








Biotechnology Bulletin ›› 2022, Vol. 38 ›› Issue (5): 240-247.doi: 10.13560/j.cnki.biotech.bull.1985.2021-1183
Previous Articles Next Articles
ZHOU Zi-qi1( ), ZHANG Yang-zi1, LAN Xin-yue1, LIU Yang-er2, ZHU Long-jiao2, XU Wen-tao2(
), ZHANG Yang-zi1, LAN Xin-yue1, LIU Yang-er2, ZHU Long-jiao2, XU Wen-tao2( )
)
Received:2021-09-14
Online:2022-05-26
Published:2022-06-10
Contact:
XU Wen-tao
E-mail:zhouziqi214@163.com;xuwentao@cau.edu.cn
ZHOU Zi-qi, ZHANG Yang-zi, LAN Xin-yue, LIU Yang-er, ZHU Long-jiao, XU Wen-tao. Selection and Application of Light-up Nucleic Acid Aptamers[J]. Biotechnology Bulletin, 2022, 38(5): 240-247.
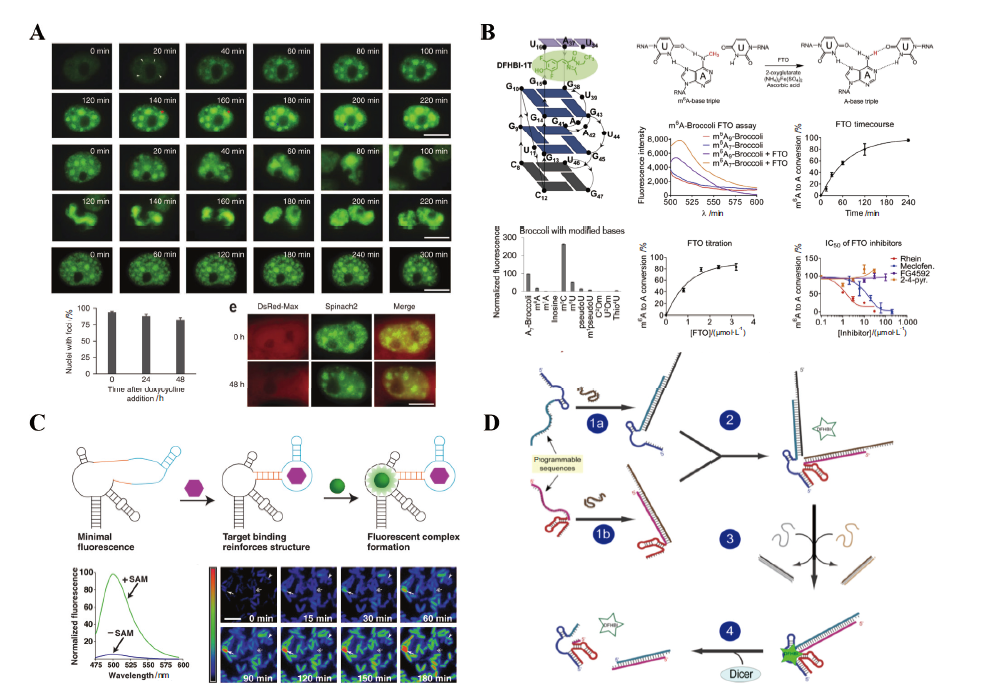
Fig. 2 Application of LNAs system in living cells A: Monitoring the formation of (CGG)60-Spinach 2 foci in transiently transfected COS-7 cell. B: Application of m6A-modified Broccoli as a fluorometric substrate in the FTO assay. C: Imaging SAM in living cells with RNA aptamer and Spinach. D: Assembly and sensing of aptamer in split Spinach

Fig. 3 Application of LNAs system in vitro A: Schematic diagram of fluorescent bivalent metal ion sensor based on riboswitch, secondary structure of czcD-2 and anaerobic titration experiment. B: Schematic representation of the in-gel detection of vegetable aptamer-tagged RNAs and their cleavage products. C: Sensing equipment and principle of ROSALIND system
| [1] | Valeur B. Molecular fluorescence[J]. Digital Encyclopedia of Applied Physics, 2003:477-531. |
| [2] |
Urbanek MO, Galka-Marciniak P, Olejniczak M, et al. RNA imaging in living cells - methods and applications[J]. RNA Biol, 2014, 11(8):1083-1095.
