








生物技术通报 ›› 2024, Vol. 40 ›› Issue (6): 134-142.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0066
白志元1( ), 徐菲1,2, 杨午1,2, 王明贵1,2, 杨玉花1, 张海平1,2(
), 徐菲1,2, 杨午1,2, 王明贵1,2, 杨玉花1, 张海平1,2( ), 张瑞军1,2(
), 张瑞军1,2( )
)
收稿日期:2024-01-16
出版日期:2024-06-26
发布日期:2024-05-14
通讯作者:
张海平,女,博士,研究员,研究方向:大豆种质资源基础与应用;E-mail: nkyzhp@126.com;作者简介:白志元,男,博士,副研究员,研究方向:大豆杂种优势基础与应用;E-mail: bzy923@163.com
基金资助:
BAI Zhi-yuan1( ), XU Fei1,2, YANG Wu1,2, WANG Ming-gui1,2, YANG Yu-hua1, ZHANG Hai-ping1,2(
), XU Fei1,2, YANG Wu1,2, WANG Ming-gui1,2, YANG Yu-hua1, ZHANG Hai-ping1,2( ), ZHANG Rui-jun1,2(
), ZHANG Rui-jun1,2( )
)
Received:2024-01-16
Published:2024-06-26
Online:2024-05-14
摘要:
【目的】 探索大豆细胞质雄性不育弱恢复型杂种F1育性转变在RNA水平上的分子机制,以期从育性转变的角度为大豆细胞质雄性不育的分子机理提供有价值的信息。【方法】 以弱恢复型杂种F1(H3A×SXTH3)为研究对象,设置苗期短光照(植株育性不育)和正常光照(植株育性可育)处理,在盛花期分别采集不同大小的混合花芽进行转录组测序和RT-qPCR分析。【结果】 筛选出3 917个差异表达基因,苗期短光照处理后,2 134个基因下调表达,1 783个基因上调表达。对差异表达基因进行生物信息学分析,GO显著富集分析表明,碳水化合物代谢过程、跨膜转运活性和细胞外围等功能在育性转变中行使着主要生物学功能;KEGG通路显著富集分析表明,戊糖葡萄糖醛酸的相互转化、淀粉蔗糖代谢和植物昼夜节律等通路为育性转变的主要代谢通路。11个基因的RT-qPCR分析发现,大豆细胞质雄性不育相关基因和PPR基因参与了大豆细胞质雄性不育弱恢复型杂种F1育性转变过程。【结论】 推测大豆细胞质雄性不育弱恢复型杂种F1育性转变与植物昼夜节律、PPR和大豆细胞质雄性不育相关的线粒体、花粉壁发育、碳水化合物代谢、糖转运和活性氧代谢等基因的异常表达相关,当昼夜节律通路的关键基因变化,引起PPR基因和大豆细胞质雄性不育相关基因的表达水平变化,将会发生育性转变。
白志元, 徐菲, 杨午, 王明贵, 杨玉花, 张海平, 张瑞军. 大豆细胞质雄性不育弱恢复型杂种F1育性转变的转录组分析[J]. 生物技术通报, 2024, 40(6): 134-142.
BAI Zhi-yuan, XU Fei, YANG Wu, WANG Ming-gui, YANG Yu-hua, ZHANG Hai-ping, ZHANG Rui-jun. Transcriptome Analysis of Fertility Transformation in Weakly Restoring Hybrid F1 of Soybean Cytoplasmic Male Sterility[J]. Biotechnology Bulletin, 2024, 40(6): 134-142.
| 基因ID Gene_id | 基因注释 Gene annotation | 正向引物序列 Forward primer sequence(5'-3') | 反向引物序列 Reverse primer sequence(5'-3') |
|---|---|---|---|
| AFR34332.1 | COX2 | TGGCTATTCCTCACGATTGCT | GCATCATAGGTGTTGCTGCG |
| AFR34381.1 | ORF103c | AAACTCGAAAGGGTGGAAGTCT | TCCCCTGCATAGGGAATACCA |
| AFR34382.1 | ORF178 | GGCATCGGTCCGTAGAACTT | TCCATAACATAACGGGGCGG |
| Glyma.02G008300 | 果胶甲基酯酶Pectin methylesterase | CTCATTAGCAAGCAGGAGGCT | AGCATCGAAAGGAACGCCAG |
| Glyma.13G064700 | 果胶裂解酶Pectin lyase | CGACAACGCCAAGGAGATCA | ATGGGTGAGCCTGAAGTGAC |
| Glyma.07G246600 | UDP葡萄糖醛酸脱羧酶 UDP glucuronic acid decarboxylase | AACCGCACCTCAGATTCACC | TGAGGAGCCGACACTATGGA |
| Glyma.20G103900 | 糖转运蛋白11 Sugar transporter protein 11 | AGCTTGTTCTTGCAAGTGAGTC | CTGAGGCCTGTATTTTGGCG |
| Glyma.02G154400 | 谷胱甘肽S转移酶Glutathione S-transferase | CTGCATCACCTCCCAACGAT | ATATCAGCCACCCATGCACC |
| Glyma.07G225400 | L抗坏血酸氧化酶L-ascorbic acid oxidase | GTGCCCCAGCAGGTTATTCT | CCCAATGGAAAAGCAAGGGC |
