








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 345-356.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0879
YAN Xin( ), WU Xiao, HE Si-yu, DUAN Yu-huan, QIU Wu-xia, YUAN Xiao-qin, MAO Xin-fang, LIU Zhong-yuan(
), WU Xiao, HE Si-yu, DUAN Yu-huan, QIU Wu-xia, YUAN Xiao-qin, MAO Xin-fang, LIU Zhong-yuan( )
)
Received:2025-08-12
Online:2026-04-26
Published:2026-04-30
Contact:
LIU Zhong-yuan
E-mail:1992334217@qq.com;lzy1168@163.com
YAN Xin, WU Xiao, HE Si-yu, DUAN Yu-huan, QIU Wu-xia, YUAN Xiao-qin, MAO Xin-fang, LIU Zhong-yuan. Inositol Biosynthesis and Fermentation Optimization of an Inositol Biosynthesis Pathway Using Citrus sinensis Inositol Phosphate Synthase Gene Csino3[J]. Biotechnology Bulletin, 2026, 42(4): 345-356.
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) |
|---|---|
| pgi-outF | CCACTGTGCCTGAACATCGCGTTCGC |
| pgi-outR | GCAGGTCGCTTCACGACTCTTCTCCAGAGG |
| pgi-inF | GCTGCACGTAGCGCTGCGTAACCG |
| pgi-inR | GCACTTCCGCGATGTGAGTCCCATCGAC |
| gldA-outF | CCCCACCATCCGCCAGTTAAACAGCA |
| gldA-outR | CTGGACACCGCTAACGTCGCAGAAGTC |
| gldA-inF | GGAGTCACTACATGCTGTTCGGCAG |
| gldA-inR | CTACACCGATGAGGGTGAGTTTGACC |
Table 1 Primer sequences for knockout of genes pgi and gldA
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) |
|---|---|
| pgi-outF | CCACTGTGCCTGAACATCGCGTTCGC |
| pgi-outR | GCAGGTCGCTTCACGACTCTTCTCCAGAGG |
| pgi-inF | GCTGCACGTAGCGCTGCGTAACCG |
| pgi-inR | GCACTTCCGCGATGTGAGTCCCATCGAC |
| gldA-outF | CCCCACCATCCGCCAGTTAAACAGCA |
| gldA-outR | CTGGACACCGCTAACGTCGCAGAAGTC |
| gldA-inF | GGAGTCACTACATGCTGTTCGGCAG |
| gldA-inR | CTACACCGATGAGGGTGAGTTTGACC |
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 限制性酶切位点 Restriction enzyme site |
|---|---|---|
| CsIno3上游 | BamH I | |
| CsIno3下游 | TTGTACTAAAACCTTATGTTCACT | Hind III |
| suhB 上游 | Nde I | |
| suhB 下游 | GAGTTAAGCGACGCTCTGAAGCGT | Xho I |
Table 2 Cloning sites of pETDuet-1-Csino3-suhB plasmid
引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 限制性酶切位点 Restriction enzyme site |
|---|---|---|
| CsIno3上游 | BamH I | |
| CsIno3下游 | TTGTACTAAAACCTTATGTTCACT | Hind III |
| suhB 上游 | Nde I | |
| suhB 下游 | GAGTTAAGCGACGCTCTGAAGCGT | Xho I |
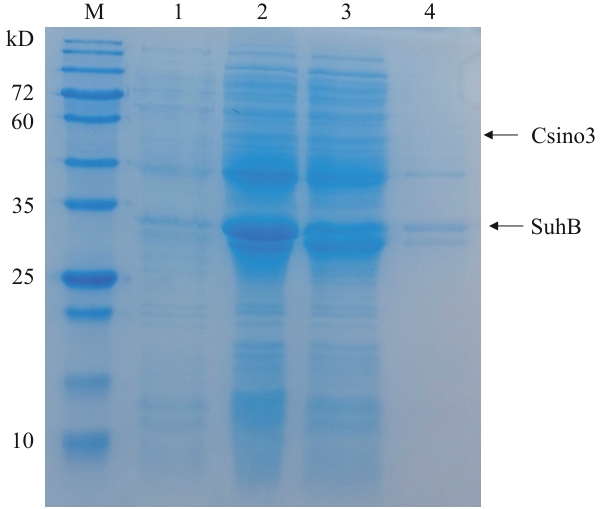
Fig. 1 Induced expression of recombinant pETDuet-1-Csino3-suhB/BL21(DE3) at 25 ℃M: Protein molecular weight marker. 1: Lysis of bacteria without induction by IPTG. 2: Lysis of bacteria after induction by IPTG. 3: Supernatant after ultrasonic crushing. 4: Sedimentation after ultrasonic crushing
