








Biotechnology Bulletin ›› 2025, Vol. 41 ›› Issue (12): 240-253.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0534
Previous Articles Next Articles
YANG Tao1,2( ), LI Lin1, MO Xiao-lian1, CHEN Xiao-long1, WANG Jian1, HUANG Yuan1, ZHAO Jie-hong1, ZOU Jie1(
), LI Lin1, MO Xiao-lian1, CHEN Xiao-long1, WANG Jian1, HUANG Yuan1, ZHAO Jie-hong1, ZOU Jie1( )
)
Received:2025-05-24
Online:2025-12-26
Published:2026-01-06
Contact:
ZOU Jie
E-mail:1586613641@qq.com;zxgh202@whu.edu.cn
YANG Tao, LI Lin, MO Xiao-lian, CHEN Xiao-long, WANG Jian, HUANG Yuan, ZHAO Jie-hong, ZOU Jie. Functional Study of DoDELLA2 in Dendrobium officinale Kimura et Migo[J]. Biotechnology Bulletin, 2025, 41(12): 240-253.
| 引物名称 Primer name | 上游引物 Forward primer (5′-3′) | 下游引物 Reverse primer (5′-3′) |
|---|---|---|
| DoDELLA2 | ACTAGGGTCTCGCACCATGAAGAGGGAGCATTTGGAGAGTGTTGGAGGAA | ACTAGGGTCTCTCGCCGTGAGCATCTGCGGCCGCGGAAGCGGA |
| 35S | CACGGGGGACTCTTGCCACC | GACACGCTGAACTTGTGG |
| eGFP | — | GACACGCTGAACTTGTGG |
| DoDELLA2-CDS | ATGAAGAGGGAGCATTTGGAG | GTGAGCATCTGCGGCCGCGGA |
| DoDELLA2-RT | CTCCTGTTGTCTTCCCTGATTT | TCGGGTTCCACTATCGATCT |
| AtActin | TCAGATGCCCAGAAGTCTTGTTCC | CCGTACAGATCCTTCCTGATATCC |
| DoDELLA1-RT | CAAGAGAGTGGAAGCGGATTAG | ATCGGCATAGCTGACGAATAAA |
| DoDELLA3-RT | CGGAGTCCTTGCATTACTACTC | CGAGAAATACCTCCGACATCAC |
| DoDELLA4-RT | CGAGGCACTGCACTTCTATT | CCGAGATAGACTTCCGACATAAAC |
| DoActin | TCCCAAGGCAAACAGAGAAA | GGCCACTAGCATATAGGGAAAG |
| AtGA2OX7 | AAACCTTGCTTGGAAGCCCT | TCACGCTTTGGTACACTCC |
| AtGASA1 | TCTCCAACTCGTCCAGGCTGATG | CTACACACGCACTCCCACAATCG |
| AtGALS2 | ACGACGGGATAGGAAGTATGCGG | TCCCCTTCCGCTCTGTGATACGTC |
| AtGA20OX2 | GCAGAAGCTTGCACCAAACA | GTGGAGAATCTGCCGGTGAA |
| AtGA20OX3 | ATCAAGACCAAGTTGGCGGT | TAGAGCCATGAAGGTGTCGC |
| AtGA3OX1 | GATCTCCTCTTCTCCGCTGC | TTTGGAAGGCACCCCAAGTT |
Table 1 Primer sequence
| 引物名称 Primer name | 上游引物 Forward primer (5′-3′) | 下游引物 Reverse primer (5′-3′) |
|---|---|---|
| DoDELLA2 | ACTAGGGTCTCGCACCATGAAGAGGGAGCATTTGGAGAGTGTTGGAGGAA | ACTAGGGTCTCTCGCCGTGAGCATCTGCGGCCGCGGAAGCGGA |
| 35S | CACGGGGGACTCTTGCCACC | GACACGCTGAACTTGTGG |
| eGFP | — | GACACGCTGAACTTGTGG |
| DoDELLA2-CDS | ATGAAGAGGGAGCATTTGGAG | GTGAGCATCTGCGGCCGCGGA |
| DoDELLA2-RT | CTCCTGTTGTCTTCCCTGATTT | TCGGGTTCCACTATCGATCT |
| AtActin | TCAGATGCCCAGAAGTCTTGTTCC | CCGTACAGATCCTTCCTGATATCC |
| DoDELLA1-RT | CAAGAGAGTGGAAGCGGATTAG | ATCGGCATAGCTGACGAATAAA |
| DoDELLA3-RT | CGGAGTCCTTGCATTACTACTC | CGAGAAATACCTCCGACATCAC |
| DoDELLA4-RT | CGAGGCACTGCACTTCTATT | CCGAGATAGACTTCCGACATAAAC |
| DoActin | TCCCAAGGCAAACAGAGAAA | GGCCACTAGCATATAGGGAAAG |
| AtGA2OX7 | AAACCTTGCTTGGAAGCCCT | TCACGCTTTGGTACACTCC |
| AtGASA1 | TCTCCAACTCGTCCAGGCTGATG | CTACACACGCACTCCCACAATCG |
| AtGALS2 | ACGACGGGATAGGAAGTATGCGG | TCCCCTTCCGCTCTGTGATACGTC |
| AtGA20OX2 | GCAGAAGCTTGCACCAAACA | GTGGAGAATCTGCCGGTGAA |
| AtGA20OX3 | ATCAAGACCAAGTTGGCGGT | TAGAGCCATGAAGGTGTCGC |
| AtGA3OX1 | GATCTCCTCTTCTCCGCTGC | TTTGGAAGGCACCCCAAGTT |
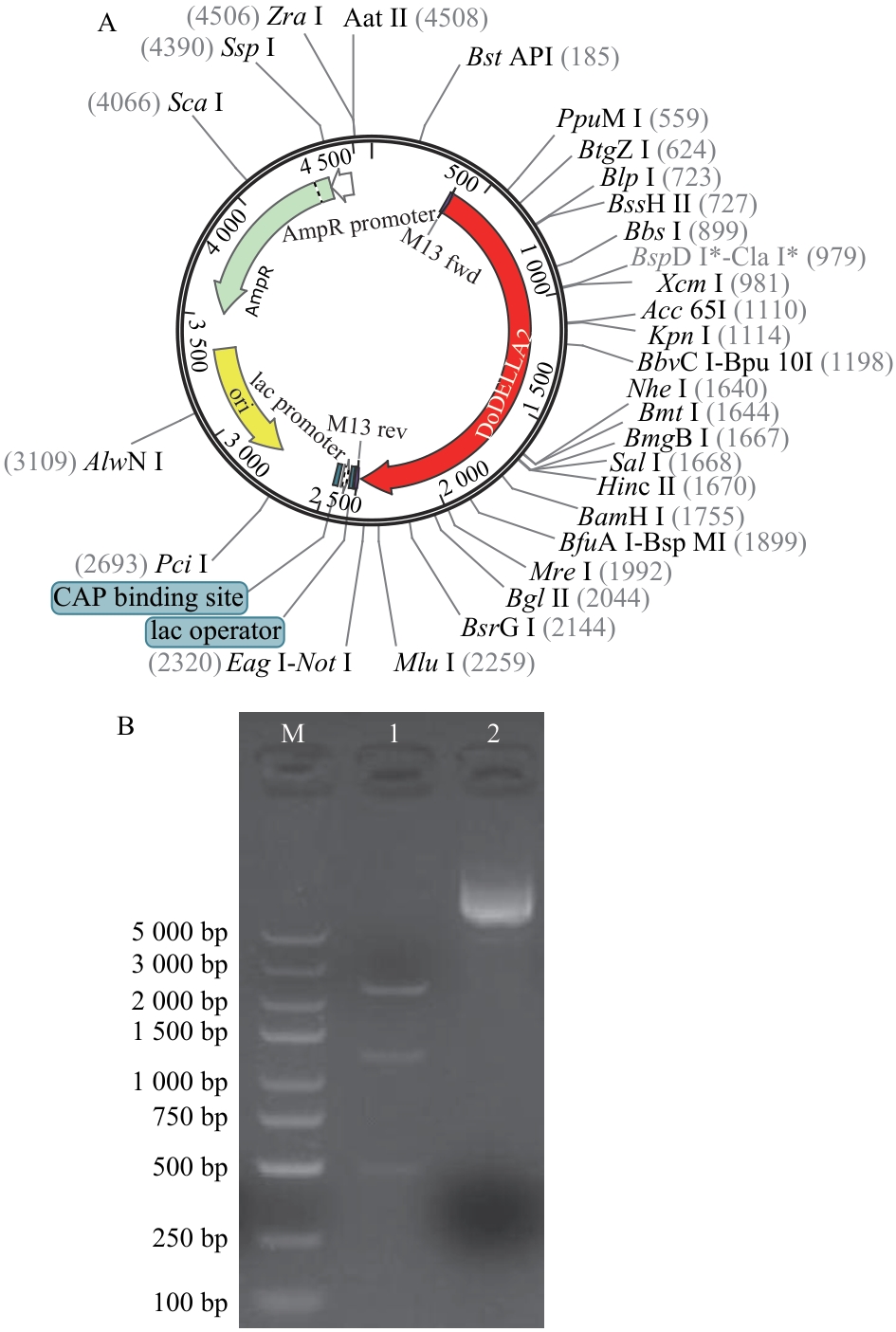
Fig. 3 Enzyme digestion verification of pUC57-DoDELLA2 plasmidA: Plasmid map of pUC57-DoDELLA2. B: Plasmid enzyme digestion verification (1: pUC57-DoDELLA2 plasmid digested with ApaL I; 2: pUC57-DoDELLA2 plasmid; M: DL 5 000 marker)