| Glyma.01G011300 | 含五肽重复序列的蛋白质At1g31920 PPR At1g31920 | GACCGTAATCGCTTCCACCA | AAGATGCTCAACATACCACAATG |
| Glyma.08G038200 | 含五肽重复序列的蛋白质At5g21222异构体X1 PPR At5g21222 isoform X1 | TGGGGATCACTTCTTGGTGC | TCAGTTTCCTGACCTGACGC |
表1 11个相关基因及其引物序列
Table 1 11 related genes and their primer sequences
| 基因ID Gene_id | 基因注释 Gene annotation | 正向引物序列 Forward primer sequence(5'-3') | 反向引物序列 Reverse primer sequence(5'-3') |
|---|---|---|---|
| AFR34332.1 | COX2 | TGGCTATTCCTCACGATTGCT | GCATCATAGGTGTTGCTGCG |
| AFR34381.1 | ORF103c | AAACTCGAAAGGGTGGAAGTCT | TCCCCTGCATAGGGAATACCA |
| AFR34382.1 | ORF178 | GGCATCGGTCCGTAGAACTT | TCCATAACATAACGGGGCGG |
| Glyma.02G008300 | 果胶甲基酯酶Pectin methylesterase | CTCATTAGCAAGCAGGAGGCT | AGCATCGAAAGGAACGCCAG |
| Glyma.13G064700 | 果胶裂解酶Pectin lyase | CGACAACGCCAAGGAGATCA | ATGGGTGAGCCTGAAGTGAC |
| Glyma.07G246600 | UDP葡萄糖醛酸脱羧酶 UDP glucuronic acid decarboxylase | AACCGCACCTCAGATTCACC | TGAGGAGCCGACACTATGGA |
| Glyma.20G103900 | 糖转运蛋白11 Sugar transporter protein 11 | AGCTTGTTCTTGCAAGTGAGTC | CTGAGGCCTGTATTTTGGCG |
| Glyma.02G154400 | 谷胱甘肽S转移酶Glutathione S-transferase | CTGCATCACCTCCCAACGAT | ATATCAGCCACCCATGCACC |
| Glyma.07G225400 | L抗坏血酸氧化酶L-ascorbic acid oxidase | GTGCCCCAGCAGGTTATTCT | CCCAATGGAAAAGCAAGGGC |
| Glyma.01G011300 | 含五肽重复序列的蛋白质At1g31920 PPR At1g31920 | GACCGTAATCGCTTCCACCA | AAGATGCTCAACATACCACAATG |
| Glyma.08G038200 | 含五肽重复序列的蛋白质At5g21222异构体X1 PPR At5g21222 isoform X1 | TGGGGATCACTTCTTGGTGC | TCAGTTTCCTGACCTGACGC |
| 样品 Sample | Reads总数 Total reads | 碱基总数 Total bases | Q20/% | Q30/% | 过滤读数 Clean reads | 过滤碱基数Clean bases | 过滤数百分比 Clean percentage/% | 比对总数 Total_ mapped | 比对总数百分比Total_ mapped percentage/% | 单个比对数 Uniquely_mapped | 单个比对数百分比Uniquely_mapped percentage/% |
|---|---|---|---|---|---|---|---|---|---|---|---|
| A_T_1 | 51 129 282 | 7 669 392 300 | 98.01 | 94.68 | 47 150 672 | 7 072 600 800 | 92.21 | 45 134 613 | 95.72 | 44 178 583 | 97.88 |
| A_T_2 | 51 432 310 | 7 714 846 500 | 98.03 | 94.73 | 47 431 430 | 7 114 714 500 | 92.22 | 45 474 740 | 95.87 | 44 424 415 | 97.69 |
| A_T_3 | 43 683 610 | 6 552 541 500 | 97.93 | 94.56 | 40 201 490 | 6 030 223 500 | 92.02 | 38 478 493 | 95.71 | 37 620 308 | 97.77 |
| A_CK_1 | 51 196 992 | 7 679 548 800 | 97.98 | 94.65 | 47 273 686 | 7 091 052 900 | 92.33 | 45 349 845 | 95.93 | 44 350 422 | 97.80 |
| A_CK_2 | 46 737 644 | 7 010 646 600 | 98.18 | 94.95 | 43 318 284 | 6 497 742 600 | 92.68 | 41 632 540 | 96.11 | 40 757 276 | 97.90 |
| A_CK_3 | 49 420 632 | 7 413 094 800 | 98.05 | 94.75 | 45 611 544 | 6 841 731 600 | 92.29 | 43 802 214 | 96.03 | 42 854 974 | 97.84 |