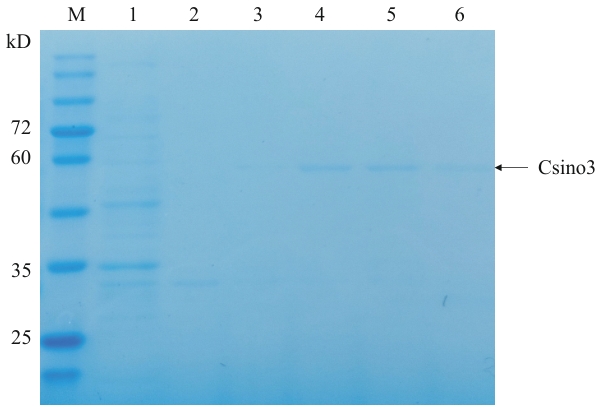
Fig. 2 Csino3 protein purificationM: Protein molecular weight standard. 1: Flow-through; 2-6: 10, 50, 100, 200, 400 mmol/L imidazole elution, respectively
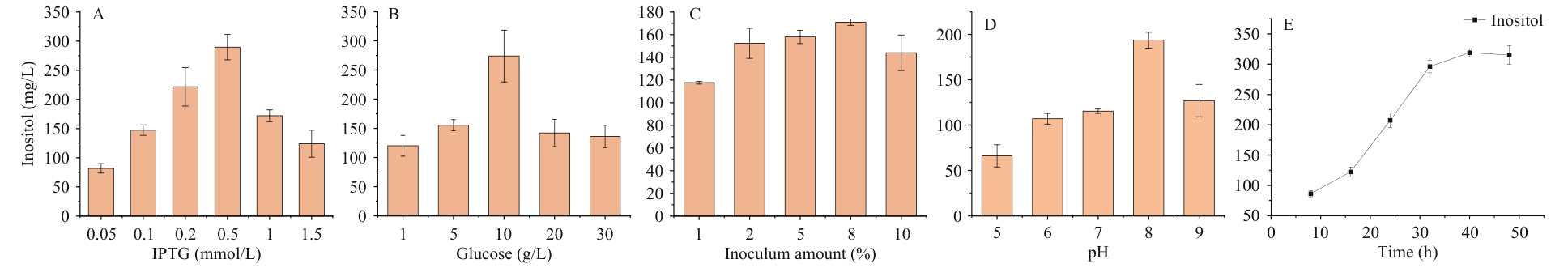
Fig. 5 Optimization of single-factor fermentationA: Final concentration of IPTG. B: Glucose concentration. C: Inoculation amount. D: Initial pH. E: Time
| 因素 Factor | Level | Low level | High level |
|---|---|---|---|
| Glucose (g/L) | 10.0 | 5.0 | 15.0 |
| IPTG (mmol/L) | 0.5 | 0.2 | 0.8 |
| Inoculation amount (%) | 8.0 | 6.0 | 10.0 |
| Initial pH | 8.0 | 7.0 | 9.0 |
Table 3 Factor and level table of Box-Behnken design
| 因素 Factor | Level | Low level | High level |
|---|---|---|---|
| Glucose (g/L) | 10.0 | 5.0 | 15.0 |
| IPTG (mmol/L) | 0.5 | 0.2 | 0.8 |
| Inoculation amount (%) | 8.0 | 6.0 | 10.0 |
| Initial pH | 8.0 | 7.0 | 9.0 |
| Number | A (g/L) | B (mmol/L) | C (%) | D | Inositol (mg/L) |
|---|---|---|---|---|---|
| 1 | 5 | 0.2 | 8 | 8 | 151.00 |
| 2 | 15 | 0.2 | 8 | 8 | 117.33 |
| 3 | 5 | 0.8 | 8 | 8 | 159.67 |
| 4 | 15 | 0.8 | 8 | 8 | 140.33 |
| 5 | 10 | 0.5 | 6 | 7 | 149.33 |
| 6 | 10 | 0.5 | 10 | 7 | 146.00 |
| 7 | 10 | 0.5 | 6 | 9 | 156.00 |
| 8 | 10 | 0.5 | 10 | 9 | 178.33 |
| 9 | 5 | 0.5 | 8 | 7 | 121.00 |
| 10 | 15 | 0.5 | 8 | 7 | 114.33 |
| 11 | 5 | 0.5 | 8 | 9 | 152.67 |
| 12 | 15 | 0.5 | 8 | 9 | 143.00 |
| 13 | 10 | 0.2 | 6 | 8 | 152.33 |
| 14 | 10 | 0.8 | 6 | 8 | 166.67 |
| 15 | 10 | 0.2 | 10 | 8 | 145.33 |
| 16 | 10 | 0.8 | 10 | 8 | 144.33 |
| 17 | 5 | 0.5 | 6 | 8 | 144.67 |
| 18 | 15 | 0.5 | 6 | 8 | 138.00 |
| 19 | 5 | 0.5 | 10 | 8 | 176.67 |
| 20 | 15 | 0.5 | 10 | 8 | 137.33 |
| 21 | 10 | 0.2 | 8 | 7 | 174.33 |
| 22 | 10 | 0.8 | 8 | 7 | 153.00 |
| 23 | 10 | 0.2 | 8 | 9 | 191.67 |
| 24 | 10 | 0.8 | 8 | 9 | 177.00 |
| 25 | 10 | 0.5 | 8 | 8 | 288.33 |
| 26 | 10 | 0.5 | 8 | 8 | 272.33 |
| 27 | 10 | 0.5 | 8 | 8 | 287.00 |
| 28 | 10 | 0.5 | 8 | 8 | 295.67 |
| 29 | 10 | 0.5 | 8 | 8 | 298.00 |
Table 4 Box-Behnken response surface experimental design and results
| Number | A (g/L) | B (mmol/L) | C (%) | D | Inositol (mg/L) |
|---|---|---|---|---|---|
| 1 | 5 | 0.2 | 8 | 8 | 151.00 |
| 2 | 15 | 0.2 | 8 | 8 | 117.33 |
| 3 | 5 | 0.8 | 8 | 8 | 159.67 |
| 4 | 15 | 0.8 | 8 | 8 | 140.33 |
| 5 | 10 | 0.5 | 6 | 7 | 149.33 |
| 6 | 10 | 0.5 | 10 | 7 | 146.00 |
| 7 | 10 | 0.5 | 6 | 9 | 156.00 |
