








Biotechnology Bulletin ›› 2022, Vol. 38 ›› Issue (6): 81-92.doi: 10.13560/j.cnki.biotech.bull.1985.2021-1102
Previous Articles Next Articles
ZHONG Hui1,2( ), LIU Ya-jun1,2, WANG Bin-hua1,2, HE Meng-jie1,2, WU Lan1,2(
), LIU Ya-jun1,2, WANG Bin-hua1,2, HE Meng-jie1,2, WU Lan1,2( )
)
Received:2021-08-27
Online:2022-06-26
Published:2022-07-11
Contact:
WU Lan
E-mail:udaio@qq.com;ncusk724@hotmail.com
ZHONG Hui, LIU Ya-jun, WANG Bin-hua, HE Meng-jie, WU Lan. Effects of Analysis Methods on the Analyzed Results of 16S rRNA Gene Amplicon Sequencing in Bacterial Communities[J]. Biotechnology Bulletin, 2022, 38(6): 81-92.
| 样本 Sample | 样本类型 Sample type | 采样时间 Sampling time | 酸碱度 pH | 总有机碳 Total organic carbon(TOC)/(g·kg-1) | 总氮 Total nitrogen(TN)/(g·kg-1) | 总磷 Total phosphorus(TP)/(g·kg-1) | 氨态氮 NH4+ N/ (mg·kg-1) | 硝态氮 NO3- N/ (mg·kg-1) |
|---|---|---|---|---|---|---|---|---|
| FS1 | 森林土壤 | 2016年8月 | 4.57±0.24 | 48.07±1.17 | 3.98±0.26 | 0.49±0.09 | 9.01±0.64 | 45.14±11.26 |
| FS2 | 森林土壤 | 2016年8月 | 4.64±0.1 | 58.29±9.12 | 3.86±0.62 | 0.68±0.17 | 46.73±4.21 | 12.19±2.08 |
| FS3 | 森林土壤 | 2016年8月 | 4.70±0.12 | 55.17±4.91 | 3.93±0.83 | 0.56±0.11 | 13.24±1.48 | 27.90±3.41 |
| FS4 | 森林土壤 | 2016年8月 | 5.35±0.67 | 22.96±0.86 | 2.68±0.54 | 0.80±0.14 | 9.06±1.05 | 32.48±7.65 |
| WS1 | 湿地土壤 | 2015年1月 | 5.29±0.14 | 7.31±1.68 | 0.92±0.25 | 0.60±0.18 | 2.30±0.27 | 0.71±0.06 |
| WS2 | 湿地土壤 | 2015年1月 | 4.95±0.58 | 6.89±1.78 | 0.97±0.26 | 0.64±0.19 | 2.85±0.42 | 0.42±0.04 |
| WS3 | 湿地土壤 | 2015年1月 | 5.19±0.25 | 8.82±3.33 | 0.98±0.23 | 0.54±0.06 | 2.79±0.33 | 0.75±0.42 |
| WS4 | 湿地土壤 | 2015年1月 | 5.72±0.39 | 6.64±2.81 | 1.02±0.24 | 0.41±0.06 | 2.65±0.47 | 1.07±0.36 |
| CS1 | 农田土壤 | 2017年5月 | 4.71±0.54 | 36.60±4.66 | 3.95±0.15 | 0.45±0.19 | 0.91±0.11 | 2.54±1.18 |
| CS2 | 农田土壤 | 2017年5月 | 5.22±0.02 | 66.34±3.13 | 7.86±1.15 | 0.55±0.09 | 1.05±0.11 | 5.81±0.25 |
| CS3 | 农田土壤 | 2017年5月 | 5.17±0.17 | 41.11±1.62 | 4.13±0.36 | 0.26±0.01 | 0.87±0.12 | 3.37±0.80 |
| CS4 | 农田土壤 | 2017年5月 | 5.06±0.07 | 44.49±2.11 | 4.97±0.13 | 0.52±0.13 | 0.96±0.08 | 6.52±0.32 |
| LS1 | 湖泊沉积物 | 2018年5月 | 6.60±0.35 | 9.78±1.61 | 1.01±0.14 | 0.96±0.24 | 15.72±9.03 | 1.18±0.68 |
| LS2 | 湖泊沉积物 | 2018年5月 | 7.30±0.22 | 9.97±5.50 | 14.40±2.07 | 1.15±0.16 | 22.10±7.40 | 1.75±0.57 |
| LS3 | 湖泊沉积物 | 2018年5月 | 6.63±0.03 | 7.47±6.21 | 6.96±2.41 | 0.71±0.11 | 5.51±1.08 | 1.68±1.84 |
| LS4 | 湖泊沉积物 | 2018年5月 | 8.08±0.08 | 5.32±3.72 | 14.37±8.57 | 0.90±0.22 | 16.83±2.47 | 1.79±1.04 |
