








生物技术通报 ›› 2024, Vol. 40 ›› Issue (6): 114-125.doi: 10.13560/j.cnki.biotech.bull.1985.2023-1184
张迪1( ), 鞠睿1, 李丽梅2, 王煜倩3, 陈瑞3, 王新一1(
), 鞠睿1, 李丽梅2, 王煜倩3, 陈瑞3, 王新一1( )
)
收稿日期:2023-12-15
出版日期:2024-06-26
发布日期:2024-04-19
通讯作者:
王新一,男,博士,教授,研究方向:分析化学、纳米生物学;E-mail: wangxinyi@dlu.edu.cn作者简介:张迪,女,博士,研究方向:环境化学;E-mail: deezhang163@163.com
基金资助:
ZHANG Di1( ), JU Rui1, LI Li-mei2, WANG Yu-qian3, CHEN Rui3, WANG Xin-yi1(
), JU Rui1, LI Li-mei2, WANG Yu-qian3, CHEN Rui3, WANG Xin-yi1( )
)
Received:2023-12-15
Published:2024-06-26
Online:2024-04-19
摘要:
转录因子是一类在细胞中通过引导RNA聚合酶与特定DNA序列结合调控基因表达的蛋白质。转录因子是生物体适应自然环境过程中的一类重要调节性蛋白,可专一性的感知环境介质中的配体分子。利用转录因子作为识别元件构建生物传感体系并与不同的信号转导和放大系统耦合,可在细胞、无细胞和体外等不同反应体系构建信号输出形式灵活多样的传感策略。基于转录因子构建的生物传感器(TFBBs),因其灵敏度高、体积小、价格低廉以及可应用于现场监测等优势,在环境分析领域显示出巨大应用潜力。文中着重介绍了TFBBs的传感体系类型、核心元件组成和工作原理,列举了其在重金属离子、芳香族化合物及抗生素等环境污染物检测中的应用进展,在此基础上讨论了TFBBs在环境污染物检测中可能面临的机遇和挑战,并对其发展趋势进行了展望,指出合成生物学、人工智能等多学科新兴技术的高速发展将促进TFBBs人工设计和传感性能提升,使其应用于更广泛的领域。
张迪, 鞠睿, 李丽梅, 王煜倩, 陈瑞, 王新一. 基于转录因子生物传感器在环境分析中的应用[J]. 生物技术通报, 2024, 40(6): 114-125.
ZHANG Di, JU Rui, LI Li-mei, WANG Yu-qian, CHEN Rui, WANG Xin-yi. Application of Transcription Factor-based Biosensors in Environmental Analysis[J]. Biotechnology Bulletin, 2024, 40(6): 114-125.
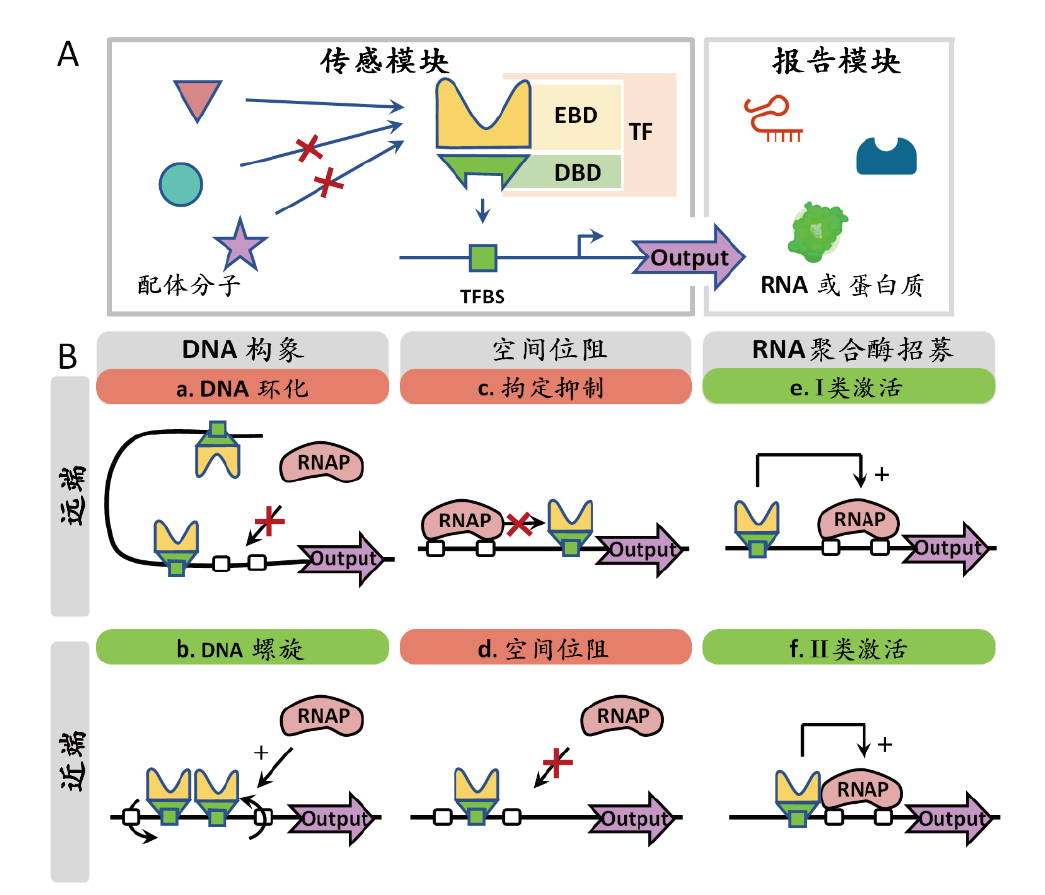
图1 TFBBs结构及工作原理图 A:生物传感器结构示意图;B:转录因子激活及抑制机制示意图
Fig. 1 Principle of transcription factor-based biosensors A: Schematic diagram of biosensor structure; B: schematic description of transcription factor activation and repression mechanisms
| 重金属 Heavy metal ion | 转录因子/家族 TFs/families | PDB | 反应平台 Reaction platform | 报告元件 Report component | 检测限 Limit of detection | 参考文献 Reference |
|---|---|---|---|---|---|---|
| 汞Hg(II) | MerR/MerR | - | 全细胞平台 | mCherry | 0.1 nmol/L | [ |
| 全细胞平台 | GFP | 4.4 nmol/L | [ | |||
| 全细胞平台 | RFP | 1 mg/kg | [ | |||
| 无细胞平台 | 3WJdB | 0.5 nmol/L | [ | |||
| 全细胞平台 | sfCherry3C | 2.24 µmol/L | [ | |||
| CadR/MerR | - | 全细胞平台 | GFP、RFP | 0.01 µg/mL | [ | |
| 镉Cd(II) | ZntR/MerR | 1Q08 | 全细胞平台 | sGFP | 35 nmol/L | [ |
| 全细胞平台 | eGFP | 1 000 nmol/L | [ | |||
| 全细胞平台 | Lux | 100 nmol/L | [ | |||
| MerR/MerR | - | 全细胞平台 | GFP | 283.9 nmol/L | [ | |
| 全细胞平台 | eGFP | 3 µmol/L | [ | |||
| CadC/ArsR/SmtB | 1U2W | 全细胞平台 | GFP | 10 µg/mL | [ | |
| CadR/MerR | - | 全细胞平台 | mCherry | 0.1 µg/mL | [ | |
| ZntR/MerR | 1Q08 | 全细胞平台 | GFP | 5 µmol/L | [ | |
| 砷As(III) | ArsR/ArsR/SmtB | 1R23 | 邻近效应驱动 | Lac Z | 8 ppb | [ |
| ArsR/ArsR/SmtB | - | 全细胞平台 | GFP | 0.1 µmol/L | [ | |
| CadR/MerR | - | 全细胞平台 | GFP、RFP | 0.01 µg/mL | [ | |
| 铜Cu(II) | CopR/CopY | - | 全细胞平台 | RFP | 24.3 µmol/L | [ |
| CueR/MerR | 1Q05 | 全细胞平台 | eGFP | 0.08 µmol/L | [ | |
| CusR | - | 全细胞平台 | GFP | 12 µmol/L | [ | |
| 锌Zn(II) | SmtB/ArsR/SmtB | 1R23 | 全细胞平台 | eGFP | 1 µmol/L | [ |
| ZntR/MerR | 1Q08 | 全细胞平台 | GFP、mCherry | 5 mg/L | [ | |
| MerRZntR/MerR | - | 全细胞平台 | eGFP、mCherry | 10 µg/mL | [ | |
| 全细胞平台 | Lux | 0.03 mmol/L | [ | |||
