








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 197-206.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0890
王尚封1( ), 程斌2,3, 王若若3,4, 丁延庆2,3, 徐建霞2,3, 曹宁2,3, 高旭2,3, 李文贞2,3, 张立异2,3(
), 程斌2,3, 王若若3,4, 丁延庆2,3, 徐建霞2,3, 曹宁2,3, 高旭2,3, 李文贞2,3, 张立异2,3( )
)
收稿日期:2025-08-18
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
张立异,女,博士,研究员,研究方向 :作物分子遗传育种;E-mail: lyzhang1997@hotmail.com作者简介:王尚封,女,硕士研究生,研究方向 :作物分子遗传育种;E-mail: 619684541@qq.com基金资助:
WANG Shang-feng1( ), CHENG Bin2,3, WANG Ruo-ruo3,4, DING Yan-qing2,3, XU Jian-xia2,3, CAO Ning2,3, GAO Xu2,3, LI Wen-zhen2,3, ZHANG Li-yi2,3(
), CHENG Bin2,3, WANG Ruo-ruo3,4, DING Yan-qing2,3, XU Jian-xia2,3, CAO Ning2,3, GAO Xu2,3, LI Wen-zhen2,3, ZHANG Li-yi2,3( )
)
Received:2025-08-18
Published:2026-02-26
Online:2026-03-17
摘要:
目的 开花时间作为高粱品种选育过程中的关键目标性状之一,对其生态适应性和高产潜力具有重要影响。 方法 以晚开花品种红缨子与早开花品种SAP001杂交构建的F2群体为材料进行开花时间遗传定位分析。 结果 表型调查显示,F2群体中晚花和早花单株的分离比符合3∶1(χ2=0.225,P>0.05),表明该性状受到一对主效基因的控制。通过对极端早花和晚花株系的BSA-Seq分析,在第3染色体59.73‒60.44 Mb区间检测到一个与开花时间显著关联的遗传位点。结合双亲重测序和转录组数据,鉴定到一个编码光系统Ⅱ叶绿素结合蛋白的关键候选基因(SbiHYZ.03G218800)。该基因启动子区存在Myb/SANT、TATA结合蛋白(TBP)结合位点突变,在亲本间表达差异显著,可能为高粱中调控开花时间的新基因。基于该候选区段中的SNP和InDel变异,开发了一个可有效区分早花与晚花植株连锁分子标记。 结论 揭示了中国酒用高粱开花时间的遗传基础,为开展高粱生育期分子育种提供了重要基因资源和分子标记。
王尚封, 程斌, 王若若, 丁延庆, 徐建霞, 曹宁, 高旭, 李文贞, 张立异. 基于BSA-seq的高粱开花时间基因定位及候选基因预测[J]. 生物技术通报, 2026, 42(2): 197-206.
WANG Shang-feng, CHENG Bin, WANG Ruo-ruo, DING Yan-qing, XU Jian-xia, CAO Ning, GAO Xu, LI Wen-zhen, ZHANG Li-yi. Gene Mapping of Sorghum Flowering Time and Prediction of Candidate Genes Based on BSA-seq[J]. Biotechnology Bulletin, 2026, 42(2): 197-206.
| 引物名称 Primer name | orward primer (5′‒3′) 正向引物 F | everse primer (5′‒3′) 反向引物 R |
|---|---|---|
| V1f1 | CAACTTCAGTGCAACGGTCG | AGCAAGGATTGTGTGGTGCT |
| V423 | TCAGTGCAACGGTCGACAAA | TGAATGGTGAACAGCCGACC |
| Hd8800 | CACATCAGTGTGTGCCGTTG | TTAGGGGCGTTCTTCTGCTC |
| V5f1 | CATCCACGGGGTTGAAGTCT | AGCGGTCCATTGCTCAAAGT |
表1 4个InDel标记的引物信息
Table 1 Primer information of the 4 InDel markers
| 引物名称 Primer name | orward primer (5′‒3′) 正向引物 F | everse primer (5′‒3′) 反向引物 R |
|---|---|---|
| V1f1 | CAACTTCAGTGCAACGGTCG | AGCAAGGATTGTGTGGTGCT |
| V423 | TCAGTGCAACGGTCGACAAA | TGAATGGTGAACAGCCGACC |
| Hd8800 | CACATCAGTGTGTGCCGTTG | TTAGGGGCGTTCTTCTGCTC |
| V5f1 | CATCCACGGGGTTGAAGTCT | AGCGGTCCATTGCTCAAAGT |
| 样本Sample | 平均测序深度Mean depth (X) | 短片段总数Number of total Reads | 总碱基读数Clean bases (bp) | GC (%) | Q30 (%) | 比对Mapped reads (%) |
|---|---|---|---|---|---|---|
| 红缨子 | 67.04 | 369 858 258 | 55 478 738 700 | 44.63 | 91.96 | 99.41 |