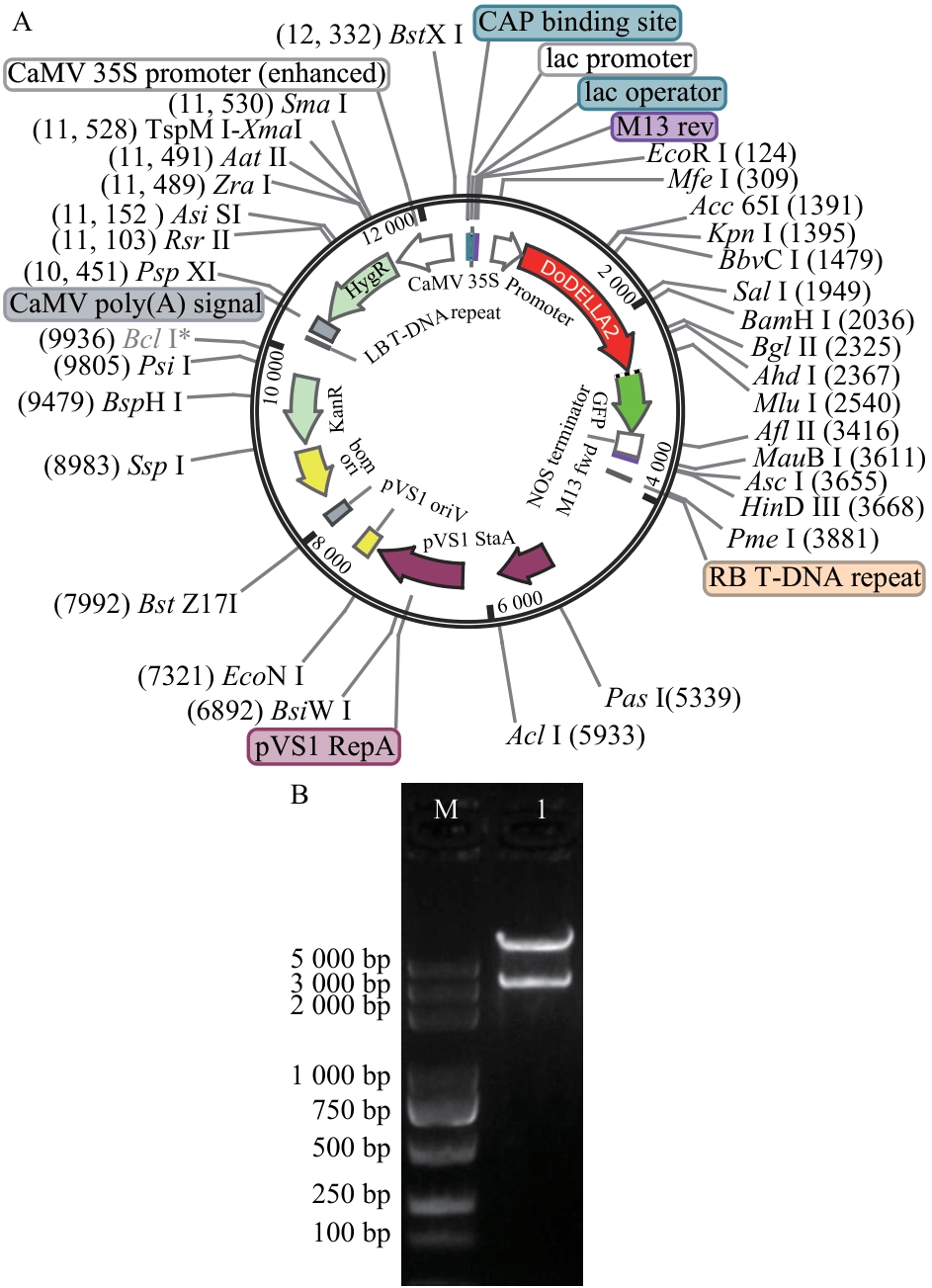
Fig. 4 Enzyme digestion verification of pEGOEP35S-H-DoDELLA2-GFP plasmidA: Plasmid map of pEGOEP35S-H-DoDELLA2-GFP. B: Restriction enzyme digestion verification of plasmid pEGOEP35S-H-DoDELLA2-GFP with EcoR I/Hind III. M: DL 5 000 marker
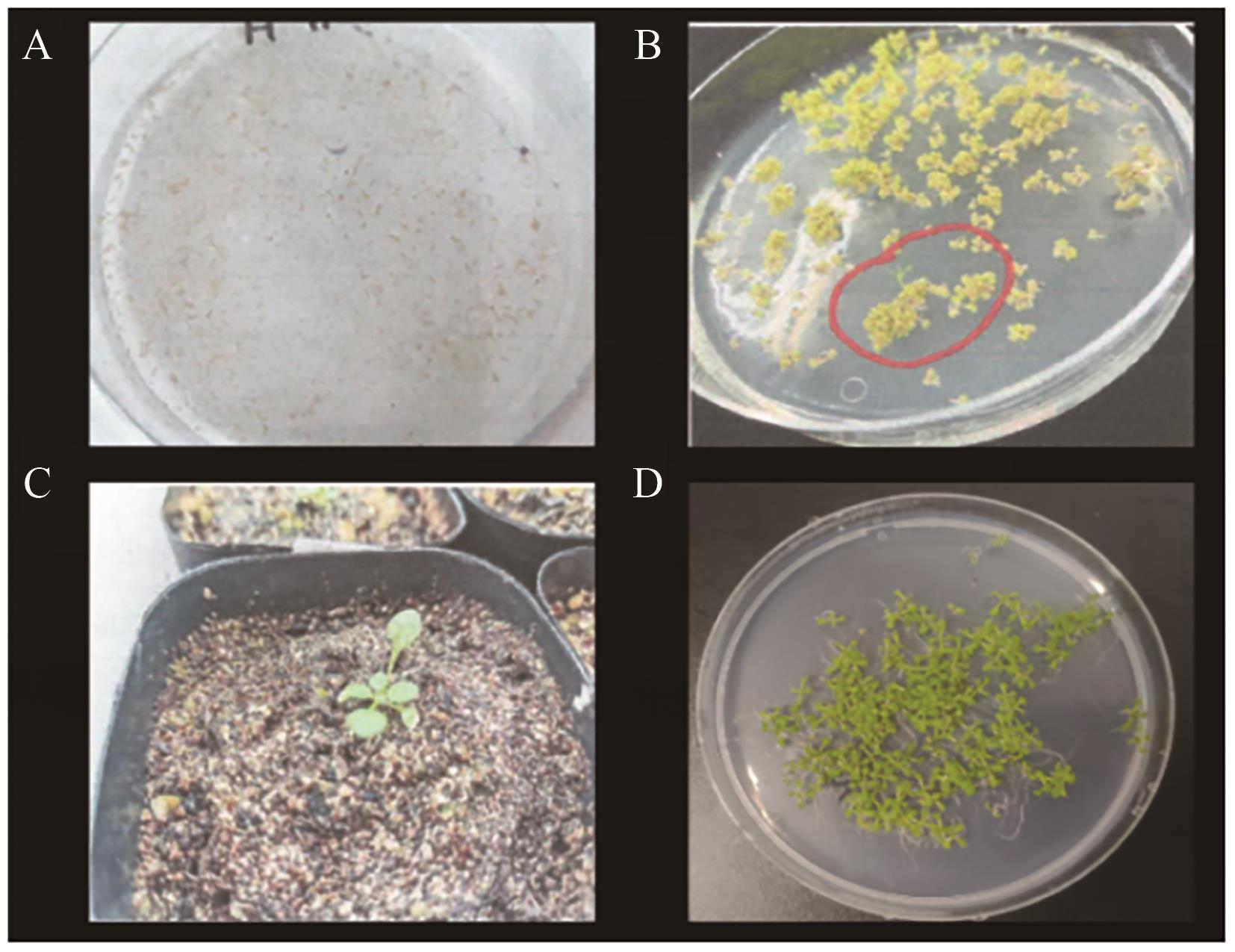

Fig. 5 Screening of positive seedlings for A. thaliana overexpressing DoDELLA2A: Seeds of the T0 generation. B: Screening of positive seedlings. C: Transplantation of positive seedlings. D: Screening of segregation ratio
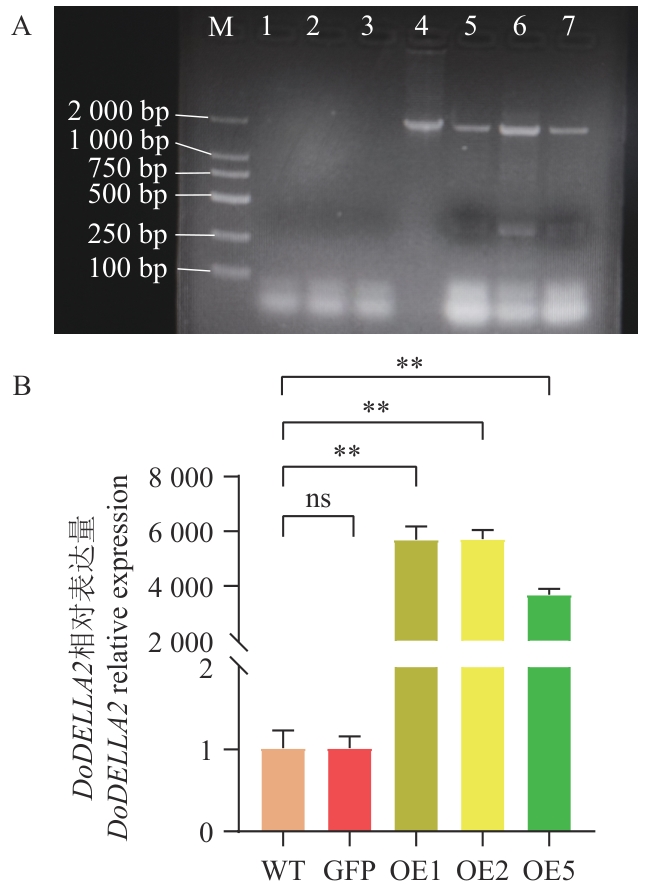
Fig. 6 PCR identification of A. thaliana overexpressing DoDELLA2 and expression analysis of DoDELLA2A: cDNA PCR electrophoresis profile of A. thaliana overexpressing DoDELLA2. (1: Water; 2: WT; 3: A. thaliana with empty GFP vector; 4: positive plasmid; 5‒7: OE1, OE2 and OE5, respectively; M: DL 2 000 marker). B: Relative expression of DoDELLA2 in A. thaliana overexpressing DoDELLA2 (WT: Wild-type A. thaliana; GFP: A. thaliana with empty vector; OE1/2/5: homozygous lines of A. thaliana overexpressing DoDELLA2 respectively). **P<0.01. The same below
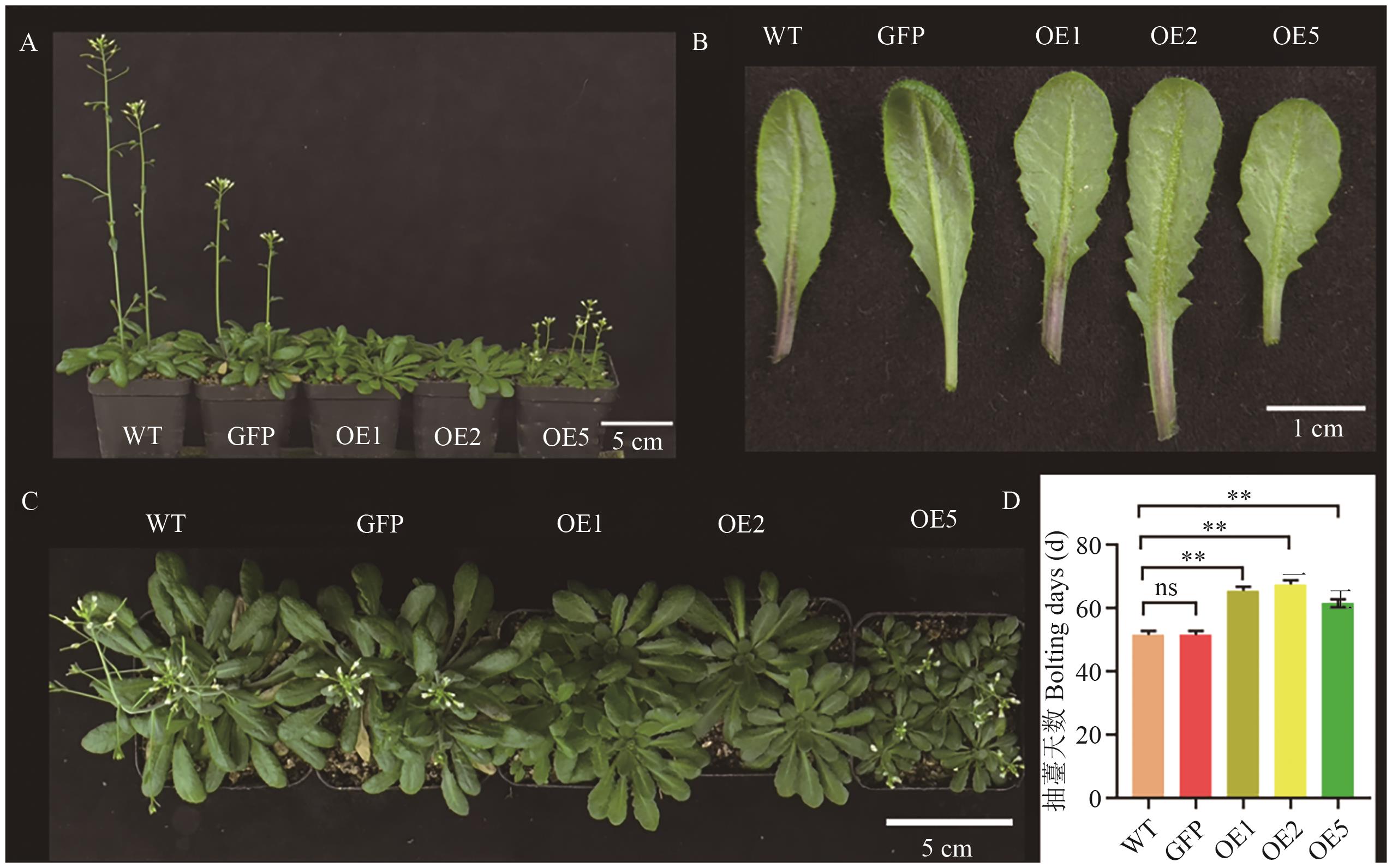

Fig. 7 Bolting and leaf phenotypic observation of A. thaliana overexpressing DoDELLA2A: Front view of 8-week-old A. thaliana. B: Leaves of 8-week-old A. thaliana. C: Top view of 8-week-old A. thaliana. D: Bolting days of A. thaliana. n=12
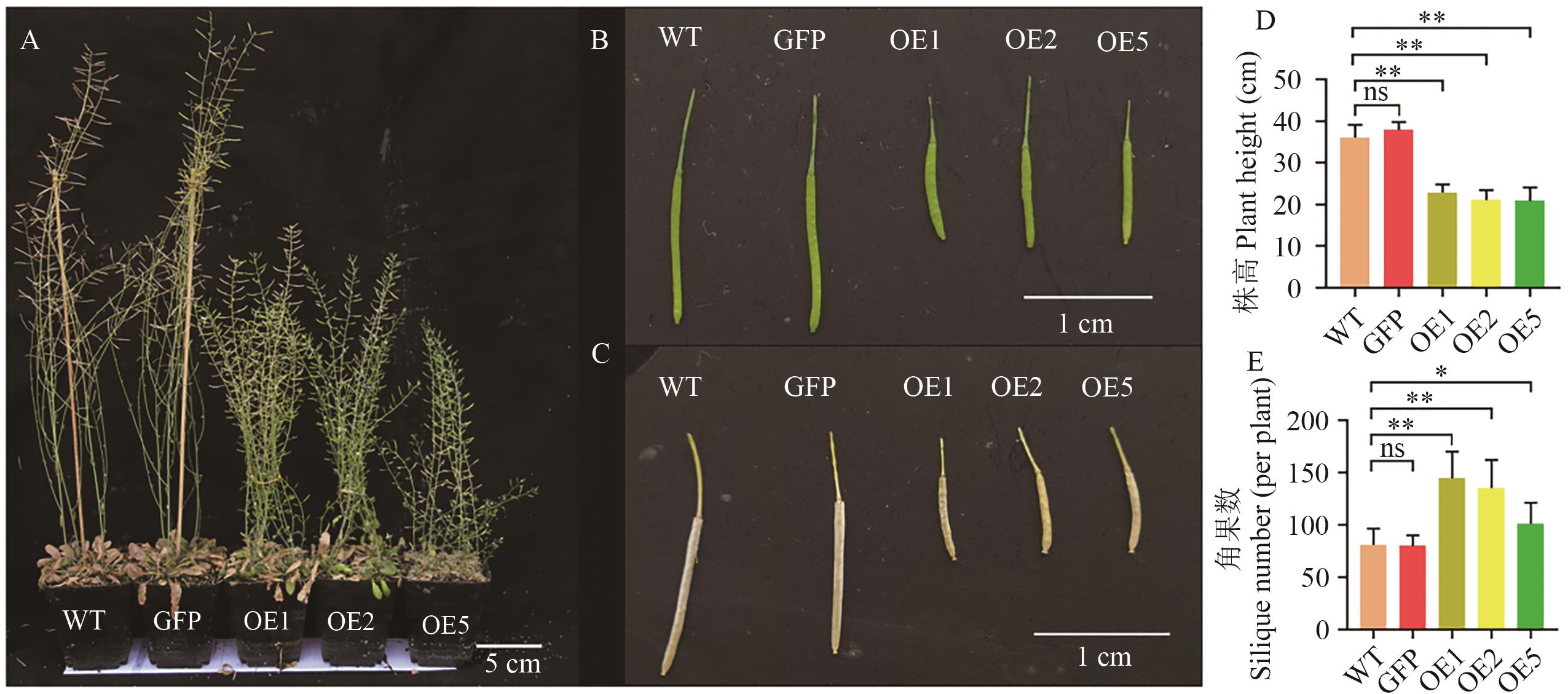

Fig. 8 Plant height and silique phenotypic observation of A. thaliana overexpressing DoDELLA2A: 12-week-old A. thaliana. B: Siliques of 9-week-old A. thaliana. C: Siliques of 12-week-old A. thaliana. D: Plant height of 12-week-old A. thaliana. E: Number of siliques of 12-week-old A. thaliana. n=12. *P<0.05
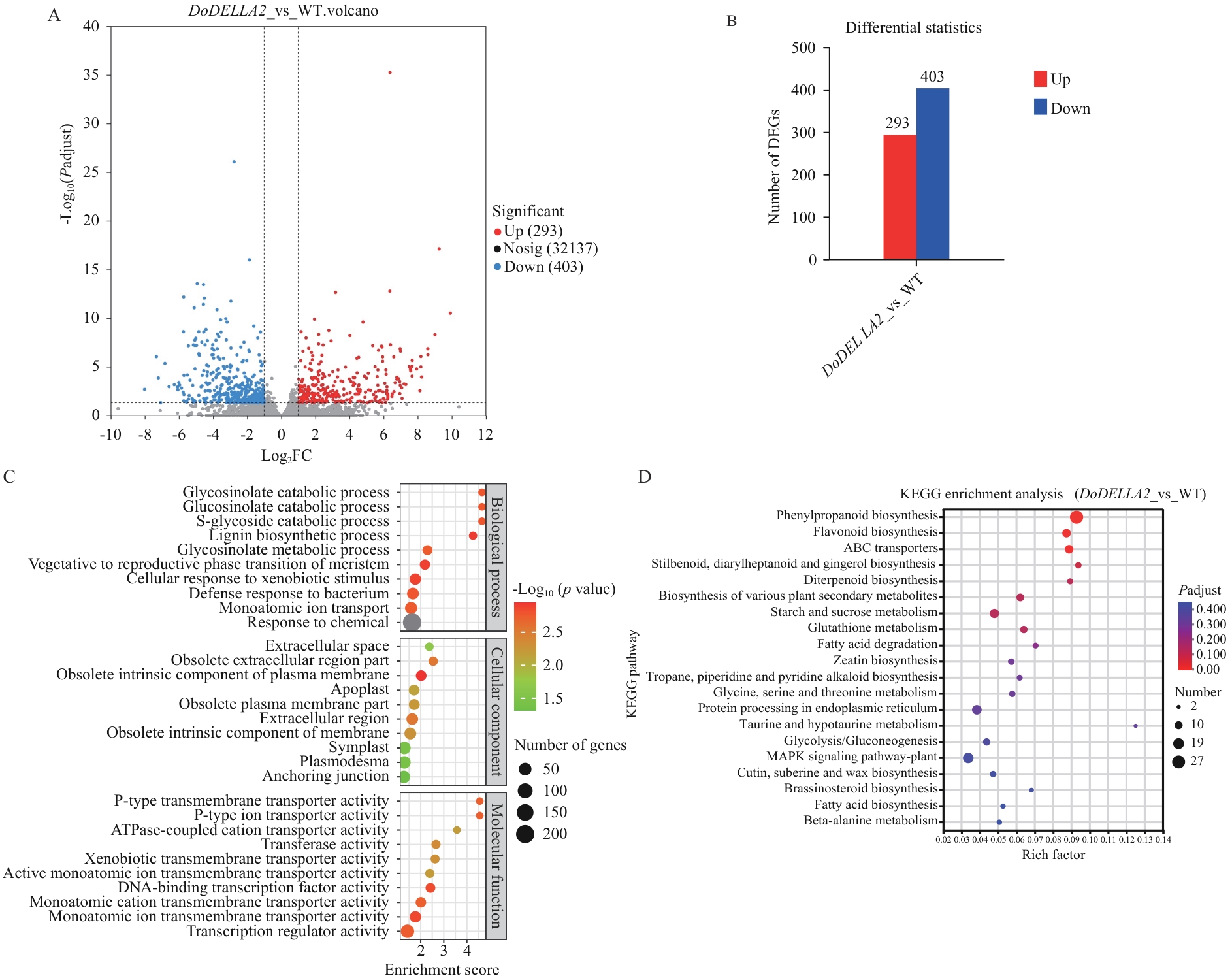
Fig. 9 Differential gene data analysis of heterologous transformation of D. officinaleDoDELLA2 into A. thalianaA: Volcano plot of DoDELLA2_vs_WT. B: Statistical histogram of differentially expressed genes in DoDELLA2_vs_WT. C: GO annotation of differentially expressed genes in DoDELLA2_vs_WT. D: KEGG enrichment analysis of differentially expressed genes in DoDELLA2_vs_WT
基因名 Gene name | 基因描述 Gene description | 调控 Regulate |
|---|---|---|
| AT1G71680 | Transmembrane amino acid transporter family protein | Up |
| STP6 | Sugar transporter 6 | Up |
| PMT2 | Polyol/monosaccharide transporter 2 | Up |
| ACA7 | Cation transporter/E1-E2 ATPase family protein | Up |
| STP9 | Sugar transporter 9 | Up |
| CAT7 | Cationic amino acid transporter 7 | Up |
| AST91 | Sulfate transporter 91 | Up |
| PHT3;2 | Phosphate transporter 3;2 | Up |
| LHT1 | Lysine histidine transporter 1 | Up |
| ABCG1 | ABC-2 type transporter family protein | Up |
| GA20OX3 | Gibberellin 20-oxidase 3 | Up |
| GA20OX2 | Gibberellin 20 oxidase 2 | Up |
| GA3OX1 | Gibberellin 3-oxidase 1 | Up |
| AT3G21970 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT3G22000 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT3G21920 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT3G29040 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT3G21945 | Cysteine-rich repeat secretory protein | Up |
