








生物技术通报 ›› 2024, Vol. 40 ›› Issue (9): 282-290.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0213
收稿日期:2024-03-06
出版日期:2024-09-26
发布日期:2024-10-12
通讯作者:
赵斌,男,博士,副教授,研究方向:新农药作用机理;E-mail: bdzhaobin@126.com;作者简介:侯智涵,男,硕士,研究方向:植物保护;E-mail: 15630858209@163.com
基金资助:
HOU Zhi-han( ), HAO Nan, LI Jia-qi, ZHAO Bin(
), HAO Nan, LI Jia-qi, ZHAO Bin( ), LIU Ying-chao(
), LIU Ying-chao( )
)
Received:2024-03-06
Published:2024-09-26
Online:2024-10-12
摘要:
【目的】拟轮枝镰孢(Fusarium verticillioides)是一种危害严重的植物病原真菌,极大降低了粮食产量,还产生2B类致癌物伏马毒素,威胁人畜健康。探究RNA甲基化修饰与伏马毒素之间的联系,解析RNA修饰甲基化转移酶在伏马毒素合成中的作用。【方法】利用HPLC检测了不同地区的拟轮枝镰孢菌株的伏马毒素合成能力,采用QuEChERS前处理结合超高效液相色谱-串联质谱(UPLC-MS/MS)技术建立了m6A、mlA、m5C、Gm、m7G和Um等6种RNA甲基化修饰检测方法,继而对不同产毒菌株的RNA甲基化修饰进行了检测,并采用生物信息学和RT-qPCR方法确定了与伏马毒素合成相关的RNA甲基化修饰基因。【结果】不同地区的拟轮枝镰孢菌株伏马毒素合成能力具有显著差异,成功建立了RNA甲基化修饰检测方法,并确定mlA和m5C RNA甲基化修饰与伏马毒素合成负相关,RT-qPCR发现Fvalyref基因负调控伏马毒素生物合成。【结论】RNA m5C甲基化修饰与伏马毒素生物合成呈负相关且其Reader基因Fvalyref负调控伏马毒素生物合成。
侯智涵, 郝楠, 李佳琪, 赵斌, 刘颖超. RNA m1A和m5C甲基化修饰在拟轮枝镰孢伏马毒素生物合成中的作用[J]. 生物技术通报, 2024, 40(9): 282-290.
HOU Zhi-han, HAO Nan, LI Jia-qi, ZHAO Bin, LIU Ying-chao. Roles of RNA m1A and m5C Methylation Modifications in the Fumonisin Biosynthesis of Fusarium verticillioides[J]. Biotechnology Bulletin, 2024, 40(9): 282-290.
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') |
|---|---|
| 18S ribosomal RNA-F 18S ribosomal RNA-R Fvnsun2-F Fvnsun2-R Fvnsun4-F Fvnsun4-R Fvalyref-F Fvalyref-R | GGCCGTTCTTAGTTGGTGGA TGCGGCCCAGAACATCTAAG AGACCTTCCGCAAGCTTCTC CGCTCTCTAAAGTCGGGGTC TGGGTGGTTTCGATCGTGTT TAAACAATGTAGCCGCCCGT AGGCCGTGCTATTGAAGTCC AAAGGGTTAGGAGGGACGGA |
表1 试验所用引物
Table 1 Primers used in the experiment
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') |
|---|---|
| 18S ribosomal RNA-F 18S ribosomal RNA-R Fvnsun2-F Fvnsun2-R Fvnsun4-F Fvnsun4-R Fvalyref-F Fvalyref-R | GGCCGTTCTTAGTTGGTGGA TGCGGCCCAGAACATCTAAG AGACCTTCCGCAAGCTTCTC CGCTCTCTAAAGTCGGGGTC TGGGTGGTTTCGATCGTGTT TAAACAATGTAGCCGCCCGT AGGCCGTGCTATTGAAGTCC AAAGGGTTAGGAGGGACGGA |
| 菌株编号 Strain number | 菌株来源 Strain source | FB1含量 Content of FB1/(mg·g-1) |
|---|---|---|
| 87 | 14-SD-94-6 | 145.20 |