| 8 | 10 | 0.5 | 10 | 9 | 178.33 |
| 9 | 5 | 0.5 | 8 | 7 | 121.00 |
| 10 | 15 | 0.5 | 8 | 7 | 114.33 |
| 11 | 5 | 0.5 | 8 | 9 | 152.67 |
| 12 | 15 | 0.5 | 8 | 9 | 143.00 |
| 13 | 10 | 0.2 | 6 | 8 | 152.33 |
| 14 | 10 | 0.8 | 6 | 8 | 166.67 |
| 15 | 10 | 0.2 | 10 | 8 | 145.33 |
| 16 | 10 | 0.8 | 10 | 8 | 144.33 |
| 17 | 5 | 0.5 | 6 | 8 | 144.67 |
| 18 | 15 | 0.5 | 6 | 8 | 138.00 |
| 19 | 5 | 0.5 | 10 | 8 | 176.67 |
| 20 | 15 | 0.5 | 10 | 8 | 137.33 |
| 21 | 10 | 0.2 | 8 | 7 | 174.33 |
| 22 | 10 | 0.8 | 8 | 7 | 153.00 |
| 23 | 10 | 0.2 | 8 | 9 | 191.67 |
| 24 | 10 | 0.8 | 8 | 9 | 177.00 |
| 25 | 10 | 0.5 | 8 | 8 | 288.33 |
| 26 | 10 | 0.5 | 8 | 8 | 272.33 |
| 27 | 10 | 0.5 | 8 | 8 | 287.00 |
| 28 | 10 | 0.5 | 8 | 8 | 295.67 |
| 29 | 10 | 0.5 | 8 | 8 | 298.00 |
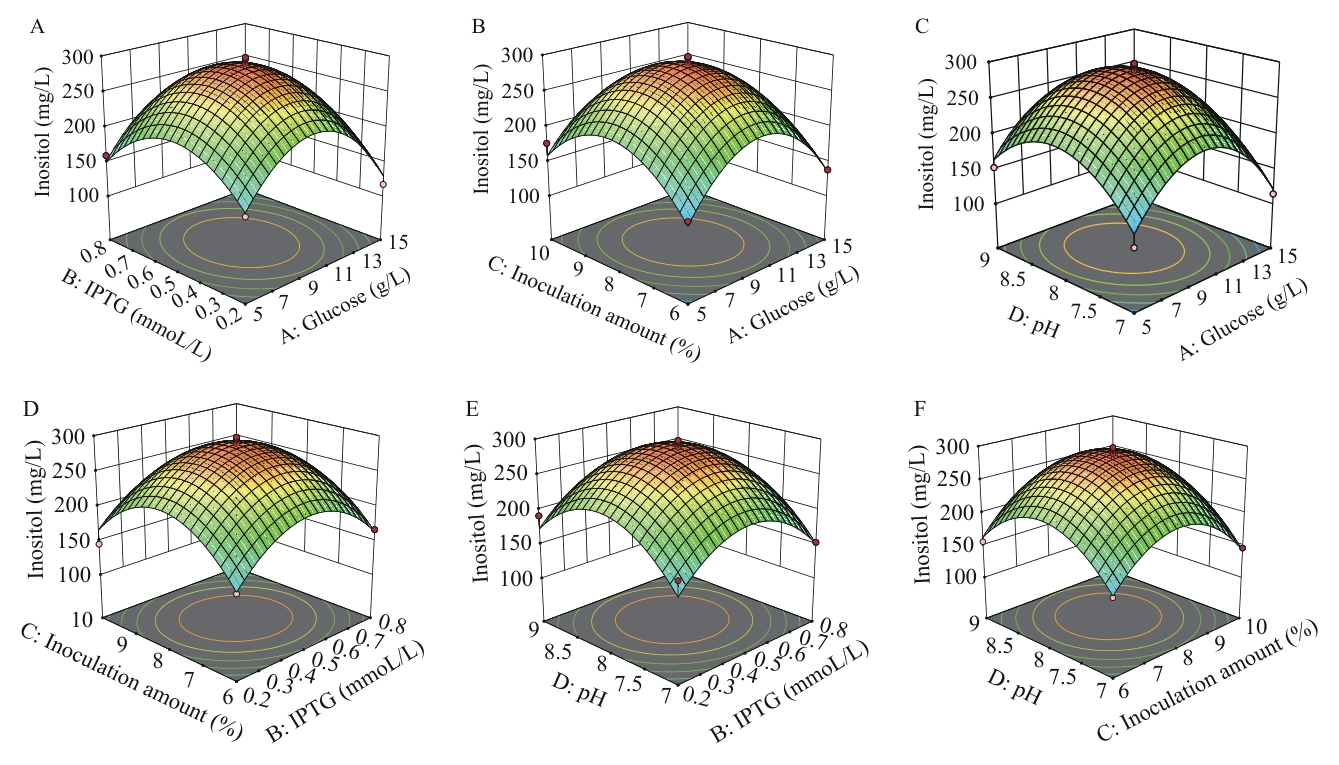
Fig. 6 Response surface on inositol production optimized by strain pETDuet-1-Csino3-suhB/BL21(DE3)A: IPTG and glucose. B: Inoculation amount and glucose. C: pH and glucose. D: Inoculation amount and IPTG. E: pH and IPTG. F: pH and inoculation amount
变异来源 Source of variation | 平方和 SS | 自由度 df | 均方 MS | F值 F-value | P值 P-value | 备注 Note |
|---|---|---|---|---|---|---|
| Model | 83 481.48 | 14 | 5 962.96 | 25.45 | <0.000 1 | ** |
| A- Glucose | 1 108.48 | 1 | 1 108.48 | 4.73 | 0.047 2 | * |
| B- IPTG | 6.75 | 1 | 6.75 | 0.028 8 | 0.867 6 | Not significant |
| C- Inoculation amount | 36.75 | 1 | 36.75 | 0.156 9 | 0.698 0 | Not significant |
| D- Initial pH | 1 648.93 | 1 | 1 648.93 | 7.04 | 0.018 9 | ** |
| AB | 51.36 | 1 | 51.36 | 0.219 2 | 0.646 8 | Not significant |
| AC | 266.78 | 1 | 266.78 | 1.14 | 0.304 0 | Not significant |
| AD | 2.25 | 1 | 2.25 | 0.009 6 | 0.923 3 | Not significant |
| BC | 58.78 | 1 | 58.78 | 0.250 9 | 0.624 2 | Not significant |
| BD | 11.11 | 1 | 11.11 | 0.047 4 | 0.830 7 | Not significant |
| CD | 164.69 | 1 | 164.69 | 0.703 0 | 0.415 9 | Not significant |
| A2 | 45 114.14 | 1 | 45 114.14 | 192.57 | <0.000 1 | ** |
| B2 | 24 352.12 | 1 | 24 352.12 | 103.94 | <0.000 1 | ** |
| C2 | 28 274.17 | 1 | 28 274.17 | 120.69 | <0.000 1 | ** |
| D2 | 26 002.04 | 1 | 26 002.04 | 110.99 | <0.000 1 | ** |
| Residual | 3 279.91 | 14 | 234.28 | |||
| Lack of fit | 2 874.94 | 10 | 287.49 | 2.84 | 0.1632 | Not significant |
| Pure error | 404.98 | 4 | 101.24 | |||
| Cor total | 86 761.39 | 28 |
Table 5 Analysis of variance (ANOVA) for Box-Behnken response surface methodology
变异来源 Source of variation | 平方和 SS | 自由度 df | 均方 MS | F值 F-value | P值 P-value | 备注 Note |