| AT3G22050 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT4G20680 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT3G21990 | Cysteine-rich repeat secretory-like protein (DUF26) | Up |
| LCR5 | Low-molecular-weight cysteine-rich 5 | Up |
| LCR1 | Low-molecular-weight cysteine-rich 1 | Up |
| LCR33 | Low-molecular-weight cysteine-rich 33 | Up |
| LCR4 | Low-molecular-weight cysteine-rich 4 | Up |
| LCR32 | Low-molecular-weight cysteine-rich 32 | Up |
| LCR25 | Low-molecular-weight cysteine-rich 25 | Up |
| LCR10 | Low-molecular-weight cysteine-rich 10 | Up |
| AT2G33690 | Late embryosis abundant protein, group 6 | Up |
| AT4G13230 | Late embryosis abundant protein (LEA) family protein | Up |
| YLS9 | Late embryosis abundant (LEA) hydroxyproline-rich glycoprotein family | Up |
| AT1G15415 | Late embryosis abundant-like protein | Up |
| AT5G53820 | Late embryosis abundant protein (LEA) family protein | Up |
| AT2G38390 | Peroxidase superfamily protein | Down |
| AT5G64110 | Peroxidase superfamily protein | Down |
| AT2G38380 | Peroxidase superfamily protein | Down |
| AT3G32980 | Peroxidase superfamily protein | Down |
| AT2G35380 | Peroxidase superfamily protein | Down |
| AT1G68850 | Peroxidase superfamily protein | Down |
| AT2G18150 | Peroxidase superfamily protein | Down |
| AT3G21770 | Peroxidase superfamily protein | Down |
| AT4G36430 | Peroxidase superfamily protein | Down |
| AT5G58390 | Peroxidase superfamily protein | Down |
| AT5G06730 | Peroxidase superfamily protein | Down |
| PRXR1 | Peroxidase superfamily protein | Down |
| CYP702A6 | Cytochrome P450, family 702, subfamily A, polypeptide 6 | Down |
| CYP81F4 | Cytochrome P450, family 81, subfamily F, polypeptide 4 | Down |
| CYP708A2 | Cytochrome P450, family 708, subfamily A, polypeptide 2 | Down |
| CYP71A19 | Cytochrome P450, family 71, subfamily A, polypeptide 19 | Down |
| CYP86A1 | Cytochrome P450, family 86, subfamily A, polypeptide 1 | Down |
| CYP86B1 | Cytochrome P450, family 86, subfamily B, polypeptide 1 | Down |
| CYP72A14 | Cytochrome P450, family 72, subfamily A, polypeptide 14 | Down |
| AT1G50060 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT4G33710 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT4G33720 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT2G19970 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT1G52100 | Mannose-binding lectin superfamily protein | Down |
| AT5G38550 | Mannose-binding lectin superfamily protein | Down |
| JAL33 | Mannose-binding lectin superfamily protein | Down |
| AT1G52120 | Mannose-binding lectin superfamily protein | Down |
| AT1G52000 | Mannose-binding lectin superfamily protein | Down |
| AT5G49850 | Mannose-binding lectin superfamily protein | Down |
Table 2 Statistical table of key genes with significant differential expression and enrichment in up and down genes
基因名 Gene name | 基因描述 Gene description | 调控 Regulate |
|---|---|---|
| AT1G71680 | Transmembrane amino acid transporter family protein | Up |
| STP6 | Sugar transporter 6 | Up |
| PMT2 | Polyol/monosaccharide transporter 2 | Up |
