








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 149-157.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0974
Previous Articles Next Articles
WANG Jing1( ), LIU Zhuo-yan2, ZHANG Xiao-lei1, LIU Bao-hai1, GUAN Hai-tao1(
), LIU Zhuo-yan2, ZHANG Xiao-lei1, LIU Bao-hai1, GUAN Hai-tao1( ), WEN Hong-tao1(
), WEN Hong-tao1( )
)
Received:2025-09-10
Online:2026-02-26
Published:2026-03-17
Contact:
GUAN Hai-tao, WEN Hong-tao
E-mail:buyijingjing@163.com;ght20080808@126.com;wen0891@163.com
WANG Jing, LIU Zhuo-yan, ZHANG Xiao-lei, LIU Bao-hai, GUAN Hai-tao, WEN Hong-tao. Rapid Detection of Transgenic Maize DBN9501 Event-specificity Based on LAMP-CRISPR-Cas12a Technology[J]. Biotechnology Bulletin, 2026, 42(2): 149-157.
| 名称 Name | 序列 Sequence (5′-3′) |
|---|---|
| DBN9501-LB-F3 | CCAGCCACTATTAAGTTTCATG |
| DBN9501-LB-B3 | AATCTGTACCTAGTTTAGCTAGT |
| DBN9501-LB-FIP | TGGCAAAAAGGCATCCTGTTTTAAAAAATCCAAAGAAACCATACG |
| DBN9501-LB-BIP | TTCAGGAAAAACAGGGGAAAAGTAGGACGTTAACAGGGACG |
| DBN9501-RB-F3 | TTAATACAAATCCACCTGCG |
| DBN9501-RB-B3 | GAACATTTCTTTATTCGGAAACA |
| DBN9501-RB-FIP | GAGTCACGTTATGACCCCCGTTTTGCGGTTCTGTCAGTTC |
| DBN9501-RB-BIP | AATTCTCCGCTCATGATCAGATTGTAATCCGGACTACTATACCATT |
| crRNA-LB | AAUUUCUACUGUUGUAGAUAAAAAAUCCAAAGAAACCAUACG |
| crRNA-RB | AAUUUCUACUGUUGUAGAUGCGGUUCUGUCAGUUCCAAACG |
| FB-reporter | 6FAM-TTTTTTTTTT-Biotin |
Table 1 Sequence information of primers, probe reporters, and CrRNA
| 名称 Name | 序列 Sequence (5′-3′) |
|---|---|
| DBN9501-LB-F3 | CCAGCCACTATTAAGTTTCATG |
| DBN9501-LB-B3 | AATCTGTACCTAGTTTAGCTAGT |
| DBN9501-LB-FIP | TGGCAAAAAGGCATCCTGTTTTAAAAAATCCAAAGAAACCATACG |
| DBN9501-LB-BIP | TTCAGGAAAAACAGGGGAAAAGTAGGACGTTAACAGGGACG |
| DBN9501-RB-F3 | TTAATACAAATCCACCTGCG |
| DBN9501-RB-B3 | GAACATTTCTTTATTCGGAAACA |
| DBN9501-RB-FIP | GAGTCACGTTATGACCCCCGTTTTGCGGTTCTGTCAGTTC |
| DBN9501-RB-BIP | AATTCTCCGCTCATGATCAGATTGTAATCCGGACTACTATACCATT |
| crRNA-LB | AAUUUCUACUGUUGUAGAUAAAAAAUCCAAAGAAACCAUACG |
| crRNA-RB | AAUUUCUACUGUUGUAGAUGCGGUUCUGUCAGUUCCAAACG |
| FB-reporter | 6FAM-TTTTTTTTTT-Biotin |
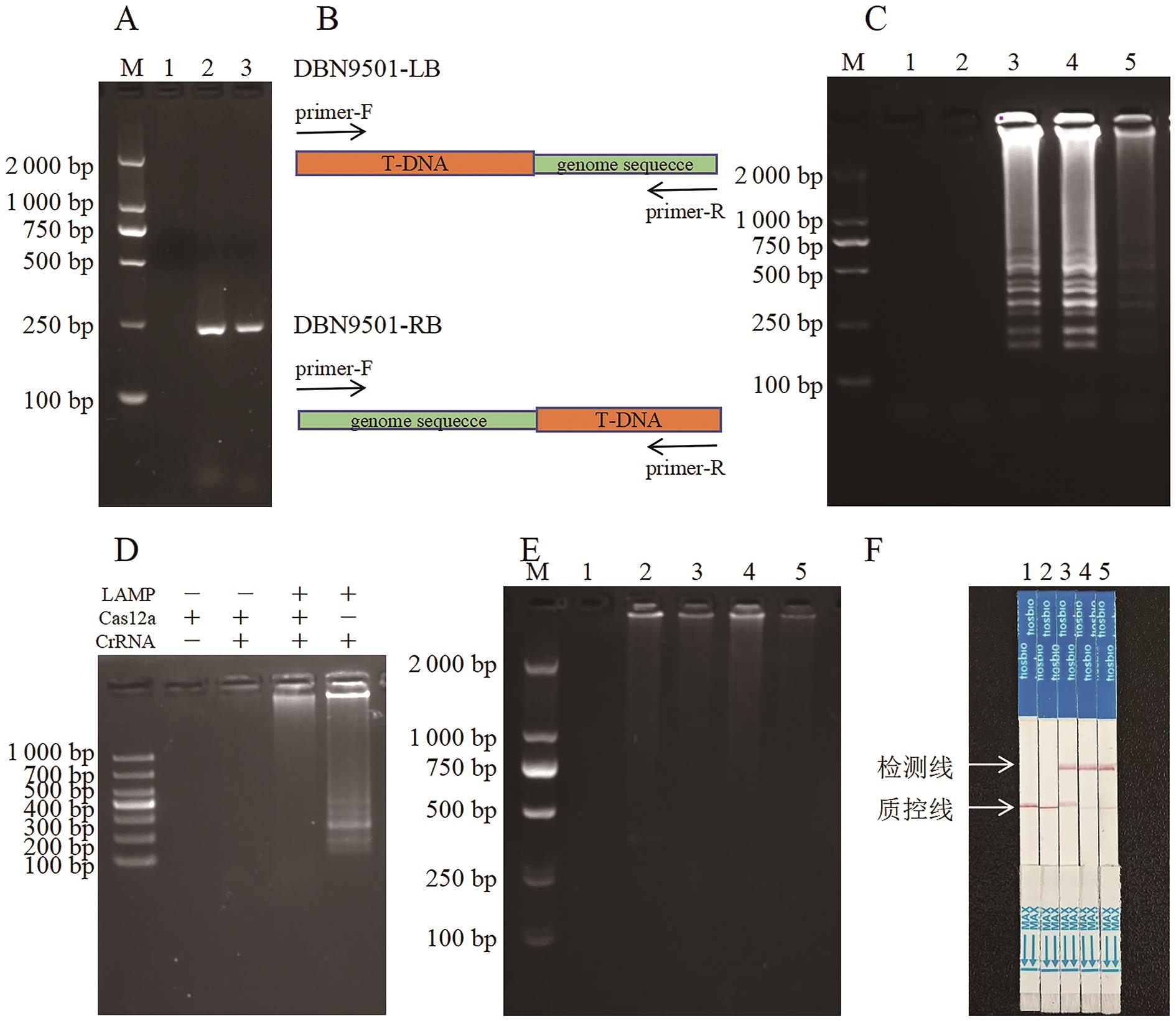
