








生物技术通报 ›› 2026, Vol. 42 ›› Issue (4): 345-356.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0879
• 研究报告 • 上一篇
颜鑫( ), 吴筱, 何思宇, 段玉欢, 仇伍霞, 袁晓琴, 毛新芳, 刘忠渊(
), 吴筱, 何思宇, 段玉欢, 仇伍霞, 袁晓琴, 毛新芳, 刘忠渊( )
)
收稿日期:2025-08-12
出版日期:2026-04-26
发布日期:2026-04-30
通讯作者:
刘忠渊,男,博士,教授,研究方向 :重组蛋白及合成生物学;E-mail: lzy1168@163.com作者简介:颜鑫,男,硕士,研究方向 :合成生物学;E-mail: 1992334217@qq.com
基金资助:
YAN Xin( ), WU Xiao, HE Si-yu, DUAN Yu-huan, QIU Wu-xia, YUAN Xiao-qin, MAO Xin-fang, LIU Zhong-yuan(
), WU Xiao, HE Si-yu, DUAN Yu-huan, QIU Wu-xia, YUAN Xiao-qin, MAO Xin-fang, LIU Zhong-yuan( )
)
Received:2025-08-12
Published:2026-04-26
Online:2026-04-30
摘要:
目的 原核共表达甜橙(Citrus sinensis)肌醇磷酸合酶(myo-inositol phosphate synthase)基因Csino3与大肠杆菌MG1655(Escherichia coli MG1655)肌醇-1-单磷酸酶(inositol-1-monophosphatas)基因suhB,构建高效的肌醇生物合成途径,并通过优化发酵条件和敲除大肠杆菌的pgi和gldA基因以提高肌醇产量。 方法 构建原核表达载体pETDuet-1-Csino3-suhB,将其转化至大肠杆菌E. coli BL21(DE3)进行异源表达。通过IPTG诱导表达,并采用SDS-PAGE鉴定可溶性蛋白表达情况。对葡萄糖浓度、IPTG浓度、接种量和初始pH值4个发酵参数进行优化,在单因素实验基础上,通过响应面法进一步优化发酵条件。随后通过敲除pgi(葡萄糖-6-磷酸异构酶)、gldA(甘油脱氢酶)改造大肠杆菌代谢途径,进一步提高肌醇产量。 结果 通过在E.coli BL21(DE3)中异源共表达重组质粒pETDuet-1-Csino3-suhB,证实Csino3和SuhB蛋白在25 ℃下均以可溶性形式存在。采用单因素实验和响应面分析法对目的菌株优化发酵条件,得出最佳发酵条件为:葡萄糖浓度9.7 g/L、IPTG浓度0.5 mmol/L、接种量8%、初始pH值8.1,此时肌醇发酵产量为309 mg/L。在上述发酵条件的基础上对敲除菌株ΔgldAΔpgi-pETDuet-1-Csino3-suhB/E. coli BL21(DE3)进行发酵优化,在添加20 g/L的初始甘油下,肌醇产量达到最大值3.97 g/L,比敲除前提高了12.8倍左右。 结论 重组菌株pETDuet-1-Csino3-suhB /E.coli BL21(DE3)能够有效产生肌醇,通过敲除pgi、gldA基因改变大肠杆菌代谢通路能极大提高肌醇产量。
颜鑫, 吴筱, 何思宇, 段玉欢, 仇伍霞, 袁晓琴, 毛新芳, 刘忠渊. 基于甜橙肌醇磷酸合酶基因Csino3的肌醇生物合成及发酵优化[J]. 生物技术通报, 2026, 42(4): 345-356.
YAN Xin, WU Xiao, HE Si-yu, DUAN Yu-huan, QIU Wu-xia, YUAN Xiao-qin, MAO Xin-fang, LIU Zhong-yuan. Inositol Biosynthesis and Fermentation Optimization of an Inositol Biosynthesis Pathway Using Citrus sinensis Inositol Phosphate Synthase Gene Csino3[J]. Biotechnology Bulletin, 2026, 42(4): 345-356.