| SAP001 | 70.27 | 355 144 466 | 53 271 669 900 | 43.54 | 96.28 | 98.88 |
| 晚熟池 | 77.31 | 387 856 226 | 58 178 433 900 | 43.67 | 97.29 | 99.34 |
| 早熟池 | 86.91 | 443 492 672 | 66 523 900 800 | 43.53 | 97.36 | 98.95 |
表2 不同样品测序比对结果
Table 2 Sequencing and mapping results of different samples
| 样本Sample | 平均测序深度Mean depth (X) | 短片段总数Number of total Reads | 总碱基读数Clean bases (bp) | GC (%) | Q30 (%) | 比对Mapped reads (%) |
|---|---|---|---|---|---|---|
| 红缨子 | 67.04 | 369 858 258 | 55 478 738 700 | 44.63 | 91.96 | 99.41 |
| SAP001 | 70.27 | 355 144 466 | 53 271 669 900 | 43.54 | 96.28 | 98.88 |
| 晚熟池 | 77.31 | 387 856 226 | 58 178 433 900 | 43.67 | 97.29 | 99.34 |
| 早熟池 | 86.91 | 443 492 672 | 66 523 900 800 | 43.53 | 97.36 | 98.95 |
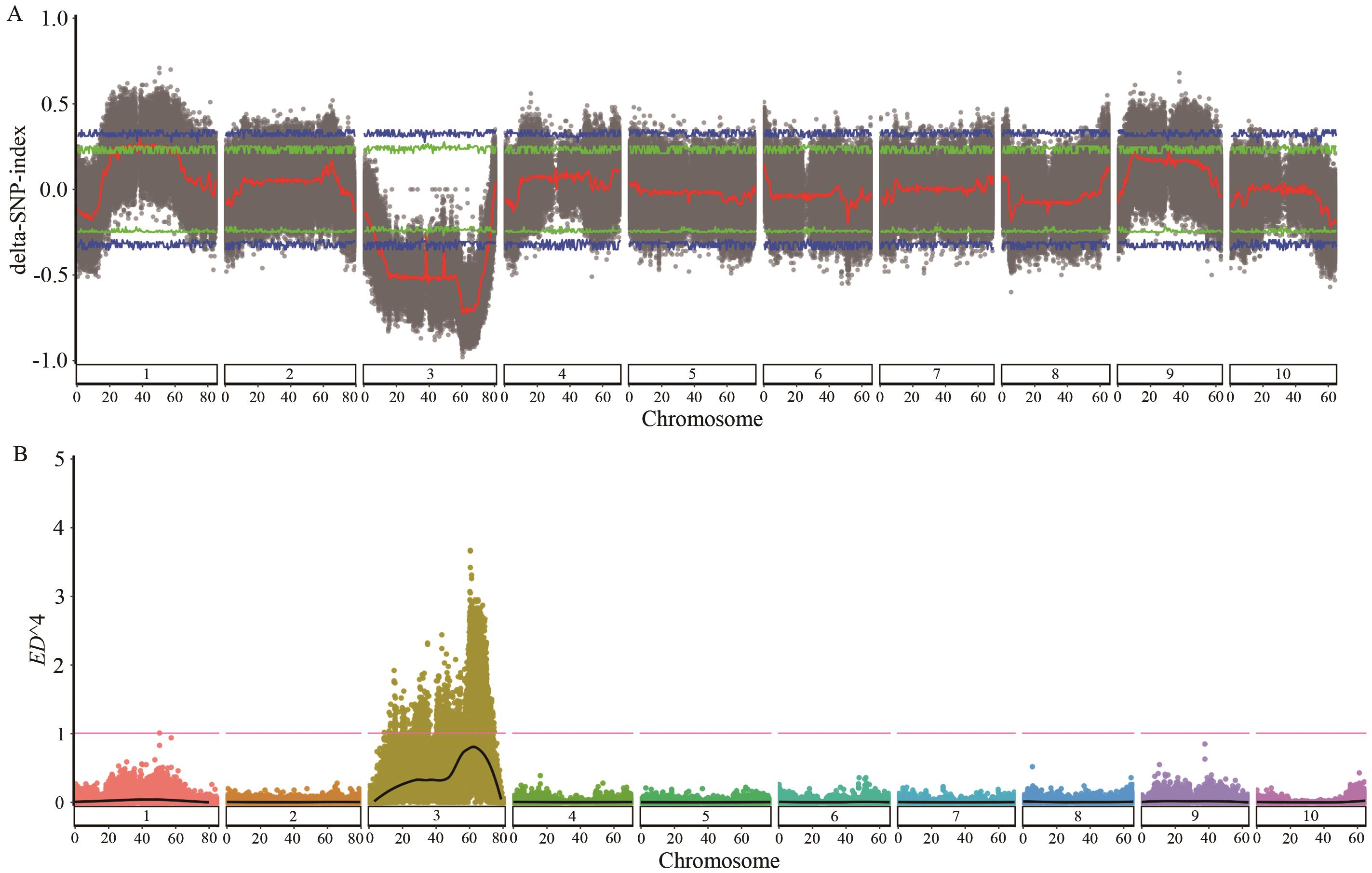
图1 SNP-index和ED4关联值在第3染色体上的分布A:基于SNP-index算法生成的曼哈顿图;蓝色和绿色线条分别表示99%和95%置信区间;黑色圆点表示每个SNP的ΔSNP指数值,红色线条为通过滑动窗口分析得到的ΔSNP指数平均值拟合曲线;B:基于ED算法生成的曼哈顿图;红色线条表示99%置信阈值;黑色线条为ED4值的拟合曲线
Fig. 1 Distribution of SNP-index and ED4 association values on chromosome 3A: Manhattan plot generated based on SNP-index algorithm. The blue and green lines indicate the 99% and 95% confidence intervals, respectively. The black dots denote the ΔSNP-index for each SNP, and the red line is the fitted curve of the mean values of ΔSNP-index derived from the sliding window analysis. B: Manhattan plot generated based on ED algorithm. The red line indicates the 99% threshold; and the black line is the fitted curve of the ED4 values
基因 Gene | 变异位置 Variation position (bp) | 参考基因组 Reference | 等位变异Alternative | CDS突变类型Type of CDS mutation | 启动子突变Promoter mutation |
|---|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 930 660 | A | - | - | 碱基缺失 |
| SbiHYZ.03G216700 | 59 972 784 | C | A | 非同义突变 | |
| SbiHYZ.03G216800 | 59 974 642 | C | A | - | 碱基突变 |
| SbiHYZ.03G216900 | 59 980 499 | A | - | 缺失突变/移码 | |
| 59 980 657 | G | A | 非同义突变 | ||
| 59 980 792 | G | T | 非同义突变 | ||
| 59 980 816 | G | C | 非同义突变 | ||
| SbiHYZ.03G217000 | 60 060 314 | A | - | - | 碱基缺失 |
| SbiHYZ.03G217200 | 60 093 586 | A | C | 非同义突变 | |
| SbiHYZ.03G217700 | 60 140 432 | G | A | 非同义突变 | |
| SbiHYZ.03G218800 | 60 365 936 | C | T | - | 碱基突变 |
| SbiHYZ.03G218900 | 60 375 442 | C | T | 非同义突变 |
表3 九个候选基因序列变异分析
Table 3 Sequence variation analysis of nine candidate genes
基因 Gene | 变异位置 Variation position (bp) | 参考基因组 Reference | 等位变异Alternative | CDS突变类型Type of CDS mutation | 启动子突变Promoter mutation |
|---|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 930 660 | A | - | - | 碱基缺失 |
| SbiHYZ.03G216700 | 59 972 784 | C | A | 非同义突变 | |