| 64 | HK12-5 | 116.90 |
| 114 | 14-17-15 | 228.20 |
| 126 | 14-SD-91-13 | 116.80 |
| 81 | HN 22-2 | 112.30 |
| 104 | 25-4 | 89.55 |
| 50 | LZ-2-81 | 103.57 |
| 80 | NMG-1-2 | 92.44 |
| 84 | 124-3 | 64.73 |
| 86 | HN 24-2 | 99.39 |
| 63 | S-9-7 | 101.82 |
| 6 | M-I1-46 | 59.29 |
| 38 | M-10-48 | ND |
| 1 | LZ-2-77甘 | 2.18 |
| 49 | P-HB-T3-4 | 2.19 |
| 55 | NMG-3-3 | ND |
| 109 | M-3-20 | ND |
| 36 | 26-4 | ND |
| 18 | 14-SD-22-20 | ND |
| 37 22 | 14-SD-91-9 M-4-78 | ND ND |
| 40 139 | ZL-1-6 14-SD-91-9 | ND 1.81 |
| 33 | P-C1-32-1 | ND |
表2 不同地区拟轮枝镰孢菌株FB1含量测定
Table 2 Determination of FB1 content of F. verticillioides strain in different regions
| 菌株编号 Strain number | 菌株来源 Strain source | FB1含量 Content of FB1/(mg·g-1) |
|---|---|---|
| 87 | 14-SD-94-6 | 145.20 |
| 64 | HK12-5 | 116.90 |
| 114 | 14-17-15 | 228.20 |
| 126 | 14-SD-91-13 | 116.80 |
| 81 | HN 22-2 | 112.30 |
| 104 | 25-4 | 89.55 |
| 50 | LZ-2-81 | 103.57 |
| 80 | NMG-1-2 | 92.44 |
| 84 | 124-3 | 64.73 |
| 86 | HN 24-2 | 99.39 |
| 63 | S-9-7 | 101.82 |
| 6 | M-I1-46 | 59.29 |
| 38 | M-10-48 | ND |
| 1 | LZ-2-77甘 | 2.18 |
| 49 | P-HB-T3-4 | 2.19 |
| 55 | NMG-3-3 | ND |
| 109 | M-3-20 | ND |
| 36 | 26-4 | ND |
| 18 | 14-SD-22-20 | ND |
| 37 22 | 14-SD-91-9 M-4-78 | ND ND |
| 40 139 | ZL-1-6 14-SD-91-9 | ND 1.81 |
| 33 | P-C1-32-1 | ND |
| 核苷Nucleotide | 母离子Precursorion(m/z) | 子离子Production(m/z) | 保留时间Residence time/ ms | 去簇电压DP/V | 碰撞电压CE /V |
|---|---|---|---|---|---|
| Um | 259.3 | 113.1 | 0.94 | 17 | 14 |
| Gm | 298.2 | 152.1 | 1.10 | 8 | 16 |
| m1A | 282.1 | 150.1 | 0.72 | 36 | 30 |
| m6A | 282.2 | 150.2 | 2.40 | 39 | 20 |
| m5C | 258.3 | 126.1 | 0.68 | 24 | 18 |
| m7G | 298.1 | 166.1 | 0.76 | 20 | 18 |
表3 6种目标核苷的保留时间和质谱参数
Table 3 Retention time and mass spectrum parameters of 6 target nucleosides
| 核苷Nucleotide | 母离子Precursorion(m/z) | 子离子Production(m/z) | 保留时间Residence time/ ms | 去簇电压DP/V | 碰撞电压CE /V |
|---|---|---|---|---|---|
| Um | 259.3 | 113.1 | 0.94 | 17 | 14 |
| Gm | 298.2 | 152.1 | 1.10 | 8 | 16 |
| m1A | 282.1 | 150.1 | 0.72 | 36 | 30 |
| m6A | 282.2 | 150.2 | 2.40 | 39 | 20 |