| 铅Pb(II) | CadR/MerR | - | 全细胞平台 | GFP、RFP | 0.01 µg/mL | [ |
| MerR/MerR | - | 全细胞平台 | GFP | 39.6 nmol/L | [ | |
| PbrR/MerR | - | 全细胞平台 | 靛蓝合成酶Indigoidine synthetase | 0.065 mol/L | [ | |
| 无细胞平台 | 3WJdB | 0.1 nmol/L | [ | |||
| 全细胞平台 | 比色法 | 2.93 nmol/L | [ | |||
| 锑Sb(III) | ArsR/ArsR/SmtB | 1R23 | 全细胞平台 | eGFP | 0.25 µmol/L | [ |
| 镍Ni(II) | RcnR/CsoR-RcnR | - | 全细胞平台 | Lux | 80 nmol/L | [ |
表1 基于转录因子构建的生物传感器对重金属离子的检测
Table 1 Detection of heavy metal ions by transcription factor-based biosensors
| 重金属 Heavy metal ion | 转录因子/家族 TFs/families | PDB | 反应平台 Reaction platform | 报告元件 Report component | 检测限 Limit of detection | 参考文献 Reference |
|---|---|---|---|---|---|---|
| 汞Hg(II) | MerR/MerR | - | 全细胞平台 | mCherry | 0.1 nmol/L | [ |
| 全细胞平台 | GFP | 4.4 nmol/L | [ | |||
| 全细胞平台 | RFP | 1 mg/kg | [ | |||
| 无细胞平台 | 3WJdB | 0.5 nmol/L | [ | |||
| 全细胞平台 | sfCherry3C | 2.24 µmol/L | [ | |||
| CadR/MerR | - | 全细胞平台 | GFP、RFP | 0.01 µg/mL | [ | |
| 镉Cd(II) | ZntR/MerR | 1Q08 | 全细胞平台 | sGFP | 35 nmol/L | [ |
| 全细胞平台 | eGFP | 1 000 nmol/L | [ | |||
| 全细胞平台 | Lux | 100 nmol/L | [ | |||
| MerR/MerR | - | 全细胞平台 | GFP | 283.9 nmol/L | [ | |
| 全细胞平台 | eGFP | 3 µmol/L | [ | |||
| CadC/ArsR/SmtB | 1U2W | 全细胞平台 | GFP | 10 µg/mL | [ | |
| CadR/MerR | - | 全细胞平台 | mCherry | 0.1 µg/mL | [ | |
| ZntR/MerR | 1Q08 | 全细胞平台 | GFP | 5 µmol/L | [ | |
| 砷As(III) | ArsR/ArsR/SmtB | 1R23 | 邻近效应驱动 | Lac Z | 8 ppb | [ |
| ArsR/ArsR/SmtB | - | 全细胞平台 | GFP | 0.1 µmol/L | [ | |
| CadR/MerR | - | 全细胞平台 | GFP、RFP | 0.01 µg/mL | [ | |
| 铜Cu(II) | CopR/CopY | - | 全细胞平台 | RFP | 24.3 µmol/L | [ |
| CueR/MerR | 1Q05 | 全细胞平台 | eGFP | 0.08 µmol/L | [ | |
| CusR | - | 全细胞平台 | GFP | 12 µmol/L | [ | |
| 锌Zn(II) | SmtB/ArsR/SmtB | 1R23 | 全细胞平台 | eGFP | 1 µmol/L | [ |
| ZntR/MerR | 1Q08 | 全细胞平台 | GFP、mCherry | 5 mg/L | [ | |
| MerRZntR/MerR | - | 全细胞平台 | eGFP、mCherry | 10 µg/mL | [ | |
| 全细胞平台 | Lux | 0.03 mmol/L | [ | |||
| 铅Pb(II) | CadR/MerR | - | 全细胞平台 | GFP、RFP | 0.01 µg/mL | [ |
| MerR/MerR | - | 全细胞平台 | GFP | 39.6 nmol/L | [ | |
| PbrR/MerR | - | 全细胞平台 | 靛蓝合成酶Indigoidine synthetase | 0.065 mol/L | [ | |
| 无细胞平台 | 3WJdB | 0.1 nmol/L | [ | |||
| 全细胞平台 | 比色法 | 2.93 nmol/L | [ | |||
| 锑Sb(III) | ArsR/ArsR/SmtB | 1R23 | 全细胞平台 | eGFP | 0.25 µmol/L | [ |
| 镍Ni(II) | RcnR/CsoR-RcnR | - | 全细胞平台 | Lux | 80 nmol/L | [ |
| 芳香族化合物 Aromatic compound | 转录因子/家族 TFs/families | PDB | 反应平台 Reaction platform | 报告元件 Report component | 检测限 Limit of detection | 参考文献 Reference |
|---|---|---|---|---|---|---|
| 苯甲酸 | BenR/XylS-AraC | - | 无细胞平台 | sfGFP | 1 µmol/L | [ |
| - | 无细胞平台 | GFP | 21 702.8 nmol/L | [ | ||
| 苯 | DmpR/XylR-NtrC | 5KBE | 体外酶放大系统 | ATP酶 | 0.3 ppm | [ |
| 氯苯酚 | DmpR/XylR-NtrC | 全细胞平台 | Lac Z | 100 µmol/L | [ | |
| 苯并芘 | DmpR/XylR-NtrC | 全细胞平台 | eGFP | 2.5 ppb | [ | |
| 甲苯 | XylR/XylR-NtrC | 4FE4 | 全细胞平台 | 荧光素酶 | 1 µmol/L | [ |
| 2,4-二硝基甲苯 | XylR/XylR-NtrC | 全细胞平台 | GFP/lux | ~µmol/L | [ | |
| XylRv17/XylR-NtrC | - | 全细胞平台 | 荧光素酶/lacZ | - | [ | |