| ACA7 | Cation transporter/E1-E2 ATPase family protein | Up |
| STP9 | Sugar transporter 9 | Up |
| CAT7 | Cationic amino acid transporter 7 | Up |
| AST91 | Sulfate transporter 91 | Up |
| PHT3;2 | Phosphate transporter 3;2 | Up |
| LHT1 | Lysine histidine transporter 1 | Up |
| ABCG1 | ABC-2 type transporter family protein | Up |
| GA20OX3 | Gibberellin 20-oxidase 3 | Up |
| GA20OX2 | Gibberellin 20 oxidase 2 | Up |
| GA3OX1 | Gibberellin 3-oxidase 1 | Up |
| AT3G21970 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT3G22000 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT3G21920 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT3G29040 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT4G20530 | Cysteine-rich repeat secretory-like protein | Up |
| AT3G21945 | Cysteine-rich repeat secretory protein | Up |
| AT3G22050 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT4G20680 | Cysteine-rich repeat secretory protein, putative (DUF26) | Up |
| AT3G21990 | Cysteine-rich repeat secretory-like protein (DUF26) | Up |
| LCR5 | Low-molecular-weight cysteine-rich 5 | Up |
| LCR1 | Low-molecular-weight cysteine-rich 1 | Up |
| LCR33 | Low-molecular-weight cysteine-rich 33 | Up |
| LCR4 | Low-molecular-weight cysteine-rich 4 | Up |
| LCR32 | Low-molecular-weight cysteine-rich 32 | Up |
| LCR25 | Low-molecular-weight cysteine-rich 25 | Up |
| LCR10 | Low-molecular-weight cysteine-rich 10 | Up |
| AT2G33690 | Late embryosis abundant protein, group 6 | Up |
| AT4G13230 | Late embryosis abundant protein (LEA) family protein | Up |
| YLS9 | Late embryosis abundant (LEA) hydroxyproline-rich glycoprotein family | Up |
| AT1G15415 | Late embryosis abundant-like protein | Up |
| AT5G53820 | Late embryosis abundant protein (LEA) family protein | Up |
| AT2G38390 | Peroxidase superfamily protein | Down |
| AT5G64110 | Peroxidase superfamily protein | Down |
| AT2G38380 | Peroxidase superfamily protein | Down |
| AT3G32980 | Peroxidase superfamily protein | Down |
| AT2G35380 | Peroxidase superfamily protein | Down |
| AT1G68850 | Peroxidase superfamily protein | Down |
| AT2G18150 | Peroxidase superfamily protein | Down |
| AT3G21770 | Peroxidase superfamily protein | Down |
| AT4G36430 | Peroxidase superfamily protein | Down |
| AT5G58390 | Peroxidase superfamily protein | Down |
| AT5G06730 | Peroxidase superfamily protein | Down |
| PRXR1 | Peroxidase superfamily protein | Down |
| CYP702A6 | Cytochrome P450, family 702, subfamily A, polypeptide 6 | Down |
| CYP81F4 | Cytochrome P450, family 81, subfamily F, polypeptide 4 | Down |
| CYP708A2 | Cytochrome P450, family 708, subfamily A, polypeptide 2 | Down |
| CYP71A19 | Cytochrome P450, family 71, subfamily A, polypeptide 19 | Down |
| CYP86A1 | Cytochrome P450, family 86, subfamily A, polypeptide 1 | Down |
| CYP86B1 | Cytochrome P450, family 86, subfamily B, polypeptide 1 | Down |
| CYP72A14 | Cytochrome P450, family 72, subfamily A, polypeptide 14 | Down |
| AT1G50060 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT4G33710 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT4G33720 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT2G19970 | CAP (Cysteine-rich secretory proteins, Antigen 5, and Pathosis-related 1 protein) superfamily protein | Down |