| m5C | 258.3 | 126.1 | 0.68 | 24 | 18 |
| m7G | 298.1 | 166.1 | 0.76 | 20 | 18 |
| 核苷 Nucleotide | 检出限LOD /(μg·kg-1) | 基质效应 Matrix effect/% | 线性方程 Linearity equation | 决定系数R2 Determination coefficient | 平均回收率/相对标准偏差 Average recovery(%)/ RSD | |||
|---|---|---|---|---|---|---|---|---|
| 1 μg/kg | 20 μg/kg | 500 μg/kg | ||||||
| Um | 0.002 | 5.41 | y =4E+06x - 2964.3 | 0.9979 | 91.9/4.7 | 84.4/4.5 | 91.9/3 | |
| Gm | 0.001 | 2.27 | y =2E+06x - 5329.46 | 0.9999 | 97.6/6.6 | 86.2/7.8 | 109.7/4.9 | |
| m1A | 0.0009 | 2.16 | y = 7E+07x - 161526 | 0.9990 | 72.5/4.9 | 75.1/1.9 | 74.4/7.6 | |
| m6A | 0.0006 | 2.13 | y = 7E+07x - 30294 | 0.9981 | 88.5/7.0 | 78.7/5.9 | 116.5/1.4 | |
| m5C | 0.0003 | 2.51 | y = 3E+06x - 1896.4 | 0.9993 | 70.3/6.5 | 71.0/4.7 | 74.1/7.2 | |
| m7G | 0.0006 | 1.95 | y = 6E+07x - 141064 | 0.9995 | 74.0/8.9 | 72.1/7.7 | 70.9/9.5 | |
表4 6种核苷的线性方程、决定系数、平均回收率以及相对标准偏差
Table 4 Linear equation, determination coefficient, average recovery, and relative standard deviation of 6 nucleosides
| 核苷 Nucleotide | 检出限LOD /(μg·kg-1) | 基质效应 Matrix effect/% | 线性方程 Linearity equation | 决定系数R2 Determination coefficient | 平均回收率/相对标准偏差 Average recovery(%)/ RSD | |||
|---|---|---|---|---|---|---|---|---|
| 1 μg/kg | 20 μg/kg | 500 μg/kg | ||||||
| Um | 0.002 | 5.41 | y =4E+06x - 2964.3 | 0.9979 | 91.9/4.7 | 84.4/4.5 | 91.9/3 | |
| Gm | 0.001 | 2.27 | y =2E+06x - 5329.46 | 0.9999 | 97.6/6.6 | 86.2/7.8 | 109.7/4.9 | |
| m1A | 0.0009 | 2.16 | y = 7E+07x - 161526 | 0.9990 | 72.5/4.9 | 75.1/1.9 | 74.4/7.6 | |
| m6A | 0.0006 | 2.13 | y = 7E+07x - 30294 | 0.9981 | 88.5/7.0 | 78.7/5.9 | 116.5/1.4 | |
| m5C | 0.0003 | 2.51 | y = 3E+06x - 1896.4 | 0.9993 | 70.3/6.5 | 71.0/4.7 | 74.1/7.2 | |
| m7G | 0.0006 | 1.95 | y = 6E+07x - 141064 | 0.9995 | 74.0/8.9 | 72.1/7.7 | 70.9/9.5 | |
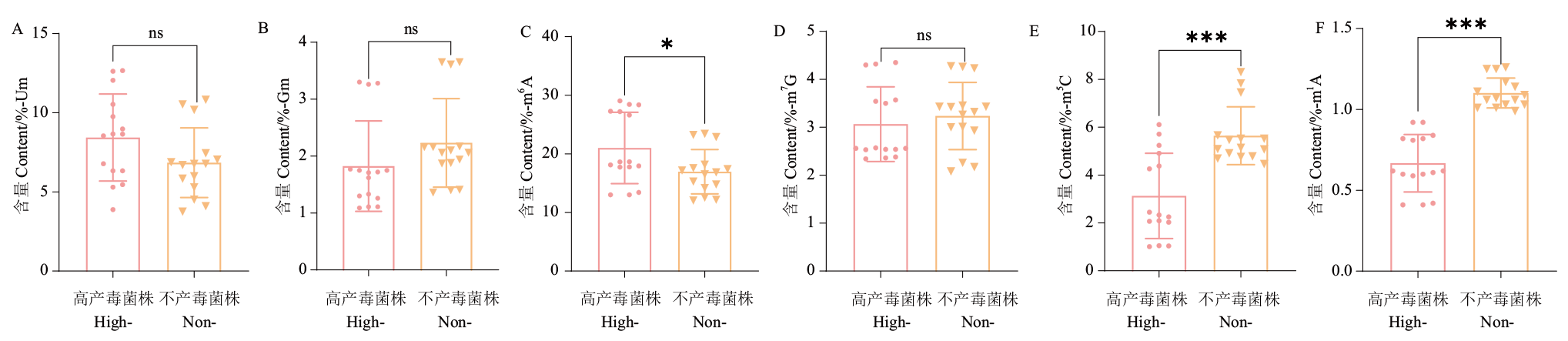
图2 产毒差异菌株中RNA修饰的变化 A:Um修饰含量;B:Gm修饰含量;C:m6A修饰含量;D:m7G修饰含量;E:m5C修饰含量;F:m1A修饰含量。High-toxicity-producing strains缩写为High-;Non-toxicity-producing strains缩写为Non-。图中数据为平均值±标准误差,ns:无显著性差异,*(P<0.05),**(P<0.01),***(P<0.001)。下同
