








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 19-36.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1434
Previous Articles Next Articles
LI Cheng-quan1( ), SHI Qing-hua1,2, YANG Xiao-yu1,2(
), SHI Qing-hua1,2, YANG Xiao-yu1,2( )
)
Received:2025-12-28
Online:2026-03-26
Published:2026-04-23
Contact:
YANG Xiao-yu
E-mail:lcq2434115933@163.com;xiaoyu.yang@sdau.edu.cn
LI Cheng-quan, SHI Qing-hua, YANG Xiao-yu. microRNA-based Regulatory Network for Fruit Development of Horticultural Crops: From Molecular Mechanism to Germplasm Innovation[J]. Biotechnology Bulletin, 2026, 42(3): 19-36.
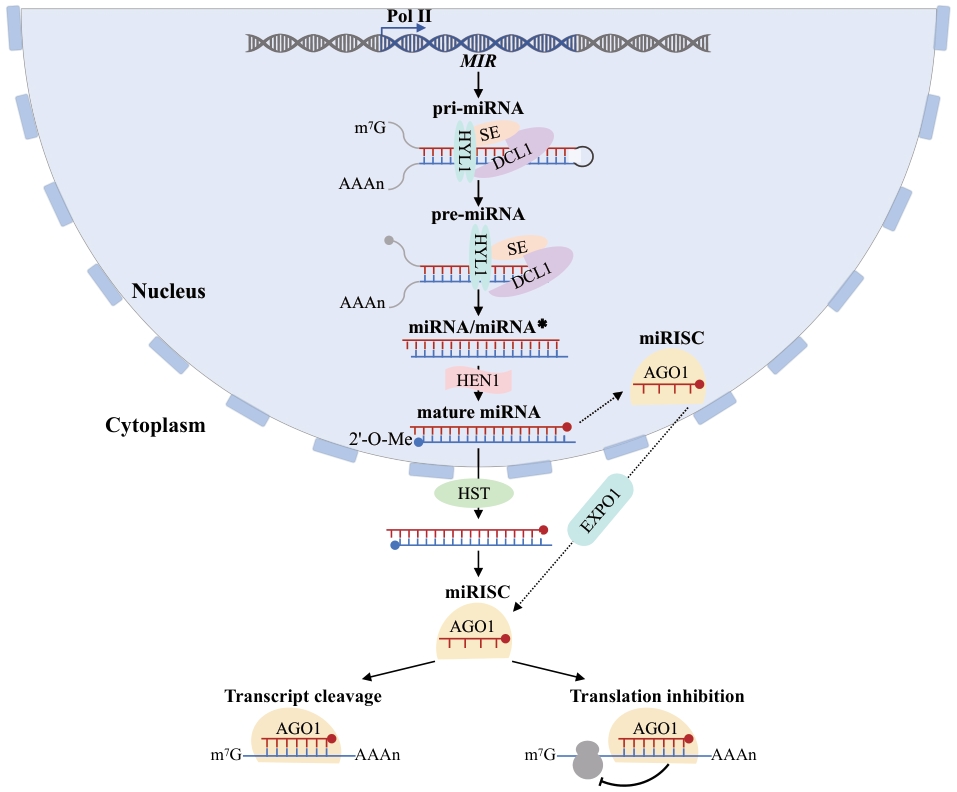
Fig. 1 Synthetic pathways and action modes of microRNAs in plantsPlant miRNAs are transcribed and processed in the nucleus, exported to the cytoplasm, and loaded into miRISC to regulate target gene expression through cleavage or translational inhibition. The schematic flow from top to bottom illustrates this process: MIR denotes the miRNA gene locus, which is transcribed by RNA polymerase II (Pol II) to generate primary miRNAs (pri-miRNAs) featuring a 5' m⁷G cap and a 3' poly(A) tail. Within the nucleus, pri-miRNAs are recognized and cleaved by a processing complex comprising DCL1 (Dicer-like 1), HYL1 (HYPONASTIC LEAVES 1), and SE (SERRATE), yielding precursor miRNAs (pre-miRNAs) and subsequently miRNA/miRNA* duplexes. HEN1 (HUA ENHANCER 1) mediates 2'-O-methylation (2'-O-Me) at the 3' ends of the duplex to enhance stability. The nuclear export factor HASTY (HST) facilitates the translocation of mature miRNA duplexes from the nucleus to the cytoplasm. AGO1 (ARGONAUTE 1) serves as the core effector protein, loading the mature miRNA to form the miRNA-induced silencing complex (miRISC). Dashed arrows indicate an alternative pathway in which miRISC may undergo preliminary assembly within the nucleus prior to nuclear export; solid arrows denote the directionality of molecular processing, transport, and loading. The two branches at the bottom indicate the primary modes of miRNA-mediated regulation, namely transcript cleavage and translational inhibition
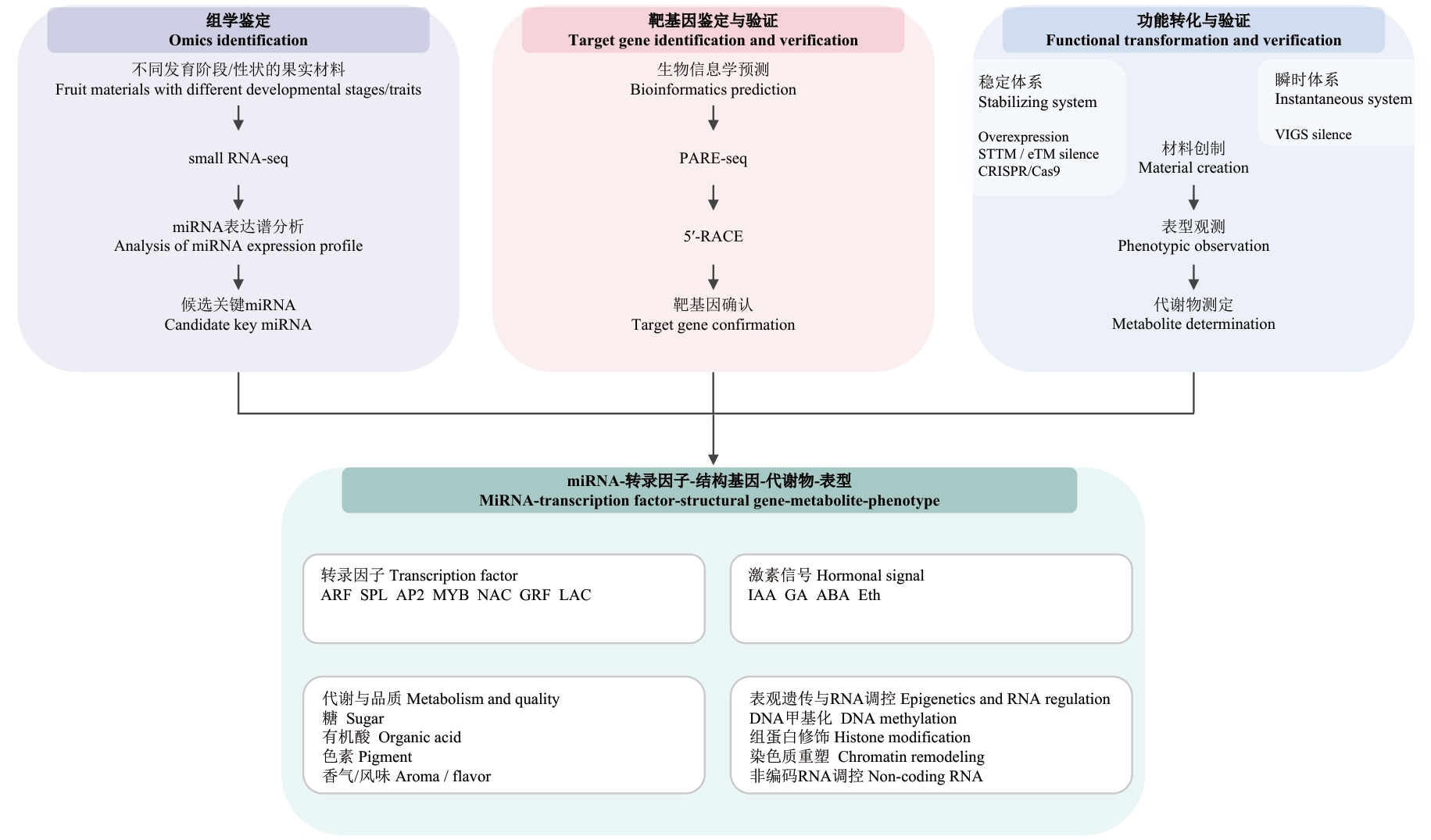
Fig. 2 Research routine of miRNA-mediated fruit trait regulation mechanismThis diagram presents the comprehensive technical outline for fruit miRNA research, progressing from “omics-based identification-target gene validation-functional characterization and translational verification” to the integrated dissection of the “miRNA-transcription factor-structural gene-metabolite-phenotype” regulatory network. The upper panel shows three major stages: Omics-based identification (Small RNA-seq is performed on fruit samples representing distinct developmental stages or phenotypic traits to profile miRNA expression and prioritize candidate miRNAs), target gene identification and validation (Putative targets are predicted via bioinformatics and empirically validated using PARE-seq and 5'-RACE, thereby confirming miRNA-target interactions), and functional characterization and translational validation (Stable or transient functional systems are established through genetic materials, including overexpression, STTM/eTM-mediated silencing, CRISPR/Cas9, and VIGS silencing, and causal relationships are subsequently built by integrating phenotypic observation, target gene validation, and metabolite profiling). The lower panel illustrates the architecture of the “miRNA-transcription factor-structural gene-metabolite-phenotype” regulatory network, incorporating key regulatory layers of transcription factors (e.g., ARF, SPL, AP2, MYB, NAC, GRF, and LAC), hormone signaling (IAA, GA, ABA, and ethylene), metabolic and quality traits (sugars, organic acids, pigments, and aroma/flavor compounds), as well as epigenetic/RNA-based regulatory mechanisms (DNA methylation, histone modification, chromatin remodeling, and non-coding RNA regulation)
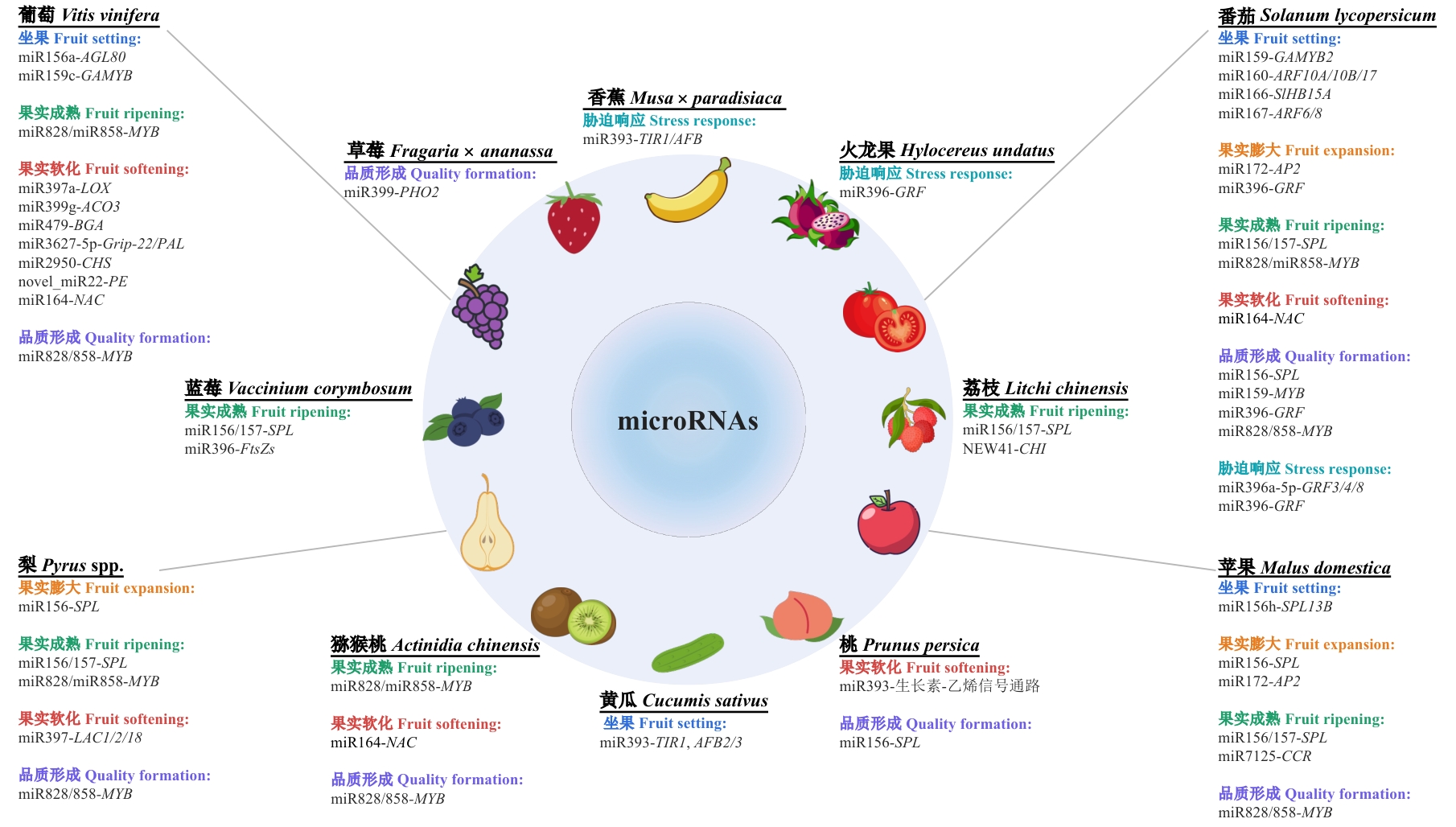
