendrobium officinale during floral development[J]. Plant Sci, 2021, 309: 110952.
doi: 10.1016/j.plantsci.2021.110952 URL |
| [28] |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT Method[J]. Methods, 2001, 25(4): 402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [29] |
Liang YF, Ma F, Zhang RL, et al. Genome-wide identification and characterization of tomato 14-3-3(SlTFT)genes and functional analysis of SlTFT6 under heat stress[J]. Physiol Plant, 2023, 175(2): e13888.
doi: 10.1111/ppl.v175.2 URL |
| [30] |
Zuo XY, Wang SX, Xiang W, et al. Genome-wide identification of the 14-3-3 gene family and its participation in floral transition by interacting with TFL1/FT in apple[J]. BMC Genomics, 2021, 22(1): 41.
doi: 10.1186/s12864-020-07330-2 |
| [31] |
Liu ZY, Jia YX, Ding YL, et al. Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during cold response[J]. Mol Cell, 2017, 66(1): 117-128.
doi: S1097-2765(17)30131-4 pmid: 28344081 |
| [32] |
Wang NN, Shi YY, Jiang Q, et al. A 14-3-3 protein positively regulates rice salt tolerance by stabilizing phospholipase C1[J]. Plant Cell Environ, 2023, 46(4): 1232-1248.
doi: 10.1111/pce.14520 URL |
| [1] | ZHANG Yu, SHI Lei, GONG Lei, NIE Feng-jie, YANG Jiang-wei, LIU Xuan, YANG Wen-jing, ZHANG Guo-hui, XIE Rui-xia, ZHANG Li. Genome-wide Identification of Potato WOX Gene Family and Its Expression Analysis in in vitro Regeneration and Abiotic Stress [J]. Biotechnology Bulletin, 2024, 40(3): 170-180. |
| [2] | WU Xing-xing, HONG Hai-bo, GAN Zhi-cheng, LI Rui-ning, HUANG Xian-zhong. Cloning and Preliminary Functional Analysis of CaPI Gene in Capsicum annuum L. [J]. Biotechnology Bulletin, 2024, 40(3): 193-201. |
| [3] | ZHOU Hong-dan, LUO Xiao-ping, TU Mi-xue, LI Zhong-guang. Phytomelatonin: An Emerging Signal Molecule Responding to Abiotic Stress [J]. Biotechnology Bulletin, 2024, 40(3): 41-51. |
| [4] | WU Cui-cui, XIAO Shui-ping. Genome-wide Identification of HD-Zip Gene Family in Gossypium hirsutum L. and Expression Analysis in Response to Abiotic Stress [J]. Biotechnology Bulletin, 2024, 40(2): 130-145. |
| [5] | XIN Qi, LI Ya-fan, YIN Zheng, ZHANG Xiao-dan, CHEN Ting, LIU Xiao-hua. Identification and Expression Analysis of the CBL-CIPK Gene Family in Sugarcane [J]. Biotechnology Bulletin, 2024, 40(2): 197-211. |
| [6] | LI Ya-nan, ZHANG Hao-jie, LIANG Meng-jing, LUO Tao, LI Wang-ning, ZHANG Chun-hui, JI Chun-li, LI Run-zhi, XUE Jin-ai, CUI Hong-li. Identification and Expression Analysis of Calcium-dependent Protein Kinase(CDPK)Family in Haematococcus pluvialis [J]. Biotechnology Bulletin, 2024, 40(2): 300-312. |
| [7] | ZOU Xiu-wei, YUE Jia-ni, LI Zhi-yu, DAI Liang-ying, LI Wei. Functional Analysis of Rice Heat Shock Transcription Factor HsfA2b Regulating the Resistance to Abiotic Stresses [J]. Biotechnology Bulletin, 2024, 40(2): 90-98. |
| [8] | ZHANG Yi, ZHANG Xin-ru, ZHANG Jin-ke, HU Li-zong, SHANGGUAN Xin-xin, ZHENG Xiao-hong, HU Juan-juan, ZHANG Cong-cong, MU Gui-qing, LI Cheng-wei. Functional Analysis of TaMYB1 Gene in Wheat Under Cadmium Stress [J]. Biotechnology Bulletin, 2024, 40(1): 194-206. |
| [9] | WU Zhen, ZHANG Ming-Ying, YAN Feng, LI Yi-min, GAO Jing, YAN Yong-Gang, ZHANG Gang. Identification and Analysis of WRKY Gene Family in Rheum palmatum L. [J]. Biotechnology Bulletin, 2024, 40(1): 250-261. |
| [10] | YANG Zhi-xiao, HOU Qian, LIU Guo-quan, LU Zhi-gang, CAO Yi, GOU Jian-yu, WANG Yi, LIN Ying-chao. Responses of Rubisco and Rubisco Activase in Different Resistant Tobacco Strains to Brown Spot Stress [J]. Biotechnology Bulletin, 2023, 39(9): 202-212. |
| [11] | CHEN Zhong-yuan, WANG Yu-hong, DAI Wei-jun, ZHANG Yan-min, YE Qian, LIU Xu-ping, TAN Wen-Song, ZHAO Liang. Mechanism Investigation of Ferric Ammonium Citrate on Transfection for Suspended HEK293 Cells [J]. Biotechnology Bulletin, 2023, 39(9): 311-318. |
| [12] | ZHAO Xue-ting, GAO Li-yan, WANG Jun-gang, SHEN Qing-qing, ZHANG Shu-zhen, LI Fu-sheng. Cloning and Expression of AP2/ERF Transcription Factor Gene ShERF3 in Sugarcane and Subcellular Localization of Its Encoded Protein [J]. Biotechnology Bulletin, 2023, 39(6): 208-216. |
| [13] | LI Yuan-hong, GUO Yu-hao, CAO Yan, ZHU Zhen-zhou, WANG Fei-fei. Research Progress in the Microalgal Growth and Accumulation of Target Products Regulated by Exogenous Phytohormone [J]. Biotechnology Bulletin, 2023, 39(6): 61-72. |
| [14] | FENG Shan-shan, WANG Lu, ZHOU Yi, WANG You-ping, FANG Yu-jie. Research Progresses on WOX Family Genes in Regulating Plant Development and Abiotic Stress Response [J]. Biotechnology Bulletin, 2023, 39(5): 1-13. |
| [15] | LI Zhi-qi, YUAN Yue, MIAO Rong-qing, PANG Qiu-ying, ZHANG Ai-qin. Melatonin Contents in Eutrema salsugineum and Arabidopsis thaliana Under Salt Stress, and Expression Pattern Analysis of Synthesis Related Genes [J]. Biotechnology Bulletin, 2023, 39(5): 142-151. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||