| SbiHYZ.03G216800 | 59 974 642 | C | A | - | 碱基突变 |
| SbiHYZ.03G216900 | 59 980 499 | A | - | 缺失突变/移码 | |
| 59 980 657 | G | A | 非同义突变 | ||
| 59 980 792 | G | T | 非同义突变 | ||
| 59 980 816 | G | C | 非同义突变 | ||
| SbiHYZ.03G217000 | 60 060 314 | A | - | - | 碱基缺失 |
| SbiHYZ.03G217200 | 60 093 586 | A | C | 非同义突变 | |
| SbiHYZ.03G217700 | 60 140 432 | G | A | 非同义突变 | |
| SbiHYZ.03G218800 | 60 365 936 | C | T | - | 碱基突变 |
| SbiHYZ.03G218900 | 60 375 442 | C | T | 非同义突变 |
基因 Gene | 物理位置 Physical position (bp) | 同源基因 Paralogous gene | 功能注释 Functional annotation | 相似率 Similarity (%) |
|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 931 181‒59 935 674 | LOC_Os01g41250 | F-box/FBD结构域 F-box/FBD domain | 66 |
| SbiHYZ.03G216700 | 59 971 690‒59 974 228 | LOC_Os01g41260 | F-box/FBD结构域 F-box/FBD domain | 63 |
| SbiHYZ.03G216800 | 59 974 820‒59 977 137 | Zm00001d011276 | F-box/FBD结构域 F-box/FBD domain | 80 |
| SbiHYZ.03G216900 | 59 978 600‒59 981 036 | LOC_Os01g41310 | F-box/FBD结构域 F-box/FBD domain | 51 |
| SbiHYZ.03G217000 | 60 061 333‒60 063 755 | Zm00001d044385 | F-box/FBD结构域 F-box/FBD domain | 83 |
| SbiHYZ.03G217200 | 60 090 312‒60 093 766 | Zm00001d044386 | F-box/FBD结构域 F-box/FBD domain | 75 |
| SbiHYZ.03G217700 | 60 139 306‒60 141 819 | LOC_Os01g41530 | F-box/FBD结构域 F-box/FBD domain | 50 |
| SbiHYZ.03G218800 | 60 366 296‒60 367 982 | ZM00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡchlorophyll a/b-binding protein | 97 |
| SbiHYZ.03G218900 | 60 374 509‒60 375 642 | Zm00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡ chlorophyll a/b-binding protein | 98 |
表4 九个候选基因信息
Table 4 Information of nine candidate genes
基因 Gene | 物理位置 Physical position (bp) | 同源基因 Paralogous gene | 功能注释 Functional annotation | 相似率 Similarity (%) |
|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 931 181‒59 935 674 | LOC_Os01g41250 | F-box/FBD结构域 F-box/FBD domain | 66 |
| SbiHYZ.03G216700 | 59 971 690‒59 974 228 | LOC_Os01g41260 | F-box/FBD结构域 F-box/FBD domain | 63 |
| SbiHYZ.03G216800 | 59 974 820‒59 977 137 | Zm00001d011276 | F-box/FBD结构域 F-box/FBD domain | 80 |
| SbiHYZ.03G216900 | 59 978 600‒59 981 036 | LOC_Os01g41310 | F-box/FBD结构域 F-box/FBD domain | 51 |
| SbiHYZ.03G217000 | 60 061 333‒60 063 755 | Zm00001d044385 | F-box/FBD结构域 F-box/FBD domain | 83 |
| SbiHYZ.03G217200 | 60 090 312‒60 093 766 | Zm00001d044386 | F-box/FBD结构域 F-box/FBD domain | 75 |
| SbiHYZ.03G217700 | 60 139 306‒60 141 819 | LOC_Os01g41530 | F-box/FBD结构域 F-box/FBD domain | 50 |
| SbiHYZ.03G218800 | 60 366 296‒60 367 982 | ZM00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡchlorophyll a/b-binding protein | 97 |