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) |
|---|---|
| pgi-outF | CCACTGTGCCTGAACATCGCGTTCGC |
| pgi-outR | GCAGGTCGCTTCACGACTCTTCTCCAGAGG |
| pgi-inF | GCTGCACGTAGCGCTGCGTAACCG |
| pgi-inR | GCACTTCCGCGATGTGAGTCCCATCGAC |
| gldA-outF | CCCCACCATCCGCCAGTTAAACAGCA |
| gldA-outR | CTGGACACCGCTAACGTCGCAGAAGTC |
| gldA-inF | GGAGTCACTACATGCTGTTCGGCAG |
| gldA-inR | CTACACCGATGAGGGTGAGTTTGACC |
表1 敲除基因pgi、gldA所用引物序列
Table 1 Primer sequences for knockout of genes pgi and gldA
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) |
|---|---|
| pgi-outF | CCACTGTGCCTGAACATCGCGTTCGC |
| pgi-outR | GCAGGTCGCTTCACGACTCTTCTCCAGAGG |
| pgi-inF | GCTGCACGTAGCGCTGCGTAACCG |
| pgi-inR | GCACTTCCGCGATGTGAGTCCCATCGAC |
| gldA-outF | CCCCACCATCCGCCAGTTAAACAGCA |
| gldA-outR | CTGGACACCGCTAACGTCGCAGAAGTC |
| gldA-inF | GGAGTCACTACATGCTGTTCGGCAG |
| gldA-inR | CTACACCGATGAGGGTGAGTTTGACC |
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 限制性酶切位点 Restriction enzyme site |
|---|---|---|
| CsIno3上游 | BamH I | |
| CsIno3下游 | TTGTACTAAAACCTTATGTTCACT | Hind III |
| suhB 上游 | Nde I | |
| suhB 下游 | GAGTTAAGCGACGCTCTGAAGCGT | Xho I |
表2 pETDuet-1- Csino3-suhB质粒克隆位点
Table 2 Cloning sites of pETDuet-1-Csino3-suhB plasmid
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 限制性酶切位点 Restriction enzyme site |
|---|---|---|
| CsIno3上游 | BamH I | |
| CsIno3下游 | TTGTACTAAAACCTTATGTTCACT | Hind III |
| suhB 上游 | Nde I | |
| suhB 下游 | GAGTTAAGCGACGCTCTGAAGCGT | Xho I |
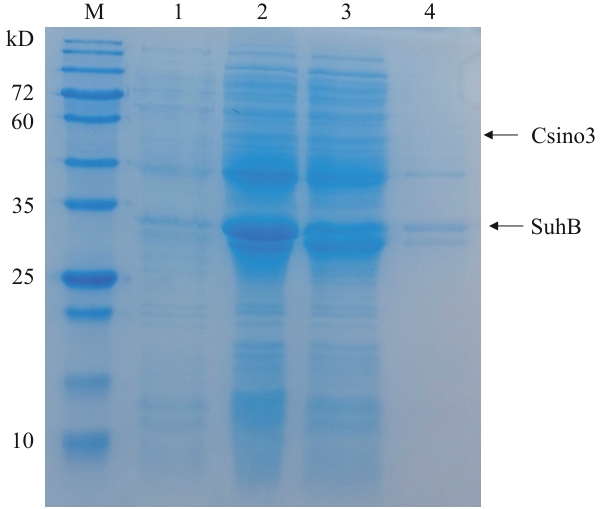
图1 重组pETDuet-1-Csino3-suhB/BL21(DE3)在25 ℃的诱导表达M:蛋白质分子量标准;1:IPTG诱导前细菌裂解液;2:IPTG诱导后的细菌裂解液;3:超声破碎后上清液;4:超声破碎后沉淀
Fig. 1 Induced expression of recombinant pETDuet-1-Csino3-suhB/BL21(DE3) at 25 ℃M: Protein molecular weight marker. 1: Lysis of bacteria without induction by IPTG. 2: Lysis of bacteria after induction by IPTG. 3: Supernatant after ultrasonic crushing. 4: Sedimentation after ultrasonic crushing
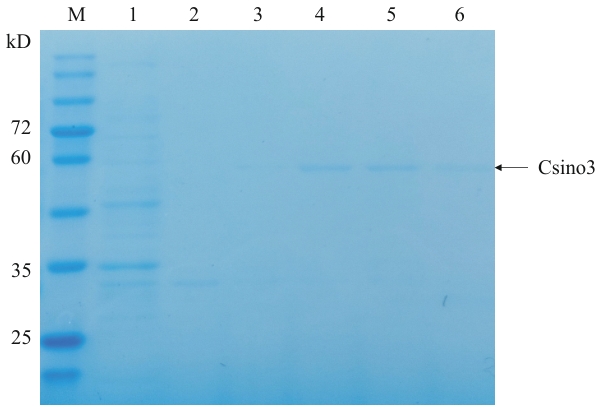
图2 Csino3蛋白纯化M:蛋白质分子量标准;1:流穿液;2-6:分别为10 、50、100 、200 、400 mmol/L咪唑洗脱液
Fig. 2 Csino3 protein purificationM: Protein molecular weight standard. 1: Flow-through; 2-6: 10, 50, 100, 200, 400 mmol/L imidazole elution, respectively
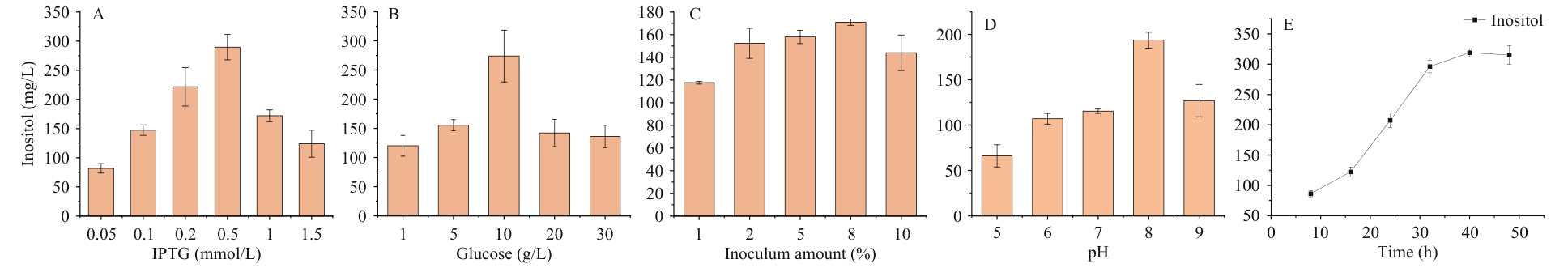
图5 单因素发酵工艺优化A:IPTG终浓度;B:葡萄糖浓度;C:接种量;D:初始pH;E:时间
Fig. 5 Optimization of single-factor fermentationA: Final concentration of IPTG. B: Glucose concentration. C: Inoculation amount. D: Initial pH. E: Time
| 因素 Factor | Level | Low level | High level |
|---|---|---|---|
| Glucose (g/L) | 10.0 | 5.0 | 15.0 |