Fig. 1 Rapid detection workflow of DBN9501 event-specificityA: PCR products of LB and RB sequences of DBN9501 event amplified by LAMP-F3/B3 (1: No template control (NTC) ; 2: LB PCR product; 3:RB PCR product); B: Illustration of DBN9501-LB plasmid sequencing results; C: LAMP amplified results. D: Verification of cis-enzyme digestion condition. E: Trans-enzyme digestion results. F: Strip test assay. In Fig.C-F, 1: NTC; 2: Non-transgenic maize. 3: DBN9501 event. 4: DBN9501-LB plasmid. 5: DBN9501-RB plasmid
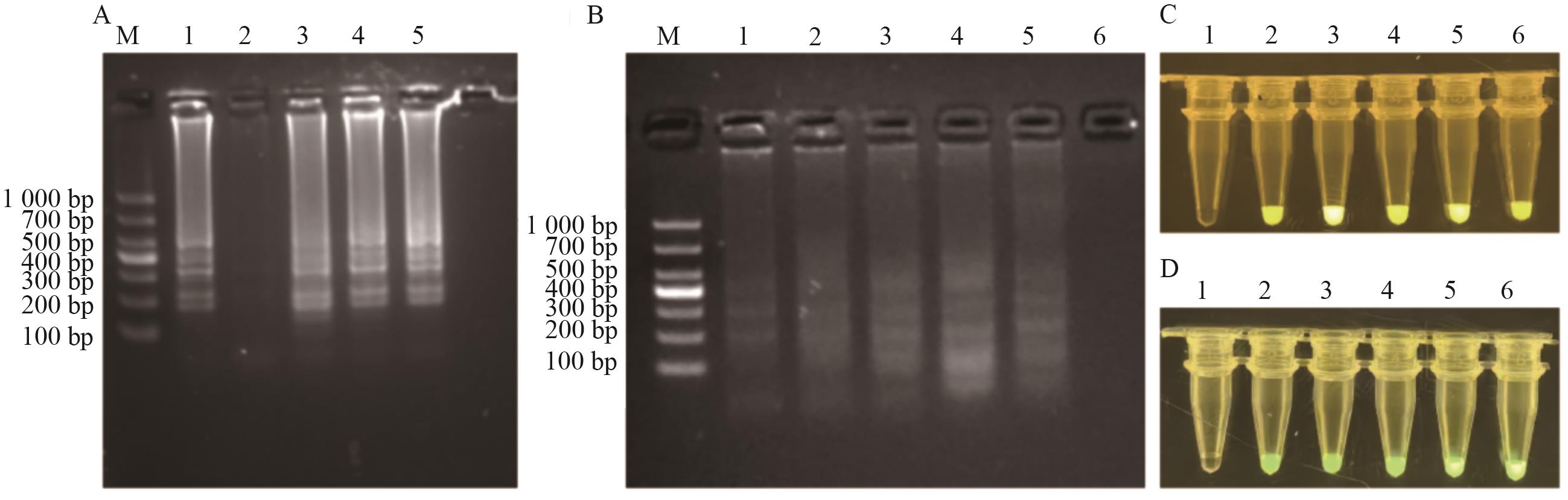
Fig. 2 Optimized results of reaction conditions for the LAMP-CRISPR/Cas12aA: LAMP primer FIP/BIP∶F3/B3 Concentration optimization (1: Positive control (2∶1): 2: NTC; 3: 6∶1; 4: 8∶1; 5: 10∶1). B: Optimization of LAMP reaction temperature (1: 62 ℃; 2: 63 ℃; 3: 64 ℃; 4: 65 ℃; 5: 66 ℃; 6: NTC). C: Optimization of Cas12a/crRNA ratio (1: H2O; 2: 0.1∶0.5; 3: 0.2∶0.4; 4: 0.3∶0.3; 5: 0.4∶0.2; 6: 0.5∶0.1). D: Optimization of FB reporter (1: H2O; 2: 62.5; 3: 50.0; 4: 37.5; 5: 25.0; 6: 12.5 nmol/L)
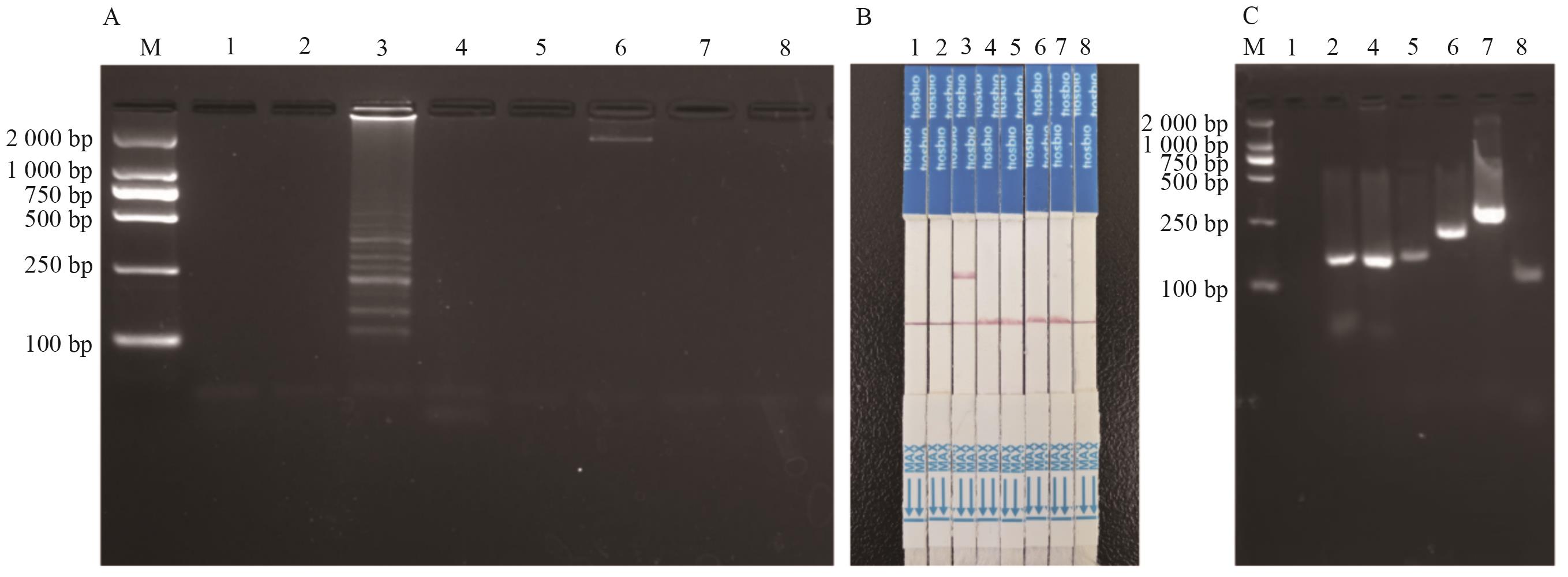
Fig. 3 Specificity detection for LAMP-CRISPR/Cas12a methodA: LAMP amplified results; B: strip test assay results; C: internal standard gene PCR amplified results. 1: Blank control; 2: non-transgenic maize; 3: DBN9501 event; 4: transgenic maize (excluding DBN9501); 5:transgenic rape; 6: transgenic soybean; 7: transgenic rice; 8: transgenic cotton
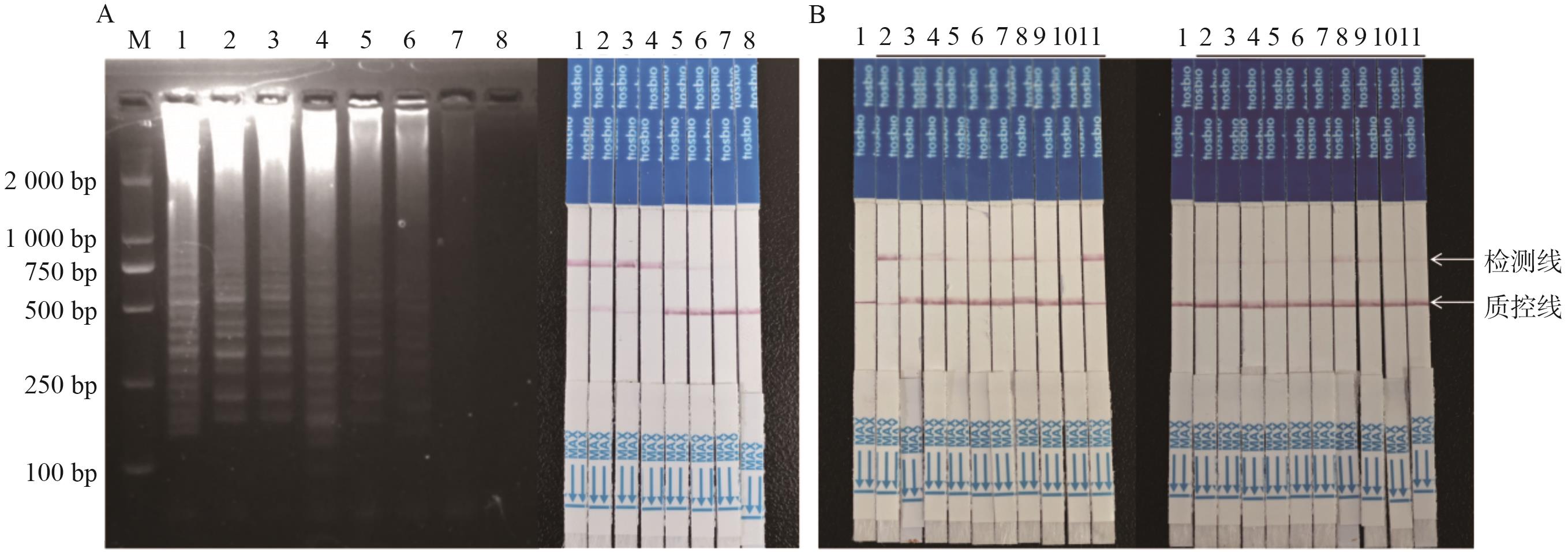

Fig. 4 Assay of sensitivity and limit of detection for LAMP-CRISPR/Cas12a methodA: Sensitivity assay, the left picture is LAMP amplified results; the right picture is strip detection assay (1: 8.0×103; 2: 8.0×102; 3: 8.0×10; 4: 4.0×10; 5: 2.0×10; 7: 1.0×10; 7: 1.0; 8: NTC). B: Assay of detection limit, left picture is the assay of detection limit for plasmid, and right picture is DBN9501 event by strip test assay (1: NTC; 2-11: 20 copies DBN9501-LB plasmid)
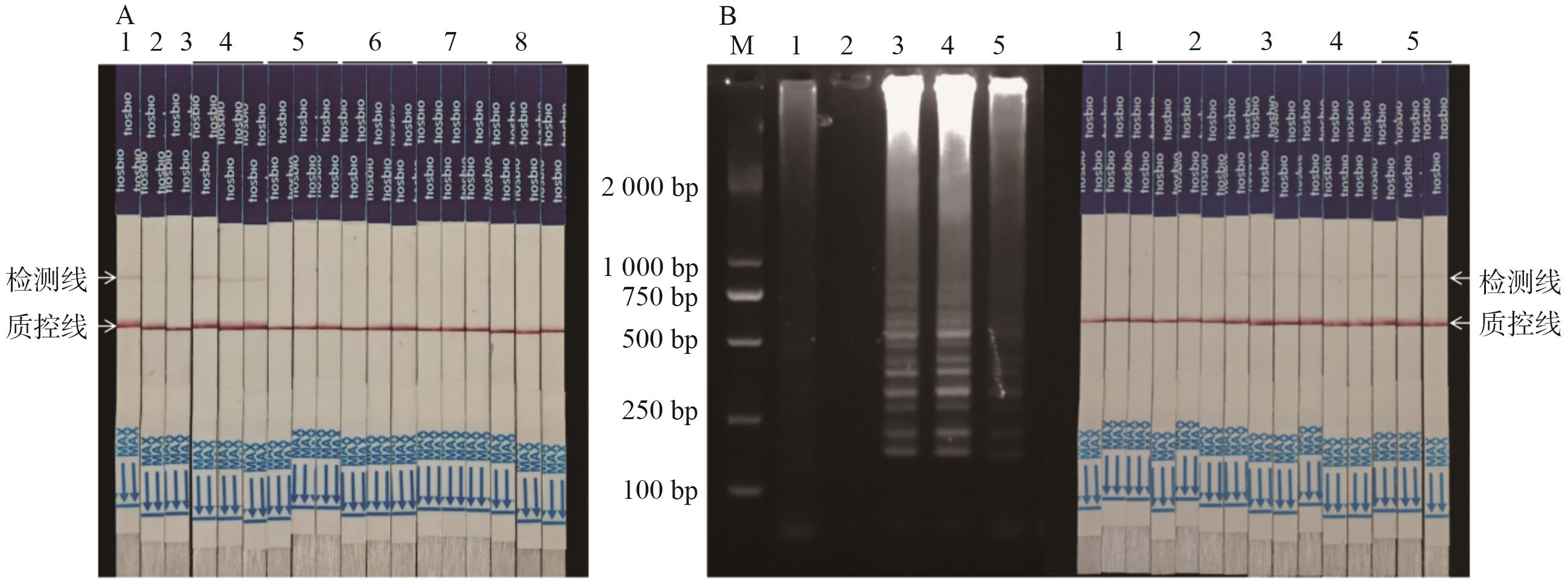