|---|---|---|---|---|---|---|
| Model | 83 481.48 | 14 | 5 962.96 | 25.45 | <0.000 1 | ** |
| A- Glucose | 1 108.48 | 1 | 1 108.48 | 4.73 | 0.047 2 | * |
| B- IPTG | 6.75 | 1 | 6.75 | 0.028 8 | 0.867 6 | Not significant |
| C- Inoculation amount | 36.75 | 1 | 36.75 | 0.156 9 | 0.698 0 | Not significant |
| D- Initial pH | 1 648.93 | 1 | 1 648.93 | 7.04 | 0.018 9 | ** |
| AB | 51.36 | 1 | 51.36 | 0.219 2 | 0.646 8 | Not significant |
| AC | 266.78 | 1 | 266.78 | 1.14 | 0.304 0 | Not significant |
| AD | 2.25 | 1 | 2.25 | 0.009 6 | 0.923 3 | Not significant |
| BC | 58.78 | 1 | 58.78 | 0.250 9 | 0.624 2 | Not significant |
| BD | 11.11 | 1 | 11.11 | 0.047 4 | 0.830 7 | Not significant |
| CD | 164.69 | 1 | 164.69 | 0.703 0 | 0.415 9 | Not significant |
| A2 | 45 114.14 | 1 | 45 114.14 | 192.57 | <0.000 1 | ** |
| B2 | 24 352.12 | 1 | 24 352.12 | 103.94 | <0.000 1 | ** |
| C2 | 28 274.17 | 1 | 28 274.17 | 120.69 | <0.000 1 | ** |
| D2 | 26 002.04 | 1 | 26 002.04 | 110.99 | <0.000 1 | ** |
| Residual | 3 279.91 | 14 | 234.28 | |||
| Lack of fit | 2 874.94 | 10 | 287.49 | 2.84 | 0.1632 | Not significant |
| Pure error | 404.98 | 4 | 101.24 | |||
| Cor total | 86 761.39 | 28 |
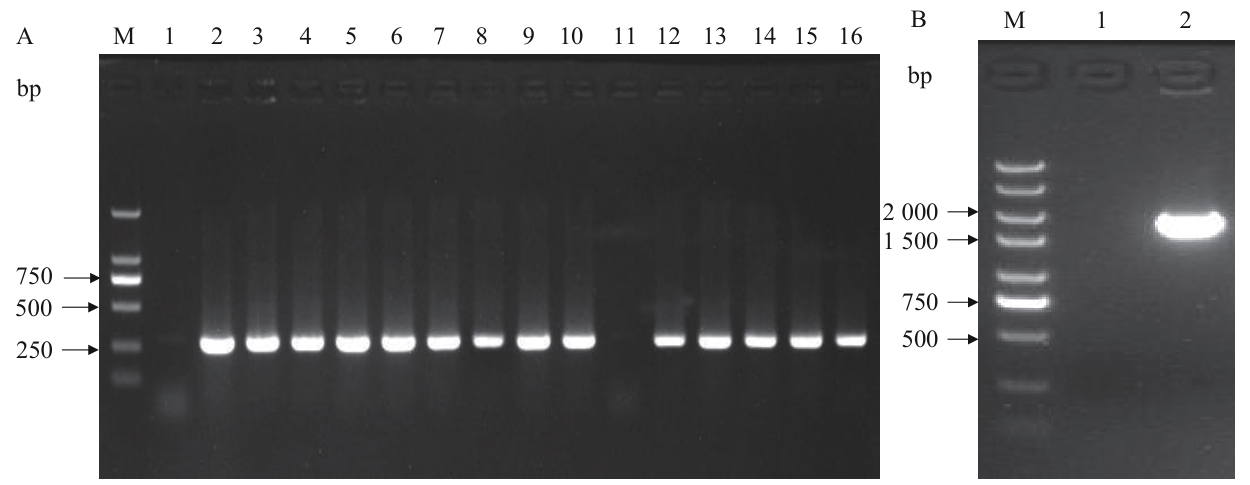
Fig. 7 Colony PCR verification of BL21(DE3)/ΔpgiM: DNA Marker. A-1: No-template negative control amplified result. A-2: Internal primer amplified result of the original strain. A-(3-16): Internal primer amplification results of clone 1-14. B-1: No-template negative control amplified result. B-2: External primer amplified result of negative clone
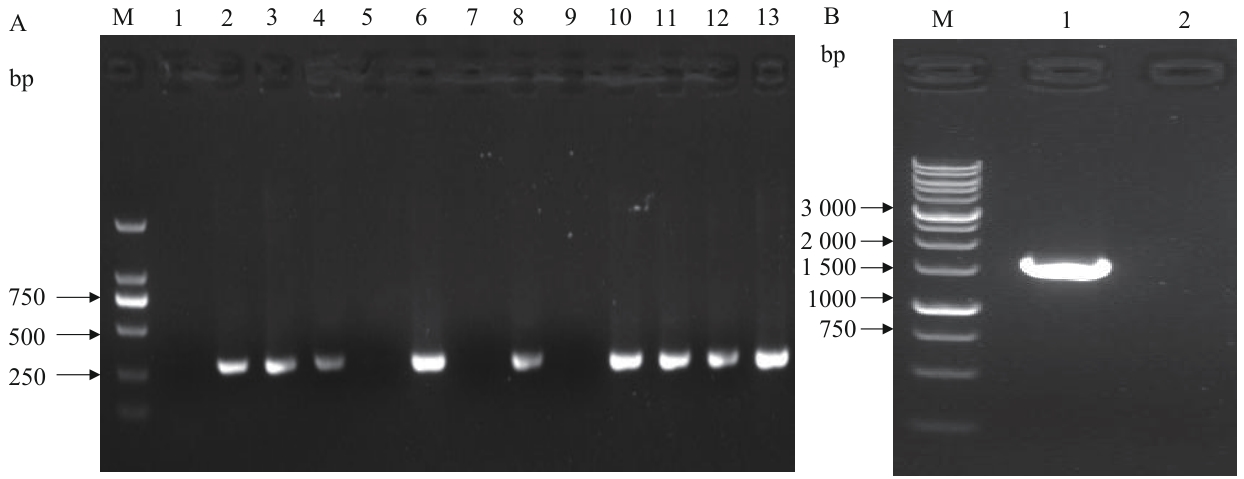
Fig. 8 Colony PCR verification of BL21(DE3)/ΔgldAM: DNA Marker. A-1: No-template negative control amplified result. A-2: Internal primer amplified result of the original strain. A-(3-13): Internal primer amplified results of clone 1-11. B-1: External primer amplified result of negative clone. B-2: No-template negative control amplified result
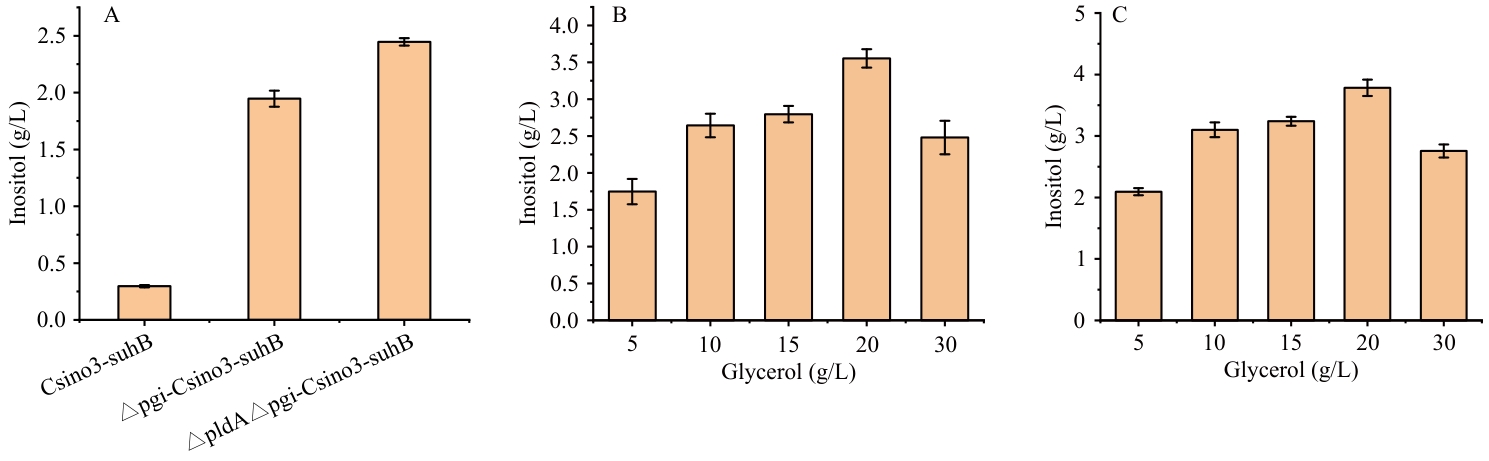
Fig. 9 Inositol-synthezing capacity of the pgi- and gldA-knockout strainA: Inositol production content by three strains. B: Effect of different glycerol concentrations on inositol fermentation by △pgi-pETDuet-1-Csino3-suhB/BL21(DE3). C: Effect of different glycerol concentrations on inositol fermentation by △gldA△pgi-pETDuet-1-Csino3-suhB/BL21(DE3)