| 平均 Average | 48 933 412 | 7 340 011 750 | 98.03 | 94.72 | 45 164 518 | 6 774 677 650 | 92.29 | 43 312 074 | 95.90 | 42 364 330 | 97.81 |
表2 转录组测序数据的整理、过滤与比对统计
Table 2 Sorting, filtering and comparison statistics of transcriptome sequencing data
| 样品 Sample | Reads总数 Total reads | 碱基总数 Total bases | Q20/% | Q30/% | 过滤读数 Clean reads | 过滤碱基数Clean bases | 过滤数百分比 Clean percentage/% | 比对总数 Total_ mapped | 比对总数百分比Total_ mapped percentage/% | 单个比对数 Uniquely_mapped | 单个比对数百分比Uniquely_mapped percentage/% |
|---|---|---|---|---|---|---|---|---|---|---|---|
| A_T_1 | 51 129 282 | 7 669 392 300 | 98.01 | 94.68 | 47 150 672 | 7 072 600 800 | 92.21 | 45 134 613 | 95.72 | 44 178 583 | 97.88 |
| A_T_2 | 51 432 310 | 7 714 846 500 | 98.03 | 94.73 | 47 431 430 | 7 114 714 500 | 92.22 | 45 474 740 | 95.87 | 44 424 415 | 97.69 |
| A_T_3 | 43 683 610 | 6 552 541 500 | 97.93 | 94.56 | 40 201 490 | 6 030 223 500 | 92.02 | 38 478 493 | 95.71 | 37 620 308 | 97.77 |
| A_CK_1 | 51 196 992 | 7 679 548 800 | 97.98 | 94.65 | 47 273 686 | 7 091 052 900 | 92.33 | 45 349 845 | 95.93 | 44 350 422 | 97.80 |
| A_CK_2 | 46 737 644 | 7 010 646 600 | 98.18 | 94.95 | 43 318 284 | 6 497 742 600 | 92.68 | 41 632 540 | 96.11 | 40 757 276 | 97.90 |
| A_CK_3 | 49 420 632 | 7 413 094 800 | 98.05 | 94.75 | 45 611 544 | 6 841 731 600 | 92.29 | 43 802 214 | 96.03 | 42 854 974 | 97.84 |
| 平均 Average | 48 933 412 | 7 340 011 750 | 98.03 | 94.72 | 45 164 518 | 6 774 677 650 | 92.29 | 43 312 074 | 95.90 | 42 364 330 | 97.81 |
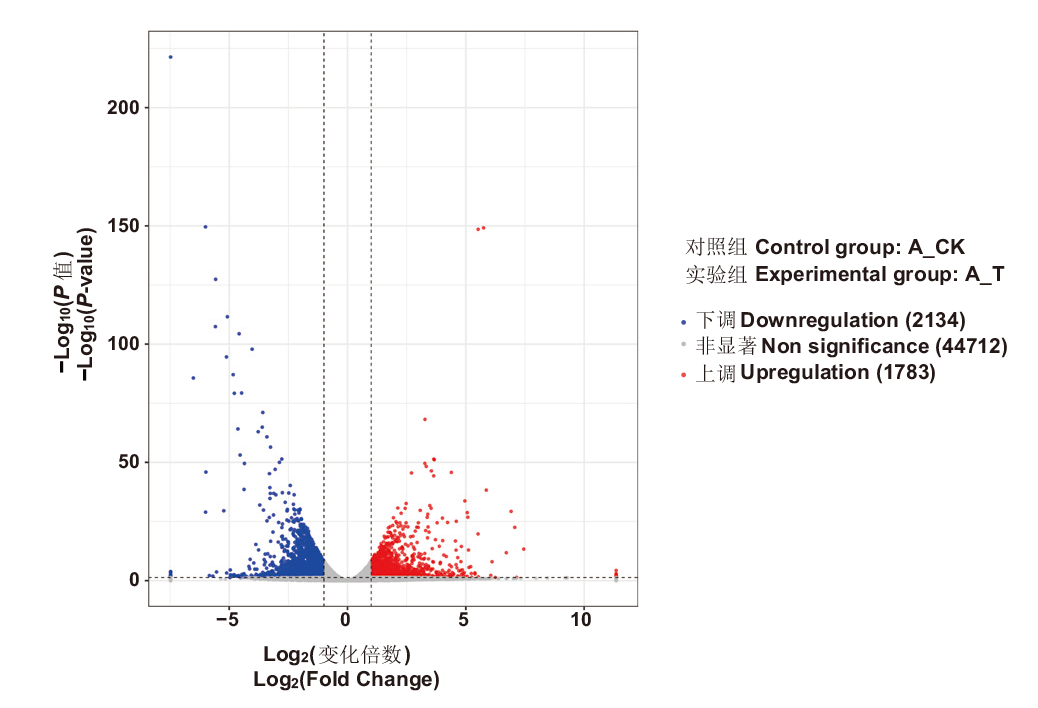
图1 大豆细胞质雄性不育弱恢复型杂种F1(H3A×SX-TH3)育性转变间差异表达基因火山图 蓝点表示差异表达基因下调,红点表示差异表达基因上调,灰点表示差异表达基因不显著
Fig. 1 Volcano map of differentially expressed genes during the fertility transformation of weakly restoring cytoplasmic male sterile hybrid F1(H3A×SXTH3)in soybean Blue dots indicate downregulation of differentially expressed genes, red dots indicate upregulation of differentially expressed genes, and gray dots indicated non-significance of differentially expressed genes
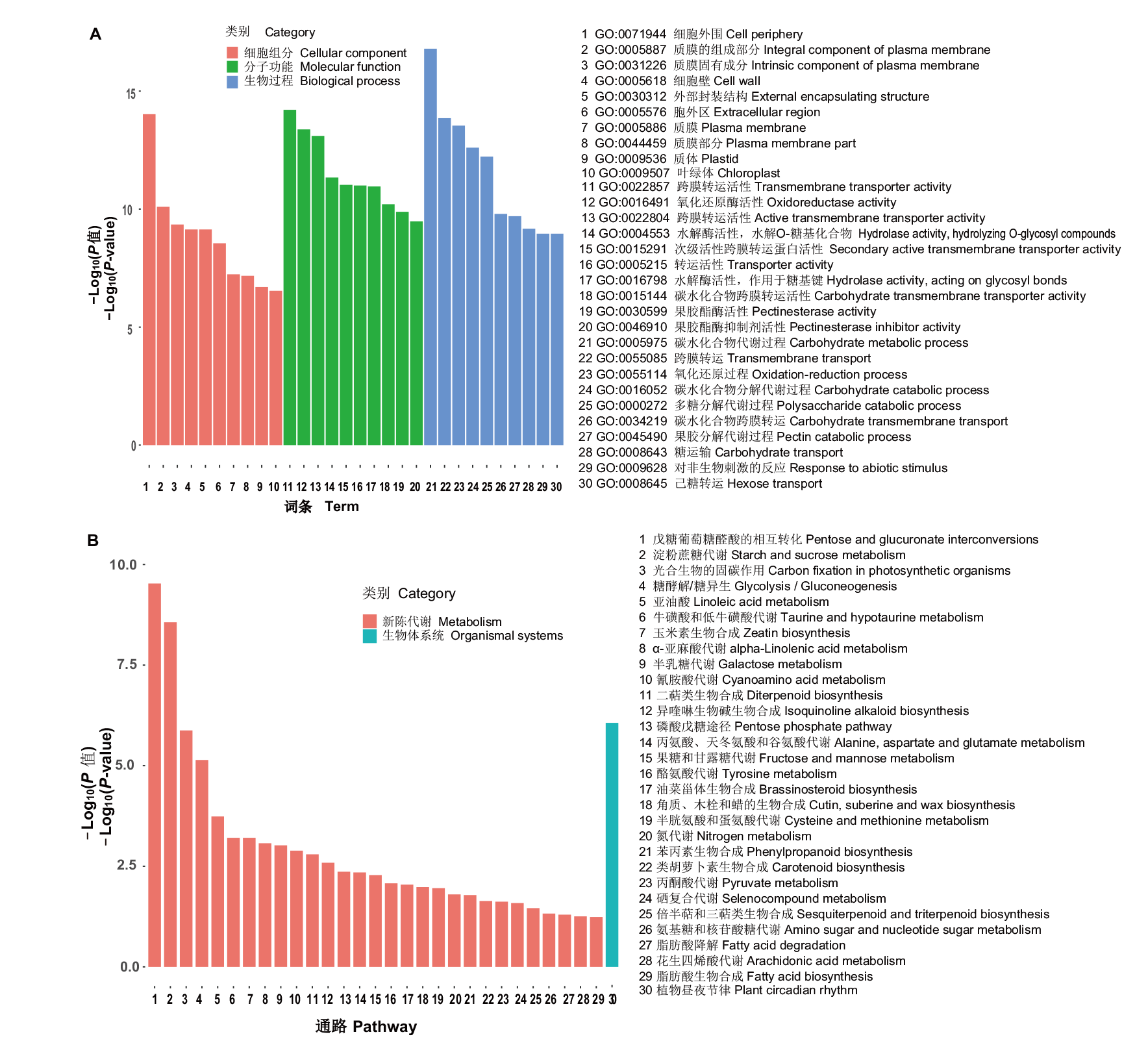
图2 大豆细胞质雄性不育弱恢复型杂种F1(H3A×SXTH3)育性转变间差异表达基因的功能富集分析 A:差异表达基因的GO富集分析;B:差异表达基因的KEGG富集分析
Fig. 2 Functional enrichment analysis of differentially expressed genes during the fertility transformation of weakly restoring cytoplasmic male sterile hybrid F1(H3A×SXTH3)in soybean A: GO enrichment analysis of differentially expressed genes. B: KEGG enrichment analysis of differentially expressed genes
| 基因ID Gene ID | A_CK平均值 baseMean_A_CK | A_T平均值 baseMean_A_T | 基因注释 Gene annotation | Log2(变化倍数) log2(FC) | P值 P value |