| 2-羟基-3',4'-二氯联苯 | HbpR/XylR-NtrC | - | 全细胞平台 | 荧光素酶 | 10 nmol/L | [ |
| 苯酚 | MopR/XylR-NtrC | 5KBE | 全细胞平台 | 荧光素酶 | 1 ppb | [ |
| 4-硝基苯酚 | Dm01/Dm12/XylR-NtrC | - | 全细胞平台 | RFP | 10 µmol/L | [ |
| 水杨酸 | SAR2349/NahR-LysR | 4EM0 | 无细胞平台 | 3WJdB荧光适配体/sfGFP | ~µmol/L | [ |
| 苯并芘 | SalR/NahR-LysR | - | 全细胞平台 | 荧光素酶 | 0.01 µmol/L | [ |
| 萘 | NahR/NahR-LysR | - | 全细胞平台 | 荧光素酶 | 50 nmol/L | [ |
| 水杨酸 | CmeR/TetR | - | 全细胞平台 | eGFP | ~µmol/L | [ |
| 苯扎氯氨 | QacR/TetR/CamR | 1JT0 | 无细胞平台 | 3WJdB荧光适配体 | ~µmol/L | [ |
| 苯甲酸盐 | MarR/MarR | 1JGS | 全细胞平台 | yEGFP | ~ fmol/L | [ |
| 苯甲醛 | BldR/MarR | 3F3X | 全细胞平台 | eGFP | ~µmol/L | [ |
| 3-羟基苯甲酸 | MobR/MarR | - | 无细胞平台 | 3WJdB荧光适配体/ | - | [ |
| 苯亚胂酸盐 | ArsR/ArsR | 6J05 | 全细胞平台 | GFP | 1 µmol/L | [ |
| 丙酮酸盐 | PdhR/GntR | - | 全细胞平台 | yEGFP | ~ fmol/L | [ |
表2 基于转录因子构建的生物传感器对芳香族化合物的检测
Table 2 Detection of aromatic compounds by transcription factor-based biosensors
| 芳香族化合物 Aromatic compound | 转录因子/家族 TFs/families | PDB | 反应平台 Reaction platform | 报告元件 Report component | 检测限 Limit of detection | 参考文献 Reference |
|---|---|---|---|---|---|---|
| 苯甲酸 | BenR/XylS-AraC | - | 无细胞平台 | sfGFP | 1 µmol/L | [ |
| - | 无细胞平台 | GFP | 21 702.8 nmol/L | [ | ||
| 苯 | DmpR/XylR-NtrC | 5KBE | 体外酶放大系统 | ATP酶 | 0.3 ppm | [ |
| 氯苯酚 | DmpR/XylR-NtrC | 全细胞平台 | Lac Z | 100 µmol/L | [ | |
| 苯并芘 | DmpR/XylR-NtrC | 全细胞平台 | eGFP | 2.5 ppb | [ | |
| 甲苯 | XylR/XylR-NtrC | 4FE4 | 全细胞平台 | 荧光素酶 | 1 µmol/L | [ |
| 2,4-二硝基甲苯 | XylR/XylR-NtrC | 全细胞平台 | GFP/lux | ~µmol/L | [ | |
| XylRv17/XylR-NtrC | - | 全细胞平台 | 荧光素酶/lacZ | - | [ | |
| 2-羟基-3',4'-二氯联苯 | HbpR/XylR-NtrC | - | 全细胞平台 | 荧光素酶 | 10 nmol/L | [ |
| 苯酚 | MopR/XylR-NtrC | 5KBE | 全细胞平台 | 荧光素酶 | 1 ppb | [ |
| 4-硝基苯酚 | Dm01/Dm12/XylR-NtrC | - | 全细胞平台 | RFP | 10 µmol/L | [ |
| 水杨酸 | SAR2349/NahR-LysR | 4EM0 | 无细胞平台 | 3WJdB荧光适配体/sfGFP | ~µmol/L | [ |
| 苯并芘 | SalR/NahR-LysR | - | 全细胞平台 | 荧光素酶 | 0.01 µmol/L | [ |
| 萘 | NahR/NahR-LysR | - | 全细胞平台 | 荧光素酶 | 50 nmol/L | [ |
| 水杨酸 | CmeR/TetR | - | 全细胞平台 | eGFP | ~µmol/L | [ |
| 苯扎氯氨 | QacR/TetR/CamR | 1JT0 | 无细胞平台 | 3WJdB荧光适配体 | ~µmol/L | [ |
| 苯甲酸盐 | MarR/MarR | 1JGS | 全细胞平台 | yEGFP | ~ fmol/L | [ |
| 苯甲醛 | BldR/MarR | 3F3X | 全细胞平台 | eGFP | ~µmol/L | [ |
| 3-羟基苯甲酸 | MobR/MarR | - | 无细胞平台 | 3WJdB荧光适配体/ | - | [ |
| 苯亚胂酸盐 | ArsR/ArsR | 6J05 | 全细胞平台 | GFP | 1 µmol/L | [ |
| 丙酮酸盐 | PdhR/GntR | - | 全细胞平台 | yEGFP | ~ fmol/L | [ |
| 抗生素 Antibiotic | 转录因子/家族 TFs/Families | PDB | 反应平台 Reaction platform | 报告元件 Report component | 检测限 Limit of detection | 参考文献 Reference |
|---|---|---|---|---|---|---|
| 四环素 | TetR/TetR | 1QPI | 限制性内切酶 | 荧光探针 | 25 nmol/L | [ |
| 全细胞平台 | EGFP和mCherry | 0.1 ng/mL | [ | |||
| aTF-NAST | RPA | 0.005 nmol/L | [ | |||
| 无细胞平台 | 荧光适配体 | ~125 nmol/L | [ | |||
| 无细胞平台 | G-四链体 | 18.08 ng/mL | [ | |||
| 无细胞平台 | 荧光适配体 | 10 nmol/L | [ | |||
| 无细胞平台 | CRISPR-Cas12a | - | [ | |||
| 无细胞平台 | 荧光素酶 | 45 nmol/L | [ | |||
| 脱水四环素 | QD-FRET | 光致发光 | 6 nmol/L | [ | ||