Table1 Information and environmental parameters of soil and sediment samples
| 样本 Sample | 样本类型 Sample type | 采样时间 Sampling time | 酸碱度 pH | 总有机碳 Total organic carbon(TOC)/(g·kg-1) | 总氮 Total nitrogen(TN)/(g·kg-1) | 总磷 Total phosphorus(TP)/(g·kg-1) | 氨态氮 NH4+ N/ (mg·kg-1) | 硝态氮 NO3- N/ (mg·kg-1) |
|---|---|---|---|---|---|---|---|---|
| FS1 | 森林土壤 | 2016年8月 | 4.57±0.24 | 48.07±1.17 | 3.98±0.26 | 0.49±0.09 | 9.01±0.64 | 45.14±11.26 |
| FS2 | 森林土壤 | 2016年8月 | 4.64±0.1 | 58.29±9.12 | 3.86±0.62 | 0.68±0.17 | 46.73±4.21 | 12.19±2.08 |
| FS3 | 森林土壤 | 2016年8月 | 4.70±0.12 | 55.17±4.91 | 3.93±0.83 | 0.56±0.11 | 13.24±1.48 | 27.90±3.41 |
| FS4 | 森林土壤 | 2016年8月 | 5.35±0.67 | 22.96±0.86 | 2.68±0.54 | 0.80±0.14 | 9.06±1.05 | 32.48±7.65 |
| WS1 | 湿地土壤 | 2015年1月 | 5.29±0.14 | 7.31±1.68 | 0.92±0.25 | 0.60±0.18 | 2.30±0.27 | 0.71±0.06 |
| WS2 | 湿地土壤 | 2015年1月 | 4.95±0.58 | 6.89±1.78 | 0.97±0.26 | 0.64±0.19 | 2.85±0.42 | 0.42±0.04 |
| WS3 | 湿地土壤 | 2015年1月 | 5.19±0.25 | 8.82±3.33 | 0.98±0.23 | 0.54±0.06 | 2.79±0.33 | 0.75±0.42 |
| WS4 | 湿地土壤 | 2015年1月 | 5.72±0.39 | 6.64±2.81 | 1.02±0.24 | 0.41±0.06 | 2.65±0.47 | 1.07±0.36 |
| CS1 | 农田土壤 | 2017年5月 | 4.71±0.54 | 36.60±4.66 | 3.95±0.15 | 0.45±0.19 | 0.91±0.11 | 2.54±1.18 |
| CS2 | 农田土壤 | 2017年5月 | 5.22±0.02 | 66.34±3.13 | 7.86±1.15 | 0.55±0.09 | 1.05±0.11 | 5.81±0.25 |
| CS3 | 农田土壤 | 2017年5月 | 5.17±0.17 | 41.11±1.62 | 4.13±0.36 | 0.26±0.01 | 0.87±0.12 | 3.37±0.80 |
| CS4 | 农田土壤 | 2017年5月 | 5.06±0.07 | 44.49±2.11 | 4.97±0.13 | 0.52±0.13 | 0.96±0.08 | 6.52±0.32 |
| LS1 | 湖泊沉积物 | 2018年5月 | 6.60±0.35 | 9.78±1.61 | 1.01±0.14 | 0.96±0.24 | 15.72±9.03 | 1.18±0.68 |
| LS2 | 湖泊沉积物 | 2018年5月 | 7.30±0.22 | 9.97±5.50 | 14.40±2.07 | 1.15±0.16 | 22.10±7.40 | 1.75±0.57 |
| LS3 | 湖泊沉积物 | 2018年5月 | 6.63±0.03 | 7.47±6.21 | 6.96±2.41 | 0.71±0.11 | 5.51±1.08 | 1.68±1.84 |
| LS4 | 湖泊沉积物 | 2018年5月 | 8.08±0.08 | 5.32±3.72 | 14.37±8.57 | 0.90±0.22 | 16.83±2.47 | 1.79±1.04 |
| 样本 Sample | 样本类型 Sample type | 采样时间 Sampling time | 酸碱度 pH | 总有机碳 TOC/(mg·kg-1) | 总氮 TN/(mg·kg-1) | 总磷 TP/(mg·kg-1) | 氨态氮 NH4+ N/(mg·kg-1) | 硝态氮 NO3--N/(mg·kg-1) |
|---|---|---|---|---|---|---|---|---|
| LW1 | 湖泊水体 | 2017年7月 | 7.87±0.50 | 16.39±6.93 | 2.40±0.66 | 0.13±0.03 | 0.26±0.10 | 0.66±0.12 |
| LW2 | 湖泊水体 | 2017年7月 | 7.3±0.16 | 12.40±5.44 | 3.10±0.63 | 0.13±0.02 | 0.14±0.05 | 0.26±0.04 |
| LW3 | 湖泊水体 | 2017年7月 | 6.79±0.41 | 11.09±5.51 | 1.38±0.36 | 0.11±0.04 | 0.34±0.15 | 0.48±0.04 |