Fig. 5 Blind sample testing and method applicationA: Blind sample test (1: DBN9501-LB; 2: non-transgenic maize; 3: NTC; 4: blind sample (DBN9501). 5: Blind samples (non-transgenic maize); 6: blind sample (Bt11); 7: blind sample (DBN9936); 8: blind sample (MON810)). B: Application of assay method, the left is LAMP amplification results, the right is a test strip test (M: marker; 1: non-transgenic maize; 2: NTC; 3: DBN9501-LB; 4: DBN9501 leaves; 5: DBN9501 seeds, 3 replicates for strip test assay)
方法 Method | 操作步骤 Procedure | 检测对象 Test object | 灵敏度 Sensitivity | 检测时间 Testing time | 参考文献 References |
|---|---|---|---|---|---|
| RPA-Cas12a-FS | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | P-CaMV35S,T-nos | 10拷贝 | ~45 min | [ |
| MR-DCA | (1) 多重RPA (2) CRISPR/Cas13a或Cas12a+荧光/试纸条显示 | CaMV35S,NOS | 20拷贝 | ~35 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas (3) G-四链体比色测定 | CaMV35 | 10 amol/L和0.01% | ~45 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | CP4-EPSPS,Cry1Ab/Ac | 10 拷贝,0.5% | ~50 min | [ |
| RAA-CRISPR-Cas12a | (1) 重组酶辅助扩增(RAA); (2) CRISPR/Cas12a (3) 裸眼观察金纳米棒颜色变化 | NOS | 1 amol/L和0.1% | ~60 min | [ |
| LAMP-CRISPR/Cas12b | (1) LAMP (2) CRISPR/Cas12b (3) 荧光显示 | CaMV35S | 10拷贝,0.05% | ~40 min | [ |
| LAMP-CRISPR/Cas12a | (1) LAMP (2) CRISPR/Cas 12a (3) 试纸条显示 | DBN9501 | 20拷贝 | ~60 min | 本方法 |
Table 2 Comparison of transgenic detecting time based on CRISPR/Cas technology
方法 Method | 操作步骤 Procedure | 检测对象 Test object | 灵敏度 Sensitivity | 检测时间 Testing time | 参考文献 References |
|---|---|---|---|---|---|
| RPA-Cas12a-FS | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | P-CaMV35S,T-nos | 10拷贝 | ~45 min | [ |
| MR-DCA | (1) 多重RPA (2) CRISPR/Cas13a或Cas12a+荧光/试纸条显示 | CaMV35S,NOS | 20拷贝 | ~35 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas (3) G-四链体比色测定 | CaMV35 | 10 amol/L和0.01% | ~45 min | [ |
| RPA-CRISPR/Cas12a | (1) RPA (2) CRISPR/Cas12a (3) 荧光/试纸条显示 | CP4-EPSPS,Cry1Ab/Ac | 10 拷贝,0.5% | ~50 min | [ |
| RAA-CRISPR-Cas12a | (1) 重组酶辅助扩增(RAA); (2) CRISPR/Cas12a (3) 裸眼观察金纳米棒颜色变化 | NOS | 1 amol/L和0.1% | ~60 min | [ |
| LAMP-CRISPR/Cas12b | (1) LAMP (2) CRISPR/Cas12b (3) 荧光显示 | CaMV35S | 10拷贝,0.05% | ~40 min | [ |
| LAMP-CRISPR/Cas12a | (1) LAMP (2) CRISPR/Cas 12a (3) 试纸条显示 | DBN9501 | 20拷贝 | ~60 min | 本方法 |