| QD-FRET | 光致发光 | 80 nmol/L | [ | |||
| 强力霉素 | 全细胞平台 | yEmRFP | 0.3 µg/mL | [ | ||
| 无细胞平台 | 荧光适配体 | ~0.2 µmol/L | [ | |||
| 放线菌素 | ActR/TetR | 3B6C | 全细胞平台 | - | [ | |
| 金霉素 | MphR/MphR | 3FRQ | 无细胞平台 | 荧光适配体 | ~125 nmol/L | [ |
| 放线菌素 | 全细胞平台 | GFP | - | [ | ||
| 红霉素 | 全细胞平台 | GFP | 13 nmol/L | [ | ||
| 无细胞平台 | 荧光适配体 | ~2.5 µmol/L | [ | |||
| 无细胞平台 | 荧光素酶 | 7.3 nmol/L | [ | |||
| 罗红霉素 | 无细胞平台 | 荧光适配体 | ~2.5 µmol/L | [ | ||
| 阿奇霉素 | 荧光适配体 | 荧光适配体 | ~2.5 µmol/L | [ | ||
| 叠氮红霉素 | 全细胞平台 | 荧光素酶 | - | [ | ||
| 竹桃霉素 | 全细胞平台 | 荧光素酶 | - | [ | ||
| 苦霉素 | 全细胞平台 | 荧光素酶 | - | [ | ||
| 克拉霉素 | MphR/MphR | 6U18 | 全细胞平台 | GFP | 1 µmol/L | [ |
| 无细胞平台 | 荧光适配体 | ~2.5 µmol/L | [ | |||
| 土霉素 | CtcS/MarR | 无细胞平台 | 荧光适配体 | ~125 nmol/L | [ | |
| 美罗培南 | AmpR/LysR | 全细胞平台 | mCherry | 8 pg/mL | [ | |
| 亚胺培南 | 全细胞平台 | mCherry | 40 pg/mL | [ |
表3 基于转录因子生物传感器对抗生素的检测
Table 3 Detection of antibiotic by transcription factor-based biosensors
| 抗生素 Antibiotic | 转录因子/家族 TFs/Families | PDB | 反应平台 Reaction platform | 报告元件 Report component | 检测限 Limit of detection | 参考文献 Reference |
|---|---|---|---|---|---|---|
| 四环素 | TetR/TetR | 1QPI | 限制性内切酶 | 荧光探针 | 25 nmol/L | [ |
| 全细胞平台 | EGFP和mCherry | 0.1 ng/mL | [ | |||
| aTF-NAST | RPA | 0.005 nmol/L | [ | |||
| 无细胞平台 | 荧光适配体 | ~125 nmol/L | [ | |||
| 无细胞平台 | G-四链体 | 18.08 ng/mL | [ | |||
| 无细胞平台 | 荧光适配体 | 10 nmol/L | [ | |||
| 无细胞平台 | CRISPR-Cas12a | - | [ | |||
| 无细胞平台 | 荧光素酶 | 45 nmol/L | [ | |||
| 脱水四环素 | QD-FRET | 光致发光 | 6 nmol/L | [ | ||
| QD-FRET | 光致发光 | 80 nmol/L | [ | |||
| 强力霉素 | 全细胞平台 | yEmRFP | 0.3 µg/mL | [ | ||
| 无细胞平台 | 荧光适配体 | ~0.2 µmol/L | [ | |||
| 放线菌素 | ActR/TetR | 3B6C | 全细胞平台 | - | [ | |
| 金霉素 | MphR/MphR | 3FRQ | 无细胞平台 | 荧光适配体 | ~125 nmol/L | [ |
| 放线菌素 | 全细胞平台 | GFP | - | [ | ||
| 红霉素 | 全细胞平台 | GFP | 13 nmol/L | [ | ||
| 无细胞平台 | 荧光适配体 | ~2.5 µmol/L | [ | |||
| 无细胞平台 | 荧光素酶 | 7.3 nmol/L | [ | |||
| 罗红霉素 | 无细胞平台 | 荧光适配体 | ~2.5 µmol/L | [ | ||
| 阿奇霉素 | 荧光适配体 | 荧光适配体 | ~2.5 µmol/L | [ | ||
| 叠氮红霉素 | 全细胞平台 | 荧光素酶 | - | [ | ||
| 竹桃霉素 | 全细胞平台 | 荧光素酶 | - | [ | ||
| 苦霉素 | 全细胞平台 | 荧光素酶 | - | [ | ||
| 克拉霉素 | MphR/MphR | 6U18 | 全细胞平台 | GFP | 1 µmol/L | [ |
| 无细胞平台 | 荧光适配体 | ~2.5 µmol/L | [ | |||
| 土霉素 | CtcS/MarR | 无细胞平台 | 荧光适配体 | ~125 nmol/L | [ | |
| 美罗培南 | AmpR/LysR | 全细胞平台 | mCherry | 8 pg/mL | [ | |
| 亚胺培南 | 全细胞平台 | mCherry | 40 pg/mL | [ |
| [1] |
杨克恭. 细菌有转录因子吗? —“转录因子” 概念的形成和发展浅析[J]. 中国生物化学与分子生物学报, 2021, 37(6): 691-696.
doi: 10.13865/j.cnki.cjbmb.2021.05.1130 |
| Yang KG. Do bacteria have the transcription factors? The formation and development of the concept of “transcription factor”[J]. Chin J Biochem Mol Biol, 2021, 37(6): 691-696. | |
| [2] | 卜恺璇, 周翠霞, 路福平, 等. 细菌转录起始调控机制[J]. 中国生物工程杂志, 2021, 41(11): 89-99. |
| Bu KX, Zhou CX, Lu FP, et al. Research on the regulation mechanism of bacterial transcription initiation[J]. China Biotechnol, 2021, 41(11): 89-99. | |
| [3] |
杨璐, 吴楠, 白茸茸, 等. 基因回路型全细胞微生物传感器的设计、优化与应用[J]. 合成生物学, 2022, 3(6): 1061-1080.