| LW4 | 湖泊水体 | 2017年7月 | 7.12±0.13 | 14.06±6.00 | 1.14±0.67 | 0.11±0.04 | 0.25±0.06 | 0.02±0.03 |
Table2 Water samples information and environmental parameters
| 样本 Sample | 样本类型 Sample type | 采样时间 Sampling time | 酸碱度 pH | 总有机碳 TOC/(mg·kg-1) | 总氮 TN/(mg·kg-1) | 总磷 TP/(mg·kg-1) | 氨态氮 NH4+ N/(mg·kg-1) | 硝态氮 NO3--N/(mg·kg-1) |
|---|---|---|---|---|---|---|---|---|
| LW1 | 湖泊水体 | 2017年7月 | 7.87±0.50 | 16.39±6.93 | 2.40±0.66 | 0.13±0.03 | 0.26±0.10 | 0.66±0.12 |
| LW2 | 湖泊水体 | 2017年7月 | 7.3±0.16 | 12.40±5.44 | 3.10±0.63 | 0.13±0.02 | 0.14±0.05 | 0.26±0.04 |
| LW3 | 湖泊水体 | 2017年7月 | 6.79±0.41 | 11.09±5.51 | 1.38±0.36 | 0.11±0.04 | 0.34±0.15 | 0.48±0.04 |
| LW4 | 湖泊水体 | 2017年7月 | 7.12±0.13 | 14.06±6.00 | 1.14±0.67 | 0.11±0.04 | 0.25±0.06 | 0.02±0.03 |
| 群落生境 Biotope | 多样性指数 Diversity index | 97 OTU | 98 OTU | 99 OTU | 100 OTU | ASV | F | P |
|---|---|---|---|---|---|---|---|---|
| FS | Shannon | 8.89±0.59c | 9.44±0.57c | 10.06±0.53b | 11.35±0.34a | 9.36±0.36c | 3.76 | 0.01 |
| Faith’s phylog-enetic diversity | 200.65±30.09a | 206.76±30.33a | 205.45±31.21a | 189.98±30.05a | 76.57±17.53b | 3.59 | 0.01 | |
| Chao1 | 6 197.73±681.64d | 7 715.24±864.03c | 9 783.86±1121.87b | 15 783.82±1 896.16a | 1 348.3±250.55e | 64.05 | <0.01 | |
| WS | Shannon | 9.15±0.44cd | 9.51±0.40c | 9.93±0.33b | 10.76±0.22a | 9.10±0.25d | 45.32 | <0.01 |
| Faith’s phylog-enetic diversity | 131.82±20.02ab | 135.38±20.39a | 131.70±21.02ab | 112.17±17.56b | 59.95±11.00c | 46.78 | <0.01 | |
| Chao1 | 3 432.91±489.63c | 4 119.94±523.16bc | 4 834.80±605.21b | 6 259.35±1 102.91a | 909.03±166.82d | 272.21 | <0.01 | |
| CS | Shannon | 8.99±0.88b | 9.27±0.93b | 9.60±1.01ab | 10.40±1.00a | 9.42±0.92ab | 3.51 | 0.01 |
| Faith’s phylog-enetic diversity | 180.30±33.33ab | 169.08±31.10ab | 202.30±38.74a | 199.45±36.96a | 159.09±30.77b | 1.1 | 0.37 | |
| Chao1 | 3 672.12±892.42c | 4 271.85±1 029.13bc | 4 993.04±1 280.13b | 9 069.09±1 554.95a | 2 186.44±570.72d | 51.61 | <0.01 | |
| LS | Shannon | 6.82±1.06b | 6.96±1.15b | 7.21±1.24ab | 8.32±1.09a | 7.20±0.90ab | 49.53 | <0.01 |
| Faith’s phylog-enetic diversity | 97.39±35.21a | 97.87±34.88a | 99.98±35.44a | 110.06±32.85a | 83.22±15.54a | 35.66 | <0.01 | |
| Chao1 | 1 429.92±867.84bc | 1 646.30±1 010.35bc | 1 916.62±1 242.16b | 5 343.72±481.48a | 934.21±263.76c | 110.54 | <0.01 | |
| LW | Shannon | 6.71±1.12b | 6.98±1.14b | 7.36±1.10b | 9.04±0.71a | 7.50±0.80b | 10.01 | <0.01 |