| [1] | 王晶, 张晓磊, 白玉, 等. 不同玉米转化体通用PCR检测体系建立 [J]. 生物技术通报, 2024, 40(12): 34-44. |
| Wang J, Zhang XL, Bai Y, et al. Establishment of a universal PCR detection system for different maize events [J]. Biotechnol Bull, 2024, 40(12): 34-44. | |
| [2] | 王颢潜, 高鸿飞, 王梦雨, 等. 转基因生物成分快速检测技术研究进展 [J]. 中国油料作物学报, 2022, 44(3): 491-496. |
| Wang HQ, Gao HF, Wang MY, et al. Research progress on the rapid detection technologies for composition of genetically modified organisms [J]. Chin J Oil Crop Sci, 2022, 44(3): 491-496. | |
| [3] | Notomi T, Okayama H, Masubuchi H, et al. Loop-mediated isothermal amplification of DNA [J]. Nucleic Acids Res, 2000, 28(12): e63. |
| [4] | Han X, Lu MH, Zhang YR, et al. A thermostable Cas12b-powered bioassay coupled with loop-mediated isothermal amplification in a customized "one-pot" vessel for visual, rapid, sensitive, and on-site detection of genetically modified crops [J]. J Agric Food Chem, 2024, 72(19): 11195-11204. |
| [5] | Liang Y, He YH, Yang SF, et al. A novel cross-priming amplification technique combined with lateral flow strips for rapid and visual detection of zoonotic Toxoplasma gondii [J]. Vet Parasitol, 2025, 334: 110402. |
| [6] | Zhu XY, Yang H, Wang M, et al. Label-free detection of transgenic crops using an isothermal amplification reporting CRISPR/Cas12 assay [J]. ACS Synth Biol, 2022, 11(1): 317-324. |
| [7] | Ding L, Wang XF, Chen XY, et al. Development of a novel Cas13a/Cas12a-mediated ‘one-pot’ dual detection assay for genetically modified crops [J]. J Adv Res, 2025, 72: 97-106. |
| [8] | Park JW. Principles and applications of loop-mediated isothermal amplification to point-of-care tests [J]. Biosensors (Basel), 2022, 12(10): 857. |
| [9] | 中华人民共和国农业农村部. 转基因植物及其产品成分检测 环介导等温扩增方法制定指南: 农业农村部公告第323号-9—2020 [S]. 北京:中国农业出版社, 2020: 3. |
| Ministry of Agriculture and Rural Affairs of the People’s Republic of China. Detection of genetically modified plants and derived products-Guide for establishing loop-mediated isothermal amplification method: Announcement No. 323-9-2020 of the Ministry of Agriculture and Rural Affairs of the People’s Republic of China [S]. Beijing: China Agriculture Press, 2020: 3. | |