| SbiHYZ.03G218900 | 60 374 509‒60 375 642 | Zm00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡ chlorophyll a/b-binding protein | 98 |
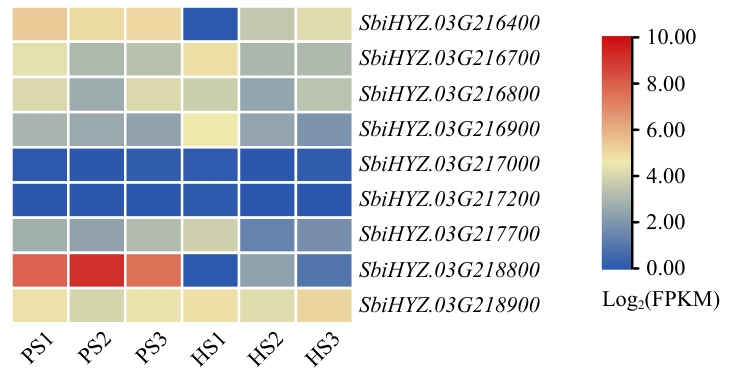
图3 亲本中候选基因在不同时期不同组织中的表达量候选基因的表达量基于RNA测序分析获得的FPKM值,取log2绘制热图;S1:孕穗期;S2:抽穗5 cm;S3:开花第1天;P:SAP001;H:红缨子
Fig. 3 Expressions of candidate genes in the parent in different tissues at different timesHeatmap of candidate gene expression based on FPKM values from RNA-seq analysis after log2 transformation.S1: Booting stage. S2: 5 cm heading. S3: First flowering day. P: SAP001. H: Hongyingzi
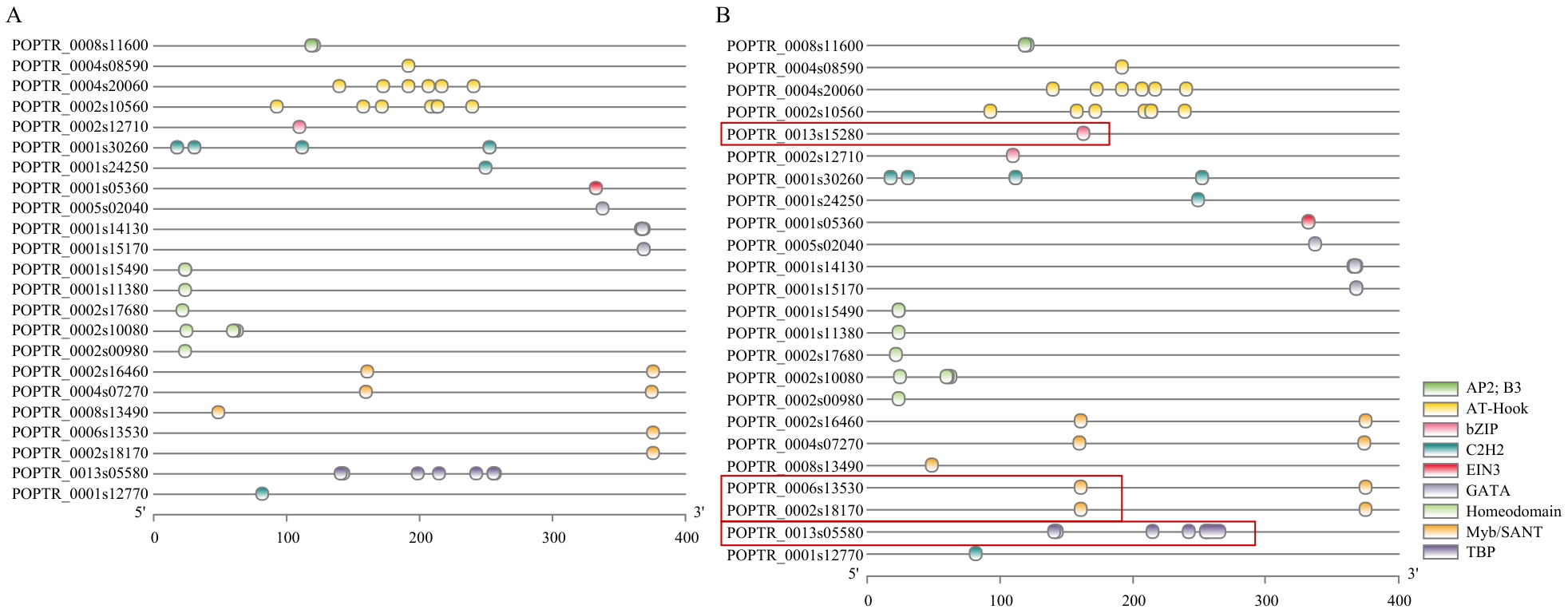
图4 亲本中关键候选基因启动子区域顺式元件对比A:红缨子启动子区域顺式元件可视图;B:SAP001启动子区域顺式元件可视图,红色方框表示元件差异
Fig. 4 Comparison of cis-elements in the promoter regions of key candidate genes in parentA: Visualized cis-acting elements in the promoter region of Hongyingzi. B: Visualized cis-acting elements in the promoter region of SAP001, with red boxes highlighting element divergence
| [1] | 王艳秋, 张飞, 朱凯, 等. 抗旱型高粱开花前期和后期应对水分亏缺的生理调节研究 [J]. 山西农业大学学报: 自然科学版, 2020, 40(3): 37-44. |
| Wang YQ, Zhang F, Zhu K, et al. Physiological response to water-deficience of drought-tolerant sorghum variety prior to and post of flowering [J]. J Shanxi Agric Univ Nat Sci Ed, 2020, 40(3): 37-44. | |
| [2] | 高士杰, 陈冰嬬, 李继洪. 对粒用高粱育种的思考 [J]. 现代农业科技, 2013(10): 31-32, 37. |
| Gao SJ, Chen BR, Li JH. Thoughts on breeding of grain sorghum [J]. Mod Agric Sci Technol, 2013(10): 31-32, 37. | |
| [3] | 陈冰嬬, 李继洪, 王阳, 等. 高粱(Sorghum bicolor (L.) Moench)种质资源研究进展 [J]. 西北农林科技大学学报: 自然科学版, 2013, 41(1): 67-72, 77. |
| Chen BR, Li JH, Wang Y, et al. Advances in germplasm resources of sorghum (Sorghum bicolor (L.) Moench) [J]. J Northwest A&F Univ Nat Sci Ed, 2013, 41(1): 67-72, 77. | |
| [4] | 丁延庆, 徐建霞, 汪灿, 等. 基于Super-GBS技术的高粱籽粒酿造相关性状QTL定位 [J]. 核农学报, 2023, 37(2): 241-250. |
| Ding YQ, Xu JX, Wang C, et al. QTL mapping of grain traits related to brewing in sorghum based on super-GBS technology [J]. J Nucl Agric Sci, 2023, 37(2): 241-250. | |
| [5] | 阳志锐, 冉红艳. 贵州省酱香型白酒全产业链发展现状及能级提升路径研究 [J]. 中国酿造, 2025, 44(3): 285-291. |
| Yang ZR, Ran HY. Development status and capacity enhancement path of the whole industry chain of sauce-flavor Baijiu in Guizhou province [J]. China Brew, 2025, 44(3): 285-291. | |
| [6] | Mace ES, Hunt CH, Jordan DR. Supermodels: sorghum and maize provide mutual insight into the genetics of flowering time [J]. Theor Appl Genet, 2013, 126(5): 1377-1395. |