| Faith’s phylog-enetic diversity | 79.77±30.70a | 88.65±32.64a | 91.25±32.18a | 85.25±24.93a | 59.01±18.95a | 2.48 | 0.05 | |
| Chao1 | 1 709.26±868.06bc | 2 043.13±914.34bc | 2 674.50±1 066.06b | 8 847.05±1 696.64a | 837.88±332.8c | 108.4 | <0.01 |
Table 3 Alpha diversity of bacterial community
| 群落生境 Biotope | 多样性指数 Diversity index | 97 OTU | 98 OTU | 99 OTU | 100 OTU | ASV | F | P |
|---|---|---|---|---|---|---|---|---|
| FS | Shannon | 8.89±0.59c | 9.44±0.57c | 10.06±0.53b | 11.35±0.34a | 9.36±0.36c | 3.76 | 0.01 |
| Faith’s phylog-enetic diversity | 200.65±30.09a | 206.76±30.33a | 205.45±31.21a | 189.98±30.05a | 76.57±17.53b | 3.59 | 0.01 | |
| Chao1 | 6 197.73±681.64d | 7 715.24±864.03c | 9 783.86±1121.87b | 15 783.82±1 896.16a | 1 348.3±250.55e | 64.05 | <0.01 | |
| WS | Shannon | 9.15±0.44cd | 9.51±0.40c | 9.93±0.33b | 10.76±0.22a | 9.10±0.25d | 45.32 | <0.01 |
| Faith’s phylog-enetic diversity | 131.82±20.02ab | 135.38±20.39a | 131.70±21.02ab | 112.17±17.56b | 59.95±11.00c | 46.78 | <0.01 | |
| Chao1 | 3 432.91±489.63c | 4 119.94±523.16bc | 4 834.80±605.21b | 6 259.35±1 102.91a | 909.03±166.82d | 272.21 | <0.01 | |
| CS | Shannon | 8.99±0.88b | 9.27±0.93b | 9.60±1.01ab | 10.40±1.00a | 9.42±0.92ab | 3.51 | 0.01 |
| Faith’s phylog-enetic diversity | 180.30±33.33ab | 169.08±31.10ab | 202.30±38.74a | 199.45±36.96a | 159.09±30.77b | 1.1 | 0.37 | |
| Chao1 | 3 672.12±892.42c | 4 271.85±1 029.13bc | 4 993.04±1 280.13b | 9 069.09±1 554.95a | 2 186.44±570.72d | 51.61 | <0.01 | |
| LS | Shannon | 6.82±1.06b | 6.96±1.15b | 7.21±1.24ab | 8.32±1.09a | 7.20±0.90ab | 49.53 | <0.01 |
| Faith’s phylog-enetic diversity | 97.39±35.21a | 97.87±34.88a | 99.98±35.44a | 110.06±32.85a | 83.22±15.54a | 35.66 | <0.01 | |
| Chao1 | 1 429.92±867.84bc | 1 646.30±1 010.35bc | 1 916.62±1 242.16b | 5 343.72±481.48a | 934.21±263.76c | 110.54 | <0.01 | |
| LW | Shannon | 6.71±1.12b | 6.98±1.14b | 7.36±1.10b | 9.04±0.71a | 7.50±0.80b | 10.01 | <0.01 |
| Faith’s phylog-enetic diversity | 79.77±30.70a | 88.65±32.64a | 91.25±32.18a | 85.25±24.93a | 59.01±18.95a | 2.48 | 0.05 | |
| Chao1 | 1 709.26±868.06bc | 2 043.13±914.34bc | 2 674.50±1 066.06b | 8 847.05±1 696.64a | 837.88±332.8c | 108.4 | <0.01 |
| 样地 Sampling site | 门总数 Number of phyla | 差异门 Differential phylum | 差异门丰度Abundance of differential phylum/% | 属总数 Number of genera | 差异属 Differential genus | 差异属丰度Abundance of differential genus/% |
|---|---|---|---|---|---|---|
| FS | 40 | 2 | 3.7 | 888 | 75 | 2.9 |