| IPTG (mmol/L) | 0.5 | 0.2 | 0.8 |
| Inoculation amount (%) | 8.0 | 6.0 | 10.0 |
| Initial pH | 8.0 | 7.0 | 9.0 |
表3 Box-Behnken design设计因素及水平表
Table 3 Factor and level table of Box-Behnken design
| 因素 Factor | Level | Low level | High level |
|---|---|---|---|
| Glucose (g/L) | 10.0 | 5.0 | 15.0 |
| IPTG (mmol/L) | 0.5 | 0.2 | 0.8 |
| Inoculation amount (%) | 8.0 | 6.0 | 10.0 |
| Initial pH | 8.0 | 7.0 | 9.0 |
| Number | A (g/L) | B (mmol/L) | C (%) | D | Inositol (mg/L) |
|---|---|---|---|---|---|
| 1 | 5 | 0.2 | 8 | 8 | 151.00 |
| 2 | 15 | 0.2 | 8 | 8 | 117.33 |
| 3 | 5 | 0.8 | 8 | 8 | 159.67 |
| 4 | 15 | 0.8 | 8 | 8 | 140.33 |
| 5 | 10 | 0.5 | 6 | 7 | 149.33 |
| 6 | 10 | 0.5 | 10 | 7 | 146.00 |
| 7 | 10 | 0.5 | 6 | 9 | 156.00 |
| 8 | 10 | 0.5 | 10 | 9 | 178.33 |
| 9 | 5 | 0.5 | 8 | 7 | 121.00 |
| 10 | 15 | 0.5 | 8 | 7 | 114.33 |
| 11 | 5 | 0.5 | 8 | 9 | 152.67 |
| 12 | 15 | 0.5 | 8 | 9 | 143.00 |
| 13 | 10 | 0.2 | 6 | 8 | 152.33 |
| 14 | 10 | 0.8 | 6 | 8 | 166.67 |
| 15 | 10 | 0.2 | 10 | 8 | 145.33 |
| 16 | 10 | 0.8 | 10 | 8 | 144.33 |
| 17 | 5 | 0.5 | 6 | 8 | 144.67 |
| 18 | 15 | 0.5 | 6 | 8 | 138.00 |
| 19 | 5 | 0.5 | 10 | 8 | 176.67 |
| 20 | 15 | 0.5 | 10 | 8 | 137.33 |
| 21 | 10 | 0.2 | 8 | 7 | 174.33 |
| 22 | 10 | 0.8 | 8 | 7 | 153.00 |
| 23 | 10 | 0.2 | 8 | 9 | 191.67 |
| 24 | 10 | 0.8 | 8 | 9 | 177.00 |
| 25 | 10 | 0.5 | 8 | 8 | 288.33 |
| 26 | 10 | 0.5 | 8 | 8 | 272.33 |
| 27 | 10 | 0.5 | 8 | 8 | 287.00 |
| 28 | 10 | 0.5 | 8 | 8 | 295.67 |
| 29 | 10 | 0.5 | 8 | 8 | 298.00 |
表4 Box-Behnken 响应面试验设计及结果
Table 4 Box-Behnken response surface experimental design and results
| Number | A (g/L) | B (mmol/L) | C (%) | D | Inositol (mg/L) |
|---|---|---|---|---|---|
| 1 | 5 | 0.2 | 8 | 8 | 151.00 |
| 2 | 15 | 0.2 | 8 | 8 | 117.33 |
| 3 | 5 | 0.8 | 8 | 8 | 159.67 |
| 4 | 15 | 0.8 | 8 | 8 | 140.33 |
| 5 | 10 | 0.5 | 6 | 7 | 149.33 |
| 6 | 10 | 0.5 | 10 | 7 | 146.00 |
| 7 | 10 | 0.5 | 6 | 9 | 156.00 |
| 8 | 10 | 0.5 | 10 | 9 | 178.33 |
| 9 | 5 | 0.5 | 8 | 7 | 121.00 |
| 10 | 15 | 0.5 | 8 | 7 | 114.33 |
| 11 | 5 | 0.5 | 8 | 9 | 152.67 |
| 12 | 15 | 0.5 | 8 | 9 | 143.00 |
| 13 | 10 | 0.2 | 6 | 8 | 152.33 |
| 14 | 10 | 0.8 | 6 | 8 | 166.67 |
| 15 | 10 | 0.2 | 10 | 8 | 145.33 |
| 16 | 10 | 0.8 | 10 | 8 | 144.33 |
| 17 | 5 | 0.5 | 6 | 8 | 144.67 |
| 18 | 15 | 0.5 | 6 | 8 | 138.00 |
| 19 | 5 | 0.5 | 10 | 8 | 176.67 |
| 20 | 15 | 0.5 | 10 | 8 | 137.33 |
| 21 | 10 | 0.2 | 8 | 7 | 174.33 |
| 22 | 10 | 0.8 | 8 | 7 | 153.00 |
| 23 | 10 | 0.2 | 8 | 9 | 191.67 |