doi: 10.12211/2096-8280.2021-021 |
| Yang L, Wu N, Bai RR, et al. Design, optimization and application of whole-cell microbial biosensors with engineered genetic circuits[J]. Synth Biol J, 2022, 3(6): 1061-1080. | |
| [4] | 赵晓蕊, 陈慧芳, 雷春阳, 等. 无细胞生物传感技术在环境污染物检测方面的应用进展[J]. 分析化学, 2023, 51(8): 1223-1231. |
| Zhao XR, Chen HF, Lei CY, et al. Advances in application of cell-free biosensing technologies for detection of environmental pollutant[J]. Chin J Anal Chem, 2023, 51(8): 1223-1231. | |
| [5] | Gu YQ, Fan F, Liu Y, et al. Cell-free protein synthesis system for bioanalysis: advances in methods and applications[J]. Trac Trends Anal Chem, 2023, 161: 117015. |
| [6] | Li SS, Li ZL, Tan GY, et al. In vitro allosteric transcription factor-based biosensing[J]. Trends Biotechnol, 2023, 41(8): 1080-1095. |
| [7] | Li SS, Zhou L, Yao YP, et al. A platform for the development of novel biosensors by configuring allosteric transcription factor recognition with amplified luminescent proximity homogeneous assays[J]. Chem Commun, 2017, 53(1): 99-102. |
| [8] | Nguyen TT, Chern M, Baer RC, et al. A Förster resonance energy transfer-based ratiometric sensor with the allosteric transcription factor TetR[J]. Small, 2020, 16(17): e1907522. |
| [9] | Yao YP, Li SS, Cao JQ, et al. Development of small molecule biosensors by coupling the recognition of the bacterial allosteric transcription factor with isothermal strand displacement amplification[J]. Chem Commun, 2018, 54(38): 4774-4777. |
| [10] |
Yao YP, Li SS, Cao JQ, et al. A novel signal transduction system for development of uric acid biosensors[J]. Appl Microbiol Biotechnol, 2018, 102(17): 7489-7497.
doi: 10.1007/s00253-018-9056-8 pmid: 29961098 |
| [11] | Cao JQ, Yao YP, Fan KQ, et al. Harnessing a previously unidentified capability of bacterial allosteric transcription factors for sensing diverse small molecules in vitro[J]. Sci Adv, 2018, 4(11): eaau4602. |
| [12] | Iwasaki RS, Batey RT. SPRINT: a Cas13a-based platform for detection of small molecules[J]. Nucleic Acids Res, 2020, 48(17): e101. |
| [13] |
Eddy SR. Computational genomics of noncoding RNA genes[J]. Cell, 2002, 109(2): 137-140.
pmid: 12007398 |
| [14] | Gajiwala KS, Burley SK. Winged helix proteins[J]. Curr Opin Struct Biol, 2000, 10(1): 110-116. |
| [15] |
Schreiter ER, Drennan CL. Ribbon-helix-helix transcription factors: variations on a theme[J]. Nat Rev Microbiol, 2007, 5(9): 710-720.
doi: 10.1038/nrmicro1717 pmid: 17676053 |
| [16] |
Yu QK, Ren KW, You MX. Genetically encoded RNA nanodevices for cellular imaging and regulation[J]. Nanoscale, 2021, 13(17): 7988-8003.
doi: 10.1039/d0nr08301a pmid: 33885099 |
| [17] |
Paige JS, Wu KY, Jaffrey SR. RNA mimics of green fluorescent protein[J]. Science, 2011, 333(6042): 642-646.
doi: 10.1126/science.1207339 pmid: 21798953 |
| [18] | Jung JK, Alam KK, Verosloff MS, et al. Cell-free biosensors for rapid detection of water contaminants[J]. Nat Biotechnol, 2020, 38(12): 1451-1459. |
| [19] | Qiu CX, Zhai HT, Hou J. Biosensors design in yeast and applications in metabolic engineering[J]. FEMS Yeast Res, 2019, 19(8): foz082. |
| [20] |
Mukherjee K, Bhattacharyya S, Peralta-Yahya P. GPCR-based chemical biosensors for medium-chain fatty acids[J]. ACS Synth Biol, 2015, 4(12): 1261-1269.
doi: 10.1021/sb500365m pmid: 25992593 |
| [21] |
Dong CY, Ly C, Dunlap LE, et al. Psychedelic-inspired drug discovery using an engineered biosensor[J]. Cell, 2021, 184(10): 2779-2792.e18.
doi: 10.1016/j.cell.2021.03.043 pmid: 33915107 |
| [22] | Dhakal S, Macreadie I. The use of yeast in biosensing[J]. Microorganisms, 2022, 10(9): 1772. |
| [23] | Wahid E, Ocheja OB, Marsili E, et al. Biological and technical challenges for implementation of yeast-based biosensors[J]. Microb Biotechnol, 2023, 16(1): 54-66. |
| [24] | Cerminati S, Soncini FC, Checa SK. A sensitive whole-cell biosensor for the simultaneous detection of a broad-spectrum of toxic heavy metal ions[J]. Chem Commun, 2015, 51(27): 5917-5920. |
| [25] | Mendoza JI, Soncini FC, Checa SK. Engineering of a Au-sensor to develop a Hg-specific, sensitive and robust whole-cell biosensor for on-site water monitoring[J]. Chem Commun, 2020, 56(48): 6590-6593. |
| [26] |
Chong HQ, Ching CB. Development of colorimetric-based whole-cell biosensor for organophosphorus compounds by engineering transcription regulator DmpR[J]. ACS Synth Biol, 2016, 5(11): 1290-1298.
pmid: 27346389 |
| [27] | Jha RK, Bingen JM, Johnson CW, et al. A protocatechuate biosensor for Pseudomonas putida KT2440 via promoter and protein evolution[J]. Metab Eng Commun, 2018, 6: 33-38. |
| [28] | Pu W, Chen JZ, Liu P, et al. Directed evolution of linker helix as an efficient strategy for engineering LysR-type transcriptional regulators as whole-cell biosensors[J]. Biosens Bioelectron, 2023, 222: 115004. |
| [29] | Zhang P, Yang MJ, Lan JJ, et al. Water quality degradation due to heavy metal contamination: health impacts and eco-friendly approaches for heavy metal remediation[J]. Toxics, 2023, 11(10): 828. |
| [30] | 朱振宇, 华垚堃, 胡婷婷, 等. 微生物金属响应蛋白研究进展[J]. 微生物学通报, 2018, 45(8): 1794-1803. |
| Zhu ZY, Hua YK, Hu TT, et al. Advances in microbial metal response proteins[J]. Microbiol China, 2018, 45(8): 1794-1803. | |
| [31] | Guo Y, Hui CY, Liu LS, et al. Development of a bioavailable Hg(II)sensing system based on MerR-regulated visual pigment biosynthesis[J]. Sci Rep, 2021, 11(1): 13516. |
| [32] | Guo MZ, Wang JL, Du RX, et al. A test strip platform based on a whole-cell microbial biosensor for simultaneous on-site detection of total inorganic mercury pollutants in cosmetics without the need for predigestion[J]. Biosens Bioelectron, 2020, 150: 111899. |
| [33] | Wang D, Zheng YN, Xu LN, et al. Engineered cells for selective detection and remediation of Hg2+ based on transcription factor MerR regulated cell surface displayed systems[J]. Biochem Eng J, 2019, 150: 107289. |
| [34] |
Bereza-Malcolm L, Aracic S, Kannan RB, et al. Functional characterization of Gram-negative bacteria from different Genera as multiplex cadmium biosensors[J]. Biosens Bioelectron, 2017, 94: 380-387.