| WS | 52 | 0 | 0 | 854 | 30 | 0.35 |
| CS | 58 | 0 | 0 | 1 227 | 38 | 3.21 |
| LS | 54 | 0 | 0 | 1 304 | 15 | 14.9 |
| LW | 51 | 0 | 0 | 1 179 | 18 | 1.32 |
Table 4 Effects of the minimum taxonomy unit division method on the abundances of bacterial community phylum and genus
| 样地 Sampling site | 门总数 Number of phyla | 差异门 Differential phylum | 差异门丰度Abundance of differential phylum/% | 属总数 Number of genera | 差异属 Differential genus | 差异属丰度Abundance of differential genus/% |
|---|---|---|---|---|---|---|
| FS | 40 | 2 | 3.7 | 888 | 75 | 2.9 |
| WS | 52 | 0 | 0 | 854 | 30 | 0.35 |
| CS | 58 | 0 | 0 | 1 227 | 38 | 3.21 |
| LS | 54 | 0 | 0 | 1 304 | 15 | 14.9 |
| LW | 51 | 0 | 0 | 1 179 | 18 | 1.32 |
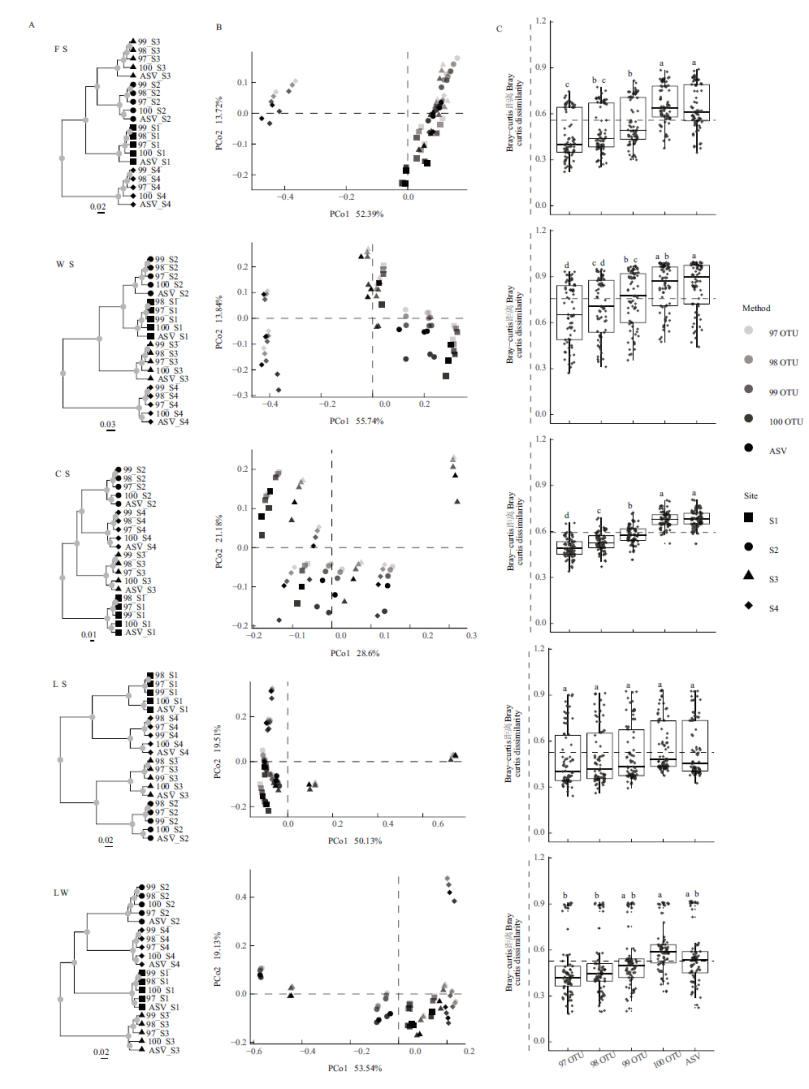
Fig.2 Analysis of β diversity based on the bacterial community(genus level)bray-curtis dissimility A:Cluster analysis. B:Principal co-ordinates analysis(PCoA). C:Differences in beta-diversity among the division methods
| [1] |
Locey KJ, Lennon JT. Scaling laws predict global microbial diversity[J]. PNAS, 2016, 113(21):5970-5975.
doi: 10.1073/pnas.1521291113 URL |
| [2] |
高贵锋, 褚海燕. 微生物组学的技术和方法及其应用[J]. 植物生态学报, 2020, 44(4):395-408.
doi: 10.17521/cjpe.2019.0222 |
|
Gao GF, Chu HY. Techniques and methods of microbiomics and their applications[J]. Chin J Plant Ecol, 2020, 44(4):395-408.
doi: 10.17521/cjpe.2019.0222 URL |
|
| [3] |
Sanz JL, Köchling T. Molecular biology techniques used in wastewater treatment:an overview[J]. Process Biochem, 2007, 42(2):119-133.
doi: 10.1016/j.procbio.2006.10.003 URL |
| [4] | Caruso V, Song XB, Asquith M, et al. Performance of microbiome sequence inference methods in environments with varying biomass[J]. mSystems, 2019, 4(1):e00163-e00118. |
| [5] |
Yang B, Wang Y, Qian PY. Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis[J]. BMC Bioinformatics, 2016, 17:135.
doi: 10.1186/s12859-016-0992-y pmid: 27000765 |
| [6] |
Loman NJ, Misra RV, Dallman TJ, et al. Performance comparison of benchtop high-throughput sequencing platforms[J]. Nat Biotechnol, 2012, 30(5):434-439.
doi: 10.1038/nbt.2198 URL |
| [7] |
Allali I, Arnold JW, Roach J, et al. A comparison of sequencing platforms and bioinformatics pipelines for compositional analysis of the gut microbiome[J]. BMC Microbiol, 2017, 17(1):194.
doi: 10.1186/s12866-017-1101-8 pmid: 28903732 |
| [8] |
Pedrós-Alió C. Marine microbial diversity:can it be determined?[J]. Trends Microbiol, 2006, 14(6):257-263.