| [1] | Kılbaş B, Balci M. Recent advances in inositol chemistry: synthesis and applications [J]. Tetrahedron, 2011, 67(13): 2355-2389. |
| [2] | 陈庆伟. 基因工程改造大肠杆菌发酵混合糖生产肌醇的研究 [D]. 杭州: 浙江工商大学, 2023. |
| Chen QW. Study on the production of inositol by fermentation of mixed sugar with genetically engineered Escherichia coli [D]. Hangzhou: Zhejiang Gongshang University, 2023. | |
| [3] | Omoruyi FO, Stennett D, Foster S, et al. New frontiers for the use of IP6 and inositol combination in treating diabetes mellitus: a review [J]. Molecules, 2020, 25(7): 1720. |
| [4] | Ma AJ, Cui WX, Wang XN, et al. Osmoregulation by the myo-inositol biosynthesis pathway in turbot Scophthalmus maximus and its regulation by anabolite and c-Myc [J]. Comp Biochem Physiol Part A Mol Integr Physiol, 2020, 242: 110636. |
| [5] | Jiang WD, Li SG, Mi HF, et al. Myo-inositol prevents the gill rot in fish caused by Flavobacterium columnare infection [J]. Aquaculture, 2022, 546: 737393. |
| [6] | Shimada M, Hibino M, Takeshita A. Dietary supplementation with myo-inositol reduces hepatic triglyceride accumulation and expression of both fructolytic and lipogenic genes in rats fed a high-fructose diet [J]. Nutr Res, 2017, 47: 21-27. |
| [7] | 杨连生, 扶雄, 孙由芳, 等. 生物酶水解植酸钙提取肌醇研究 [J]. 食品工业科技, 1997, 18(5): 48-48, 53. |
| Yang LS, Fu X, Sun YF, et al. Study on drawing inositol from phytin by hydrolysis enzyme method[J]. Sci Technol Food Ind, 1997, 18(5): 48-48, 53. | |
| [8] | 王建玲, 王敏, 王春霞, 等. 利用酶法生产肌醇初步研究 [J]. 天津轻工业学院学报, 1998, 13(2): 15-19. |
| Wang JL, Wang M, Wang CX, et al. Primary study on inositol production by hydrolyst[J]. J Tianjin Univ Sci Technol, 1998, 13(2): 15-19. | |
| [9] | 郭育涛, 陈坚, 伦世仪. 微生物发酵法生产肌醇的研究 [J]. 西北轻工业学院学报, 1998, 16(1): 117-120. |
| Guo YT, Chen J, Lun SY. Microbiol synthesis of inositol by fermentation [J]. J Shaanxi Univ Sci Technol, 1998, 16(1): 117-120. | |
| [10] | Li YJ, Han PP, Wang J, et al. Production of myo-inositol: Recent advance and prospective [J]. Biotechnol Appl Biochem, 2022, 69(3): 1101-1111. |
| [11] | Dinicola S, Unfer V, Facchinetti F, et al. Inositols: from established knowledge to novel approaches [J]. Int J Mol Sci, 2021, 22(19): 10575. |
| [12] | Lu YP, Wang L, Teng F, et al. Production of myo-inositol from glucose by a novel trienzymatic cascade of polyphosphate glucokinase, inositol 1-phosphate synthase and inositol monophosphatase [J]. Enzyme Microb Technol, 2018, 112: 1-5. |
| [13] | 黄贞杰, 陈由强, 陈丽霞, 等. 代谢工程改造酿酒酵母合成肌醇 [J]. 微生物学通报, 2017, 44(10): 2289-2296. |