|---|---|---|---|---|---|
| Glyma.03G261800 | 9 951.556 4 | 55.757 5 | MYB转录因子MYB114 MYB transcription factor MYB114 | -7.479 6 | 4.09E-222 |
| Glyma.19G260900 | 3 490.330 4 | 54.321 4 | LHY蛋白 LHY protein | -6.005 7 | 2.61E-150 |
| Glyma.16G017400 | 14 705.494 2 | 897.924 7 | 晚期伸长下胚轴/昼夜节律时钟相关蛋白 Late elongation hypocotyl/ circadian clock related proteins | -4.033 6 | 1.45E-98 |
| Glyma.02G267800 | 662.352 9 | 165.158 2 | E3泛素蛋白连接酶COP1 E3 ubiquitin-protein ligase COP1 | -2.003 8 | 5.92E-20 |
| Glyma.14G049700 | 1 034.788 7 | 263.650 1 | E3泛素蛋白连接酶COP1 E3 ubiquitin-protein ligase COP1 | -1.972 6 | 2.56E-22 |
| Glyma.07G048500 | 1 425.609 8 | 402.471 1 | LHY1蛋白 LHY1 protein | -1.824 6 | 5.89E-21 |
| Glyma.07G013500 | 264.329 0 | 87.150 1 | ELF3蛋白2亚型X1 ELF3 protein 2 isoform X1 | -1.600 8 | 1.67E-09 |
| Glyma.03G225000 | 801.462 1 | 309.803 6 | 转录因子PIF3 Transcription factor PIF3 | -1.371 3 | 9.81E-12 |
| Glyma.14G174200 | 959.094 4 | 386.228 3 | 隐色素-1 Cryptochrome-1 | -1.312 2 | 2.63E-11 |
| Glyma.16G092700 | 205.123 5 | 102.488 5 | bZIP转录因子bZIP69 bZIP transcription factor bZIP69 | -1.001 0 | 0.000 170 8 |
| Glyma.04G228300 | 520.821 0 | 5 515.352 4 | 双组分响应调节器样APRR9亚型X2 Two component response regulator like APRR9 subtype X2 | 3.404 6 | 1.95E-23 |
| Glyma.16G150700 | 1.939 3 | 16.919 3 | 磷脂酰乙醇胺结合蛋白FT2a Phosphatidyl ethanolamine binding protein FT2a | 3.125 1 | 0.043 659 5 |
| Glyma.06G136600 | 232.560 3 | 1 063.196 7 | 双组分响应调节器,如PRR95 Two component response regulator-like PRR95 | 2.192 7 | 3.16E-05 |
| Glyma.07G058200 | 340.731 0 | 1 475.765 4 | PHYA-105 1亚型X2的蛋白抑制剂 Protein inhibitor of PHYA-105 1 isoform X2 | 2.114 8 | 4.15E-05 |
| Glyma.10G221500 | 2 158.991 6 | 6 635.026 4 | GIGANTEA核蛋白 GIGANTEA nuclear protein | 1.619 7 | 4.07E-10 |
| Glyma.16G027200 | 342.509 0 | 880.504 9 | PHYA-105蛋白抑制剂 Protein suppressor of PHYA-105 | 1.362 2 | 7.15E-12 |
| Glyma.01G192100 | 426.999 5 | 1 089.072 6 | 酪蛋白激酶II亚单位α-2 Casein kinase II subunit α-2 | 1.350 8 | 2.99E-12 |
| Glyma.04G050200 | 542.052 4 | 1 120.864 8 | 早花相关蛋白 Ealrly flower related proteins | 1.048 1 | 0.002 186 9 |
| Glyma.19G260400 | 586.539 7 | 1 207.006 2 | 双组分响应调节器样PRR95亚型X1 Two component response regulator-like PRR95 isoform X1 | 1.041 1 | 0.018 590 2 |
| Glyma.13G135900 | 1 456.119 6 | 2 977.895 5 | 双组分响应调节器APRR7亚型X1 Two component response regulator-like APRR7 isoform X1 | 1.032 2 | 4.15E-09 |
表3 大豆细胞质雄性不育弱恢复型杂种F1(H3A×SXTH3)育性转变间KEGG通路植物昼夜节律的差异表达基因分析