doi: 10.4161/rna.35506 URL |
| [3] |
Wang YX, Shyy JYJ, Chien S. Fluorescence proteins, live-cell imaging, and mechanobiology:seeing is believing[J]. Annu Rev Biomed Eng, 2008, 10:1-38.
doi: 10.1146/annurev.bioeng.010308.161731 URL |
| [4] |
Giepmans BNG, Adams SR, Ellisman MH, et al. The fluorescent toolbox for assessing protein location and function[J]. Science, 2006, 312(5771):217-224.
doi: 10.1126/science.1124618 pmid: 16614209 |
| [5] |
Schnell U, Dijk F, Sjollema KA, et al. Immunolabeling artifacts and the need for live-cell imaging[J]. Nat Methods, 2012, 9(2):152-158.
doi: 10.1038/nmeth.1855 pmid: 22290187 |
| [6] |
Santangelo PJ. Molecular beacons and related probes for intracellular RNA imaging[J]. WIREs Nanomed Nanobiotechnol, 2010, 2(1):11-19.
doi: 10.1002/wnan.52 URL |
| [7] | Ouellet J. RNA fluorescence with light-up aptamers[J]. Front Chem, 2016, 4:29. |
| [8] |
Fusco D, Accornero N, Lavoie B, et al. Single mRNA molecules demonstrate probabilistic movement in living mammalian cells[J]. Curr Biol, 2003, 13(2):161-167.
doi: 10.1016/S0960-9822(02)01436-7 URL |
| [9] |
Babendure JR, Adams SR, Tsien RY. Aptamers switch on fluorescence of triphenylmethane dyes[J]. J Am Chem Soc, 2003, 125(48):14716-14717.
pmid: 14640641 |
| [10] |
Bouhedda F, Fam KT, Collot M, et al. A dimerization-based fluorogenic dye-aptamer module for RNA imaging in live cells[J]. Nat Chem Biol, 2020, 16(1):69-76.
doi: 10.1038/s41589-019-0381-8 pmid: 31636432 |
| [11] |
Chen X, Zhang D, Su N, et al. Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs[J]. Nat Biotechnol, 2019, 37(11):1287-1293.
doi: 10.1038/s41587-019-0249-1 URL |
| [12] |
Gao T, Luo Y, Li W, et al. Progress in the isolation of aptamers to light-up the dyes and the applications[J]. Analyst, 2020, 145(3):701-718.
doi: 10.1039/C9AN01825E URL |
| [13] |
Swetha P, Fan Z, Wang FL, et al. Genetically encoded light-up RNA aptamers and their applications for imaging and biosensing[J]. J Mater Chem B, 2020, 8(16):3382-3392.
doi: 10.1039/C9TB02668A URL |
| [14] |
Stoltenburg R, Reinemann C, Strehlitz B. SELEX——a(r)evolutionary method to generate high-affinity nucleic acid ligands[J]. Biomol Eng, 2007, 24(4):381-403.
pmid: 17627883 |
| [15] |
Islam MM, Ghielmetti VM, Allen PB. Graphene oxide assisted light-up aptamer selection against Thioflavin T for label-free detection of microRNA[J]. Sci Rep, 2021, 11(1):4291.
doi: 10.1038/s41598-021-83640-z URL |
| [16] |
Chen Y, Wang J, Zhang Y, et al. Selection and characterization of a DNA aptamer to crystal violet[J]. Photochem Photobiol Sci, 2018, 17(6):800-806.
doi: 10.1039/C7PP00457E URL |
| [17] | Sando S, Narita A, Hayami M, et al. Transcription monitoring using fused RNA with a dye-binding light-up aptamer as a tag:a blue fluorescent RNA[J]. Chem Commun:Camb, 2008(33):3858-3860. |
| [18] |
Sando S, Narita A, Aoyama Y. Light-up Hoechst-DNA aptamer pair:generation of an aptamer-selective fluorophore from a conventional DNA-staining dye[J]. Chembiochem, 2007, 8(15):1795-1803.