doi: S0956-5663(17)30180-X pmid: 28319906 |
| [35] | Tang TC, Tham E, Liu XY, et al. Hydrogel-based biocontainment of bacteria for continuous sensing and computation[J]. Nat Chem Biol, 2021, 17(6): 724-731. |
| [36] |
Wang BJ, Barahona M, Buck M. A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals[J]. Biosens Bioelectron, 2013, 40(1): 368-376.
doi: 10.1016/j.bios.2012.08.011 pmid: 22981411 |
| [37] | Cayron J, Prudent E, Escoffier C, et al. Pushing the limits of nickel detection to nanomolar range using a set of engineered bioluminescent Escherichia coli[J]. Environ Sci Pollut Res Int, 2017, 24(1): 4-14. |
| [38] |
Kim HJ, Jeong H, Lee SJ. Synthetic biology for microbial heavy metal biosensors[J]. Anal Bioanal Chem, 2018, 410(4): 1191-1203.
doi: 10.1007/s00216-017-0751-6 pmid: 29184994 |
| [39] | Zhao XY, Dong T. A microfluidic device for continuous sensing of systemic acute toxicants in drinking water[J]. Int J Environ Res Public Health, 2013, 10(12): 6748-6763. |
| [40] | Zhang YK, Zhao C, Bi HX, et al. A cell-free paper-based biosensor dependent on allosteric transcription factors(aTFs)for on-site detection of harmful metals Hg2+ and Pb2+ in water[J]. J Hazard Mater, 2022, 438: 129499. |
| [41] | Lin PH, Tsai ST, Chang YC, et al. Harnessing split fluorescent proteins in modular protein logic for advanced whole-cell detection[J]. Anal Chim Acta, 2023, 1275: 341593. |
| [42] | Elcin E, Öktem HA. Inorganic cadmium detection using a fluorescent whole-cell bacterial bioreporter[J]. Anal Lett, 2020, 53(17): 2715-2733. |
| [43] | Kim H, Lee W, Yoon Y. Heavy metal(loid)biosensor based on split-enhanced green fluorescent protein: development and characterization[J]. Appl Microbiol Biotechnol, 2019, 103(15): 6345-6352. |
| [44] | Hou QH, Ma AZ, Wang T, et al. Detection of bioavailable cadmium, lead, and arsenic in polluted soil by tailored multiple Escherichia coli whole-cell sensor set[J]. Anal Bioanal Chem, 2015, 407(22): 6865-6871. |
| [45] |
Kim HJ, Lim JW, Jeong H, et al. Development of a highly specific and sensitive cadmium and lead microbial biosensor using synthetic CadC-T7 genetic circuitry[J]. Biosens Bioelectron, 2016, 79: 701-708.
doi: 10.1016/j.bios.2015.12.101 pmid: 26773374 |
| [46] | Kumar S, Verma N, Singh AK. Development of cadmium specific recombinant biosensor and its application in milk samples[J]. Sens Actuat B Chem, 2017, 240: 248-254. |
| [47] | Jia XQ, Liu T, Ma YB, et al. Construction of cadmium whole-cell biosensors and circuit amplification[J]. Appl Microbiol Biotechnol, 2021, 105(13): 5689-5699. |
| [48] | Fernández M, Morel B, Ramos JL, et al. Paralogous regulators ArsR1 and ArsR2 of Pseudomonas putida KT2440 as a basis for arsenic biosensor development[J]. Appl Environ Microbiol, 2016, 82(14): 4133-4144. |
| [49] | Chen PH, Lin C, Guo KH, et al. Development of a pigment-based whole-cell biosensor for the analysis of environmental copper[J]. RSC Adv, 2017, 7(47): 29302-29305. |
| [50] | Kang Y, Lee W, Kim S, et al. Enhancing the copper-sensing capability of Escherichia coli-based whole-cell bioreporters by genetic engineering[J]. Appl Microbiol Biotechnol, 2018, 102(3): 1513-1521. |
| [51] |
Date A, Pasini P, Daunert S. Construction of spores for portable bacterial whole-cell biosensing systems[J]. Anal Chem, 2007, 79(24): 9391-9397.
pmid: 18020369 |
| [52] | Yoon Y, Kim S, Chae Y, et al. Use of tunable whole-cell bioreporters to assess bioavailable cadmium and remediation performance in soils[J]. PLoS One, 2016, 11(5): e0154506. |
| [53] |
Yoon Y, Kim S, Chae Y, et al. Simultaneous detection of bioavailable arsenic and cadmium in contaminated soils using dual-sensing bioreporters[J]. Appl Microbiol Biotechnol, 2016, 100(8): 3713-3722.
doi: 10.1007/s00253-016-7338-6 pmid: 26852408 |
| [54] |
Ghataora JS, Gebhard S, Reeksting BJ. Chimeric MerR-family regulators and logic elements for the design of metal sensitive genetic circuits in Bacillus subtilis[J]. ACS Synth Biol, 2023, 12(3): 735-749.
doi: 10.1021/acssynbio.2c00545 pmid: 36629785 |
| [55] | Hui CY, Guo Y, Li LM, et al. Indigoidine biosynthesis triggered by the heavy metal-responsive transcription regulator: a visual whole-cell biosensor[J]. Appl Microbiol Biotechnol, 2021, 105(14-15): 6087-6102. |
| [56] | Hui CY, Guo Y, Zhu DL, et al. Metabolic engineering of the violacein biosynthetic pathway toward a low-cost, minimal-equipment lead biosensor[J]. Biosens Bioelectron, 2022, 214: 114531. |
| [57] |
Lee W, Kim H, Jang G, et al. Antimony sensing whole-cell bioreporters derived from ArsR genetic engineering[J]. Appl Microbiol Biotechnol, 2020, 104(6): 2691-2699.
doi: 10.1007/s00253-020-10413-5 pmid: 32002600 |
| [58] |
Muir T, Michalek JE, Palmer RF. Determination of safe levels of persistent organic pollutants in toxicology and epidemiology[J]. Rev Environ Health, 2022, 38(3): 401-408.