Fig. 2 Changes of RNA modification in toxicity-yielding differential strains A: Um modification content; B: Gm modification content; C: m6A modification content; D: m7G modification content; E: m5C modification content; F: m1A modification content. High-toxicity-producing strains abbreviated as High-. Non-toxicity-producing strains abbreviated as Non-. Data in the figure are mean ± SE. ns: No significant difference,* (P<0.05), ** (P<0.05), *** (P<0.001). The same below
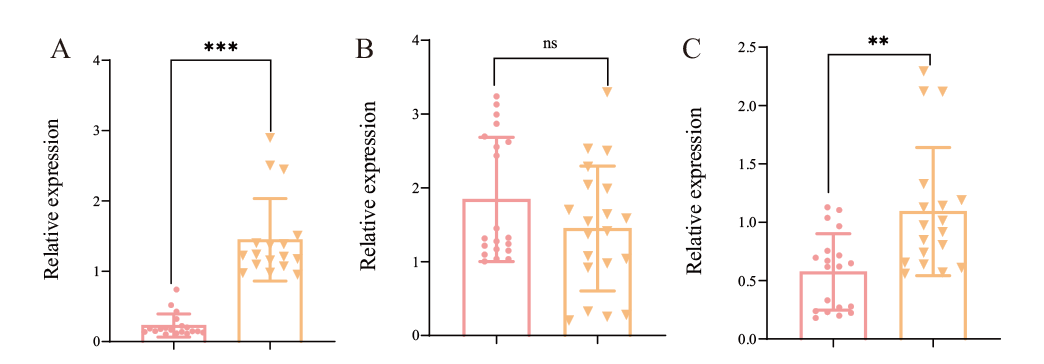
图4 m5C修饰相关基因在不同产毒菌株中的表达 A:Fvalyref在不同含量伏马毒素菌株表达量;B-C:Fvnsun2与Fvnsun4在不同含量伏马毒素菌株表达量
Fig. 4 Expression of m5C modification-related genes in different toxin-producing strains A: Expression of Fvalyref in the strains with different fumonisin contents; B-C: Expressions of Fvnsun2 and Fvnsun4 in the strains with different fumonisin content
| [1] | Deepa N, Achar PN, Sreenivasa MY. Current perspectives of biocontrol agents for management of Fusarium verticillioides and its fumonisin in cereals-a review[J]. J Fungi, 2021, 7(9): 776. |
| [2] |
Han SL, Wang MM, Ma ZY, et al. Fusarium diversity associated with diseased cereals in China, with an updated phylogenomic assessment of the genus[J]. Stud Mycol, 2023, 104: 87-148.
doi: 10.3114/sim.2022.104.02 pmid: 37351543 |
| [3] | Iqbal N, Czékus Z, Poór P, et al. Plant defence mechanisms against mycotoxin Fumonisin B1[J]. Chem Biol Interact, 2021, 343: 109494. |
| [4] | Carvajal-Moreno M. Mycotoxin challenges in maize production and possible control methods in the 21st century[J]. J Cereal Sci, 2022, 103: 103293. |
| [5] |
Winter G, Pereg L. A review on the relation between soil and mycotoxins: effect of aflatoxin on field, food and finance[J]. Eur J Soil Sci, 2019, 70(4): 882-897.
doi: 10.1111/ejss.12813 |
| [6] | Desmond J. A modest proposal: a response to the marketing challenges presented by the crisis confronting humanity in respect to the requirement to feed nine billion by 2050[J]. J Mark Manag, 2013, 29(13/14): 1631-1643. |
| [7] |
Dai YQ, Huang KL, Zhang BY, et al. Aflatoxin B1-induced epigenetic alterations: an overview[J]. Food Chem Toxicol, 2017, 109(Pt 1): 683-689.