doi: 10.1002/cbic.200700325 URL |
| [19] |
Wang HY, Wang JN, Sun N, et al. Selection and characterization of malachite green aptamers for the development of light-up probes[J]. ChemistrySelect, 2016, 1(8):1571-1574.
doi: 10.1002/slct.201600154 URL |
| [20] |
Wang JN, Zhang YJ, Wang HY, et al. Selection and analysis of DNA aptamers to berberine to develop a label-free light-up fluorescent probe[J]. New J Chem, 2016, 40(11):9768-9773.
doi: 10.1039/C6NJ02290A URL |
| [21] |
Wang HY, Wang JN, Xu LJ, et al. Selection and characterization of thioflavin T aptamers for the development of light-up probes[J]. Anal Methods, 2016, 8(48):8461-8465.
doi: 10.1039/C6AY02890J URL |
| [22] |
Kaur H. Recent developments in cell-SELEX technology for aptamer selection[J]. Biochim Biophys Acta Gen Subj, 2018, 1862(10):2323-2329.
doi: 10.1016/j.bbagen.2018.07.029 URL |
| [23] |
Paige JS, Wu KY, Jaffrey SR. RNA mimics of green fluorescent protein[J]. Science, 2011, 333(6042):642-646.
doi: 10.1126/science.1207339 URL |
| [24] |
Wang H, Wang J, Wang Q, et al. Selection and characterization of dimethylindole red DNA aptamers for the development of light-up fluorescent probes[J]. Talanta, 2017, 168:217-221.
doi: 10.1016/j.talanta.2017.03.041 URL |
| [25] |
Zou J, Huang X, Wu L, et al. Selection of intracellularly functional RNA mimics of green fluorescent protein using fluorescence-activated cell sorting[J]. J Mol Evol, 2015, 81(5/6):172-178.
doi: 10.1007/s00239-015-9718-4 URL |
| [26] |
Duchardt-Ferner E, Juen M, Bourgeois B, et al. Structure of an RNA aptamer in complex with the fluorophore tetramethylrhodamine[J]. Nucleic Acids Res, 2020, 48(2):949-961.
doi: 10.1093/nar/gkz1113 pmid: 31754719 |
| [27] |
Filonov GS, Moon JD, Svensen N, et al. Broccoli:rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution[J]. J Am Chem Soc, 2014, 136(46):16299-16308.
doi: 10.1021/ja508478x URL |
| [28] |
Lee J, Lee KH, Jeon J, et al. Combining SELEX screening and rational design to develop light-up fluorophore-RNA aptamer pairs for RNA tagging[J]. ACS Chem Biol, 2010, 5(11):1065-1074.
doi: 10.1021/cb1001894 URL |
| [29] |
Ryckelynck M, Baudrey S, Rick C, et al. Using droplet-based microfluidics to improve the catalytic properties of RNA under multiple-turnover conditions[J]. RNA, 2015, 21(3):458-469.
doi: 10.1261/rna.048033.114 pmid: 25605963 |
| [30] |
Autour A, Westhof E, Ryckelynck M. iSpinach:a fluorogenic RNA aptamer optimized for in vitro applications[J]. Nucleic Acids Res, 2016, 44(6):2491-2500.
doi: 10.1093/nar/gkw083 URL |
| [31] |
Armstrong-Price DE, Deore PS, Manderville RA. Intrinsic “turn-on” aptasensor detection of ochratoxin A using energy-transfer fluorescence[J]. J Agric Food Chem, 2020, 68(7):2249-2255.
doi: 10.1021/acs.jafc.9b07391 URL |
| [32] |
Dolgosheina EV, Jeng SC, Panchapakesan SS, et al. RNA mango aptamer-fluorophore:a bright, high-affinity complex for RNA labeling and tracking[J]. ACS Chem Biol, 2014, 9(10):2412-2420.
doi: 10.1021/cb500499x pmid: 25101481 |
| [33] | Okuda M, Fourmy D, Yoshizawa S. Use of Baby Spinach and Broccoli for imaging of structured cellular RNAs[J]. Nucleic Acids Res, 2017, 45(3):1404-1415. |
| [34] |
Strack RL, Disney MD, Jaffrey SR. A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat-containing RNA[J]. Nat Methods, 2013, 10(12):1219-1224.