pmid: 16679014 |
| [9] |
Lu HP, Yeh YC, Sastri AR, et al. Evaluating community-environment relationships along fine to broad taxonomic resolutions reveals evolutionary forces underlying community assembly[J]. Isme J, 2016, 10(12):2867-2878.
doi: 10.1038/ismej.2016.78 URL |
| [10] |
Capunitan DC, Johnson O, Terrill RS, et al. Evolutionary signal in the gut microbiomes of 74 bird species from Equatorial Guinea[J]. Mol Ecol, 2020, 29(4):829-847.
doi: 10.1111/mec.15354 pmid: 31943484 |
| [11] |
Edgar RC. UPARSE:highly accurate OTU sequences from microbial amplicon reads[J]. Nat Methods, 2013, 10(10):996-998.
doi: 10.1038/nmeth.2604 URL |
| [12] |
Gevers D, Cohan FM, Lawrence JG, et al. Re-evaluating prokaryotic species[J]. Nat Rev Microbiol, 2005, 3(9):733-739.
pmid: 16138101 |
| [13] | Erko S, Ebers J. Taxonomic parameters revisited:tarnished gold standards[J]. Microbiol Today, 2006, 33:152-155. |
| [14] |
Edgar RC. Updating the 97% identity threshold for 16S ribosomal RNA OTUs[J]. Bioinformatics, 2018, 34(14):2371-2375.
doi: 10.1093/bioinformatics/bty113 URL |
| [15] |
Callahan BJ, McMurdie PJ, Rosen MJ, et al. DADA2:High-resolution sample inference from Illumina amplicon data[J]. Nat Methods, 2016, 13(7):581-583.
doi: 10.1038/NMETH.3869 |
| [16] |
Berg G, Rybakova D, et al. Microbiome definition re-visited:old concepts and new challenges[J]. Microbiome, 2020, 8(1):103.
doi: 10.1186/s40168-020-00875-0 URL |
| [17] |
Scarlett K, Denman S, Clark DR, et al. Relationships between nitrogen cycling microbial community abundance and composition reveal the indirect effect of soil pH on oak decline[J]. Isme J, 2021, 15(3):623-635.
doi: 10.1038/s41396-020-00801-0 pmid: 33067585 |
| [18] |
Joos L, Beirinckx S, Haegeman A, et al. Daring to be differential:metabarcoding analysis of soil and plant-related microbial communities using amplicon sequence variants and operational taxonomical units[J]. BMC Genom, 2020, 21(1):733.
doi: 10.1186/s12864-020-07126-4 URL |
| [19] | 刘亚军, 蔡润发, 李赟璟, 等. 湿地土壤微生物碳源代谢活性对不同水分条件的动态响应——以鄱阳湖为例[J]. 土壤, 2018, 50(4):705-711. |
| Liu YJ, Cai RF, Li YJ, et al. Functional response of wetland soil microbial carbon source metabolic activity to different water conditions—A case of lake Poyang[J]. Soils, 2018, 50(4):705-711. | |
| [20] | 何世耀. 鄱阳湖饶河入湖口水体和底泥微生物对氮、磷及重金属污染物输入的响应[D]. 南昌: 南昌大学, 2019. |
| He SY. The response of microbial in sediment and water of Poyang lake to the inputs of N, P and heavy metals pollutions in Raohe river[D]. Nanchang: Nanchang University, 2019. | |
| [21] |
Bolyen E, Rideout JR, Dillon MR, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2[J]. Nat Biotechnol, 2019, 37(8):852-857.
doi: 10.1038/s41587-019-0209-9 pmid: 31341288 |
| [22] | Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads[J]. EMBnet J, 2011, 17(1):10. |
| [23] | Rognes T, Flouri T, Nichols B, et al. VSEARCH:a versatile open source tool for metagenomics[J]. PeerJ, 2016, 4:e2584. |
| [24] | Quast C, Pruesse E, Yilmaz P, et al. The SILVA ribosomal RNA gene database project:improved data processing and web-based tools[J]. Nucleic Acids Res, 2013, 41(database issue):D590-D596. |
| [25] |
Bokulich NA, Kaehler BD, Rideout JR, et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin[J]. Microbiome, 2018, 6(1):90.
doi: 10.1186/s40168-018-0470-z URL |
| [26] |
Katoh K, Misawa K, Kuma K, et al. MAFFT:a novel method for rapid multiple sequence alignment based on fast Fourier transform[J]. Nucleic Acids Res, 2002, 30(14):3059-3066.
doi: 10.1093/nar/gkf436 URL |
| [27] | Price MN, Dehal PS, Arkin AP. FastTree 2——approximately maximum-likelihood trees for large alignments[J]. PLoS One, 2010, 5(3):e9490. |
| [28] | Oksanen J, Blanchet FG, Friendly M, et al. vegan:Community Ecology Package[J]. R package version 2. 5-7, 2020. |
| [29] |
Hothorn T, Bretz F, Westfall P. Simultaneous inference in general parametric models[J]. Biom J, 2008, 50(3):346-363.