| [10] | Chylinski K, Makarova KS, Charpentier E, et al. Classification and evolution of type II CRISPR-Cas systems [J]. Nucleic Acids Res, 2014, 42(10): 6091-6105. |
| [11] | Zhu ZB, Li R, Zhang HW, et al. PAM-free loop-mediated isothermal amplification coupled with CRISPR/Cas12a cleavage (Cas-PfLAMP) for rapid detection of rice pathogens [J]. Biosens Bioelectron, 2022, 204: 114076. |
| [12] | 王渭霞, 朱廷恒, 赖凤香, 等. CRISPR/Cas系统在基因修饰植物及其产品检测应用中的原理和进展 [J]. 中国稻米, 2023, 29(6): 21-27. |
| Wang WX, Zhu TH, Lai FX, et al. Principle and progress of CRISPR/cas system in the detection of genetically modified plants and their products [J]. China Rice, 2023, 29(6): 21-27. | |
| [13] | Wang HM, Su AL, Chang JJ, et al. Sensitive detection of genetically modified maize based on a CRISPR/Cas12a system [J]. Analyst, 2024, 149(3): 836-845. |
| [14] | Li GY, Li SN, Li XH, et al. A novel electrochemical aptasensor based on NrGO-H-Mn3O4 NPs integrated CRISPR/Cas12a system for ultrasensitive low-density lipoprotein determination [J]. Microchim Acta, 2024, 191(9): 547. |
| [15] | Wang YH, Yang TM, Liu GF, et al. Application of CRISPR/Cas12a in the rapid detection of pathogens [J]. Clin Chim Acta, 2023, 548: 117520. |
| [16] | Xue PP, Peng YB, Wang RJ, et al. Advances, challenges, and opportunities for food safety analysis in the isothermal nucleic acid amplification/CRISPR-Cas12a era [J]. Crit Rev Food Sci Nutr, 2025, 65(13): 2473-2488. |
| [17] | Cao GH, Dong JB, Chen XL, et al. Simultaneous detection of CaMV35S and T-nos utilizing CRISPR/Cas12a and Cas13a with multiplex-PCR (MPT-Cas12a/13a) [J]. Chem Commun, 2022, 58(43): 6328-6331. |
| [18] | Huang D, Qian JJ, Shi ZW, et al. CRISPR-Cas12a-assisted multicolor biosensor for semiquantitative point-of-use testing of the nopaline synthase Terminator in genetically modified crops by unaided eyes [J]. ACS Synth Biol, 2020, 9(11): 3114-3123. |
| [19] | Wang Z, Huang CM, Wei S, et al. A CRISPR/Cas12a-mediated sensitive DNA detection system for gene-edited rice [J]. J AOAC Int, 2023, 106(3): 558-567. |