Fig. 3 Schematic diagram of the core miRNA regulatory network underlying key traits during fruit development in horticultural cropsCentered on microRNAs, this integrative diagram illustrates key miRNA-target gene modules and their associated traits across various horticultural crops during fruit development and stress responses. The outer ring features representative crop species, each accompanied by experimentally validated miRNA-target gene (or pathway) regulatory modules. These modules are systematically categorized according to fruit developmental stages and trait categories: fruit setting, fruit expansion, fruit ripening, fruit softening, quality formation, and stress response. Gray connecting lines indicate associations between each crop and its corresponding regulatory modules
| 发育阶段Developmental stage | miRNA | 靶基因 Target gene | 物种 Species | 靶向表型/通路 Target phenotype/pathway | 参考文献Reference |
|---|---|---|---|---|---|
坐果与早期果实形成 Fruit setting and early formation | miR156a | AGL80 | 葡萄 Vitis vinifera | 调控开花、坐果等发育过程 | [ |
| miR156h | SPL13B | 苹果 Malus domestica | 调控赤霉素(GA)代谢/信号、促进单性结实/坐果 | [ | |
| miR159 | GAMYB2 | 番茄 Solanum lycopersicum和拟南芥 Arabidopsis thaliana | GA信号与坐果启动,以及幼果早期形态建成/果形调控 | [ | |
| miR159c | GAMYB | 葡萄 Vitis vinifera | 调控GA-DELLA相关通路、及单性结实/坐果等发育过程 | [ | |
| miR160 | ARF10A/10B/17 | 番茄 Solanum lycopersicum | 调控生长素(IAA)信号、坐果与幼果早期膨大 | [ | |
| miR166 | SlHB15A | 番茄 Solanum lycopersicum | 平衡生长素-乙烯信号、调控温度胁迫下的坐果/单性结实 | [ | |
| miR167 | ARF6/8 | 番茄 Solanum lycopersicum | 花器官发育与坐果/单性结实;影响子房壁细胞伸长 | [ | |
| miR390 | ARF2/3/4 | 拟南芥 Arabidopsis thaliana | 调控TAS3-tasiARF通路及早期果实形成与器官生长 | [ | |
| miR393 | TIR1, AFB2/3 | 拟南芥 Arabidopsis thaliana和 黄瓜 Cucumis sativus | 调控生长素通路、及坐果和结实等过程 | [ | |
果实膨大 Fruit expansion | miR156 | SPL | 苹果 Malus domestica和梨 Pyrus spp. | 调控细胞分裂和膨大及果实大小 | [ |
| miR172 | AP2 | 苹果 Malus domestica、番茄 Solanum lycopersicum 拟南芥 Arabidopsis thaliana | 调控组织分化与膨大及果实大小/形态 | [ | |
| miR396 | GRF | 番茄 Solanum lycopersicum | 调控细胞增殖与器官大小,及果实大小和形状 | [ | |
果实成熟 Fruit ripening | miR156/157 | SPL | 蓝莓 Vaccinium corymbosum、梨 Pyrus spp.、荔枝 Litchi chinensis、番茄 Solanum lycopersicum和苹果 Malus domestica | 调控果实成熟和转色,及花青素/叶绿素代谢 | [ |
| miR396 | FtsZs | 蓝莓 Vaccinium corymbosum | 调控色素体/叶绿体发育,及成熟转色和色素积累 | [ | |
| miR828/miR858 | MYB | 番茄 Solanum lycopersicum、葡萄 Vitis vinifera、梨 Pyrus spp.和猕猴桃 Actinidia chinensis | 调控花青素/黄酮合成通路,影响着色与抗氧化品质 | [ | |
| miR7125 | CCR | 苹果 Malus domestica | 调控木质素-花青素平衡及光诱导着色 | [ | |
| NEW41 | CHI | 荔枝 Litchi chinensis | 调控黄酮/花青素合成结构基因,影响着色 | [ | |
果实软化 Fruit softening | miR393 | 生长素-乙烯信号通路 | 桃 Prunus persica | 调控激素互作引发的软化进程,影响贮藏期 | [ |
| miR397 | LAC1/2/18 | 梨 Pyrus spp. | 调控木质素/细胞壁木质化与硬度,影响质地与货架期 | [ | |
| miR397a | LOX | 葡萄 Vitis vinifera | 调控脂质代谢/膜脂过氧化、软化及与风味相关过程 | [ | |
| miR399g | ACO3 | 葡萄 Vitis vinifera | 调控乙烯生物合成、果实成熟和软化 | [ | |
| miR479 | BGA | 葡萄 Vitis vinifera | 调控β-半乳糖苷酶等细胞壁多糖降解基因,影响软化 | [ | |
| miR3627-5p | Grip-22/PAL | 葡萄 Vitis vinifera | 调控苯丙烷代谢/细胞壁相关基因,影响果实软化进程 | [ | |
| miR2950 | CHS | 葡萄 Vitis vinifera | 调控黄酮合成相关基因及软化伴随的次生代谢途径 | [ | |
| novel_miR22 | PE | 葡萄 Vitis vinifera | 调控果胶酯酶等促进软化 | [ | |
| miR164 | NAC | 番茄 Solanum lycopersicum、葡萄 Vitis vinifera和猕猴桃 Actinidia chinensis | 乙烯合成和响应及色泽、质地相关基因表达调控 | [ | |
品质形成 Quality formation | miR156 | SPL | 桃 Prunus persica和番茄Solanum lycopersicum | 糖酸代谢/品质形成相关通路调控及代谢重塑 | [ |
| miR159 | MYB | 番茄 Solanum lycopersicum | 色素/次生代谢与风味品质相关通路的转录调控 | [ | |
| miR396 | GRF | 番茄 Solanum lycopersicum | 果实大小-品质耦合调控,影响代谢/可溶性固形物等 | [ | |
| miR399 | PHO2 | 草莓 Fragaria × ananassa | 磷稳态‒糖积累耦合影响品质形成 | [ | |
| miR828/858 | MYB | 番茄 Solanum lycopersicum、葡萄 Vitis vinifera、梨 Pyrus spp.、猕猴桃Actinidia chinensis和苹果 Malus domestica | 调控花青素/黄酮合成,影响着色与抗氧化品质 | [ | |
胁迫响应 Stress response | miR393 | TIR1/AFB | 香蕉 Musa × paradisiaca | 逆境下生长素信号调控与适应性响应 | [ |
| miR396a-5p | GRF3/4/8 | 番茄 Solanum lycopersicum | 干旱与高温下生长素信号调控与适应性响应 | [ | |
| miR396 | GRF | 番茄 Solanum lycopersicum和火龙果 Hylocereus undatus | 调控逆境下的生长/细胞增殖和抗逆响应 | [ |
Table 1 Summary of miRNAs and their target genes involved in fruit developmental regulation
| 发育阶段Developmental stage | miRNA | 靶基因 Target gene | 物种 Species | 靶向表型/通路 Target phenotype/pathway | 参考文献Reference |
|---|---|---|---|---|---|
坐果与早期果实形成 Fruit setting and early formation | miR156a | AGL80 | 葡萄 Vitis vinifera | 调控开花、坐果等发育过程 | [ |
| miR156h | SPL13B | 苹果 Malus domestica | 调控赤霉素(GA)代谢/信号、促进单性结实/坐果 | [ | |
| miR159 | GAMYB2 | 番茄 Solanum lycopersicum和拟南芥 Arabidopsis thaliana | GA信号与坐果启动,以及幼果早期形态建成/果形调控 | [ | |
| miR159c | GAMYB | 葡萄 Vitis vinifera | 调控GA-DELLA相关通路、及单性结实/坐果等发育过程 | [ | |
| miR160 | ARF10A/10B/17 | 番茄 Solanum lycopersicum | 调控生长素(IAA)信号、坐果与幼果早期膨大 | [ | |
| miR166 | SlHB15A | 番茄 Solanum lycopersicum | 平衡生长素-乙烯信号、调控温度胁迫下的坐果/单性结实 | [ | |