| AT1G52100 | Mannose-binding lectin superfamily protein | Down |
| AT5G38550 | Mannose-binding lectin superfamily protein | Down |
| JAL33 | Mannose-binding lectin superfamily protein | Down |
| AT1G52120 | Mannose-binding lectin superfamily protein | Down |
| AT1G52000 | Mannose-binding lectin superfamily protein | Down |
| AT5G49850 | Mannose-binding lectin superfamily protein | Down |
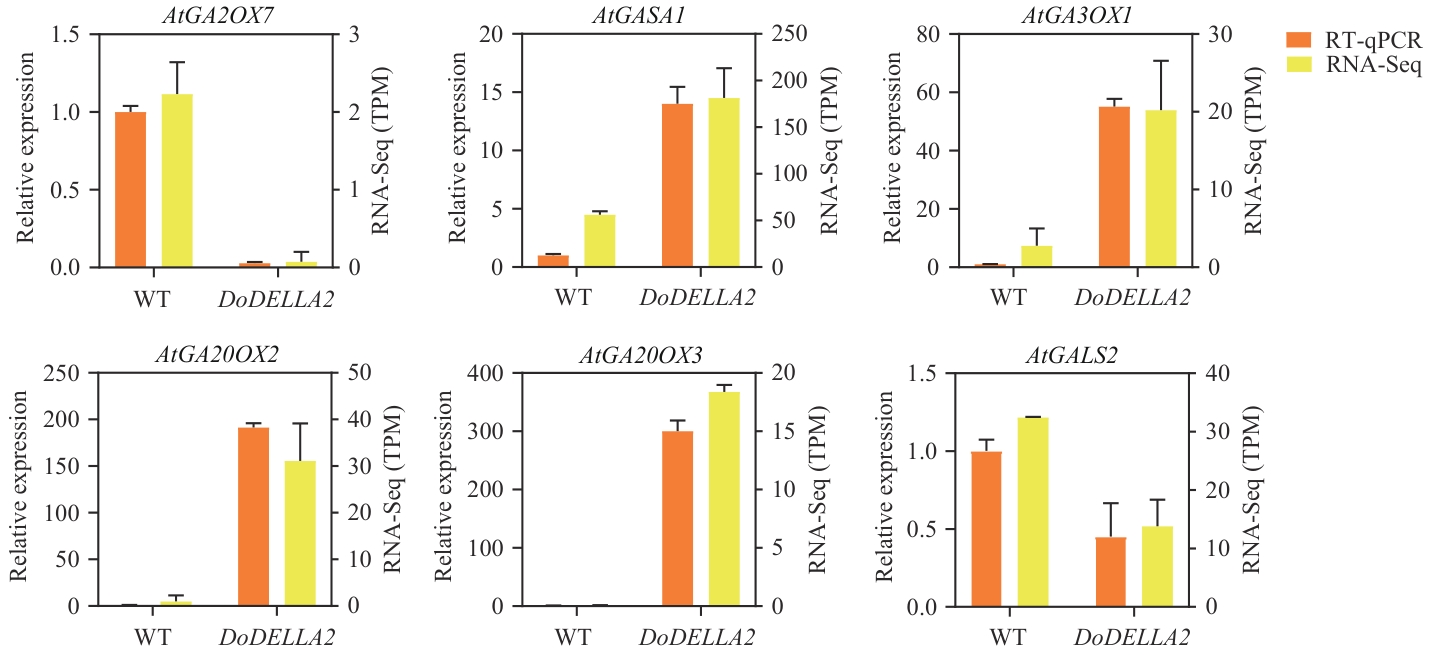
Fig. 10 Validation of RT-qPCR expression levels of differentially expressed genes related to the gibberellin signaling pathwayWT: Wild-type A. thaliana. DoDELLA2: A. thaliana overexpressing DoDELLA2
| [1] | Reinecke DM, Wickramarathna AD, Ozga JA, et al. Gibberellin 3-oxidase gene expression patterns influence gibberellin biosynthesis, growth, and development in pea [J]. Plant Physiol, 2013, 163(2): 929-945. |
| [2] | Eriksson S, Böhlenius H, Moritz T, et al. GA4 is the active gibberellin in the regulation of LEAFY transcription and Arabidopsis floral initiation [J]. Plant Cell, 2006, 18(9): 2172-2181. |
| [3] | Webster AD. Vigour mechanisms in dwarfing rootstocks for temperate fruit trees [J]. Acta Hortic, 2004(658): 29-41. |
| [4] | Peng JR, Harberd NP. The role of GA-mediated signalling in the control of seed germination [J]. Curr Opin Plant Biol, 2002, 5(5): 376-381. |
| [5] | Zhang N, Xie YD, Guo HJ, et al. Gibberellins regulate the stem elongation rate without affecting the mature plant height of a quick development mutant of winter wheat (Triticum aestivum L.) [J]. Plant Physiol Biochem, 2016, 107: 228-236. |
| [6] | Wang GL, Que F, Xu ZS, et al. Exogenous gibberellin altered morphology, anatomic and transcriptional regulatory networks of hormones in carrot root and shoot [J]. BMC Plant Biol, 2015, 15: 290. |
| [7] | Appleford NE, Lenton JR. Gibberellins and leaf expansion in near-isogenic wheat lines containing Rht1 and Rht3 dwarfing alleles [J]. Planta, 1991, 183(2): 229-236. |
| [8] | Giacomelli L, Rota-Stabelli O, Masuero D, et al. Gibberellin metabolism in Vitis vinifera L. during bloom and fruit-set: functional characterization and evolution of grapevine gibberellin oxidases [J]. J Exp Bot, 2013, 64(14): 4403-4419. |
| [9] | Chen S, Wang XJ, Zhang LY, et al. Identification and characterization of tomato gibberellin 2-oxidases (GA2oxs) and effects of fruit-specific SlGA2ox1 overexpression on fruit and seed growth and development [J]. Hortic Res, 2016, 3: 16059. |
| [10] | Song PL, Li G, Xu JF, et al. Genome-wide analysis of genes involved in the GA signal transduction pathway in ‘Duli’ pear (Pyrus betulifolia Bunge) [J]. Int J Mol Sci, 2022, 23(12): 6570. |
| [11] | Achard P, Gusti A, Cheminant S, et al. Gibberellin signaling controls cell proliferation rate in Arabidopsis [J]. Curr Biol, 2009, 19(14): 1188-1193. |
| [12] | Wang Y, He SE, Wei YZ, et al. Molecular and functional characterization of two DELLA protein-coding genes in Litchi [J]. Gene, 2020, 738: 144455. |
| [13] | 王倩. 百合LiDELLA基因的克隆及初步功能分析 [D]. 长春: 吉林农业大学, 2019. |
| Wang Q. Cloning and preliminary functional analysis of LiDELLA gene from Lilium [D]. Changchun: Jilin Agricultural University, 2019. | |