| [20] | Chen L, Wang C, Zhu ZB, et al. Field-deployable detection of genetically modified organisms with an integrated method of loop-mediated isothermal amplification and CRISPR/FnCas12a [J]. J Agric Food Chem, 2025, 73(9): 5625-5634. |
| [21] | Li R, Chen JW, Zhang XJ, et al. Mini-disk capillary array coupling with LAMP for visual detection of multiple nucleic acids using genetically modified organism analysis as an example [J]. J Agric Food Chem, 2020, 68(3): 899-906. |
| [22] | Wu H, He JS, Zhang F, et al. Contamination-free visual detection of CaMV35S promoter amplicon using CRISPR/Cas12a coupled with a designed reaction vessel: Rapid, specific and sensitive [J]. Anal Chim Acta, 2020, 1096: 130-137. |
| [23] | Wang J, Xiao B, Zhang RY, et al. Inter-laboratory validation by event-specific qPCR methods for the detection of genetically modified insect and herbicide-tolerant maize DBN9501 [J]. J Consum Prot Food Saf, 2023, 18(4): 451-460. |
| [24] | Li SY, Cheng QX, Liu JK, et al. CRISPR-Cas12a has both Cis- and trans-cleavage activities on single-stranded DNA [J]. Cell Res, 2018, 28(4): 491-493. |
| [25] | Liu H, Wang JB, Zeng HJ, et al. RPA-Cas12a-FS: a frontline nucleic acid rapid detection system for food safety based on CRISPR-Cas12a combined with recombinase polymerase amplification [J]. Food Chem, 2021, 334: 127608. |
| [26] | Li XH, Liu ML, Men DH, et al. Rapid, portable, and sensitive detection of CaMV35S by RPA-CRISPR/Cas12a-G4 colorimetric assays with high accuracy deep learning object recognition and classification [J]. Talanta, 2024, 278: 126441. |
| [27] | Wang JB, Luo JW, Liu H, et al. “Blue-red-purple” multicolored lateral flow immunoassay for simultaneous detection of GM crops utilizing RPA and CRISPR/Cas12a [J]. Talanta, 2025, 282: 127010. |
| [28] | Holst-Jensen A, Bertheau Y, de Loose M, et al. Detecting un-authorized genetically modified organisms (GMOs) and derived materials [J]. Biotechnol Adv, 2012, 30(6): 1318-1335. |