| 24 | 10 | 0.8 | 8 | 9 | 177.00 |
| 25 | 10 | 0.5 | 8 | 8 | 288.33 |
| 26 | 10 | 0.5 | 8 | 8 | 272.33 |
| 27 | 10 | 0.5 | 8 | 8 | 287.00 |
| 28 | 10 | 0.5 | 8 | 8 | 295.67 |
| 29 | 10 | 0.5 | 8 | 8 | 298.00 |
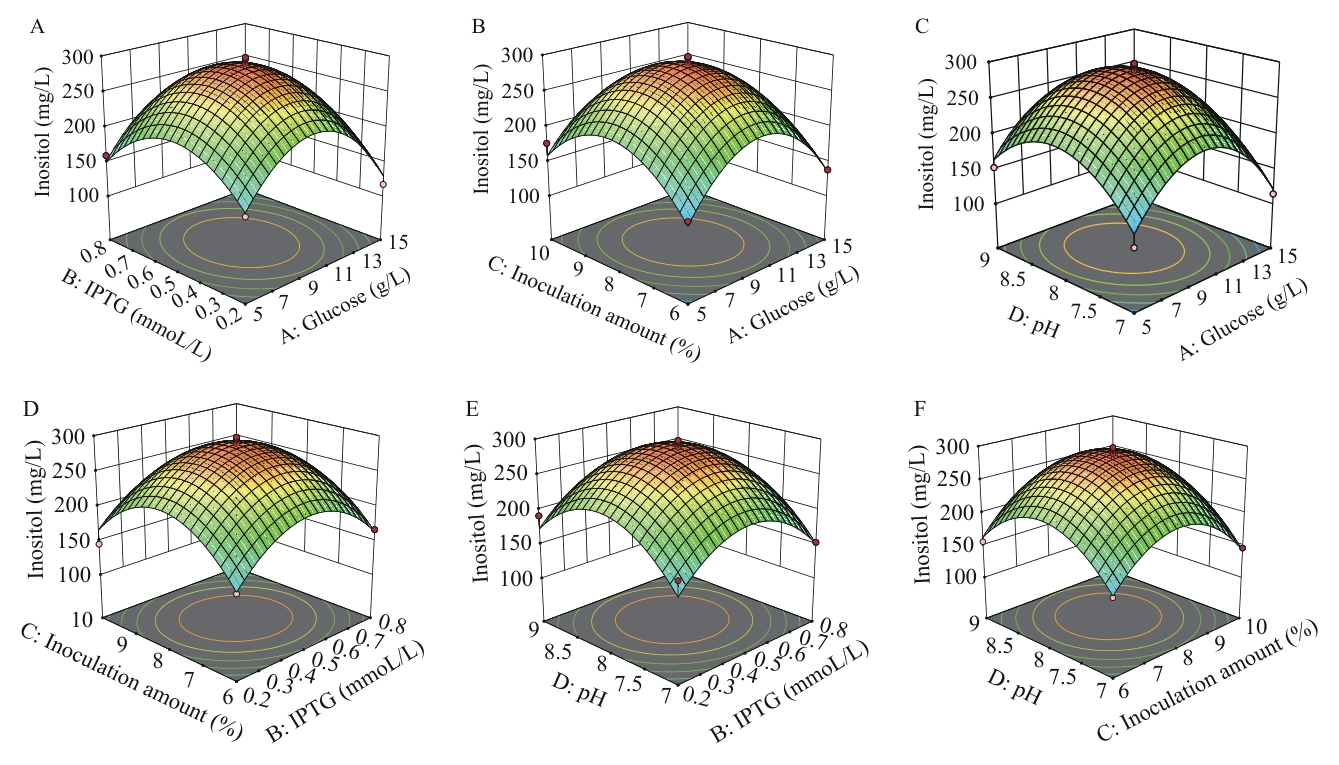
图6 响应面优化菌株pETDuet-1-Csino3-suhB/BL21(DE3)生产肌醇A:IPTG和葡萄糖;B:接种量和葡萄糖;C:pH和葡萄糖;D:接种量和IPTG;E:pH和IPTG;F:pH和接种量
Fig. 6 Response surface on inositol production optimized by strain pETDuet-1-Csino3-suhB/BL21(DE3)A: IPTG and glucose. B: Inoculation amount and glucose. C: pH and glucose. D: Inoculation amount and IPTG. E: pH and IPTG. F: pH and inoculation amount
变异来源 Source of variation | 平方和 SS | 自由度 df | 均方 MS | F值 F-value | P值 P-value | 备注 Note |
|---|---|---|---|---|---|---|
| Model | 83 481.48 | 14 | 5 962.96 | 25.45 | <0.000 1 | ** |
| A- Glucose | 1 108.48 | 1 | 1 108.48 | 4.73 | 0.047 2 | * |
| B- IPTG | 6.75 | 1 | 6.75 | 0.028 8 | 0.867 6 | Not significant |
| C- Inoculation amount | 36.75 | 1 | 36.75 | 0.156 9 | 0.698 0 | Not significant |
| D- Initial pH | 1 648.93 | 1 | 1 648.93 | 7.04 | 0.018 9 | ** |
| AB | 51.36 | 1 | 51.36 | 0.219 2 | 0.646 8 | Not significant |
| AC | 266.78 | 1 | 266.78 | 1.14 | 0.304 0 | Not significant |
| AD | 2.25 | 1 | 2.25 | 0.009 6 | 0.923 3 | Not significant |
| BC | 58.78 | 1 | 58.78 | 0.250 9 | 0.624 2 | Not significant |
| BD | 11.11 | 1 | 11.11 | 0.047 4 | 0.830 7 | Not significant |
| CD | 164.69 | 1 | 164.69 | 0.703 0 | 0.415 9 | Not significant |
| A2 | 45 114.14 | 1 | 45 114.14 | 192.57 | <0.000 1 | ** |
| B2 | 24 352.12 | 1 | 24 352.12 | 103.94 | <0.000 1 | ** |
| C2 | 28 274.17 | 1 | 28 274.17 | 120.69 | <0.000 1 | ** |
| D2 | 26 002.04 | 1 | 26 002.04 | 110.99 | <0.000 1 | ** |
| Residual | 3 279.91 | 14 | 234.28 | |||
| Lack of fit | 2 874.94 | 10 | 287.49 | 2.84 | 0.1632 | Not significant |
| Pure error | 404.98 | 4 | 101.24 | |||
| Cor total | 86 761.39 | 28 |
表5 Box-Behnken 响应面试验方差分析
Table 5 Analysis of variance (ANOVA) for Box-Behnken response surface methodology
变异来源 Source of variation | 平方和 SS | 自由度 df | 均方 MS | F值 F-value | P值 P-value | 备注 Note |
|---|---|---|---|---|---|---|
| Model | 83 481.48 | 14 | 5 962.96 | 25.45 | <0.000 1 | ** |
| A- Glucose | 1 108.48 | 1 | 1 108.48 | 4.73 | 0.047 2 | * |
| B- IPTG | 6.75 | 1 | 6.75 | 0.028 8 | 0.867 6 | Not significant |
| C- Inoculation amount | 36.75 | 1 | 36.75 | 0.156 9 | 0.698 0 | Not significant |
| D- Initial pH | 1 648.93 | 1 | 1 648.93 | 7.04 | 0.018 9 | ** |
| AB | 51.36 | 1 | 51.36 | 0.219 2 | 0.646 8 | Not significant |