| miR167 | ARF6/8 | 番茄 Solanum lycopersicum | 花器官发育与坐果/单性结实;影响子房壁细胞伸长 | [ | |
| miR390 | ARF2/3/4 | 拟南芥 Arabidopsis thaliana | 调控TAS3-tasiARF通路及早期果实形成与器官生长 | [ | |
| miR393 | TIR1, AFB2/3 | 拟南芥 Arabidopsis thaliana和 黄瓜 Cucumis sativus | 调控生长素通路、及坐果和结实等过程 | [ | |
果实膨大 Fruit expansion | miR156 | SPL | 苹果 Malus domestica和梨 Pyrus spp. | 调控细胞分裂和膨大及果实大小 | [ |
| miR172 | AP2 | 苹果 Malus domestica、番茄 Solanum lycopersicum 拟南芥 Arabidopsis thaliana | 调控组织分化与膨大及果实大小/形态 | [ | |
| miR396 | GRF | 番茄 Solanum lycopersicum | 调控细胞增殖与器官大小,及果实大小和形状 | [ | |
果实成熟 Fruit ripening | miR156/157 | SPL | 蓝莓 Vaccinium corymbosum、梨 Pyrus spp.、荔枝 Litchi chinensis、番茄 Solanum lycopersicum和苹果 Malus domestica | 调控果实成熟和转色,及花青素/叶绿素代谢 | [ |
| miR396 | FtsZs | 蓝莓 Vaccinium corymbosum | 调控色素体/叶绿体发育,及成熟转色和色素积累 | [ | |
| miR828/miR858 | MYB | 番茄 Solanum lycopersicum、葡萄 Vitis vinifera、梨 Pyrus spp.和猕猴桃 Actinidia chinensis | 调控花青素/黄酮合成通路,影响着色与抗氧化品质 | [ | |
| miR7125 | CCR | 苹果 Malus domestica | 调控木质素-花青素平衡及光诱导着色 | [ | |
| NEW41 | CHI | 荔枝 Litchi chinensis | 调控黄酮/花青素合成结构基因,影响着色 | [ | |
果实软化 Fruit softening | miR393 | 生长素-乙烯信号通路 | 桃 Prunus persica | 调控激素互作引发的软化进程,影响贮藏期 | [ |
| miR397 | LAC1/2/18 | 梨 Pyrus spp. | 调控木质素/细胞壁木质化与硬度,影响质地与货架期 | [ | |
| miR397a | LOX | 葡萄 Vitis vinifera | 调控脂质代谢/膜脂过氧化、软化及与风味相关过程 | [ | |
| miR399g | ACO3 | 葡萄 Vitis vinifera | 调控乙烯生物合成、果实成熟和软化 | [ | |
| miR479 | BGA | 葡萄 Vitis vinifera | 调控β-半乳糖苷酶等细胞壁多糖降解基因,影响软化 | [ | |
| miR3627-5p | Grip-22/PAL | 葡萄 Vitis vinifera | 调控苯丙烷代谢/细胞壁相关基因,影响果实软化进程 | [ | |
| miR2950 | CHS | 葡萄 Vitis vinifera | 调控黄酮合成相关基因及软化伴随的次生代谢途径 | [ | |
| novel_miR22 | PE | 葡萄 Vitis vinifera | 调控果胶酯酶等促进软化 | [ | |
| miR164 | NAC | 番茄 Solanum lycopersicum、葡萄 Vitis vinifera和猕猴桃 Actinidia chinensis | 乙烯合成和响应及色泽、质地相关基因表达调控 | [ | |
品质形成 Quality formation | miR156 | SPL | 桃 Prunus persica和番茄Solanum lycopersicum | 糖酸代谢/品质形成相关通路调控及代谢重塑 | [ |
| miR159 | MYB | 番茄 Solanum lycopersicum | 色素/次生代谢与风味品质相关通路的转录调控 | [ | |
| miR396 | GRF | 番茄 Solanum lycopersicum | 果实大小-品质耦合调控,影响代谢/可溶性固形物等 | [ | |
| miR399 | PHO2 | 草莓 Fragaria × ananassa | 磷稳态‒糖积累耦合影响品质形成 | [ | |
| miR828/858 | MYB | 番茄 Solanum lycopersicum、葡萄 Vitis vinifera、梨 Pyrus spp.、猕猴桃Actinidia chinensis和苹果 Malus domestica | 调控花青素/黄酮合成,影响着色与抗氧化品质 | [ | |
胁迫响应 Stress response | miR393 | TIR1/AFB | 香蕉 Musa × paradisiaca | 逆境下生长素信号调控与适应性响应 | [ |
| miR396a-5p | GRF3/4/8 | 番茄 Solanum lycopersicum | 干旱与高温下生长素信号调控与适应性响应 | [ | |
| miR396 | GRF | 番茄 Solanum lycopersicum和火龙果 Hylocereus undatus | 调控逆境下的生长/细胞增殖和抗逆响应 | [ |
| [79] | Qian MJ, Ni JB, Niu QF, et al. Response of miR156-SPL module during the red peel coloration of bagging-treated Chinese sand pear (Pyrus pyrifolia Nakai) [J]. Front Physiol, 2017, 8: 550. |
| [80] | Liu R, Lai B, Hu B, et al. Identification of microRNAs and their target genes related to the accumulation of anthocyanins in Litchi chinensis by high-throughput sequencing and degradome analysis [J]. Front Plant Sci, 2017, 7: 2059. |
| [81] | Xia R, Zhu H, An YQ, et al. Apple miRNAs and tasiRNAs with novel regulatory networks [J]. Genome Biol, 2012, 13(6): R47. |
| [82] | Pantaleo V, Szittya G, Moxon S, et al. Identification of grapevine microRNAs and their targets using high-throughput sequencing and degradome analysis: grapevine microRNAs and their targets [J]. Plant J, 2010, 62(6): 960-976. |
| [83] | Guo DL, Li Q, Lv WQ, et al. microRNA profiling analysis of developing berries for ‘Kyoho’ and its early-ripening mutant during berry ripening [J]. BMC Plant Biol, 2018, 18: 285. |
| [84] | Sunitha S, Loyola R, Alcalde JA, et al. The role of UV-B light on small RNA activity during grapevine berry development [J]. G3, 2019, 9(3): 769-787. |
| [85] | Li XB, Hong Y, Jackson A, et al. Dynamic regulation of small RNAs in anthocyanin accumulation during blueberry fruit maturation [J]. Sci Rep, 2021, 11: 15080. |
| [86] | Hou YM, Zhai LL, Li XY, et al. Comparative analysis of fruit ripening-related miRNAs and their targets in blueberry using small RNA and degradome sequencing [J]. Int J Mol Sci, 2017, 18(12): 2767. |
| [87] | Hu YJ, Cheng H, Zhang Y, et al. The MdMYB16/MdMYB1-miR7125-MdCCR module regulates the homeostasis between anthocyanin and lignin biosynthesis during light induction in apple [J]. New Phytol, 2021, 231(3): 1105-1122. |
| [88] | Xue C, Yao JL, Qin MF, et al. PbrmiR397a regulates lignification during stone cell development in pear fruit [J]. Plant Biotechnol J, 2019, 17(1): 103-117. |