| [29] | Shehata HR, Ragupathy S, Shanmughanandhan D, et al. Guidelines for validation of qualitative real-time PCR methods for molecular diagnostic identification of probiotics [J]. J Aoac Int, 2019, 102(6): 1774-1778. |
| [30] | 麦晓东, 石磊, 常彦磊, 等. 环介导等温扩增技术(LAMP)在转基因大豆DBN9004品系快速检测中的应用 [J/OL]. 现代食品科技, 2025: 1-10. . |
| Mai XD, Shi L, Chang YL, et al. Application of Loop-mediated isothermal amplification (LAMP) in rapid identification of genetically modified soybean event DBN 9004. Modern food Sci Technol [J/OL]. Mod Food Sci Technol, 2025: 1-10. | |
| [31] | 咸若彤, 缪青梅, 彭城, 等. 转基因玉米WYN17132转化体特异性实时荧光PCR检测方法的建立与应用 [J]. 浙江农业学报, 2025, 37(7): 1397-1406. |
| Xian RT, Miao QM, Peng C, et al. Establishment and application of event-specific real-time PCR detection method of transgenic maize WYN17132 [J]. Acta Agric Zhejiangensis, 2025, 37(7): 1397-1406. |
| [1] | LI Ya-qi, SUN Meng, LI Xiu-li, WEI Jing-na, ZHAO Lin-lin, ZHAO Yun-ping, LIU Zheng-hui, SU Fan. Optimization of a High-performance and Low-cost Fluorescence Detection Buffer with Broad Compatibility across Cas12a Orthologs [J]. Biotechnology Bulletin, 2026, 42(4): 83-91. |
| [2] | HUO Guan-zhong, ZHANG Xin-ru, TIAN Shi-jun, LI Jun. Current Progress and Applications of CRISPR/Cas12a Gene Editing Technology in Plants [J]. Biotechnology Bulletin, 2025, 41(6): 1-11. |
| [3] | TAN Yu-rong, CHEN Dong-liang, YANG Shou-zhen, LAI Zhen-guang, TANG Xiang-min, SUN Zu-dong, ZENG Wei-ying. Functioal Analysis on GmKTI1-like Gene of Soybean Resistance to Bean Pyralid (Lamprosema indicata) [J]. Biotechnology Bulletin, 2025, 41(6): 99-108. |
| [4] | LIU Hua, SONG Jie, ZENG Hai-juan, WANG Jin-bin, QIAN Yun-fang. Research Progress in Single-base Mutation Detection Methods and Applications [J]. Biotechnology Bulletin, 2025, 41(6): 61-70. |
| [5] | GAO Chang, ZHUANG Tian-chi, LI Ning, LIU Yun, GU Peng-fei, ZHAO Xin-yi, JI Ming-hui. Gravity-driven Microfluidic Chip Based on RPA-CRISPR/Cas12a for the Rapid Detection of Mycobacterium tuberculosis [J]. Biotechnology Bulletin, 2025, 41(5): 62-69. |
| [6] | CHEN Mo-yan, ZHU Cheng. Mechanism Study and Application of CRISPR/Cas12a-based Biosensing Platform [J]. Biotechnology Bulletin, 2024, 40(7): 90-98. |