| [7] | Upadhyaya HD, Reddy KN, Vetriventhan M, et al. Latitudinal adaptation of flowering response to photoperiod and temperature in the world collection of sorghum landraces [J]. Crop Sci, 2018, 58(3): 1265-1276. |
| [8] | Rooney WL, Aydin S. Genetic control of a photoperiod-sensitive response in Sorghum bicolor (L.) moench [J]. Crop Sci, 1999, 39(2): 397-400. |
| [9] | Smith CW, Frederiksen RA. Sorghum: Origin, History, Technology, and Production[M]. Wiley, 2000: 240-242. |
| [10] | Murphy RL, Klein RR, Morishige DT, et al. Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum [J]. Proc Natl Acad Sci USA, 2011, 108(39): 16469-16474. |
| [11] | Casto AL, Mattison AJ, Olson SN, et al. Maturity2, a novel regulator of flowering time in Sorghum bicolor, increases expression of SbPRR37 and SbCO in long days delaying flowering [J]. PLoS One, 2019, 14(4): e0212154. |
| [12] | Childs KL, Miller FR, Cordonnier-Pratt MM, et al. The sorghum photoperiod sensitivity gene, Ma3, encodes a phytochrome B [J]. Plant Physiol, 1997, 113(2): 611-619. |
| [13] | Liu HH, Liu HQ, Zhou LN, et al. Parallel domestication of the heading date 1 gene in cereals [J]. Mol Biol Evol, 2015, 32(10): 2726-2737. |
| [14] | Murphy RL, Morishige DT, Brady JA, et al. Ghd7 (Ma6) represses sorghum flowering in long days: Ghd7 alleles enhance biomass accumulation and grain production [J]. Plant Genome, 2014, 7(2): plantgenome2013.11.0040. |
| [15] | Yang SS, Murphy RL, Morishige DT, et al. Sorghum phytochrome B inhibits flowering in long days by activating expression of SbPRR37 and SbGHD7, repressors of SbEHD1, SbCN8 and SbCN12 [J]. PLoS One, 2014, 9(8): e105352. |
| [16] | Song YH, Ito S, Imaizumi T. Similarities in the circadian clock and photoperiodism in plants [J]. Curr Opin Plant Biol, 2010, 13(5): 594-603. |
| [17] | Yang MK, Lin WJ, Xu YR, et al. Flowering-time regulation by the circadian clock: From Arabidopsis to crops [J]. Crop J, 2024, 12(1): 17-27. |
| [18] | Huang XZ, Yang YF, Xu C. Biomolecular condensation programs floral transition to orchestrate flowering time and inflorescence architecture [J]. New Phytol, 2025, 245(1): 88-94. |
| [19] | Kong WQ, Kim C, Zhang D, et al. Genotyping by sequencing of 393 Sorghum bicolor BTx623×IS3620C recombinant inbred lines improves sensitivity and resolution of QTL detection [J]. G3, 2018, 8(8): 2563-2572. |
| [20] | Ding YQ, Wang YL, Xu JX, et al. A telomere-to-telomere genome assembly of Hongyingzi, a sorghum cultivar used for Chinese Baijiu production [J]. Crop J, 2024, 12(2): 635-640. |
| [21] | Bangbol Sangma H. Genetic characterization of flowering time in sorghum [D]. Queensland: The University of Queensland, 2013. |