| [89] | Xue MZ, Yi HL, Wang H. Identification of miRNAs involved in SO2 preservation in Vitis vinifera L. by deep sequencing [J]. Environ Exp Bot, 2018, 153: 218-228. |
| [90] | Alexander L, Grierson D. Ethylene biosynthesis and action in tomato: a model for climacteric fruit ripening [J]. J Exp Bot, 2002, 53(377): 2039-2055. |
| [91] | Li S, Chen KS, Grierson D. A critical evaluation of the role of ethylene and MADS transcription factors in the network controlling fleshy fruit ripening [J]. New Phytol, 2019, 221(4): 1724-1741. |
| [92] | Chung MY, Nath UK, Vrebalov J, et al. Ectopic expression of miRNA172 in tomato (Solanum lycopersicum) reveals novel function in fruit development through regulation of an AP2 transcription factor [J]. BMC Plant Biol, 2020, 20: 283. |
| [93] | Liu MC, Pirrello J, Chervin C, et al. Ethylene control of fruit ripening: revisiting the complex network of transcriptional regulation [J]. Plant Physiol, 2015, 169(4): 2380-2390. |
| [94] | Moxon S, Jing RC, Szittya G, et al. Deep sequencing of tomato short RNAs identifies microRNAs targeting genes involved in fruit ripening [J]. Genome Res, 2008, 18(10): 1602-1609. |
| [95] | Zuo JH, Zhu BZ, Fu DQ, et al. Sculpting the maturation, softening and ethylene pathway: the influences of microRNAs on tomato fruits [J]. BMC Genom, 2012, 13: 7. |
| [96] | Lin DB, Zhu XE, Qi BL, et al. SlMIR164A regulates fruit ripening and quality by controlling SlNAM2 and SlNAM3 in tomato [J]. Plant Biotechnol J, 2022, 20(8): 1456-1469. |
| [97] | Wang ZQ, Wang Q, Duan WH, et al. Overexpression of Sly-miR164a in tomato decreases expression of NAC and delays pre- and postharvest ripening of fruit [J]. Postharvest Biol Technol, 2023, 195: 112132. |
| [98] | Wang WQ, Wang J, Wu YY, et al. Genome-wide analysis of coding and non-coding RNA reveals a conserved miR164-NAC regulatory pathway for fruit ripening [J]. New Phytol, 2020, 225(4): 1618-1634. |
| [99] | Shi MY, Hu X, Wei Y, et al. Genome-wide profiling of small RNAs and degradome revealed conserved regulations of miRNAs on auxin-responsive genes during fruit enlargement in peaches [J]. Int J Mol Sci, 2017, 18(12): 2599. |
| [100] | Zhang CH, Zhang BB, Ma RJ, et al. Identification of known and novel microRNAs and their targets in peach (Prunus persica) fruit by high-throughput sequencing [J]. PLoS One, 2016, 11(7): e0159253. |
| [101] | Zhang B, Huang ZY, Wang ZD, et al. Jasmonate-responsive MdMYC2/MdMED25 complex regulates malic acid accumulation in apples through the miR858-MdMYB73 module [J]. J Integr Plant Biol, 2026, 68(1): 148-168. |
| [102] | Silva GFFE, Silva EM, da Silva Azevedo M, et al. microRNA156-targeted SPL/SBP box transcription factors regulate tomato ovary and fruit development [J]. Plant J, 2014, 78(4): 604-618. |
| [103] | Gou JY, Felippes FF, Liu CJ, et al. Negative regulation of anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor [J]. Plant Cell, 2011, 23(4): 1512-1522. |
| [104] | Luo YJ, Zhang XN, Luo ZR, et al. Identification and characterization of microRNAs from Chinese pollination constant non-astringent persimmon using high-throughput sequencing [J]. BMC Plant Biol, 2015, 15: 11. |
| [105] | Wang Y, Zhang JX, Cui WX, et al. Improvement in fruit quality by overexpressing miR399a in woodland strawberry [J]. J Agric Food Chem, 2017, 65(34): 7361-7370. |
| [106] | Zhou H, Ren F, Wang X, et al. Genome-wide identification and characterization of long noncoding RNAs during peach (Prunus persica) fruit development and ripening [J]. Sci Rep, 2022, 12: 11044. |
| [107] | Zhang B, Yang HJ, Yang YZ, et al. Mdm-miR828 participates in the feedback loop to regulate anthocyanin accumulation in apple peel [J]. Front Plant Sci, 2020, 11: 608109. |
| [108] | Ali Khan N, Owens L, Nuñez MA, et al. Complexity of combined abiotic stresses to crop plants [J]. Plant Stress, 2025, 17: 100926. |
| [109] | Nejat N, Mantri N. Emerging roles of long non-coding RNAs in plant response to biotic and abiotic stresses [J]. Crit Rev Biotechnol, 2018, 38(1): 93-105. |
| [110] | Sunkar R. microRNAs with macro-effects on plant stress responses [J]. Semin Cell Dev Biol, 2010, 21(8): 805-811. |
| [111] | Zhu H, Zhang Y, Tang RF, et al. Banana sRNAome and degradome identify microRNAs functioning in differential responses to temperature stress [J]. BMC Genom, 2019, 20: 33. |