| AC | 266.78 | 1 | 266.78 | 1.14 | 0.304 0 | Not significant |
| AD | 2.25 | 1 | 2.25 | 0.009 6 | 0.923 3 | Not significant |
| BC | 58.78 | 1 | 58.78 | 0.250 9 | 0.624 2 | Not significant |
| BD | 11.11 | 1 | 11.11 | 0.047 4 | 0.830 7 | Not significant |
| CD | 164.69 | 1 | 164.69 | 0.703 0 | 0.415 9 | Not significant |
| A2 | 45 114.14 | 1 | 45 114.14 | 192.57 | <0.000 1 | ** |
| B2 | 24 352.12 | 1 | 24 352.12 | 103.94 | <0.000 1 | ** |
| C2 | 28 274.17 | 1 | 28 274.17 | 120.69 | <0.000 1 | ** |
| D2 | 26 002.04 | 1 | 26 002.04 | 110.99 | <0.000 1 | ** |
| Residual | 3 279.91 | 14 | 234.28 | |||
| Lack of fit | 2 874.94 | 10 | 287.49 | 2.84 | 0.1632 | Not significant |
| Pure error | 404.98 | 4 | 101.24 | |||
| Cor total | 86 761.39 | 28 |
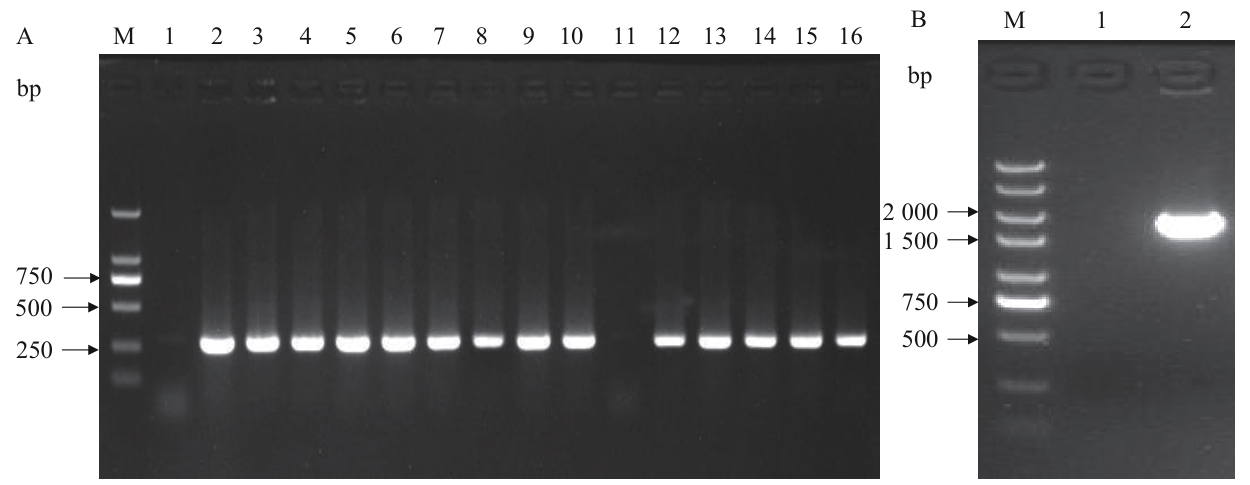
图7 BL21(DE3)/Δpgi菌落PCR验证M:DNA Marker;A-1:无模板阴性对照扩增结果;A-2:原始菌株的内部引物扩增结果;A-(3-16):第1-14号克隆的内部引物扩增结果;B-1:无模板阴性对照的扩增结果;B-2:阴性克隆外侧引物的扩增结果
Fig. 7 Colony PCR verification of BL21(DE3)/ΔpgiM: DNA Marker. A-1: No-template negative control amplified result. A-2: Internal primer amplified result of the original strain. A-(3-16): Internal primer amplification results of clone 1-14. B-1: No-template negative control amplified result. B-2: External primer amplified result of negative clone
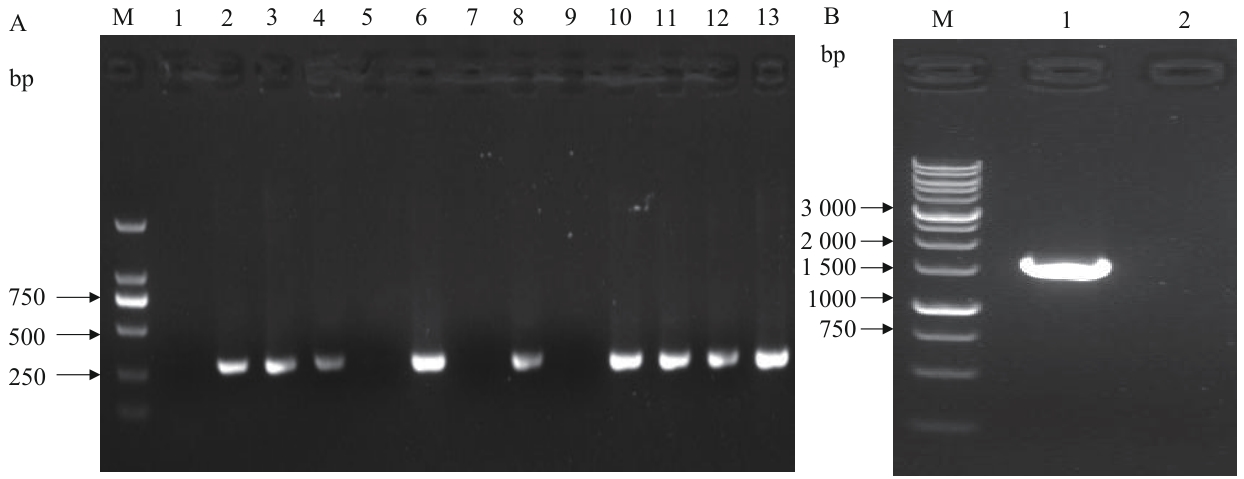
图8 BL21(DE3)/ΔgldA 菌落PCR验证M:DNA Marker;A-1:无模板阴性对照扩增结果;A-2:原始菌株的内部引物扩增结果;A-(3-13):第1-11号克隆的内部引物扩增结果;B-1:阴性克隆外侧引物的扩增结果;B-2:无模板阴性对照的扩增结果
Fig. 8 Colony PCR verification of BL21(DE3)/ΔgldAM: DNA Marker. A-1: No-template negative control amplified result. A-2: Internal primer amplified result of the original strain. A-(3-13): Internal primer amplified results of clone 1-11. B-1: External primer amplified result of negative clone. B-2: No-template negative control amplified result
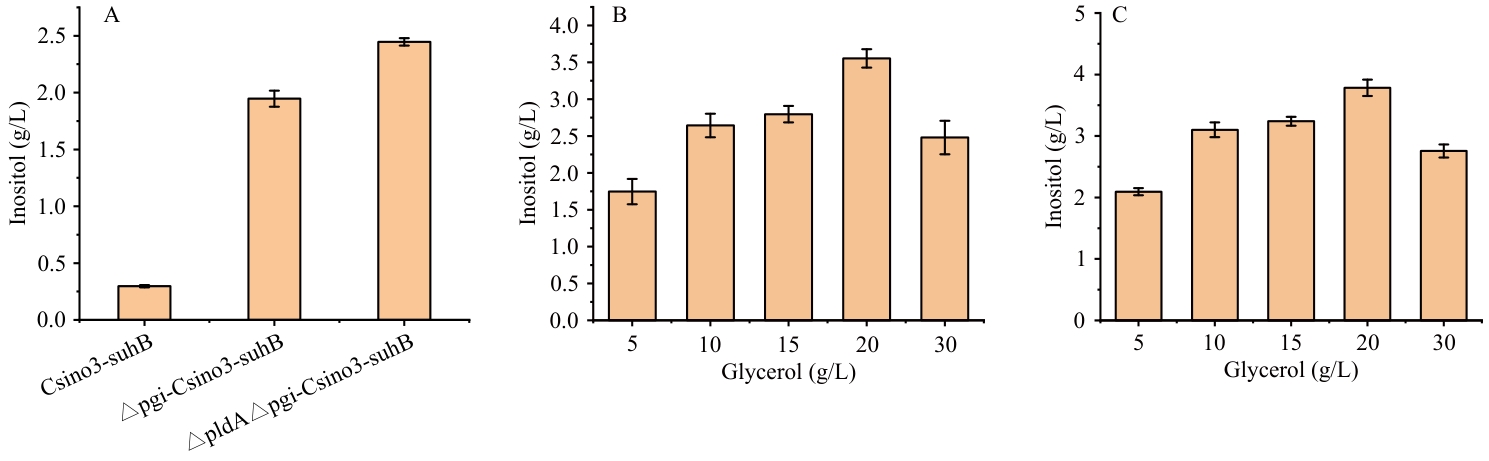
图9 敲除pgi、gldA基因菌株合成肌醇能力A:3种菌株生产肌醇含量;B:不同甘油浓度对△pgi-pETDuet-1-Csino3-suhB / BL21(DE3)发酵肌醇的影响;C:不同甘油浓度对△gldA△pgi -pETDuet-1-Csino3-suhB / BL21(DE3)发酵肌醇的影响
Fig. 9 Inositol-synthezing capacity of the pgi- and gldA-knockout strainA: Inositol production content by three strains. B: Effect of different glycerol concentrations on inositol fermentation by △pgi-pETDuet-1-Csino3-suhB/BL21(DE3). C: Effect of different glycerol concentrations on inositol fermentation by △gldA△pgi-pETDuet-1-Csino3-suhB/BL21(DE3)
| [1] | Kılbaş B, Balci M. Recent advances in inositol chemistry: synthesis and applications [J]. Tetrahedron, 2011, 67(13): 2355-2389. |
| [2] | 陈庆伟. 基因工程改造大肠杆菌发酵混合糖生产肌醇的研究 [D]. 杭州: 浙江工商大学, 2023. |