doi: 10.1002/bimj.200810425 URL |
| [30] |
Wickham H. ggplot2[J]. WIREs Comp Stat, 2011, 3(2):180-185.
doi: 10.1002/wics.147 URL |
| [31] |
Shade A. Diversity is the question, not the answer[J]. Isme J, 2017, 11(1):1-6.
doi: 10.1038/ismej.2016.118 URL |
| [32] | Louca S, Mazel F, Doebeli M, et al. A census-based estimate of Earth’s bacterial and archaeal diversity[J]. PLoS Biol, 2019, 17(2):e3000106. |
| [33] | Li XC, Huo SL, Xi BD. Updating the resolution for 16S rRNA OTUs clustering reveals the cryptic cyanobacterial genus and species[J]. Ecol Indic, 2020, 117:106695. |
| [34] |
Kunin V, Engelbrektson A, et al. Wrinkles in the rare biosphere:pyrosequencing errors can lead to artificial inflation of diversity estimates[J]. Environ Microbiol, 2010, 12(1):118-123.
doi: 10.1111/j.1462-2920.2009.02051.x URL |
| [35] |
Washburne AD, Morton JT, Sanders J, et al. Methods for phylogenetic analysis of microbiome data[J]. Nat Microbiol, 2018, 3(6):652-661.
doi: 10.1038/s41564-018-0156-0 pmid: 29795540 |
| [36] |
Barnes CJ, Rasmussen L, et al. Comparing DADA2 and OTU clustering approaches in studying the bacterial communities of atopic dermatitis[J]. J Med Microbiol, 2020, 69(11):1293-1302.
doi: 10.1099/jmm.0.001256 URL |
| [37] |
Zhou Z, Wang C, Luo Y. Meta-analysis of the impacts of global change factors on soil microbial diversity and functionality[J]. Nat Commun, 2020, 11(1):3072.
doi: 10.1038/s41467-020-16881-7 URL |
| [38] |
Berlow M, Phillips JN, Derryberry EP. Effects of urbanization and landscape on gut microbiomes in white-crowned sparrows[J]. Microb Ecol, 2021, 81(1):253-266.
doi: 10.1007/s00248-020-01569-8 URL |
| [39] |
Cottrell MT, David KL. Contribution of major bacterial groups to bacterial biomass production(thymidine and leucine incorporation)in the Delaware estuary[J]. Limnol Oceanogr, 2003, 48(1):168-178.
doi: 10.4319/lo.2003.48.1.0168 URL |
| [40] | Dell’Anno F, Rastelli E, et al. Highly contaminated marine sediments can host rare bacterial taxa potentially useful for bioremediation[J]. Front Microbiol, 2021, 12:584850. |
| [41] |
Grady EN, MacDonald J, Liu L, et al. Current knowledge and perspectives of Paenibacillus:a review[J]. Microb Cell Fact, 2016, 15(1):203.
doi: 10.1186/s12934-016-0603-7 URL |
| [42] |
McSpadden Gardener BB. Ecology of Bacillus and Paenibacillus spp. in agricultural systems[J]. Phytopathology®, 2004, 94(11):1252-1258.
doi: 10.1094/PHYTO.2004.94.11.1252 URL |
| [43] |
Monciardini P, Cavaletti L, Schumann P, et al. Conexibacter woesei gen. nov., sp. nov., a novel representative of a deep evolutionary line of descent within the class Actinobacteria[J]. Int J Syst Evol Microbiol, 2003, 53(Pt 2):569-576.
doi: 10.1099/ijs.0.02400-0 URL |
| [44] | Dong L, Cheng R, Xiao L, et al. Diversity and composition of bacterial endophytes among plant parts of Panax notoginseng[J]. Chinese medicine, 2018, 13(1):1-9. |
| [45] |
Berestovskaya JJ, Kotsyurbenko OR, Tourova TP, et al. Methylorosula polaris gen. nov., sp. nov., an aerobic, facultatively methylotrophic psychrotolerant bacterium from tundra wetland soil[J]. Int J Syst Evol Microbiol, 2012, 62(Pt 3):638-646.
doi: 10.1099/ijs.0.007005-0 URL |
| [46] |
Lozupone CA, Hamady M, Kelley ST, et al. Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities[J]. Appl Environ Microbiol, 2007, 73(5):1576-1585.
doi: 10.1128/AEM.01996-06 URL |
| [47] |
Tromas N, Fortin N, Bedrani L, et al. Characterising and predicting cyanobacterial blooms in an 8-year amplicon sequencing time course[J]. Isme J, 2017, 11(8):1746-1763.
doi: 10.1038/ismej.2017.58 URL |
| [48] |
An JX, Liu C, Wang Q, et al. Soil bacterial community structure in Chinese wetlands[J]. Geoderma, 2019, 337:290-299.