| Huang ZJ, Chen YQ, Chen LX, et al. Metabolic engineering of Saccharomyces cerevisiae for inositol production [J]. Microbiol China, 2017, 44(10): 2289-2296. | |
| [14] | Shiue E, Brockman IM, Prather KLJ. Improving product yields on D-glucose in Escherichia coli via knockout of pgi and zwf and feeding of supplemental carbon sources [J]. Biotechnol Bioeng, 2015, 112(3): 579-587. |
| [15] | Yi MH, Yang LZ, Ma J, et al. Biosynthesis of myo-inositol in Escherichia coli by engineering myo-inositol-1-phosphate pathway [J]. Biochem Eng J, 2020, 164: 107792. |
| [16] | 朱宏宇, 王晓璐, 刘亚君, 等. 产肌醇毕赤酵母细胞工厂的优化 [J]. 微生物学通报, 2023, 50(9): 3731-3746. |
| Zhu HY, Wang XL, Liu YJ, et al. Optimization of a Komagataella phaffii cell factory for producing inositol [J]. Microbiol China, 2023, 50(9): 3731-3746. | |
| [17] | Henry AS, Gaspar LM, Jesch AS. The response to inositol: Regulation of glycerolipid metabolism and stress response signaling in yeast[J]. Chemistry and Physics of Lipids,2014,180:23-43. |
| [18] | 邵帅, 凌宏志, 何平, 等. 自杀质粒同源重组技术敲除阴沟肠杆菌budC基因以提升乙偶姻的产率 [J]. 黑龙江大学自然科学学报, 2023, 40(4): 434-441. |
| Shao S, Ling HZ, He P, et al. Increase acetoin production of Enterobacter cloacae: budC gene knockout by suicide plasmid homologous recombination technology [J]. J Nat Sci Heilongjiang Univ, 2023, 40(4): 434-441. | |
| [19] | 李莉, 张赛, 何强, 等. 响应面法在试验设计与优化中的应用 [J]. 实验室研究与探索, 2015, 34(8): 41-45. |
| Li L, Zhang S, He Q, et al. Application of response surface methodology in experiment design and optimization [J]. Res Explor Lab, 2015, 34(8): 41-45. | |
| [20] | Sarkar M, Majumdar P. Application of response surface methodology for optimization of heavy metal biosorption using surfactant modified chitosan bead [J]. Chem Eng J, 2011, 175: 376-387. |
| [21] | 潘柯莉, 茅以诚, 岳圣杰, 等. pgi基因敲除对大肠杆菌利用混合碳源的影响 [J]. 高校化学工程学报, 2018, 32(1): 138-146. |
| Pan KL, Mao YC, Yue SJ, et al. Effects of pgi knockout on mixed carbon source utilization of Escherichia coli [J]. J Chem Eng Chin Univ, 2018, 32(1): 138-146. | |
| [22] | 张芹, 王芳, 陈雅蕾, 等. 肌醇生产及应用研究进展 [J]. 中国稻米, 2012(3): 19-21. |
| Zhang Q, Wang F, Chen YL, et al. Research progress of inositol production and application [J]. China Rice, 2012(3): 19-21. | |
| [23] | Villa-García MJ, Choi MS, Hinz FI, et al. Genome-wide screen for inositol auxotrophy in Saccharomyces cerevisiae implicates lipid metabolism in stress response signaling [J]. Mol Genet Genom, 2011, 285(2): 125-149. |