| Chen QW. Study on the production of inositol by fermentation of mixed sugar with genetically engineered Escherichia coli [D]. Hangzhou: Zhejiang Gongshang University, 2023. | |
| [3] | Omoruyi FO, Stennett D, Foster S, et al. New frontiers for the use of IP6 and inositol combination in treating diabetes mellitus: a review [J]. Molecules, 2020, 25(7): 1720. |
| [4] | Ma AJ, Cui WX, Wang XN, et al. Osmoregulation by the myo-inositol biosynthesis pathway in turbot Scophthalmus maximus and its regulation by anabolite and c-Myc [J]. Comp Biochem Physiol Part A Mol Integr Physiol, 2020, 242: 110636. |
| [5] | Jiang WD, Li SG, Mi HF, et al. Myo-inositol prevents the gill rot in fish caused by Flavobacterium columnare infection [J]. Aquaculture, 2022, 546: 737393. |
| [6] | Shimada M, Hibino M, Takeshita A. Dietary supplementation with myo-inositol reduces hepatic triglyceride accumulation and expression of both fructolytic and lipogenic genes in rats fed a high-fructose diet [J]. Nutr Res, 2017, 47: 21-27. |
| [7] | 杨连生, 扶雄, 孙由芳, 等. 生物酶水解植酸钙提取肌醇研究 [J]. 食品工业科技, 1997, 18(5): 48-48, 53. |
| Yang LS, Fu X, Sun YF, et al. Study on drawing inositol from phytin by hydrolysis enzyme method[J]. Sci Technol Food Ind, 1997, 18(5): 48-48, 53. | |
| [8] | 王建玲, 王敏, 王春霞, 等. 利用酶法生产肌醇初步研究 [J]. 天津轻工业学院学报, 1998, 13(2): 15-19. |
| Wang JL, Wang M, Wang CX, et al. Primary study on inositol production by hydrolyst[J]. J Tianjin Univ Sci Technol, 1998, 13(2): 15-19. | |
| [9] | 郭育涛, 陈坚, 伦世仪. 微生物发酵法生产肌醇的研究 [J]. 西北轻工业学院学报, 1998, 16(1): 117-120. |
| Guo YT, Chen J, Lun SY. Microbiol synthesis of inositol by fermentation [J]. J Shaanxi Univ Sci Technol, 1998, 16(1): 117-120. | |
| [10] | Li YJ, Han PP, Wang J, et al. Production of myo-inositol: Recent advance and prospective [J]. Biotechnol Appl Biochem, 2022, 69(3): 1101-1111. |
| [11] | Dinicola S, Unfer V, Facchinetti F, et al. Inositols: from established knowledge to novel approaches [J]. Int J Mol Sci, 2021, 22(19): 10575. |
| [12] | Lu YP, Wang L, Teng F, et al. Production of myo-inositol from glucose by a novel trienzymatic cascade of polyphosphate glucokinase, inositol 1-phosphate synthase and inositol monophosphatase [J]. Enzyme Microb Technol, 2018, 112: 1-5. |
| [13] | 黄贞杰, 陈由强, 陈丽霞, 等. 代谢工程改造酿酒酵母合成肌醇 [J]. 微生物学通报, 2017, 44(10): 2289-2296. |
| Huang ZJ, Chen YQ, Chen LX, et al. Metabolic engineering of Saccharomyces cerevisiae for inositol production [J]. Microbiol China, 2017, 44(10): 2289-2296. | |