Table 3 Differentially expressed genes analysis in plant circadian rhythm of KEGG pathway during the fertility transformation of weakly restoring cytoplasmic male sterile hybrid F1(H3A×SXTH3)in soybean
| 基因ID Gene ID | A_CK平均值 baseMean_A_CK | A_T平均值 baseMean_A_T | 基因注释 Gene annotation | Log2(变化倍数) log2(FC) | P值 P value |
|---|---|---|---|---|---|
| Glyma.03G261800 | 9 951.556 4 | 55.757 5 | MYB转录因子MYB114 MYB transcription factor MYB114 | -7.479 6 | 4.09E-222 |
| Glyma.19G260900 | 3 490.330 4 | 54.321 4 | LHY蛋白 LHY protein | -6.005 7 | 2.61E-150 |
| Glyma.16G017400 | 14 705.494 2 | 897.924 7 | 晚期伸长下胚轴/昼夜节律时钟相关蛋白 Late elongation hypocotyl/ circadian clock related proteins | -4.033 6 | 1.45E-98 |
| Glyma.02G267800 | 662.352 9 | 165.158 2 | E3泛素蛋白连接酶COP1 E3 ubiquitin-protein ligase COP1 | -2.003 8 | 5.92E-20 |
| Glyma.14G049700 | 1 034.788 7 | 263.650 1 | E3泛素蛋白连接酶COP1 E3 ubiquitin-protein ligase COP1 | -1.972 6 | 2.56E-22 |
| Glyma.07G048500 | 1 425.609 8 | 402.471 1 | LHY1蛋白 LHY1 protein | -1.824 6 | 5.89E-21 |
| Glyma.07G013500 | 264.329 0 | 87.150 1 | ELF3蛋白2亚型X1 ELF3 protein 2 isoform X1 | -1.600 8 | 1.67E-09 |
| Glyma.03G225000 | 801.462 1 | 309.803 6 | 转录因子PIF3 Transcription factor PIF3 | -1.371 3 | 9.81E-12 |
| Glyma.14G174200 | 959.094 4 | 386.228 3 | 隐色素-1 Cryptochrome-1 | -1.312 2 | 2.63E-11 |
| Glyma.16G092700 | 205.123 5 | 102.488 5 | bZIP转录因子bZIP69 bZIP transcription factor bZIP69 | -1.001 0 | 0.000 170 8 |
| Glyma.04G228300 | 520.821 0 | 5 515.352 4 | 双组分响应调节器样APRR9亚型X2 Two component response regulator like APRR9 subtype X2 | 3.404 6 | 1.95E-23 |
| Glyma.16G150700 | 1.939 3 | 16.919 3 | 磷脂酰乙醇胺结合蛋白FT2a Phosphatidyl ethanolamine binding protein FT2a | 3.125 1 | 0.043 659 5 |
| Glyma.06G136600 | 232.560 3 | 1 063.196 7 | 双组分响应调节器,如PRR95 Two component response regulator-like PRR95 | 2.192 7 | 3.16E-05 |
| Glyma.07G058200 | 340.731 0 | 1 475.765 4 | PHYA-105 1亚型X2的蛋白抑制剂 Protein inhibitor of PHYA-105 1 isoform X2 | 2.114 8 | 4.15E-05 |
| Glyma.10G221500 | 2 158.991 6 | 6 635.026 4 | GIGANTEA核蛋白 GIGANTEA nuclear protein | 1.619 7 | 4.07E-10 |
| Glyma.16G027200 | 342.509 0 | 880.504 9 | PHYA-105蛋白抑制剂 Protein suppressor of PHYA-105 | 1.362 2 | 7.15E-12 |
| Glyma.01G192100 | 426.999 5 | 1 089.072 6 | 酪蛋白激酶II亚单位α-2 Casein kinase II subunit α-2 | 1.350 8 | 2.99E-12 |
| Glyma.04G050200 | 542.052 4 | 1 120.864 8 | 早花相关蛋白 Ealrly flower related proteins | 1.048 1 | 0.002 186 9 |
| Glyma.19G260400 | 586.539 7 | 1 207.006 2 | 双组分响应调节器样PRR95亚型X1 Two component response regulator-like PRR95 isoform X1 | 1.041 1 | 0.018 590 2 |