doi: S0278-6915(17)30342-3 pmid: 28645871 |
| [8] | Teng PC, Liang YW, Yarmishyn AA, et al. RNA modifications and epigenetics in modulation of lung cancer and pulmonary diseases[J]. Int J Mol Sci, 2021, 22(19): 10592. |
| [9] | 马士清, 彭金英, 伊成器. RNA修饰检测技术[J]. 生命科学, 2018, 30(4): 440-446. |
| Ma SQ, Peng JY, Yi CQ. The detection methods of RNA modifications[J]. Chin Bull Life Sci, 2018, 30(4): 440-446. | |
| [10] | Yu HW, Raza SHA, Zhang WZ, et al. Research progress of m6A regulation during animal growth and development[J]. Mol Cell Probes, 2022, 65: 101851. |
| [11] | Ghazi T, Nagiah S, Chuturgoon AA. Fusaric acid induces hepatic global m6A RNA methylation and differential expression of m6A regulatory genes in vivo - a pilot study[J]. Epigenetics, 2022, 17(6): 695-703. |
| [12] |
Růžička K, Zhang M, Campilho A, et al. Identification of factors required for m6 A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI[J]. New Phytol, 2017, 215(1): 157-172.
doi: 10.1111/nph.14586 pmid: 28503769 |
| [13] |
Zou MJ, Mu Y, Chai X, et al. The critical function of the plastid rRNA methyltransferase, CMAL, in ribosome biogenesis and plant development[J]. Nucleic Acids Res, 2020, 48(6): 3195-3210.
doi: 10.1093/nar/gkaa129 pmid: 32095829 |
| [14] |
Zhang C, Zhang H, Zhu QQ, et al. Overexpression of global regulator LaeA increases secondary metabolite production in Monascus purpureus[J]. Appl Microbiol Biotechnol, 2020, 104(7): 3049-3060.
doi: 10.1007/s00253-020-10379-4 pmid: 32043189 |
| [15] | Wang B, Li XJ, Tabudravu J, et al. The chemical profile of activated secondary metabolites by overexpressing LaeA in Aspergillus niger[J]. Microbiol Res, 2021, 248: 126735. |
| [16] | Wang YS, Chen YC, Zhang J, et al. Overexpression of llm1 affects the synthesis of secondary metabolites of Aspergillus cristatus[J]. Microorganisms, 2022, 10(9): 1707. |
| [17] | Yadav PK, Rajasekharan R. The m6A methyltransferase Ime4 epitranscriptionally regulates triacylglycerol metabolism and vacuolar morphology in haploid yeast cells[J]. J Biol Chem, 2017, 292(33): 13727-13744. |
| [18] | Zhang YW, Wang YM, Fan JL, et al. Aspergillus fumigatus Elongator complex subunit 3 affects hyphal growth, adhesion and virulence through wobble uridine tRNA modification[J]. PLoS Pathog, 2022, 18(11): e1010976. |
| [19] | Ren ZY, Tang BZ, Xing JJ, et al. MTA1-mediated RNA m6 A modification regulates autophagy and is required for infection of the rice blast fungus[J]. New Phytol, 2022, 235(1): 247-262. |
| [20] |
Huang DY, Cui LQ, Sajid A, et al. The epigenetic mechanisms in Fusarium mycotoxins induced toxicities[J]. Food Chem Toxicol, 2019, 123: 595-601.