| [14] | Shiue E, Brockman IM, Prather KLJ. Improving product yields on D-glucose in Escherichia coli via knockout of pgi and zwf and feeding of supplemental carbon sources [J]. Biotechnol Bioeng, 2015, 112(3): 579-587. |
| [15] | Yi MH, Yang LZ, Ma J, et al. Biosynthesis of myo-inositol in Escherichia coli by engineering myo-inositol-1-phosphate pathway [J]. Biochem Eng J, 2020, 164: 107792. |
| [16] | 朱宏宇, 王晓璐, 刘亚君, 等. 产肌醇毕赤酵母细胞工厂的优化 [J]. 微生物学通报, 2023, 50(9): 3731-3746. |
| Zhu HY, Wang XL, Liu YJ, et al. Optimization of a Komagataella phaffii cell factory for producing inositol [J]. Microbiol China, 2023, 50(9): 3731-3746. | |
| [17] | Henry AS, Gaspar LM, Jesch AS. The response to inositol: Regulation of glycerolipid metabolism and stress response signaling in yeast[J]. Chemistry and Physics of Lipids,2014,180:23-43. |
| [18] | 邵帅, 凌宏志, 何平, 等. 自杀质粒同源重组技术敲除阴沟肠杆菌budC基因以提升乙偶姻的产率 [J]. 黑龙江大学自然科学学报, 2023, 40(4): 434-441. |
| Shao S, Ling HZ, He P, et al. Increase acetoin production of Enterobacter cloacae: budC gene knockout by suicide plasmid homologous recombination technology [J]. J Nat Sci Heilongjiang Univ, 2023, 40(4): 434-441. | |
| [19] | 李莉, 张赛, 何强, 等. 响应面法在试验设计与优化中的应用 [J]. 实验室研究与探索, 2015, 34(8): 41-45. |
| Li L, Zhang S, He Q, et al. Application of response surface methodology in experiment design and optimization [J]. Res Explor Lab, 2015, 34(8): 41-45. | |
| [20] | Sarkar M, Majumdar P. Application of response surface methodology for optimization of heavy metal biosorption using surfactant modified chitosan bead [J]. Chem Eng J, 2011, 175: 376-387. |
| [21] | 潘柯莉, 茅以诚, 岳圣杰, 等. pgi基因敲除对大肠杆菌利用混合碳源的影响 [J]. 高校化学工程学报, 2018, 32(1): 138-146. |
| Pan KL, Mao YC, Yue SJ, et al. Effects of pgi knockout on mixed carbon source utilization of Escherichia coli [J]. J Chem Eng Chin Univ, 2018, 32(1): 138-146. | |
| [22] | 张芹, 王芳, 陈雅蕾, 等. 肌醇生产及应用研究进展 [J]. 中国稻米, 2012(3): 19-21. |
| Zhang Q, Wang F, Chen YL, et al. Research progress of inositol production and application [J]. China Rice, 2012(3): 19-21. | |
| [23] | Villa-García MJ, Choi MS, Hinz FI, et al. Genome-wide screen for inositol auxotrophy in Saccharomyces cerevisiae implicates lipid metabolism in stress response signaling [J]. Mol Genet Genom, 2011, 285(2): 125-149. |
| [24] | 祁浩, 刘新利. 大肠杆菌表达系统和酵母表达系统的研究进展 [J]. 安徽农业科学, 2016, 44(17): 4-6, 52. |
| Qi H, Liu XL. Research progress of expression systems of Escherichia coli and yeast [J]. J Anhui Agric Sci, 2016, 44(17): 4-6, 52. | |