| Glyma.13G135900 | 1 456.119 6 | 2 977.895 5 | 双组分响应调节器APRR7亚型X1 Two component response regulator-like APRR7 isoform X1 | 1.032 2 | 4.15E-09 |
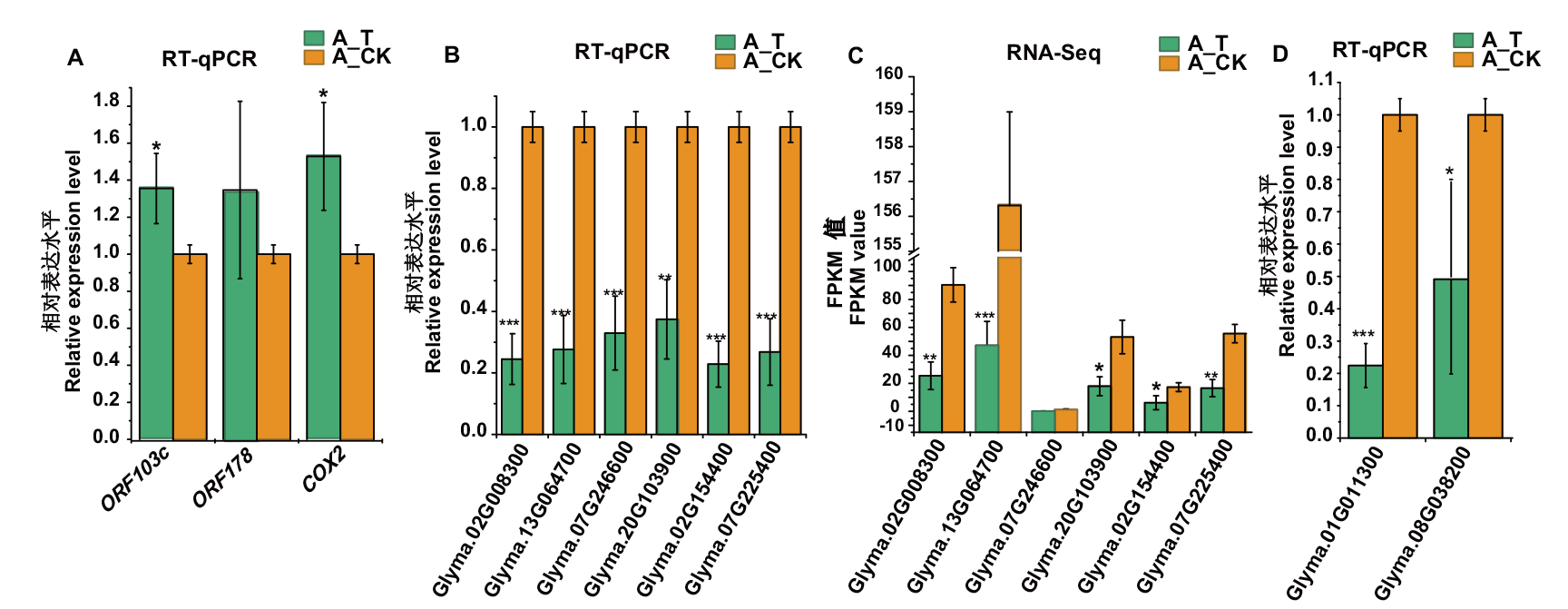
图3 大豆细胞质雄性不育弱恢复型杂种F1(H3A×SXTH3)育性转变间选择相关差异表达基因的RT-qPCR和RNA-seq分析 A:大豆细胞质雄性不育相关线粒体基因的RT-qPCR分析;B、C:大豆细胞质雄性不育相关细胞核基因的RT-qPCR和RNA-seq分析;D:PPR基因的RT-qPCR分析;*、**、***分别表示在P<0.05、P<0.01、P<0.001水平上差异显著
Fig. 3 RT-qPCR and RNA-seq analysis of selected differentially expressed genes during the fertility transformation of weakly restoring cytoplasmic male sterile hybrid F1(H3A×SXTH3)in soybean A: RT-qPCR analysis of mitochondrial genes related to cytoplasmic male sterility in soybean. B, C: RT-qPCR and RNA-seq analysis of nuclear genes related to cytoplasmic male sterility in soybean. D: RT-qPCR analysis of PPR genes. *, **, *** indicate significant difference at P<0.05, P<0.01, P<0.001 levels, respectively
| [1] | 刘杰, 黄学辉. 作物杂种优势研究现状与展望[J]. 中国科学: 生命科学, 2021, 51(10): 1396-1404. |
| Liu J, Huang XH. Advances and perspectives in crop heterosis[J]. Sci Sin Vitae, 2021, 51(10): 1396-1404. | |
| [2] |
Tester M, Langridge P. Breeding technologies to increase crop production in a changing world[J]. Science, 2010, 327(5967): 818-822.
doi: 10.1126/science.1183700 pmid: 20150489 |
| [3] | 王茜, 陈景斌, 林云, 等. 豆类作物雄性不育及杂种优势利用的研究进展[J]. 江苏农业科学, 2022, 50(4): 9-16. |
| Wang Q, Chen JB, Lin Y, et al. Research progress on male sterility and heterosis utilization in legume crops[J]. Jiangsu Agric Sci, 2022, 50(4): 9-16. | |
| [4] | 孙妍妍, 赵丽梅, 张伟, 等. 大豆杂种优势利用研究进展[J]. 大豆科技, 2021(6): 26-35. |
| Sun YY, Zhao LM, Zhang W, et al. Research progress on utilization of soybean heterosis[J]. Soybean Sci Technol, 2021(6): 26-35. | |
| [5] | 杨景华, 张明方. 线粒体反向调控介导高等植物细胞质雄性不育发生机制[J]. 遗传, 2007, 29(10): 1173-1181. |
| Yang JH, Zhang MF. Mechanism of cytoplasmic male-sterility modulated by mitochondrial retrograde regulation in higher plants[J]. Hereditas, 2007, 29(10): 1173-1181. | |
| [6] | Ji JJ, Huang W, Yin CC, et al. Mitochondrial cytochrome c oxidase and F1Fo-ATPase dysfunction in peppers(Capsicum annuum L.) with cytoplasmic male sterility and its association with orf507 and ψatp6-2 genes[J]. Int J Mol Sci, 2013, 14(1): 1050-1068. |
| [7] | Hu J, Wang K, Huang WC, et al. The rice pentatricopeptide repeat protein RF5 restores fertility in Hong-Lian cytoplasmic male-sterile lines via a complex with the glycine-rich protein GRP162[J]. Plant Cell, 2012, 24(1): 109-122. |
| [8] |
Luo DP, Xu H, Liu ZL, et al. A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice[J]. Nat Genet, 2013, 45: 573-577.