| [14] | 李露露. 谷子DELLA基因的鉴定、克隆与功能分析 [D]. 太谷: 山西农业大学, 2022. |
| Li LL. Identification, cloning and functional analysis of DELLA gene in Setaria italica [D]. Taigu: Shanxi Agricultural University, 2022. | |
| [15] | 李翠. 花生DELLA家族基因的表达及转基因研究 [D]. 广州: 仲恺农业工程学院, 2015. |
| Li C. Expression and transgenic study of DELLA family genes in peanut [D]. Guangzhou: Zhongkai University of Agriculture and Engineering, 2015. | |
| [16] | Liu Q, Wu K, Harberd NP, et al. Green revolution DELLAs: From translational reinitiation to future sustainable agriculture [J]. Mol Plant, 2021, 14(4): 547-549. |
| [17] | 倪凯, 何鹏飞, 梁志庆, 等. 铁皮石斛化学成分、药理作用及毒理学评价研究进展 [J]. 云南中医中药杂志, 2023, 44(10): 86-93. |
| Ni K, He PF, Liang ZQ, et al. Research progress on chemical constituents, pharmacological effects and toxicological evaluation of Dendrobium candidum [J]. Yunnan J Tradit Chin Med Mater Med, 2023, 44(10): 86-93. | |
| [18] | 马庆, 付强, 邹西西, 等. 铁皮石斛DELLA家族基因克隆与生物信息学分析 [J]. 贵州中医药大学学报, 2022, 44(6): 13-19. |
| Ma Q, Fu Q, Zou XX, et al. Cloning and bioinformatics analysis of DELLA family gene of Dendrobium candidum [J]. J Guizhou Univ Tradit Chin Med, 2022, 44(6): 13-19. | |
| [19] | 李琳, 付强, 杨涛, 等. 铁皮石斛种子cDNA酵母文库的构建及DELLA互作蛋白筛选 [J]. 植物科学学报, 2025, 43(2): 221-229. |
| Li L, Fu Q, Yang T, et al. cDNA yeast library construction of Dendrobium officinale Kimura et Migo seeds and screening and analysis of DELLA interacting proteins [J]. Plant Sci J, 2025, 43(2): 221-229. | |
| [20] | 王健, 杨莎, 孙庆文, 等. 金钗石斛bHLH转录因子家族全基因组鉴定及表达分析 [J]. 生物技术通报, 2024, 40(6): 203-218. |
| Wang J, Yang S, Sun QW, et al. Genome-wide identification and expression analysis of bHLH transcription factor family in Dendrobium nobile [J]. Biotechnol Bull, 2024, 40(6): 203-218. | |
| [21] | 黄龙成. 拟南芥PTED1基因影响花粉壁模式形成的机理研究 [D]. 上海: 上海师范大学, 2025. |
| Huang LC. Study on the mechanism of PTED1 gene influencing pollen wall pattern formation in Arabidopsis thaliana [D]. Shanghai: Shanghai Normal University, 2025. | |
| [22] | 陈紫竹, 庆军, 格根塔娜, 等. 基于转录组测序的枣胚败育关键基因挖掘 [J]. 种子, 2025, 44(2): 95-104. |
| Chen ZZ, Qing J, Ge G, et al. Extraction of key genes of Ziziphus jujuba embryo abortion based on transcriptome sequencing [J]. Seed, 2025, 44(2): 95-104. | |
| [23] | 陈凯, 王灏, 陈燚婷, 等. 铁皮石斛WOX家族基因在生长发育中的功能分析 [J]. 遗传, 2023, 45(8): 700-714. |
| Chen K, Wang H, Chen YT, et al. Functional analysis of WOX family genes in Dendrobium catenatum during growth and development [J]. Hereditas: Beijing, 2023, 45(8): 700-714. | |
| [24] | Wilson RN, Somerville CR. Phenotypic suppression of the gibberellin-insensitive mutant (Gai) of Arabidopsis [J]. Plant Physiol, 1995, 108(2): 495-502. |
| [25] | Silverstone AL, Ciampaglio CN, Sun T. The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway [J]. Plant Cell, 1998, 10(2): 155-169. |
| [26] | 赵静一, 吴小旭, 胡云捷, 等. 洋葱AcGAI的克隆及其在开花途径的功能分析 [J]. 园艺学报, 2024, 51(8): 1792-1802. |
| Zhao JY, Wu XX, Hu YJ, et al. Molecular cloning and functional analysis of AcGAI in onion flowering regulation [J]. Acta Hortic Sin, 2024, 51(8): 1792-1802. | |
| [27] | 贾玉凤. 小麦过氧化物酶家族基因功能分析及小麦矮腥黑粉菌效应蛋白研究 [D]. 乌鲁木齐: 新疆农业大学, 2024. |