| [24] | 祁浩, 刘新利. 大肠杆菌表达系统和酵母表达系统的研究进展 [J]. 安徽农业科学, 2016, 44(17): 4-6, 52. |
| Qi H, Liu XL. Research progress of expression systems of Escherichia coli and yeast [J]. J Anhui Agric Sci, 2016, 44(17): 4-6, 52. | |
| [25] | Moon TS, Yoon SH, Lanza AM, et al. Production of glucaric acid from a synthetic pathway in recombinant Escherichia coli [J]. Appl Environ Microbiol, 2009, 75(3): 589-595. |
| [26] | Wu YF, Shen XL, Yuan QP, et al. Metabolic engineering strategies for co-utilization of carbon sources in microbes [J]. Bioengineering, 2016, 3(1): 10. |
| [27] | 马健. 产肌醇的重组大肠杆菌工程菌构建及其发酵条件优化 [D]. 杭州: 浙江工商大学, 2022. |
| Ma J. Construction of inositol-producing recombinant Escherichia coli and optimization of fermentation conditions [D]. Hangzhou: Zhejiang Gongshang University, 2022. | |
| [28] | Zhang QQ, Wang XL, Luo HY, et al. Metabolic engineering of Pichia pastoris for myo-inositol production by dynamic regulation of central metabolism [J]. Microb Cell Fact, 2022, 21(1): 112. |
| [29] | You R, Wang L, Shi CR, et al. Efficient production of myo-inositol in Escherichia coli through metabolic engineering [J]. Microb Cell Fact, 2020, 19(1): 109. |
| [1] | WANG Ting-ting, HE Meng-ya, SHENG Jia-shun, GAO Chen, CAI Han-fang, FU Tong, SUN Yu, GAO Teng-yun, ZHANG Tian-liu. Preparation of NCAPG Knockout Bovine Fibroblast Cell Lines Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2026, 42(2): 317-324. |
| [2] | LIAN Shao-jie, TANG Sheng-shuo, KANG Chuan-li, LIU Lei, ZHENG De-qiang, DU Shuai, TANG Li-wei, ZHANG Mei-xia, LIU Qiang. Isolation, Identification, Optimization of Fermentation Conditions of High-yield Tremella fuciformis Polysaccharides Enzyme-producing Strain and Its Enzyme Characteristics Analysis [J]. Biotechnology Bulletin, 2025, 41(9): 302-313. |
| [3] | CHENG Hui-juan, WANG Xin, SHI Xiao-tao, MA Dong-xu, GONG Da-chun, HU Jun-peng, XIE Zhi-wen. Effects of Transcription Factor CREA Knockout on the Morphology and the Secretion of β-glucosidase in Aspergillus niger [J]. Biotechnology Bulletin, 2025, 41(6): 344-354. |
| [4] | WANG Xin-xin, GUAN Yu-zhu, LI Xiao-wei, HONG Wei, WU Dao-yan, KANG Ying-qian, LIU Yong-chang, CHEN Zheng-hong, CUI Gu-zhen. Function of katA in Helicobacter pylori and Its Role in the Tolerance to Oxidative Damage [J]. Biotechnology Bulletin, 2024, 40(5): 310-320. |
| [5] | GAO Deng-ke, MA Bai-rong, GUO Yi-ying, LIU Wei, LIU Tian, JIN Ya-ping, JIANG Zhou, CHEN Hua-tao. Establishment of Quaking Knockout Mouse Embryonic Fibroblast Cell Line Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2024, 40(2): 65-72. |