doi: S0278-6915(18)30798-1 pmid: 30599843 |
| [21] | 刘静. 拟轮枝镰孢ORPs家族主效蛋白鉴定及其功能解析[D]. 保定: 河北农业大学, 2023. |
| Liu J. Identification and functional analysis of major protein from ORPs family of Fusarium verticillioides[D]. Baoding: Hebei Agricultural University, 2023. | |
| [22] | Tian F, Lee SY, Woo SY, et al. Effect of plant-based compounds on the antifungal and antiaflatoxigenic efficiency of strobilurins against Aspergillus flavus[J]. J Hazard Mater, 2021, 415: 125663. |
| [23] | Bryła M, Roszko M, Szymczyk K, et al. Fumonisins in plant-origin food and fodder—a review[J]. Food Addit Contam Part A Chem Anal Control Expo Risk Assess, 2013, 30(9): 1626-1640. |
| [24] | Yang X, Gao J, Liu Q, et al. Co-occurrence of mycotoxins in maize and maize-derived food in China and estimation of dietary intake[J]. Food Addit Contam Part B Surveill, 2019, 12(2): 124-134. |
| [25] | 王燕, 董燕婕, 岳晖, 等. 山东省玉米真菌毒素污染状况调查及分析[J]. 粮油食品科技, 2016, 24(3): 69-73. |
| Wang Y, Dong YJ, Yue H, et al. Investigatin and analysis on mycotoxins contamination of maize in Shandong Province[J]. Sci Technol Cereals Oils Foods, 2016, 24(3): 69-73. | |
| [26] | Hu L, Liu HW, Yang J, et al. Free and hidden fumonisins in raw maize and maize-based products from China[J]. Food Addit Contam Part B Surveill, 2019, 12(2): 90-96. |
| [27] |
Covarelli L, Stifano S, Beccari G, et al. Characterization of Fusarium verticillioides strains isolated from maize in Italy: fumonisin production, pathogenicity and genetic variability[J]. Food Microbiol, 2012, 31(1): 17-24.
doi: 10.1016/j.fm.2012.02.002 pmid: 22475938 |
| [28] | Li LL, He ZQ, Shi Y, et al. Role of epigenetics in mycotoxin toxicity: a review[J]. Environ Toxicol Pharmacol, 2023, 100: 104154. |
| [29] | Yang C, Wu DD, Lin H, et al. Role of RNA modifications, especially m6A, in aflatoxin biosynthesis of Aspergillus flavus[J]. J Agric Food Chem, 2024, 72(1): 726-741. |
| [30] | Liang LK, Wang XY, Lan HE, et al. Comprehensive analysis of aflatoxin B1 biosynthesis in Aspergillus flavus via transcriptome-wide m6A methylome response to cycloleucine[J]. J Hazard Mater, 2024, 461: 132677. |
| [31] | Shafik AM, Zhou HQ, Lim J, et al. Dysregulated mitochondrial and cytosolic tRNA m1A methylation in Alzheimer's disease[J]. Hum Mol Genet, 2022, 31(10): 1673-1680. |
| [32] | Boo SH, Ha H, Kim YK. m1A and m6A modifications function cooperatively to facilitate rapid mRNA degradation[J]. Cell Rep, 2022, 40(10): 111317. |
| [33] | Qi ZY, Zhang C, Jian H, et al. N1-Methyladenosine modification of mRNA regulates neuronal gene expression and oxygen glucose deprivation/reoxygenation induction[J]. Cell Death Discov, 2023, 9(1): 159. |
| [34] | Trixl L, Lusser A. The dynamic RNA modification 5-methylcytosine and its emerging role as an epitranscriptomic mark[J]. Wiley Interdiscip Rev RNA, 2019, 10(1): e1510-e1527. |
| [35] | Zhao Y, Xing C, Peng HL. ALYREF(Aly/REF export factor): a potential biomarker for predicting cancer occurrence and therapeutic efficacy[J]. Life Sci, 2024, 338: 122372. |
| [36] |
Klec C, Knutsen E, Schwarzenbacher D, et al. ALYREF, a novel factor involved in breast carcinogenesis, acts through transcriptional and post-transcriptional mechanisms selectively regulating the short NEAT1 isoform[J]. Cell Mol Life Sci, 2022, 79(7): 391.
doi: 10.1007/s00018-022-04402-2 pmid: 35776213 |
| [37] | Wang N, Chen RX, Deng MH, et al. m5C-dependent cross-regulation between nuclear reader ALYREF and writer NSUN2 promotes urothelial bladder cancer malignancy through facilitating RABL6/TK1 mRNAs splicing and stabilization[J]. Cell Death Dis, 2023, 14(2): 139. |
| [38] | Berson A, Goodman LD, Sartoris AN, et al. Drosophila Ref1/ALYREF regulates transcription and toxicity associated with ALS/FTD disease etiologies[J]. Acta Neuropathol Commun, 2019, 7(1): 65. |
| [39] |
Kow RL, Black AH, Saxton AD, et al. Loss of aly/ALYREF suppresses toxicity in both tau and TDP-43 models of neurodegeneration[J]. GeroScience, 2022, 44(2): 747-761.