| [112] | Zhou R, Yu XQ, Ottosen CO, et al. Unique miRNAs and their targets in tomato leaf responding to combined drought and heat stress [J]. BMC Plant Biol, 2020, 20: 107. |
| [113] | Cai XW, Zhang LF, Xiao L, et al. Genome-wide identification of GRF gene family and their contribution to abiotic stress response in pitaya (Hylocereus polyrhizus) [J]. Int J Biol Macromol, 2022, 223: 618-635. |
| [114] | Tang YY, Du GN, Xiang J, et al. Genome-wide identification of auxin response factor (ARF) gene family and the miR160-ARF18-mediated response to salt stress in peanut (Arachis hypogaea L.) [J]. Genomics, 2022, 114(1): 171-184. |
| [1] | Slavin JL, Lloyd B. Health benefits of fruits and vegetables [J]. Adv Nutr, 2012, 3(4): 506-516. |
| [2] | Yang CP, Ying SY, Tang BB, et al. The mechanistic insights into fruit ripening: integrating phytohormones, transcription factors, and epigenetic modification [J]. J Genet Genom, 2025, 52(12): 1475-1489. |
| [3] | Giovannoni J, Nguyen C, Ampofo B, et al. The epigenome and transcriptional dynamics of fruit ripening [J]. Annu Rev Plant Biol, 2017, 68: 61-84. |
| [4] | Gapper NE, Giovannoni JJ, Watkins CB. Understanding development and ripening of fruit crops in an ‘omics’ era [J]. Hortic Res, 2014, 1: 14034. |
| [5] | Baral R, Vainer A, Melzer S, et al. ‘Bud to fruit’—hormonal interactions governing early fruit development [J]. J Exp Bot, 2025, 76(22): 6657-6673. |
| [6] | He H, Yamamuro C. Interplays between auxin and GA signaling coordinate early fruit development [J]. Hortic Res, 2022, 9: uhab078. |
| [7] | Kumar R, Khurana A, Sharma AK. Role of plant hormones and their interplay in development and ripening of fleshy fruits [J]. J Exp Bot, 2014, 65(16): 4561-4575. |
| [8] | Allan AC, Hellens RP, Laing WA. MYB transcription factors that colour our fruit [J]. Trends Plant Sci, 2008, 13(3): 99-102. |
| [9] | Espley RV, Hellens RP, Putterill J, et al. Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10 [J]. Plant J, 2007, 49(3): 414-427. |
| [10] | Kobayashi S, Goto-Yamamoto N, Hirochika H. Retrotransposon-induced mutations in grape skin color [J]. Science, 2004, 304(5673): 982. |
| [11] | Yu Y, Wang H, You CJ, et al. Plant microRNA maturation and function [J]. Nat Rev Mol Cell Biol, 2026, 27(1): 55-70. |
| [12] | Ayubov MS, Mirzakhmedov MH, Sripathi VR, et al. Role of microRNAs and small RNAs in regulation of developmental processes and agronomic traits in Gossypium species [J]. Genomics, 2019, 111(5): 1018-1025. |
| [13] | Song XW, Li Y, Cao XF, et al. microRNAs and their regulatory roles in plant-environment interactions [J]. Annu Rev Plant Biol, 2019, 70: 489-525. |
| [14] | Brant EJ, Budak H. Plant small non-coding RNAs and their roles in biotic stresses [J]. Front Plant Sci, 2018, 9: 1038. |
| [15] | Feng SJ, Xu YM, Guo CK, et al. Modulation of miR156 to identify traits associated with vegetative phase change in tobacco (Nicotiana tabacum) [J]. J Exp Bot, 2016, 67(5): 1493-1504. |
| [16] | Shriram V, Kumar V, Devarumath RM, et al. microRNAs as potential targets for abiotic stress tolerance in plants [J]. Front Plant Sci, 2016, 7: 817. |
| [17] | Budak H, Kantar M, Bulut R, et al. Stress responsive miRNAs and isomiRs in cereals [J]. Plant Sci, 2015, 235: 1-13. |
| [18] | Wu G, Park MY, Conway SR, et al. The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis [J]. Cell, 2009, 138(4): 750-759. |
| [19] | Karlova R, van Haarst JC, Maliepaard C, et al. Identification of microRNA targets in tomato fruit development using high-throughput sequencing and degradome analysis [J]. J Exp Bot, 2013, 64(7): 1863-1878. |
| [20] | Chen CJ, Zeng ZH, Liu ZR, et al. Small RNAs, emerging regulators critical for the development of horticultural traits [J]. Hortic Res, 2018, 5: 63. |
| [21] | Samadi SM, Asefpour Vakilian K, Javidan SM. A review of the most important microRNAs involved in fruit postharvest properties [J]. Food Chem Adv, 2025, 9: 101168. |
| [22] | Cedillo-Jimenez CA, Feregrino-Perez AA, Guevara-González RG, et al. microRNA regulation during the tomato fruit development and ripening: a review [J]. Sci Hortic, 2020, 270: 109435. |
| [23] | Rubio-Somoza I, Weigel D. microRNA networks and developmental plasticity in plants [J]. Trends Plant Sci, 2011, 16(5): 258-264. |