| [25] | Moon TS, Yoon SH, Lanza AM, et al. Production of glucaric acid from a synthetic pathway in recombinant Escherichia coli [J]. Appl Environ Microbiol, 2009, 75(3): 589-595. |
| [26] | Wu YF, Shen XL, Yuan QP, et al. Metabolic engineering strategies for co-utilization of carbon sources in microbes [J]. Bioengineering, 2016, 3(1): 10. |
| [27] | 马健. 产肌醇的重组大肠杆菌工程菌构建及其发酵条件优化 [D]. 杭州: 浙江工商大学, 2022. |
| Ma J. Construction of inositol-producing recombinant Escherichia coli and optimization of fermentation conditions [D]. Hangzhou: Zhejiang Gongshang University, 2022. | |
| [28] | Zhang QQ, Wang XL, Luo HY, et al. Metabolic engineering of Pichia pastoris for myo-inositol production by dynamic regulation of central metabolism [J]. Microb Cell Fact, 2022, 21(1): 112. |
| [29] | You R, Wang L, Shi CR, et al. Efficient production of myo-inositol in Escherichia coli through metabolic engineering [J]. Microb Cell Fact, 2020, 19(1): 109. |
| [1] | 苏畅, 文凤, 王业林, 邵瑞盈, 唐家杰, 夏占峰. 一株产表面活性素芽胞杆菌发酵优化及其防治应用[J]. 生物技术通报, 2026, 42(4): 332-344. |
| [2] | 王婷婷, 何孟雅, 盛家顺, 高晨, 蔡含芳, 付彤, 孙宇, 高腾云, 张天留. 基于CRISPR/Cas9技术制备NCAPG基因敲除牛成纤维细胞系[J]. 生物技术通报, 2026, 42(2): 317-324. |
| [3] | 廉少杰, 唐胜硕, 康传利, 刘磊, 郑德强, 杜帅, 汤丽伟, 张美霞, 刘蔷. 高产银耳多糖酶菌株的分离、鉴定、发酵条件优化及其酶的特性分析[J]. 生物技术通报, 2025, 41(9): 302-313. |
| [4] | 程慧娟, 王昕, 石小涛, 马东旭, 龚大春, 胡骏鹏, 谢智文. 转录因子CREA敲除对黑曲霉形态和分泌β-葡萄糖苷酶的影响[J]. 生物技术通报, 2025, 41(6): 344-354. |
| [5] | 王鑫鑫, 管玉祝, 李晓苇, 洪伟, 吴道艳, 康颖倩, 刘永畅, 陈峥宏, 崔古贞. 幽门螺杆菌katA基因功能及其在耐受氧化损伤中的作用分析[J]. 生物技术通报, 2024, 40(5): 310-320. |
| [6] | 高登科, 马白荣, 郭怡莹, 刘薇, 刘田, 靳亚平, 江舟, 陈华涛. 利用CRISPR/Cas9技术构建Quaking敲除的小鼠胚胎成纤维细胞株[J]. 生物技术通报, 2024, 40(2): 65-72. |
| [7] | 史亚楠, 王德培, 王一川, 周昊, 薛鲜丽. 敲除msn2对米曲霉生长和发酵产曲酸的影响[J]. 生物技术通报, 2022, 38(8): 188-197. |
| [8] | 王新光, 田磊, 王恩泽, 钟成, 田春杰. 玉米秸秆高效降解微生物复合菌系的构建及降解效果评价[J]. 生物技术通报, 2022, 38(4): 217-229. |
| [9] | 丁亚群, 丁宁, 谢深民, 黄梦娜, 张昱, 张勤, 姜力. Vps28基因敲除小鼠模型的构建及其对泌乳和免疫性状影响的研究[J]. 生物技术通报, 2022, 38(3): 164-172. |
| [10] | 燕炯, 冯晨毅, 高学坤, 许祥, 杨佳敏, 陈朝阳. 基于CRISPR/Cas9技术构建Plin1基因敲除小鼠模型及表型分析[J]. 生物技术通报, 2022, 38(3): 173-180. |
| [11] | 钟菁, 孙玲玲, 张姝, 蒙园, 支怡飞, 涂黎晴, 徐天鹏, 濮黎萍, 陆阳清. 应用CRISPR/Cas9技术敲除Mda5基因对新城疫及传染性法氏囊病毒复制的影响[J]. 生物技术通报, 2022, 38(11): 90-96. |
| [12] | 王海杰, 王成稷, 郭洋, 王云, 陈艳娟, 梁敏, 王珏, 龚慧, 沈如凌. 基于CRSIPR/Cas9技术构建凝血因子8基因敲除小鼠模型及表型验证[J]. 生物技术通报, 2022, 38(10): 273-280. |
| [13] | 孙靖雅, 马玉超. 假单胞菌Tw224抗砷基因簇的功能研究[J]. 生物技术通报, 2022, 38(1): 141-149. |
| [14] | 王睿, 韩烈保. CRISPR/Cas9介导的二穗短柄草bdfls2敲除突变体的获得[J]. 生物技术通报, 2022, 38(1): 70-76. |
| [15] | 刘雪丹, 杨萌, 张静, 赵东旭. 葡萄糖-木糖共利用对重组大肠杆菌合成D-1,2,4-丁三醇的影响[J]. 生物技术通报, 2021, 37(9): 171-179. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||