doi: 10.1007/s11357-022-00526-2 pmid: 35122183 |
| [40] | Sultana S, Bao WX, Shimizu M, et al. Frequency of three mutations in the fumonisin biosynthetic gene cluster ofFusarium fujikuroi that are predicted to block fumonisin production[J]. World Mycotoxin J, 2021, 14(1): 49-60. |
| [41] | Janevska S, Ferling I, Jojić K, et al. Self-protection against the sphingolipid biosynthesis inhibitor fumonisin B1 is conferred by a FUM cluster-encoded ceramide synthase[J]. mBio, 2020, 11(3): e00455-20. |
| [1] | 王超敏, 何美丹, 王文治, 袁潜华, 张树珍, 沈林波. 甘蔗条点病毒荧光定量PCR检测方法的建立及应用[J]. 生物技术通报, 2024, 40(6): 126-133. |
| [2] | 桑森骅. 基于实时动态成像系统对NK细胞毒性的检测方法[J]. 生物技术通报, 2024, 40(4): 77-84. |
| [3] | 杨伟杰, 杨周林, 朱浩东, 魏煜, 刘君, 刘训. 地衣素合成酶关键模块 LchAD 蛋白的性质和功能研究[J]. 生物技术通报, 2024, 40(3): 322-332. |
| [4] | 杨文莉, 朱梨梨, 陈健, 陈燕欣, 姚涓, 姜大刚. 我国作物病菌标准品的研究进展[J]. 生物技术通报, 2024, 40(2): 31-37. |
| [5] | 刘星雨, 李洁, 朱龙佼, 李相阳, 许文涛. 铜绿假单胞菌适配体的获得及应用[J]. 生物技术通报, 2024, 40(1): 186-193. |
| [6] | 谢宏, 周丽莹, 李舒文, 王梦迪, 艾晔, 晁跃辉. 蒺藜苜蓿MtCIM基因结构和功能分析[J]. 生物技术通报, 2024, 40(1): 262-269. |
| [7] | 张雪萍, 鲁雨晴, 张月倩, 李晓娟. 植物细胞外囊泡及其分析技术的进展[J]. 生物技术通报, 2023, 39(5): 32-43. |
| [8] | 高凯月, 郭雨婷, 杜奕谋, 郑小梅, 马欣荣, 赵伟, 郑平, 孙际宾. 黑曲霉葡萄糖吸收定量检测的方法建立及其在MstC功能研究中的应用[J]. 生物技术通报, 2023, 39(12): 71-80. |
| [9] | 陈晓琳, 刘洋儿, 许文涛, 郭明璋, 刘慧琳. 合成生物学细胞传感技术在食品安全快速检测中的应用[J]. 生物技术通报, 2023, 39(1): 137-149. |
| [10] | 胡海洋, 应婉琴, 何军, 吕芷贤, 谢小平, 邓仲良. 酶促重组等温扩增实时荧光法快速检测肺炎支原体方法的建立及应用[J]. 生物技术通报, 2022, 38(9): 264-270. |
| [11] | 刘娜, 焦京琳, 饶正华. 短链脂肪酸在动物样本中的检测方法研究进展[J]. 生物技术通报, 2022, 38(8): 84-91. |
| [12] | 朱秋雨, 段绪果. L-天冬氨酸-α-脱羧酶的重组表达、定点突变及高通量检测方法的建立[J]. 生物技术通报, 2022, 38(5): 269-278. |
| [13] | 兰欣悦, 刘宁宁, 朱龙佼, 陈旭, 褚华硕, 李相阳, 段诺, 许文涛. 四环素双价适配体非酶免标记传感器[J]. 生物技术通报, 2022, 38(3): 276-284. |
| [14] | 罗雪琮, 安梦楠, 吴元华, 夏子豪. 重组酶聚合酶扩增技术在植物病毒检测中的应用[J]. 生物技术通报, 2022, 38(2): 269-280. |
| [15] | 孔德真, 聂迎彬, 徐红军, 崔凤娟, 穆培源, 田笑明. 三系杂交小麦混播制种对杂交种产量、纯度及F1产量优势的影响[J]. 生物技术通报, 2022, 38(10): 132-139. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||