| [24] | Gao C, Ju Z, Cao DY, et al. microRNA profiling analysis throughout tomato fruit development and ripening reveals potential regulatory role of RIN on microRNAs accumulation [J]. Plant Biotechnol J, 2015, 13(3): 370-382. |
| [25] | Chen WW, Kong JH, Lai TF, et al. Tuning LeSPL-CNR expression by SlymiR157 affects tomato fruit ripening [J]. Sci Rep, 2015, 5: 7852. |
| [115] | Itaya A, Bundschuh R, Archual AJ, et al. Small RNAs in tomato fruit and leaf development [J]. Biochim Biophys Acta Gene Regul Mech, 2008, 1779(2): 99-107. |
| [116] | Song CN, Fang JG, Li XY, et al. Identification and characterization of 27 conserved microRNAs in citrus [J]. Planta, 2009, 230(4): 671-685. |
| [117] | Islam W, Tauqeer A, Waheed A, et al. microRNA mediated plant responses to nutrient stress [J]. Int J Mol Sci, 2022, 23(5): 2562. |
| [118] | Kuo HF, Chiou TJ. The role of microRNAs in phosphorus deficiency signaling [J]. Plant Physiol, 2011, 156(3): 1016-1024. |
| [119] | Kant S, Peng MS, Rothstein SJ. Genetic regulation by NLA and microRNA827 for maintaining nitrate-dependent phosphate homeostasis in Arabidopsis [J]. PLoS Genet, 2011, 7(3): e1002021. |
| [120] | Lin SI, Chiang SF, Lin WY, et al. Regulatory network of microRNA399 and PHO2 by systemic signaling [J]. Plant Physiol, 2008, 147(2): 732-746. |
| [121] | Bari R, Datt Pant B, Stitt M, et al. PHO2, microRNA399, and PHR1 define a phosphate-signaling pathway in plants [J]. Plant Physiol, 2006, 141(3): 988-999. |
| [122] | Lu YB, Yang LT, Qi YP, et al. Identification of boron-deficiency-responsive microRNAs in Citrus sinensis roots by Illumina sequencing [J]. BMC Plant Biol, 2014, 14: 123. |
| [123] | Lu YB, Qi YP, Yang LT, et al. Boron-deficiency-responsive microRNAs and their targets in Citrus sinensis leaves [J]. BMC Plant Biol, 2015, 15: 271. |
| [124] | Pei MS, Liu HN, Wei TL, et al. Identification, characterization, and verification of miR399 target gene in grape [J]. Hortic Plant J, 2024, 10(1): 91-102. |
| [125] | Liu X, Tan CC, Cheng X, et al. miR168 targets Argonaute1A mediated miRNAs regulation pathways in response to potassium deficiency stress in tomato [J]. BMC Plant Biol, 2020, 20: 477. |
| [126] | Liu X, Pei LL, Zhang LL, et al. Regulation of miR319b-targeted SlTCP10 during the tomato response to low-potassium stress [J]. Int J Mol Sci, 2023, 24(8): 7058. |
| [127] | Jahed KR, Hirst PM. Fruit growth and development in apple: a molecular, genomics and epigenetics perspective [J]. Front Plant Sci, 2023, 14: 1122397. |
| [26] | Tirumalai V, Swetha C, Nair A, et al. miR828 and miR858 regulate VvMYB114 to promote anthocyanin and flavonol accumulation in grapes [J]. J Exp Bot, 2019, 70(18): 4775-4792. |
| [27] | Pantaleo V, Vitali M, Boccacci P, et al. Novel functional microRNAs from virus-free and infected Vitis vinifera plants under water stress [J]. Sci Rep, 2016, 6: 20167. |
| [28] | Bologna NG, Iselin R, Abriata LA, et al. Nucleo-cytosolic shuttling of ARGONAUTE1 prompts a revised model of the plant microRNA pathway [J]. Mol Cell, 2018, 69(4): 709-719.e5. |
| [29] | Lobbes D, Rallapalli G, Schmidt DD, et al. SERRATE: a new player on the plant microRNA scene [J]. EMBO Rep, 2006, 7(10): 1052-1058. |
| [30] | Baumberger N, Baulcombe DC. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs [J]. Proc Natl Acad Sci U S A, 2005, 102(33): 11928-11933. |
| [31] | Xie ZX, Allen E, Fahlgren N, et al. Expression of Arabidopsis MIRNA genes [J]. Plant Physiol, 2005, 138(4): 2145-2154. |
| [32] | Han MH, Goud S, Song L, et al. The Arabidopsis double-stranded RNA-binding protein HYL1 plays a role in microRNA-mediated gene regulation [J]. Proc Natl Acad Sci U S A, 2004, 101(4): 1093-1098. |
| [33] | Deng PC, Muhammad S, Cao M, et al. Biogenesis and regulatory hierarchy of phased small interfering RNAs in plants [J]. Plant Biotechnol J, 2018, 16(5): 965-975. |
| [34] | Fei Q, Xia R, Meyers BC. Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks [J]. Plant Cell, 2013, 25(7): 2400-2415. |
| [35] | Zhang CQ, Ng DW, Lu J, et al. Roles of target site location and sequence complementarity in trans-acting siRNA formation in Arabidopsis [J]. Plant J, 2012, 69(2): 217-226. |
| [36] | Montgomery TA, Howell MD, Cuperus JT, et al. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation [J]. Cell, 2008, 133(1): 128-141. |