doi: 10.1515/reveh-2021-0105 pmid: 35506713 |
| [59] | 郭庆伟, 王倩, 张海东, 等. 新污染物检测技术研究进展[J]. 化学通报, 2024, 87(1): 78-85. |
| Guo QW, Wang Q, Zhang HD, et al. Research progress in detection technology of emerging pollutants[J]. Chemistry, 2024, 87(1): 78-85. | |
| [60] | Sun SW, Peng KL, Sun S, et al. Engineering modular and highly sensitive cell-based biosensors for aromatic contaminant monitoring and high-throughput enzyme screening[J]. ACS Synth Biol, 2023, 12(3): 877-891. |
| [61] | Darjee SM, Modi KM, Panchal U, et al. Highly selective and sensitive fluorescent sensor: Thiacalix[4]arene-1-naphthalene carboxylate for Zn2+ ions[J]. J Mol Struct, 2017, 1133: 1-8. |
| [62] |
Sun YJ, Zhao XH, Zhang DY, et al. New naphthalene whole-cell bioreporter for measuring and assessing naphthalene in polycyclic aromatic hydrocarbons contaminated site[J]. Chemosphere, 2017, 186: 510-518.
doi: S0045-6535(17)31248-1 pmid: 28810221 |
| [63] | Roy R, Ray S, Chowdhury A, et al. Tunable multiplexed whole-cell biosensors as environmental diagnostics for ppb-level detection of aromatic pollutants[J]. ACS Sens, 2021, 6(5): 1933-1939. |
| [64] | Juárez JF, Lecube-Azpeitia B, Brown SL, et al. Biosensor libraries harness large classes of binding domains for construction of allosteric transcriptional regulators[J]. Nat Commun, 2018, 9(1): 3101. |
| [65] | Voyvodic PL, Pandi A, Koch M, et al. Plug-and-play metabolic transducers expand the chemical detection space of cell-free biosensors[J]. Nat Commun, 2019, 10(1): 1697. |
| [66] |
Zhang P, Feng HB, Yang JZ, et al. Detection of inorganic ions and organic molecules with cell-free biosensing systems[J]. J Biotechnol, 2019, 300: 78-86.
doi: S0168-1656(19)30175-0 pmid: 31141711 |
| [67] | Ray S, Panjikar S, Anand R. Design of protein-based biosensors for selective detection of benzene groups of pollutants[J]. ACS Sens, 2018, 3(9): 1632-1638. |
| [68] | Campos VL, Zaror CA, Mondaca MA. Detection of chlorinated phenols in kraft pulp bleaching effluents using DmpR mutant strains[J]. Bull Environ Contam Toxicol, 2004, 73(4): 666-673. |
| [69] |
Behzadian F, Barjeste H, Hosseinkhani S, et al. Construction and characterization of Escherichia coli whole-cell biosensors for toluene and related compounds[J]. Curr Microbiol, 2011, 62(2): 690-696.
doi: 10.1007/s00284-010-9764-5 pmid: 20872219 |
| [70] |
Garmendia J, de las Heras A, Galvão TC, et al. Tracing explosives in soil with transcriptional regulators of Pseudomonas putida evolved for responding to nitrotoluenes[J]. Microb Biotechnol, 2008, 1(3): 236-246.
doi: 10.1111/j.1751-7915.2008.00027.x pmid: 21261843 |
| [71] | de Las Heras A, de Lorenzo V. Cooperative amino acid changes shift the response of the σ54-dependent regulator XylR from natural m-xylene towards xenobiotic 2, 4-dinitrotoluene[J]. Mol Microbiol, 2011, 79(5): 1248-1259. |
| [72] |
Turner K, Xu SF, Pasini P, et al. Hydroxylated polychlorinated biphenyl detection based on a genetically engineered bioluminescent whole-cell sensing system[J]. Anal Chem, 2007, 79(15): 5740-5745.
pmid: 17602671 |
| [73] | Werlen C, Jaspers MCM, van der Meer JR. van der Meer JR. Measurement of biologically available naphthalene in gas and aqueous phases by use of a Pseudomonas putida biosensor[J]. Appl Environ Microbiol, 2004, 70(1): 43-51. |
| [74] |
Nasr MA, Timmins LR, Martin VJJ, et al. A versatile transcription factor biosensor system responsive to multiple aromatic and indole inducers[J]. ACS Synth Biol, 2022, 11(4): 1692-1698.
doi: 10.1021/acssynbio.2c00063 pmid: 35316041 |
| [75] | Asemoloye MD, Marchisio MA. Synthetic metabolic transducers in Saccharomyces cerevisiae as sensors for aromatic permeant acids and bioreporters of hydrocarbon metabolism[J]. Biosens Bioelectron, 2023, 220: 114897. |
| [76] |
Fiorentino G, Ronca R, Bartolucci S. A novel E. coli biosensor for detecting aromatic aldehydes based on a responsive inducible archaeal promoter fused to the green fluorescent protein[J]. Appl Microbiol Biotechnol, 2009, 82(1): 67-77.
doi: 10.1007/s00253-008-1771-0 pmid: 18998120 |
| [77] | Chen J, Sun S, Li CZ, et al. Biosensor for organoarsenical herbicides and growth promoters[J]. Environ Sci Technol, 2014, 48(2): 1141-1147. |
| [78] | Anomaly J. Harm to others: the social cost of antibiotics in agriculture[J]. J Agric Environ Ethics, 2009, 22(5): 423-435. |
| [79] |
Founou LL, Founou RC, Essack SY. Antibiotic resistance in the food chain: a developing country-perspective[J]. Front Microbiol, 2016, 7: 1881.
pmid: 27933044 |
| [80] | Chern M, Garden PM, Baer RC, et al. Transcription factor based small-molecule sensing with a rapid cell phone enabled fluorescent bead assay[J]. Angew Chem Int Ed Engl, 2020, 59(48): 21597-21602. |
| [81] | Zhang R, Wang Y, Deng HF, et al. Fast and bioluminescent detection of antibiotic contaminants by on-demand transcription of RNA scaffold arrays[J]. Anal Chim Acta, 2023, 1273: 341538. |
| [82] | Richter MF, Drown BS, Riley AP, et al. Predictive compound accumulation rules yield a broad-spectrum antibiotic[J]. Nature, 2017, 545(7654): 299-304. |
| [83] |
Miller CA, Ho JM, Parks SE, et al. Macrolide biosensor optimization through cellular substrate sequestration[J]. ACS Synth Biol, 2021, 10(2): 258-264.
doi: 10.1021/acssynbio.0c00572 pmid: 33555859 |
| [84] | Li SL, Chen DD, Liu ZQ, et al. Directed evolution of TetR for constructing sensitive and broad-spectrum tetracycline antibiotics whole-cell biosensor[J]. J Hazard Mater, 2023, 460: 132311. |
| [85] |
Rodríguez-Serrano AF, Hsing IM. Allosteric regulation of DNA circuits enables minimal and rapid biosensors of small molecules[J]. ACS Synth Biol, 2021, 10(2): 371-378.