doi: 10.1038/nmeth.2701 pmid: 24162923 |
| [35] |
Gao S, Zhang S, Sun X, et al. Fluorescent aptasensor based on G-quadruplex-assisted structural transformation for the detection of biomarker lipocalin 1[J]. Biosens Bioelectron, 2020, 169:112607.
doi: 10.1016/j.bios.2020.112607 URL |
| [36] |
Nilaratanakul V, Hauer DA, Griffin DE. Development of encoded Broccoli RNA aptamers for live cell imaging of Alphavirus genomic and subgenomic RNAs[J]. Sci Rep, 2020, 10(1):5233.
doi: 10.1038/s41598-020-61573-3 pmid: 32251299 |
| [37] | Renuka RM, Maroli N, Achuth J, et al. Fret based aptamer assay for sensitive detection of salmonella paratyphi a and revealing its molecular interaction with DNA gyrase[J]. bioRxiv, 2020. DOI: https://doi.org/10.1101/2020.04.02.021881. |
| [38] |
Shi L, Peng P, Zheng J, et al. I-Motif/miniduplex hybrid structures bind benzothiazole dyes with unprecedented efficiencies:a generic light-up system for label-free DNA nanoassemblies and bioimaging[J]. Nucleic Acids Res, 2020, 48(4):1681-1690.
doi: 10.1093/nar/gkaa020 URL |
| [39] |
Yoon T, Kim S, Shin J, et al. Highly sensitive multiplex detection of microRNA using light-up RNA aptamers[J]. Sens Actuat B:Chem, 2021, 330:129410.
doi: 10.1016/j.snb.2020.129410 URL |
| [40] |
Zhang J, Wang L, Jäschke A, et al. A color-shifting near-infrared fluorescent aptamer-fluorophore module for live-cell RNA imaging[J]. Angew Chem Int Ed Engl, 2021, 60(39):21441-21448.
doi: 10.1002/anie.202107250 URL |
| [41] |
Zhou YY, Zuo LX, Wei YL, et al. Development of fluorescent aptasensing system for ultrasensitive analysis of kanamycin[J]. J Lumin, 2020, 222:117124.
doi: 10.1016/j.jlumin.2020.117124 URL |
| [42] |
Grate D, Wilson C. Laser-mediated, site-specific inactivation of RNA transcripts[J]. PNAS, 1999, 96(11):6131-6136.
pmid: 10339553 |
| [43] |
Rogers TA, Andrews GE, Jaeger L, et al. Fluorescent monitoring of RNA assembly and processing using the split-spinach aptamer[J]. ACS Synth Biol, 2015, 4(2):162-166.
doi: 10.1021/sb5000725 URL |
| [44] |
Song W, Filonov GS, Kim H, et al. Imaging RNA polymerase III transcription using a photostable RNA-fluorophore complex[J]. Nat Chem Biol, 2017, 13(11):1187-1194.
doi: 10.1038/nchembio.2477 URL |
| [45] |
Svensen N, Jaffrey SR. Fluorescent RNA aptamers as a tool to study RNA-modifying enzymes[J]. Cell Chem Biol, 2016, 23(3):415-425.
doi: 10.1016/j.chembiol.2015.11.018 pmid: 26877022 |
| [46] |
Paige JS, Nguyen-Duc T, Song W, et al. Fluorescence imaging of cellular metabolites with RNA[J]. Science, 2012, 335(6073):1194.
doi: 10.1126/science.1218298 URL |
| [47] |
Kim H, Jaffrey SR. A fluorogenic RNA-based sensor activated by metabolite-induced RNA dimerization[J]. Cell Chem Biol, 2019, 26(12):1725-1731.e6.
doi: 10.1016/j.chembiol.2019.09.013 URL |
| [48] |
Kellenberger CA, Wilson SC, Sales-Lee J, et al. RNA-based fluorescent biosensors for live cell imaging of second messengers cyclic di-GMP and cyclic AMP-GMP[J]. J Am Chem Soc, 2013, 135(13):4906-4909.
doi: 10.1021/ja311960g pmid: 23488798 |
| [49] | You M, Litke JL, Jaffrey SR. Imaging metabolite dynamics in living cells using a Spinach-based riboswitch[J]. PNAS, 2015, 112(21):E2756-E2765. |
| [50] |
Xu JS, Cotruvo JA. The czcD(NiCo)riboswitch responds to iron(II)[J]. Biochemistry, 2020, 59(15):1508-1516.