doi: 10.1016/j.geoderma.2018.09.035 URL |
| [1] | SUN Hai-hang, GUAN Hui-lin, WANG Xu, WANG Tong, LI Hong-lin, PENG Wen-jie, LIU Bo-zhen, FAN Fang-ling. Effects of Biochar on the Soil Properties and Fungal Community Structure under Continuous Cropping of Panax notoginseng [J]. Biotechnology Bulletin, 2023, 39(2): 221-231. |
| [2] | CHEN Tian-ci, WU Shao-lan, YANG Guo-hui, JIANG Dan-xia, JIANG Yu-ji, CHEN Bing-zhi. Effects of Ganoderma resinaceum Alcohol Extract on Sleep and Intestinal Microbiota in Mice [J]. Biotechnology Bulletin, 2022, 38(8): 225-232. |
| [3] | ZHAO Lin-yan, GUAN Hui-lin, XIANG Ping, LI Ze-cheng, BAI Yu-long, SONG Hong-chuan, SUN Shi-zhong, XU Wu-mei. Composition Features of Microbial Community in the Rhizospheric Soil of Bletilla striata with Root Rot [J]. Biotechnology Bulletin, 2022, 38(2): 67-74. |
| [4] | CHEN Yu-jie, ZHENG Hua-bao, ZHOU Xin-yan. Modified High-throughput Sequencing Reveals the Effects of Different Algicides towards Algal Community [J]. Biotechnology Bulletin, 2022, 38(11): 70-79. |
| [5] | CAO Xiu-kai, WANG Shan, GE Ling, ZHANG Wei-bo, SUN Wei. Advances in Extrachromosomal Circular DNA and Their Application in Domestic Animal Breeding [J]. Biotechnology Bulletin, 2022, 38(1): 247-257. |
| [6] | WANG Qi, WU Zhi-xuan, CHEN Zhong-ling, WU Bai-yi-la, HU Zong-fu, NIU Hua-xin. Effects of Lactobacillus paracasei on the Quality and Bacterial Diversity of Silage Alfalfa After Aerobic Exposure [J]. Biotechnology Bulletin, 2021, 37(9): 77-85. |
| [7] | MAO Ting, NIU Yong-yan, ZHENG Qun, YANG Tao, MU Yong-song, ZHU Ying, JI Bin, WANG Zhi-ye. Effects of Microbial Inoculants on the Fermentation Quality and Microbial Community Diversity of Alfalfa Silage [J]. Biotechnology Bulletin, 2021, 37(9): 86-94. |
| [8] | TANG Die, ZHOU Qian. Research Advances in Plant Genome Assembly [J]. Biotechnology Bulletin, 2021, 37(6): 1-12. |
| [9] | ZHU Bin, GAN Chen-chen, WANG Hong-cheng. Characteristics of the Complete Chloroplast Genome of Dendrobium thyrsiflorum and Its Phylogenetic Relationship Analysis [J]. Biotechnology Bulletin, 2021, 37(5): 38-47. |
| [10] | ZHANG Shu-hua, FANG Qian, JIA Hong-mei, HAN Gui-qi, YAN Zhu-yun, HE Dong-mei. Difference Analysis of Fungal Community Among Bulk Soil,Rhizosphere and Rhizomes of Ligusticum chuanxiong Hort. [J]. Biotechnology Bulletin, 2021, 37(4): 56-69. |
| [11] | GUO Yan-ping, ZHANG Hao, ZHAO Xin-gang, LUO Hai-ling, ZHANG Ying-jun. Applications of DNA Metabarcoding in Diet Identification of Herbivores [J]. Biotechnology Bulletin, 2021, 37(3): 252-260. |
| [12] | ZHENG Fang-fang, LIN Jun-sheng. Selection and Specificity of Nucleic Acid Aptamers for a Proliferation Inducing Ligand [J]. Biotechnology Bulletin, 2021, 37(10): 196-202. |
| [13] | LI Ye-qing, JING Zhang-mu, JIANG Hao, XU Quan, ZHOU Hong-jun, FENG Lu. Microbiome and Its Research Progress of Anaerobic Digestion [J]. Biotechnology Bulletin, 2021, 37(1): 90-101. |
| [14] | WANG Hong-jie, LIU Shao-dong, LIU Rui-hua, ZHANG Si-ping, YANG Jun, PANG Chao-you. Effects of Crop Rotation on Bacterial Communities in Cotton Rhizosphere Soil [J]. Biotechnology Bulletin, 2020, 36(9): 117-124. |
| [15] | ZHANG Miao, CHEN Yu-feng, CHEN Long, HUANG Piao-ling, WEI Lu-ling. Difference Analysis of the Community Diversity of Fungi in the Rhizosphere Soil of Zanthoxylum nitidum(Roxb.)DC in Different Regions [J]. Biotechnology Bulletin, 2020, 36(9): 167-179. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||