| [6] | WANG Xin-guang, TIAN Lei, WANG En-ze, ZHONG Cheng, TIAN Chun-jie. Construction of Microbial Consortium for Efficient Degradation of Corn Straw and Evaluation of Its Degradation Effect [J]. Biotechnology Bulletin, 2022, 38(4): 217-229. |
| [7] | YAN Jiong, FENG Chen-yi, GAO Xue-kun, XU Xiang, YANG Jia-min, CHEN Zhao-yang. Construction of Homozygous Plin1-knockout Mouse Model and Phenotype Analysis Based on CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2022, 38(3): 173-180. |
| [8] | WANG Hai-jie, WANG Cheng-ji, GUO Yang, WANG Yun, CHEN Yan-juan, LIANG Min, WANG Jue, GONG Hui, SHEN Ru-ling. Construction of Coagulation Factor 8 Gene Knockout Mouse Model Based on CRSIPR/Cas9 Technique and Verification of Phenotype [J]. Biotechnology Bulletin, 2022, 38(10): 273-280. |
| [9] | SUN Jing-ya, MA Yu-chao. Functions of Arsenic-resistant Gene Cluster in Pseudomonas sp. Tw224 [J]. Biotechnology Bulletin, 2022, 38(1): 141-149. |
| [10] | WANG Rui, HAN Lie-bao. Generation of bdfls2-knockout Mutant in Brachypodium distachyon Mediated by CRISPR/Cas9 [J]. Biotechnology Bulletin, 2022, 38(1): 70-76. |
| [11] | CHEN Qian, ZHANG Lu-yuan, CHEN Bo-chang, WU Hai-yan. Optimization of Fermentation Conditions of Myrothecium verrucaria ZW-2,a Biocontrol Strain Against Heterodera glycines and Analysis of Active Substances [J]. Biotechnology Bulletin, 2021, 37(7): 127-136. |
| [12] | JIANG Cheng-hui, ZENG Qiao-ying, WANG Meng, PAN Yang-yang, LIU Xu-ming, SHANG Tian-tian. Construction of srtA-Knockout Strain in Staphylococcus aureus by CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2020, 36(9): 253-265. |
| [13] | WANG Song, LI Peng-cheng, BAI Chao-hui, XU Hong-xin, YING Yan-min, ZHANG Mo, BAI Yi-chun. Generation of Stable α-ENaC Knockout Rat L2 Cell Strains by CRISPR/Cas9 [J]. Biotechnology Bulletin, 2020, 36(3): 115-123. |
| [14] | WU Hang, LI Zhi-qiang, LIU Wen-de. Functional Analysis of Secreted Protein MoCDIE2 Inducing Plant Cell-Death in Magnaporthe oryzae [J]. Biotechnology Bulletin, 2020, 36(1): 15-22. |
| [15] | CHEN He-feng, ZHU Chao-yi, LI Shuang. Expression Vector Adaptation of Valencene-producing Saccharomyces cerevisiae and Optimization of Fermentation Carbon and Nitrogen Sources [J]. Biotechnology Bulletin, 2020, 36(1): 209-219. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||