| [22] | Parh DK. DNA-based markers for ergot resistance in sorghum [D]. Queensland: The University of Queensland, 2005. |
| [23] | Srinivas G, Satish K, Madhusudhana R, et al. Identification of quantitative trait loci for agronomically important traits and their association with genic-microsatellite markers in sorghum [J]. Theor Appl Genet, 2009, 118(8): 1439-1454. |
| [24] | Wang XM, Mace E, Hunt C, et al. Two distinct classes of QTL determine rust resistance in sorghum [J]. BMC Plant Biol, 2014, 14: 366. |
| [25] | Mantilla Perez MB, Zhao J, Yin YH, et al. Association mapping of brassinosteroid candidate genes and plant architecture in a diverse panel of Sorghum bicolor [J]. Theor Appl Genet, 2014, 127(12): 2645-2662. |
| [26] | Guindo D, Teme N, Vaksmann M, et al. Quantitative trait loci for sorghum grain morphology and quality traits: Toward breeding for a traditional food preparation of West-Africa [J]. J Cereal Sci, 2019, 85: 256-272. |
| [27] | Reddy RN, Madhusudhana R, Mohan SM, et al. Mapping QTL for grain yield and other agronomic traits in post-rainy sorghum [Sorghum bicolor (L.) Moench [J]. Theor Appl Genet, 2013, 126(8): 1921-1939. |
| [28] | Upadhyaya HD, Wang YH, Sharma R, et al. SNP markers linked to leaf rust and grain mold resistance in sorghum [J]. Mol Breed, 2013, 32(2): 451-462. |
| [29] | Feltus FA, Hart GE, Schertz KF, et al. Alignment of genetic maps and QTLs between inter- and intra-specific sorghum populations [J]. Theor Appl Genet, 2006, 112(7): 1295-1305. |
| [30] | Bouchet S, Olatoye MO, Marla SR, et al. Increased power to dissect adaptive traits in global sorghum diversity using a nested association mapping population [J]. Genetics, 2017, 206(2): 573-585. |
| [31] | Shiringani AL, Frisch M, Friedt W. Genetic mapping of QTLs for sugar-related traits in a RIL population of Sorghum bicolor L. Moench [J]. Theor Appl Genet, 2010, 121(2): 323-336. |
| [32] | Olatoye MO, Marla SR, Hu ZB, et al. Dissecting adaptive traits with nested association mapping: genetic architecture of inflorescence morphology in sorghum [J]. G3, 2020, 10(5): 1785-1796. |
| [33] | Boycheva I, Vassileva V, Revalska M, et al. Cyclin-like F-box protein plays a role in growth and development of the three model species Medicago truncatula, Lotus japonicus, and Arabidopsis thaliana [J]. Res Rep Biol, 2015, 6: 117-130. |
| [34] | Lechner E, Achard P, Vansiri A, et al. F-box proteins everywhere [J]. Curr Opin Plant Biol, 2006, 9(6): 631-638. |