doi: 10.1038/ng.2570 pmid: 23502780 |
| [9] |
Wang K, Gao F, Ji YX, et al. ORFH79 impairs mitochondrial function via interaction with a subunit of electron transport chain complex III in Honglian cytoplasmic male sterile rice[J]. New Phytol, 2013, 198(2): 408-418.
doi: 10.1111/nph.12180 pmid: 23437825 |
| [10] |
Chen LT, Liu YG. Male sterility and fertility restoration in crops[J]. Annu Rev Plant Biol, 2014, 65: 579-606.
doi: 10.1146/annurev-arplant-050213-040119 pmid: 24313845 |
| [11] | Xiao SL, Zang J, Pei YR, et al. Activation of mitochondrial orf355 gene expression by a nuclear-encoded DREB transcription factor causes cytoplasmic male sterility in maize[J]. Mol Plant, 2020, 13(9): 1270-1283. |
| [12] | He TT, Ding XL, Zhang H, et al. Comparative analysis of mitochondrial genomes of soybean cytoplasmic male-sterile lines and their maintainer lines[J]. Funct Integr Genomics, 2021, 21(1): 43-57. |
| [13] | Li JJ, Han SH, Ding XL, et al. Comparative transcriptome analysis between the cytoplasmic male sterile line NJCMS1A and its maintainer NJCMS1B in soybean(Glycine max(L.)Merr.)[J]. PLoS One, 2015, 10(5): e0126771. |
| [14] | Bai ZY, Ding XL, Zhang RJ, et al. Transcriptome analysis reveals the genes related to pollen abortion in a cytoplasmic male-sterile soybean(Glycine max(L.)Merr.)[J]. Int J Mol Sci, 2022, 23(20): 12227. |
| [15] | Zhang CB, Fu FY, Lin CJ, et al. MicroRNAs involved in regulatory cytoplasmic male sterility by analysis RNA-seq and small RNA-seq in soybean[J]. Front Genet, 2021, 12: 654146. |
| [16] | 燕厚兴. 大豆RN型细胞质雄性不育相关基因挖掘与初步鉴定[D]. 长春: 长春师范大学, 2022. |
| Yan HX. Exploration and preliminary identification of genes related to RN-type cytoplasmic male sterility in soybean[D]. Changchun: Changchun Normal University, 2022. | |
| [17] |
白志元, 杨玉花, 张瑞军. 不同恢复型大豆细胞质雄性不育杂种F1的转录组分析[J]. 植物遗传资源学报, 2022, 23(6): 1847-1855.
doi: 10.13430/j.cnki.jpgr.20220510001 |
| Bai ZY, Yang YH, Zhang RJ. Transcriptomic analysis of soybean cytoplasmic male sterile F1hybrids from pollination with different restorer types[J]. J Plant Genet Resour, 2022, 23(6): 1847-1855. | |
| [18] |
Zhao Z, Ding Z, Huang JJ, et al. Copy number variation of the restorer Rf4 underlies human selection of three-line hybrid rice breeding[J]. Nat Commun, 2023, 14: 7333.
doi: 10.1038/s41467-023-43009-4 pmid: 37957162 |
| [19] | Lin YN, Yang HL, Liu HM, et al. A P-type pentatricopeptide repeat protein ZmRF5 promotes 5' region partial cleavages of atp6c transcripts to restore the fertility of CMS-C maize by recruiting a splicing factor[J]. Plant Biotechnol J, 2023, 11:38073308. |
| [20] | 张井勇, 孙寰, 赵丽梅, 等. 大豆RN不育胞质不育与恢复类型的研究[J]. 大豆科学, 2010, 29(4): 559-564. |
| Zhang JY, Sun H, Zhao LM, et al. Classification of male-sterile lines with RN sterile cytoplasm and their restorers[J]. Soybean Sci, 2010, 29(4): 559-564. | |
| [21] | 白志元, 杨玉花, 张瑞军. 苗期短光照对三系杂交大豆及亲本育性的影响[J]. 中国农业大学学报, 2021, 26(9): 11-17. |
| Bai ZY, Yang YH, Zhang RJ. Effects of short light at seedling stage on fertility of three-line hybrid soybean and its parents[J]. J China Agric Univ, 2021, 26(9): 11-17. | |
| [22] | 门中华, 李生秀. 植物生物节律性研究进展[J]. 生物学杂志, 2009, 26(5): 53-55. |
| Men ZH, Li SX. Progress in the study of plant biorhythm[J]. J Biol, 2009, 26(5): 53-55. | |
| [23] |
Harmer SL. The circadian system in higher plants[J]. Annu Rev Plant Biol, 2009, 60: 357-377.
doi: 10.1146/annurev.arplant.043008.092054 pmid: 19575587 |
| [24] |
Ye JL, Duan Y, Hu G, et al. Identification of candidate genes and biosynthesis pathways related to fertility conversion by wheat KTM3315A transcriptome profiling[J]. Front plant sci, 2017, 8: 449.