doi: 10.1021/acssynbio.0c00545 pmid: 33481567 |
| [86] | Ullrich T, Weirich S, Jeltsch A. Development of an epigenetic tetracycline sensor system based on DNA methylation[J]. PLoS One, 2020, 15(5): e0232701. |
| [87] | Liu RN, Liu X, Yang H, et al. A cell-free biosensor based on strand displacement amplification and hybridization chain reaction for fluorescence detection of tetracycline[J]. Microchem J, 2023, 185: 108239. |
| [88] | Bi HX, Zhao C, Zhang YK, et al. IVT cell-free biosensors for tetracycline and macrolide detection based on allosteric transcription factors(aTFs)[J]. Anal Methods, 2022, 14(44): 4545-4554. |
| [89] |
Mahas A, Wang QC, Marsic T, et al. Development of Cas12a-based cell-free small-molecule biosensors via allosteric regulation of CRISPR array expression[J]. Anal Chem, 2022, 94(11): 4617-4626.
doi: 10.1021/acs.analchem.1c04332 pmid: 35266687 |
| [90] | Chen MF, Nguyen TT, Varongchayakul N, et al. Surface immobilized nucleic acid-transcription factor quantum dots for biosensing[J]. Adv Healthc Mater, 2020, 9(17): e2000403. |
| [91] | Miller RA, Brown G, Barron E, et al. Development of a paper-immobilized yeast biosensor for the detection of physiological concentrations of doxycycline in technology-limited settings[J]. Anal Methods, 2020, 12(16): 2123-2132. |
| [92] | Ahn SK, Tahlan K, Yu Z, et al. Investigation of transcription repression and small-molecule responsiveness by TetR-like transcription factors using a heterologous Escherichia coli-based assay[J]. J Bacteriol, 2007, 189(18): 6655-6664. |
| [93] |
Kasey CM, Zerrad M, Li YW, et al. Development of transcription factor-based designer macrolide biosensors for metabolic engineering and synthetic biology[J]. ACS Synth Biol, 2018, 7(1): 227-239.
doi: 10.1021/acssynbio.7b00287 pmid: 28950701 |
| [94] | Möhrle V, Stadler M, Eberz G. Biosensor-guided screening for macrolides[J]. Anal Bioanal Chem, 2007, 388(5/6): 1117-1125. |
| [95] |
Li YW, Reed M, Wright HT, et al. Development of genetically encoded biosensors for reporting the methyltransferase-dependent biosynthesis of semisynthetic macrolide antibiotics[J]. ACS Synth Biol, 2021, 10(10): 2520-2531.
doi: 10.1021/acssynbio.1c00151 pmid: 34546703 |
| [96] | Higuera-Llantén S, Alcalde-Rico M, Vasquez-Ponce F, et al. A whole-cell hypersensitive biosensor for beta-lactams based on the AmpR-AmpC regulatory circuit from the Antarctic Pseudomonas sp. IB20[J]. Microb Biotechnol, 2024, 17(1): e14385. |
| [97] | Liu CJ, Yu H, Zhang BC, et al. Engineering whole-cell microbial biosensors: design principles and applications in monitoring and treatment of heavy metals and organic pollutants[J]. Biotechnol Adv, 2022, 60: 108019. |
| [1] | 胡雅丹, 伍国强, 刘晨, 魏明. MYB转录因子在调控植物响应逆境胁迫中的作用[J]. 生物技术通报, 2024, 40(6): 5-22. |
| [2] | 王秋月, 段鹏亮, 李海笑, 刘宁, 曹志艳, 董金皋. 玉米大斑病菌cDNA文库的构建及转录因子StMR1互作蛋白的筛选[J]. 生物技术通报, 2024, 40(6): 281-289. |
| [3] | 王健, 杨莎, 孙庆文, 陈宏宇, 杨涛, 黄园. 金钗石斛bHLH转录因子家族全基因组鉴定及表达分析[J]. 生物技术通报, 2024, 40(6): 203-218. |
| [4] | 王迪, 张晓宇, 宋宇鑫, 郑东然, 田静, 李玉花, 王宇, 吴昊. 细胞全能性转录因子调控植物组培再生的分子机制研究进展[J]. 生物技术通报, 2024, 40(6): 23-33. |
| [5] | 李慧, 文钰芳, 王悦, 纪超, 石国优, 罗英, 周勇, 李志敏, 吴晓玉, 杨有新, 刘建萍. 盐胁迫下辣椒CaPIF4的表达特性与功能分析[J]. 生物技术通报, 2024, 40(4): 148-158. |
| [6] | 肖雅茹, 贾婷婷, 罗丹, 武喆, 李丽霞. 黄瓜CsERF025L转录因子的克隆及表达分析[J]. 生物技术通报, 2024, 40(4): 159-166. |
| [7] | 郭纯, 宋桂梅, 闫艳, 邸鹏, 王英平. 西洋参bZIP基因家族全基因组鉴定和表达分析[J]. 生物技术通报, 2024, 40(4): 167-178. |
| [8] | 谢倩, 江来, 贺进, 刘玲玲, 丁明月, 陈清西. 不同鲜食品质橄榄果实转录组测序及酚类代谢途径相关调控基因挖掘[J]. 生物技术通报, 2024, 40(3): 215-228. |
| [9] | 陈艳梅. 蛋白质翻译后修饰之间的互作关系及其协同调控机理[J]. 生物技术通报, 2024, 40(2): 1-8. |
| [10] | 杨艳, 胡洋, 刘霓如, 殷璐, 杨锐, 王鹏飞, 穆霄鹏, 张帅, 程春振, 张建成. ‘红满堂’苹果MbbZIP43基因的克隆与功能研究[J]. 生物技术通报, 2024, 40(2): 146-159. |
| [11] | 路喻丹, 刘晓驰, 冯新, 陈桂信, 陈义挺. 猕猴桃BBX基因家族成员鉴定与转录特征分析[J]. 生物技术通报, 2024, 40(2): 172-182. |
| [12] | 邹修为, 岳佳妮, 李志宇, 戴良英, 李魏. 水稻热激转录因子HsfA2b调控非生物胁迫抗性的功能分析[J]. 生物技术通报, 2024, 40(2): 90-98. |
| [13] | 周会汶, 吴兰花, 韩德鹏, 郑伟, 余跑兰, 吴杨, 肖小军. 甘蓝型油菜种子硫苷含量全基因组关联分析[J]. 生物技术通报, 2024, 40(1): 222-230. |
| [14] | 吴圳, 张明英, 闫锋, 李依民, 高静, 颜永刚, 张岗. 掌叶大黄(Rheum palmatum L.)WRKY基因家族鉴定与分析[J]. 生物技术通报, 2024, 40(1): 250-261. |
| [15] | 王斌, 袁晓, 蒋园园, 王玉昆, 肖艳辉, 何金明. bHLH96的克隆及其在薄荷萜烯生物合成调控中的功能[J]. 生物技术通报, 2024, 40(1): 281-293. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||