| [35] | Kuroda H, Takahashi N, Shimada H, et al. Classification and expression analysis of Arabidopsis F-box-containing protein genes [J]. Plant Cell Physiol, 2002, 43(10): 1073-1085. |
| [36] | Cheng CH, Wang ZJ, Ren ZY, et al. SCFAtPP2-B11 modulates ABA signaling by facilitating SnRK2.3 degradation in Arabidopsis thaliana [J]. PLoS Genet, 2017, 13(8): e1006947. |
| [37] | 曾冰洁, 阎晋东, 杨飘, 等. 拟南芥ZTL/FKF1/LKP2蛋白家族功能研究进展 [J]. 生物信息学, 2019, 17(3): 145-150. |
| Zeng BJ, Yan JD, Yang P, et al. Progress of function studies of ZTL/FKF1/LKP2 proteins family in Arabidopsis [J]. Chin J Bioinform, 2019, 17(3): 145-150. | |
| [38] | Zoltowski BD, Imaizumi T. Structure and function of the ZTL/FKF1/LKP2 group proteins in Arabidopsis [J]. Enzymes, 2014, 35: 213-239. |
| [39] | Lee HG, Kim J, Park KH, et al. High-temperature-induced FKF1 accumulation promotes flowering through the dispersion of GI and degradation of SVP [J]. Nat Plants, 2025, 11: 1282-1297. |
| [40] | Hicks KA, Millar AJ, Carré IA, et al. Conditional circadian dysfunction of the Arabidopsis early-flowering 3 mutant [J]. Science, 1996, 274(5288): 790-792. |
| [41] | Makino S, Kiba T, Imamura A, et al. Genes encoding pseudo-response regulators: insight into his-to-asp phosphorelay and circadian rhythm in Arabidopsis thaliana [J]. Plant Cell Physiol, 2000, 41(6): 791-803. |
| [42] | Sreekantan L, Mathiason K, Grimplet J, et al. Differential floral development and gene expression in grapevines during long and short photoperiods suggests a role for floral genes in dormancy transitioning [J]. Plant Mol Biol, 2010, 73(1/2): 191-205. |
| [43] | Alabadí D, Oyama T, Yanovsky MJ, et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock [J]. Science, 2001, 293(5531): 880-883. |
| [44] | Quail PH. Phytochrome photosensory signalling networks [J]. Nat Rev Mol Cell Biol, 2002, 3(2): 85-93. |
| [45] | 彭凌涛. 控制拟南芥和水稻开花时间光周期途径的分子机制 [J]. 植物生理学通讯, 2006, 42(6): 1021-1031. |
| Peng LT. Molecular mechanism of flowering time controlling photoperiod pathway in Arabidopsis and rice [J]. Plant Physiol Commun, 2006, 42(6): 1021-1031. | |
| [46] | 贺军虎, 李唯正, 陈华蕊, 等. ‘金煌’杧果MiCAB2的克隆及表达与花期调控关系分析 [J]. 园艺学报, 2017, 44(7): 1275-1286. |
| He JH, Li WZ, Chen HR, et al. Cloning of the light harvesting chlorophyll a/b binding protein gene (MiCAB2) from ‘Jinhuang’ mango, and correlation analysis between its expression and flowering regulation [J]. Acta Hortic Sin, 2017, 44(7): 1275-1286. | |
| [47] | Zhao YG, Kong H, Guo YL, et al. Light-harvesting chlorophyll a/b-binding protein-coding genes in Jatropha and the comparison with Castor, cassava and Arabidopsis [J]. PeerJ, 2020, 8: e8465. |