doi: 10.3389/fpls.2017.00449 pmid: 28428792 |
| [25] | Yu JP, Zhao GL, Li W, et al. A single nucleotide polymorphism in an R2R3 MYB transcription factor gene triggers the male sterility in soybean ms6(Ames1)[J]. Theor Appl Genet, 2021, 134(11): 3661-3674. |
| [26] | 聂智星. 大豆新雄性不育细胞质的发掘, 核不育基因的定位和产量杂种优势的QTL-等位变异解析[D]. 南京: 南京农业大学, 2017. |
| Nie ZX. Discovery of a new cytoplasmic male-sterile source, detection of a nuclear male-sterile gene and QTL-allele constitutions of heterosis in soybean[Glycine max(L.)merr.][D]. Nanjing: Nanjing Agricultural University, 2017. | |
| [27] |
Uyttewaal M, Arnal N, Quadrado M, et al. Characterization of raphanus sativus pentatricopeptide repeat proteins encoded by the fertility restorer locus for ogura cytoplasmic male sterility[J]. Plant Cell, 2008, 20(12): 3331-3345.
doi: 10.1105/tpc.107.057208 pmid: 19098270 |
| [28] | Yang SP, Duan MP, Meng QC, et al. Inheritance and gene tagging of male fertility restoration of cytoplasmic-nuclear male-sterile line NJCMS1A in soybean[J]. Plant Breed, 2007, 126(3): 302-305. |
| [29] | Wang TL, He TT, Ding XL, et al. Confirmation of GmPPR576 as a fertility restorer gene of cytoplasmic male sterility in soybean[J]. J Exp Bot, 2021, 72(22): 7729-7742. |
| [30] | Sun YY, Zhang Y, Jia SG, et al. Identification of a candidate restorer-of-fertility gene Rf3 encoding a pentatricopeptide repeat protein for the cytoplasmic male sterility in soybean[J]. Int J Mol Sci, 2022, 23: 5388. |
| [1] | 秦健, 李振月, 何浪, 李俊玲, 张昊, 杜荣. 肌源性细胞分化的单细胞转录谱变化及细胞间通讯分析[J]. 生物技术通报, 2024, 40(6): 330-342. |
| [2] | 张震, 李清, 徐菁, 陈凯园, 张春芝, 祝光涛. 马铃薯线粒体靶向表达载体的构建与应用[J]. 生物技术通报, 2024, 40(5): 66-73. |
| [3] | 娄银, 高浩竣, 王茜, 牛景萍, 王敏, 杜维俊, 岳爱琴. 大豆GmHMGS基因的鉴定及表达模式分析[J]. 生物技术通报, 2024, 40(4): 110-121. |
| [4] | 杨淇, 魏子迪, 宋娟, 童堃, 杨柳, 王佳涵, 刘海燕, 栾维江, 马轩. 水稻组蛋白H1三突变体的创建和转录组学分析[J]. 生物技术通报, 2024, 40(4): 85-96. |
| [5] | 王帅, 冯宇梅, 白苗, 杜维俊, 岳爱琴. 大豆GmHMGR基因响应外源激素及非生物胁迫功能研究[J]. 生物技术通报, 2023, 39(7): 131-142. |
| [6] | 李文辰, 刘鑫, 康越, 李伟, 齐泽铮, 于璐, 王芳. TRV病毒诱导大豆基因沉默体系优化及应用[J]. 生物技术通报, 2023, 39(7): 143-150. |
| [7] | 赵金玲, 安磊, 任晓亮. 单细胞转录组测序技术及其在秀丽隐杆线虫中的应用[J]. 生物技术通报, 2023, 39(6): 158-170. |
| [8] | 翟莹, 李铭杨, 张军, 赵旭, 于海伟, 李珊珊, 赵艳, 张梅娟, 孙天国. 异源表达大豆转录因子GmNF-YA19提高转基因烟草抗旱性[J]. 生物技术通报, 2023, 39(5): 224-232. |
| [9] | 侯筱媛, 车郑郑, 李姮静, 杜崇玉, 胥倩, 王群青. 大豆膜系统cDNA文库的构建及大豆疫霉效应子PsAvr3a互作蛋白的筛选[J]. 生物技术通报, 2023, 39(4): 268-276. |
| [10] | 杨春洪, 董璐, 陈林, 宋丽. 大豆VAS1基因家族的鉴定及参与侧根发育的研究[J]. 生物技术通报, 2023, 39(3): 133-142. |
| [11] | 汪格格, 邱诗蕊, 张琳晗, 杨国伟, 徐小云, 汪爱羚, 曾淑华, 刘雅洁. 异源三倍体普通烟草(SST)减数分裂期的分子细胞学研究[J]. 生物技术通报, 2023, 39(2): 183-192. |
| [12] | 陈奕博, 杨万明, 岳爱琴, 王利祥, 杜维俊, 王敏. 基于SLAF标记的大豆遗传图谱构建及苗期耐盐性QTL定位[J]. 生物技术通报, 2023, 39(2): 70-79. |
| [13] | 苗淑楠, 高宇, 李昕儒, 蔡桂萍, 张飞, 薛金爱, 季春丽, 李润植. 大豆GmPDAT1参与油脂合成和非生物胁迫应答的功能分析[J]. 生物技术通报, 2023, 39(2): 96-106. |
| [14] | 白苗, 田雯青, 武帅, 王敏, 王利祥, 岳爱琴, 牛景萍, 张永坡, 高春艳, 张武霞, 郭数进, 杜维俊, 赵晋忠. 激素和逆境胁迫对大豆维生素E和γ-TMT表达的影响[J]. 生物技术通报, 2023, 39(10): 148-162. |
| [15] | 于惠林, 吴孔明. 中国转基因大豆的产业化策略[J]. 生物技术通报, 2023, 39(1): 1-15. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||