| [7] | ZHU Shao-xi, JIN Zhao-yang, GE Jian-rong, WANG Rui, WANG Feng-ge, LU Yun-cai. High-throughput Specific Detection Methods for Transgenic Maize Based on the KASP Platform [J]. Biotechnology Bulletin, 2023, 39(6): 133-140. |
| [8] | CHEN Xiao-lin, LIU Yang-er, XU Wen-tao, GUO Ming-zhang, LIU Hui-lin. Application of Synthetic Biology Based Whole-cell Biosensor Technology in the Rapid Detection of Food Safety [J]. Biotechnology Bulletin, 2023, 39(1): 137-149. |
| [9] | LI Sheng-yan, LI Xiang-yin, LI Peng-cheng, ZHANG Ming-jun, ZHANG Jie, LANG Zhi-hong. Identification of Target Traits and Genetic Stability of Transgenic Maize 2HVB5 [J]. Biotechnology Bulletin, 2023, 39(1): 21-30. |
| [10] | LI Peng-cheng, ZHANG Ming-jun, WANG Yin-xiao, LI Xiang-yin, LI Sheng-yan, LANG Zhi-hong. Insect Resistance Identification and Agronomy Traits Analysis of Transgenic Maize HGK60 with Different Genetic Backgrounds [J]. Biotechnology Bulletin, 2023, 39(1): 40-47. |
| [11] | LI Jia-le, LIN Sheng-hao, XU Wen-tao. Construction of an Ultra-sensitive Colorimetric Biosensor for Insect Resistance Genes Based on Loop-mediated Isothermal Amplification [J]. Biotechnology Bulletin, 2022, 38(8): 69-76. |
| [12] | FU Zhi-qiang, XIONG Yan. Research Progress on Portable Bio-optical Sensors [J]. Biotechnology Bulletin, 2021, 37(3): 219-226. |
| [13] | LI Xin-shen, HUANG Xiao-mei, WU Shu-xiu, HUANG Rui-rong, WEI Lin-gen, HUA Ju-ling. Rapid Detection of Plant Bacterial Wilt by Loop-mediated Isothermal Amplification [J]. Biotechnology Bulletin, 2021, 37(1): 272-281. |
| [14] | ZHAO Ying, WANG Nan, LU An-xiang, FENG Xiao-yuan, GUO Xiao-jun, LUAN Yun-xia. Application in the Detection of Fungal Toxins by Nucleic Acid Aptamer Lateral Flow Chromatography Analysis Technique [J]. Biotechnology Bulletin, 2020, 36(8): 217-227. |
| [15] | GAO Wei-fang, ZHANG Li-ping, ZHU Peng. Recent Progress on Isothermal Amplification Technology and Its Combination with CRISPR in Rapid Detection of Microorganisms [J]. Biotechnology Bulletin, 2020, 36(5): 22-31. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||