| [37] | Allen E, Xie ZX, Gustafson AM, et al. microRNA-directed phasing during trans-acting siRNA biogenesis in plants [J]. Cell, 2005, 121(2): 207-221. |
| [38] | Yoshikawa M, Peragine A, Park MY, et al. A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis [J]. Genes Dev, 2005, 19(18): 2164-2175. |
| [39] | Peragine A, Yoshikawa M, Wu G, et al. SGS3 and SGS2/SDE1/RDR6 are required for juvenile development and the production of trans-acting siRNAs in Arabidopsis [J]. Genes Dev, 2004, 18(19): 2368-2379. |
| [40] | Vazquez F, Vaucheret H, Rajagopalan R, et al. Endogenous trans-acting siRNAs regulate the accumulation of Arabidopsis mRNAs [J]. Mol Cell, 2004, 16(1): 69-79. |
| [41] | Zhao XM, Li FJ, Ali M, et al. Emerging roles and mechanisms of lncRNAs in fruit and vegetables [J]. Hortic Res, 2024, 11(4): uhae046. |
| [42] | de Oliveira Correa JP, Silva EM, Nogueira FTS. Molecular control by non-coding RNAs during fruit development: from gynoecium patterning to fruit ripening [J]. Front Plant Sci, 2018, 9: 1760. |
| [43] | Zaman F, Zhang M, Wu R, et al. Recent research advances of small regulatory RNA in fruit crops [J]. Horticulturae, 2023, 9(3): 294. |
| [44] | Chen CJ, Li JW, Feng JT, et al. sRNAanno—a database repository of uniformly annotated small RNAs in plants [J]. Hortic Res, 2021, 8: 45. |
| [45] | Kozomara A, Birgaoanu M, Griffiths-Jones S. miRBase: from microRNA sequences to function [J]. Nucleic Acids Res, 2019, 47(D1): D155-D162. |
| [46] | Kozomara A, Griffiths-Jones S. miRBase: annotating high confidence microRNAs using deep sequencing data [J]. Nucleic Acids Res, 2014, 42(D1): D68-D73. |
| [47] | Kozomara A, Griffiths-Jones S. miRBase: integrating microRNA annotation and deep-sequencing data [J]. Nucleic Acids Res, 2011, 39(): D152-D157. |
| [48] | Griffiths-Jones S, Saini HK, van Dongen S, et al. miRBase: tools for microRNA genomics [J]. Nucleic Acids Res, 2008, 36(): D154-D158. |
| [49] | Griffiths-Jones S. miRBase: microRNA sequences, targets and gene nomenclature [J]. Nucleic Acids Res, 2006, 34(90001): D140-D144. |
| [50] | Griffiths‐Jones S. The microRNA registry [J]. Nucleic Acids Res, 2004, 32(): D109-D111. |
| [51] | Wu JX, Zheng SS, Feng GZ, et al. Comparative analysis of miRNAs and their target transcripts between a spontaneous late-ripening sweet orange mutant and its wild-type using small RNA and degradome sequencing [J]. Front Plant Sci, 2016, 7: 1416. |
| [52] | Llave C, Xie ZX, Kasschau KD, et al. Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA [J]. Science, 2002, 297(5589): 2053-2056. |
| [53] | Ma L, Zhao YJ, Chen MJ, et al. The microRNA ppe-miR393 mediates auxin-induced peach fruit softening by promoting ethylene production [J]. Plant Physiol, 2023, 192(2): 1638-1655. |
| [128] | Othman SMIS, Mustaffa AF, Che-Othman MH, et al. Overview of repressive miRNA regulation by short tandem target mimic (STTM): applications and impact on plant biology [J]. Plants, 2023, 12(3): 669. |
| [129] | Shinozaki Y, Nicolas P, Fernandez-Pozo N, et al. High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening [J]. Nat Commun, 2018, 9: 364. |
| [130] | Ormancey M, Guillotin B, San Clemente H, et al. Use of microRNA-encoded peptides to improve agronomic traits [J]. Plant Biotechnol J, 2021, 19(9): 1687-1689. |
| [131] | Lauressergues D, Ormancey M, Guillotin B, et al. Characterization of plant microRNA-encoded peptides (miPEPs) reveals molecular mechanisms from the translation to activity and specificity [J]. Cell Rep, 2022, 38(6): 110339. |
| [132] | Yang RL, Zhang HM. Application of artificial intelligence technology in plant microRNA research: progress, challenges, and prospects [J]. Int J Mol Sci, 2025, 26(24): 11854. |
| [133] | Cheng Q, Wang XF. Machine learning for AI breeding in plants [J]. Genom Proteom Bioinform, 2024, 22(4): qzae051. |
| [54] | Damodharan S, Zhao DZ, Arazi T. A common miRNA160-based mechanism regulates ovary patterning, floral organ abscission and lamina outgrowth in tomato [J]. Plant J, 2016, 86(6): 458-471. |
| [55] | Liu N, Wu S, Van Houten J, et al. Down-regulation of auxin response factors 6 and 8 by microRNA167 leads to floral development defects and female sterility in tomato [J]. J Exp Bot, 2014, 65(9): 2507-2520. |