| [48] | 阳江华, 张希财, 邹智. 橡胶树捕光叶绿素a/b结合蛋白基因CAB2的克隆与分析 [J]. 西南林业大学学报: 自然科学, 2019, 39(1): 88-94. |
| Yang JH, Zhang XC, Zou Z. Molecular cloning and analysis of HbCAB2, a chlorophyll a/b-binding protein-encoding gene from Hevea brasiliensis [J]. J Southwest For Univ Nat Sci, 2019, 39(1): 88-94. | |
| [49] | Zou Z, Li MY, Jia RZ, et al. Genes encoding light-harvesting chlorophyll a/b-binding proteins in papaya (Carica papaya L.) and insight into lineage-specific evolution in Brassicaceae [J]. Gene, 2020, 748: 144685. |
| [50] | 孔德元. 光周期调控甘菊成花过程中ClRVE基因家族功能分析 [D]. 北京: 北京林业大学, 2022. |
| Kong DY. Functional analysis of ClRVE gene family during photoperiod regulation of chamomile flower formation [D]. Beijing: Beijing Forestry University, 2022. | |
| [51] | Mishal R, Luna-Arias JP. Role of the TATA-box binding protein (TBP) and associated family members in transcription regulation [J]. Gene, 2022, 833: 146581. |
| [1] | 张超超, 韩开元, 王彤, 陈仲. 毛白杨PtoYABBY2和PtoYABBY12的克隆及功能分析[J]. 生物技术通报, 2025, 41(9): 256-264. |
| [2] | 胡璐, 王凯, 徐婧仪, 叶丽慧, 王永飞, 王丽华, 李杰勤. 玉米和高粱遗传转化技术研究进展[J]. 生物技术通报, 2025, 41(9): 32-43. |
| [3] | 吴夏明, 周陈平, 杨敏, 徐泽, 邝瑞彬, 刘传和, 贺涵, 魏岳荣. 基于BSA-seq的番木瓜株高候选基因定位[J]. 生物技术通报, 2025, 41(9): 54-61. |
| [4] | 刘泽洲, 段乃彬, 岳丽昕, 王清华, 姚行浩, 高莉敏, 孔素萍. 大蒜叶片蜡质成分分析及蜡质缺失基因Ggl-1筛选[J]. 生物技术通报, 2025, 41(9): 219-231. |
| [5] | 闫梦阳, 梁晓阳, 戴君昂, 张妍, 关团, 张辉, 刘良波, 孙志华. 阿莫西林降解菌的筛选及降解机制研究[J]. 生物技术通报, 2025, 41(9): 314-325. |
| [6] | 李雅琼, 格桑拉毛, 陈启迪, 杨宇环, 何花转, 赵耀飞. 异源过表达高粱SbSnRK2.1增强拟南芥对盐胁迫的抗性[J]. 生物技术通报, 2025, 41(8): 115-123. |
| [7] | 王月琛, 韩鑫骐, 魏文敏, 崔兆兰, 罗阳美, 陈鹏如, 王海岗, 刘龙龙, 张莉, 王纶. 黍稷落粒的生物学基础研究及落粒调控基因的鉴定[J]. 生物技术通报, 2025, 41(7): 164-171. |
| [8] | 赵强, 陈思宇, 彭方丽, 汪灿, 高杰, 周棱波, 张国兵, 姜昱雯, 邵明波. 间作与施氮对高粱根际土壤细菌多样性及功能的影响[J]. 生物技术通报, 2025, 41(6): 307-316. |
| [9] | 刘卓君, 柴文婷, 任毅乐, 王新宇, 朱立勋, 赵珊珊, 杨博慧, 范佳利, 李新凤, 赵威军, 吕晋慧, 张春来. 高粱TGA基因响应孢堆黑粉菌(Sporisorium reilianum)侵染表达和DNA变异分析[J]. 生物技术通报, 2025, 41(5): 90-103. |
| [10] | 彭绍智, 王登科, 张祥, 戴雄泽, 徐昊, 邹学校. 辣椒CaFD1基因克隆、表达特征及功能验证[J]. 生物技术通报, 2025, 41(5): 153-164. |
| [11] | 李旭娟, 李纯佳, 刘洪博, 徐超华, 林秀琴, 陆鑫, 刘新龙. 甘蔗腋芽形成发育过程的转录组分析[J]. 生物技术通报, 2025, 41(3): 202-218. |
| [12] | 李欣芃, 张武汉, 张莉, 舒服, 何强, 郭杨, 邓华凤, 王悦, 孙平勇. γ射线诱变创制水稻突变体及其分子鉴定[J]. 生物技术通报, 2025, 41(3): 35-43. |
| [13] | 许聪聪, 郑美, 李翠, 赵春桥, 何玮, 侯新村, 郭强. 利用WGCNA 筛选鉴定花花柴耐盐核心基因[J]. 生物技术通报, 2025, 41(12): 177-189. |
| [14] | 杜品廷, 吴国江, 王振国, 李岩, 周伟, 周亚星. 高粱CPP基因家族鉴定及表达分析[J]. 生物技术通报, 2025, 41(1): 132-142. |
| [15] | 岳丽昕, 王清华, 刘泽洲, 孔素萍, 高莉敏. 基于转录组和WGCNA筛选大葱雄性不育相关基因[J]. 生物技术通报, 2024, 40(9): 212-224. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||