| Jia YF. Functional analysis of the wheat peroxidase gene family and research on tilletia controversa Kühn effector proteins [D]. Urumqi: Xinjiang Agricultural University, 2024. | |
| [28] | 钟春梅, 王小菁. 富含半胱氨酸的GASA小分子蛋白研究进展 [J]. 植物学报, 2016, 51(1): 1-8. |
| Zhong CM, Wang XJ. Progress in cysteine-rich gibberellic acid-stimulated Arabidopsis protein [J]. Chin Bull Bot, 2016, 51(1): 1-8. | |
| [29] | Khalifa MAS, Zhang Q, Du YY, et al. Functional characterisation of GmGASA1-like gene in Glycine max (L.) Merr. overexpression promotes growth, development and stress responses [J]. Life, 2024, 14(11): 1436. |
| [30] | 宋建播. 拟南芥局部及系统获得抗性的脂质代谢组学研究及GASA1基因的抗病功能验证 [D]. 西安: 西北大学, 2022. |
| Song JB. Lipid metabonomics study on local and systemic acquired resistance in Arabidopsis thaliana and verification of disease resistance function of GASA1 gene [D]. Xi’an: Northwest University, 2022. | |
| [31] | Wang L, Ma CR, Wang SH, et al. Ethylene and jasmonate signaling converge on gibberellin catabolism during thigmomorphogenesis in Arabidopsis [J]. Plant Physiol, 2024, 194(2): 758-773. |
| [32] | Gao XM, Lou SL, Han Y, et al. Allelic variations in GA20ox3 regulate fruit length and seed germination timing for high-altitude adaptation in Arabidopsis thaliana [J]. Nat Commun, 2025, 16(1): 5053. |
| [33] | Paciorek T, Chiapelli BJ, Wang JY, et al. Targeted suppression of gibberellin biosynthetic genes ZmGA20ox3 and ZmGA20ox5 produces a short stature maize ideotype [J]. Plant Biotechnol J, 2022, 20(6): 1140-1153. |
| [34] | Cheng JY, Hill C, Han Y, et al. New semi-dwarfing alleles with increased coleoptile length by gene editing of gibberellin 3-oxidase 1 using CRISPR-Cas9 in barley (Hordeum vulgare L.) [J]. Plant Biotechnol J, 2023, 21(4): 806-818. |
| [35] | Chakraborty P, Biswas A, Dey S, et al. Cytochrome P450 gene families: role in plant secondary metabolites production and plant defense [J]. J Xenobiot, 2023, 13(3): 402-423. |
| [36] | Tan HJ, Man C, Xie Y, et al. A crucial role of GA-regulated flavonol biosynthesis in root growth of Arabidopsis [J]. Mol Plant, 2019, 12(4): 521-537. |
| [37] | Xue HD, Gao X, He P, et al. Origin, evolution, and molecular function of DELLA proteins in plants [J]. Crop J, 2022, 10(2): 287-299. |
| [38] | Achard P, Cheng H, De Grauwe L, et al. Integration of plant responses to environmentally activated phytohormonal signals [J]. Science, 2006, 311(5757): 91-94. |
| [1] | WANG Fang, SHAO Hui-ru, LYU Lin-long, ZHAO Dian, HU Zhen, LYU Jian-zhen, JIANG Liang. Establishment of TurboID Proximity Labeling Technology in Plants and Bacteria [J]. Biotechnology Bulletin, 2025, 41(9): 44-53. |
| [2] | LIU Jian-guo, LIU Ge-er, GUO Ying-xin, WANG Bin, WANG Yu-kun, LU Jin-feng, HUANG Wen-ting, ZHU Yun-na. Integrate Transcriptomic and Metabolomic Analysis of Fruits Quality Differences between ‘Guiyou No. 1’ and ‘Shatianyou’ Pomelo (Citrus maxima) [J]. Biotechnology Bulletin, 2025, 41(9): 168-181. |
| [3] | LIU Ze-zhou, DUAN Nai-bin, YUE Li-xin, WANG Qing-hua, YAO Xing-hao, GAO Li-min, KONG Su-ping. Analysis of Wax Components and Screening of Wax-deficient Gene Ggl-1 in Garlic (Allium sativum L.) [J]. Biotechnology Bulletin, 2025, 41(9): 219-231. |