doi: 10.1021/acs.biochem.0c00074 URL |
| [51] |
Filonov GS, Kam CW, Song W, et al. In-gel imaging of RNA processing using broccoli reveals optimal aptamer expression strategies[J]. Chem Biol, 2015, 22(5):649-660.
doi: 10.1016/j.chembiol.2015.04.018 URL |
| [52] |
Ponchon L, Dardel F. Recombinant RNA technology:the tRNA scaffold[J]. Nat Methods, 2007, 4(7):571-576.
pmid: 17558412 |
| [53] |
Jung JK, Alam KK, Verosloff MS, et al. Cell-free biosensors for rapid detection of water contaminants[J]. Nat Biotechnol, 2020, 38(12):1451-1459.
doi: 10.1038/s41587-020-0571-7 URL |
| [54] |
Roxo C, Kotkowiak W, Pasternak A. G-quadruplex-forming aptamers—characteristics, applications, and perspectives[J]. Molecules, 2019, 24(20):3781.
doi: 10.3390/molecules24203781 URL |
| [55] |
Xiao XN, Zhu LJ, He WC, et al. Functional nucleic acids tailoring and its application[J]. Trac Trends Anal Chem, 2019, 118:138-157.
doi: 10.1016/j.trac.2019.05.027 URL |
| [56] |
Xu WT, He WC, Du ZH, et al. Functional nucleic acid nanomaterials:development, properties, and applications[J]. Angew Chem Int Ed, 2021, 60(13):6890-6918.
doi: 10.1002/anie.201909927 URL |
| [57] |
Sunbul M, Jäschke A. SRB-2:a promiscuous rainbow aptamer for live-cell RNA imaging[J]. Nucleic Acids Res, 2018, 46(18):e110.
doi: 10.1093/nar/gky543 URL |
| [58] |
Song W, Strack RL, Jaffrey SR. Imaging bacterial protein expression using genetically encoded RNA sensors[J]. Nat Methods, 2013, 10(9):873-875.
doi: 10.1038/nmeth.2568 URL |
| [1] | LI Tian-shun, LI Chen-wei, WANG Jia, ZHU Long-Jiao, XU Wen-tao. Efficient Generation of Secondary Libraries During Functional Nucleic Acids Screening [J]. Biotechnology Bulletin, 2023, 39(3): 116-122. |
| [2] | YANG Min, LI Shu-ting, YANG Wen-ping, LI Xiang-yang, XU Wen-tao. Research Progress on Functional Nucleic Acid Biosensors Mediated by DNA/Silver Nanoclusters [J]. Biotechnology Bulletin, 2020, 36(6): 245-254. |
| [3] | XIE Yin-xia, WANG Wei-ran, CHENG Nan, XU Wen-tao. Research Progress on Electrical Signal Molecules in Electrochemical Functional Nucleic Acids Biosensors [J]. Biotechnology Bulletin, 2019, 35(5): 157-169. |
| [4] | LI Chen-wei, DU Zai-hui, LIN Shao-hua, LUO Yun-bo, XU Wen-tao. Research Progress on Functional Nucleic Acids for Detecting Pb2+ [J]. Biotechnology Bulletin, 2019, 35(1): 131-139. |
| [5] | HE Wan-chong, HUANG Kun-lun, XU Wen-tao. Research Advances on Apurinic/Apyrimidinic Endonuclease 1 Mediated Functional Nucleic Acids Biosensors [J]. Biotechnology Bulletin, 2018, 34(9): 48-54. |
| [6] | WANG Lian-zhen, ZHANG Qian, LI Kai, HE Wan-chong, LUO Yun-bo, HUANG Kun-lun, XU Wen-tao. Research Progress on Biosensor Detection Technology for Mg2+ Functional Nucleic Acids [J]. Biotechnology Bulletin, 2018, 34(9): 104-115. |
| [7] | LI Shu-ting, HE Wan-chong, HUANG Kun-lun, XU Wen-tao. Research Progress on Mesoporous Silica Mediated Functional Nucleic Acids-based Detection Technologies [J]. Biotechnology Bulletin, 2018, 34(9): 139-148. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||