| [4] | YAN Meng-yang, LIANG Xiao-yang, DAI Jun-ang, ZHANG Yan, GUAN Tuan, ZHANG Hui, LIU Liang-bo, SUN Zhi-hua. Screening of Amoxicillin-degrading Bacteria and Study on Its Degradation Mechanisms [J]. Biotechnology Bulletin, 2025, 41(9): 314-325. |
| [5] | LIU Jia-li, SONG Jing-rong, ZHAO Wen-yu, ZHANG Xin-yuan, ZHAO Zi-yang, CAO Yi-bo, ZHANG Ling-yun. Identification of the R2R3-MYB Gene and Expression Analysis of Flavonoid Regulatory Genes in Blueberry [J]. Biotechnology Bulletin, 2025, 41(9): 124-138. |
| [6] | LI Yu-zhen, LI Meng-dan, ZHANG Wei, PENG Ting. Functional Study of RmEXPB2 Genein Rosa multiflora Based on the Identification of the Expansin Gene Family in Rosa sp. [J]. Biotechnology Bulletin, 2025, 41(9): 182-194. |
| [7] | LI Ya-qiong, GESANG La-mao, CHEN Qi-di, YANG Yu-huan, HE Hua-zhuan, ZHAO Yao-fei. Heterologous Overexpression of Sorghum SbSnRK2.1 Enhances the Resistance to Salt Stress in Arabidopsis [J]. Biotechnology Bulletin, 2025, 41(8): 115-123. |
| [8] | ZENG Dan, HUANG Yuan, WANG Jian, ZHANG Yan, LIU Qing-xia, GU Rong-hui, SUN Qing-wen, CHEN Hong-yu. Genome-wide Identification and Expression Analysis of bZIP Transcription Factor Family in Dendrobium officinale [J]. Biotechnology Bulletin, 2025, 41(8): 197-210. |
| [9] | BAI Yu-guo, LI Wan-di, LIANG Jian-ping, SHI Zhi-yong, LU Geng-long, LIU Hong-jun, NIU Jing-ping. Growth-promoting Mechanism of Trichoderma harzianum T9131 on Astragalus membranaceus Seedlings [J]. Biotechnology Bulletin, 2025, 41(8): 175-185. |
| [10] | WANG Yue-chen, HAN Xin-qi, WEI Wen-min, CUI Zhao-lan, LUO Yang-mei, CHEN Peng-ru, WANG Hai-gang, LIU long-long, ZHANG Li, WANG Lun. Biological Basis Study for Grain Shattering in Proso Millet and Identification of Genes Regulating Grain Shattering [J]. Biotechnology Bulletin, 2025, 41(7): 164-171. |
| [11] | HUANG Xu-sheng, ZHOU Ya-li, CHAI Xu-dong, WEN Jing, WANG Ji-ping, JIA Xiao-yun, LI Run-zhi. Cloning of Plastidial PfLPAT1B Gene from Perilla frutescens and Its Functional Analysis in Oil Biosynthesis [J]. Biotechnology Bulletin, 2025, 41(7): 226-236. |
| [12] | ZHANG Yue, BI Yu, MU Xue-nan, ZHENG Zi-wei, WANG Zhi-gang, XU Wei-hui. Biocontrol Characteristics of Strain JB7 against Fusarium graminearum [J]. Biotechnology Bulletin, 2025, 41(7): 261-271. |
| [13] | LI Cheng-hua, DOU Fei-fei, REN Yu-zhao, LIU Cai-xia, LIU Feng-lou, WANG Zhang-jun, LI Qing-feng. Effect of Exogenous Salicylic Acid on Wheat Infested with Blumeria graminis f. sp. tritici and Its Transcriptome Analysis [J]. Biotechnology Bulletin, 2025, 41(7): 272-280. |
| [14] | GUO Xiu-juan, FENG Yu, WU Rui-xiang, WANG Li-qin, YANG Jian-chun. Transcriptome Analysis of the Effect of Ca 2+ Treatment on the Seed Germination of Flax [J]. Biotechnology Bulletin, 2025, 41(7): 139-149. |
| [15] | HU Ruo-qun, ZENG Jing-jing, LIANG Wan-feng, CAO Jia-yu, HUANG Xiao-wei, LIANG Xiao-ying, QIU Ming-yue, CHEN Ying. Integrated Transcriptome and Metabolome Analysis to Explore the Carotenoid Synthesis and Metabolism Mechanism in Anoectochilus roxburghii under Different Shading Conditions [J]. Biotechnology Bulletin, 2025, 41(5): 231-243. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||