| [56] | Goetz M, Hooper LC, Johnson SD, et al. Expression of aberrant forms of AUXIN RESPONSE FACTOR8 stimulates parthenocarpy in Arabidopsis and tomato [J]. Plant Physiol, 2007, 145(2): 351-366. |
| [57] | Vidal EA, Araus V, Lu C, et al. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana [J]. Proc Natl Acad Sci U S A, 2010, 107(9): 4477-4482. |
| [58] | Xu J, Li J, Cui L, et al. New insights into the roles of cucumber TIR1 homologs and miR393 in regulating fruit/seed set development and leaf morphogenesis [J]. BMC Plant Biol, 2017, 17: 130. |
| [59] | Sarkar Das S, Yadav S, Singh A, et al. Expression dynamics of miRNAs and their targets in seed germination conditions reveals miRNA-ta-siRNA crosstalk as regulator of seed germination [J]. Sci Rep, 2018, 8: 1233. |
| [60] | Clepet C, Devani RS, Boumlik R, et al. The miR166-SlHB15A regulatory module controls ovule development and parthenocarpic fruit set under adverse temperatures in tomato [J]. Mol Plant, 2021, 14(7): 1185-1198. |
| [61] | da Silva EM, Silva GFFE, Bidoia DB, et al. microRNA159-targeted SlGAMYB transcription factors are required for fruit set in tomato [J]. Plant J, 2017, 92(1): 95-109. |
| [62] | Allen RS, Li JY, Stahle MI, et al. Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family [J]. Proc Natl Acad Sci U S A, 2007, 104(41): 16371-16376. |
| [63] | Zhao PP, Wang FP, Deng YJ, et al. Sly-miR159 regulates fruit morphology by modulating GA biosynthesis in tomato [J]. Plant Biotechnol J, 2022, 20(5): 833-845. |
| [64] | Wang C, Jogaiah S, Zhang WY, et al. Spatio-temporal expression of miRNA159 family members and their GAMYB target gene during the modulation of gibberellin-induced grapevine parthenocarpy [J]. J Exp Bot, 2018, 69(15): 3639-3650. |
| [65] | Su ZW, Xuan XX, Sheng ZL, et al. Characterization and regulatory mechanism analysis of VvmiR156a-VvAGL80 pair during grapevine flowering and parthenocarpy process induced by gibberellin [J]. Plant Genome, 2022, 15: e20181. |
| [66] | Wang DR, Zhang CL, Wang X, et al. microRNA156-SPL13B module induces parthenocarpy through the gibberellin pathway [J]. Plant Biotechnol J, 2025, 23(12): 5536-5549. |
| [67] | Cao DY, Wang J, Ju Z, et al. Regulations on growth and development in tomato Cotyledon, flower and fruit via destruction of miR396 with short tandem target mimic [J]. Plant Sci, 2016, 247: 1-12. |
| [68] | Peng T, Qiao MM, Liu HP, et al. A resource for inactivation of microRNAs using short tandem target mimic technology in model and crop plants [J]. Mol Plant, 2018, 11(11): 1400-1417. |
| [69] | Khatun K, Robin AHK, Park JI, et al. Molecular characterization and expression profiling of tomato GRF transcription factor family genes in response to abiotic stresses and phytohormones [J]. Int J Mol Sci, 2017, 18(5): 1056. |
| [70] | Rodriguez RE, Mecchia MA, Debernardi JM, et al. Control of cell proliferation in Arabidopsis thaliana by microRNA miR396 [J]. Development, 2010, 137(1): 103-112. |
| [71] | Wang MM, Li TX, Wu Y, et al. Genome-wide identification of microRNAs involved in the regulation of fruit ripening in apple (Malus domestica) [J]. Sci Hortic, 2021, 289: 110416. |
| [72] | Yao JL, Tomes S, Xu J, et al. How microRNA172 affects fruit growth in different species is dependent on fruit type [J]. Plant Signal Behav, 2016, 11(4): e1156833. |
| [73] | Yao JL, Xu J, Cornille A, et al. A microRNAallele that emerged prior to apple domestication may underlie fruit size evolution [J]. Plant J, 2015, 84(2): 417-427. |
| [74] | Xia B, Feng JQ, Wang YR, et al. Silencing microRNA172 delays fruit ripening in apple (Malus domestica) through an AP2-ERF transcriptional regulation mechanism [J]. Plant Sci, 2025, 360: 112719. |
| [75] | José Ripoll J, Bailey LJ, Mai QA, et al. microRNA regulation of fruit growth [J]. Nat Plants, 2015, 1(4): 15036. |
| [76] | Li XY, Hou YM, Xie X, et al. A blueberry MIR156a-SPL12 module coordinates the accumulation of chlorophylls and anthocyanins during fruit ripening [J]. J Exp Bot, 2020, 71(19): 5976-5989. |
| [77] | Jia XY, Shen J, Liu H, et al. Small tandem target mimic-mediated blockage of microRNA858 induces anthocyanin accumulation in tomato [J]. Planta, 2015, 242(1): 283-293. |
| [78] | Li YK, Cui W, Qi XJ, et al. microRNA858 negatively regulates anthocyanin biosynthesis by repressing AaMYBC1 expression in kiwifruit (Actinidia arguta) [J]. Plant Sci, 2020, 296: 110476. |
| [1] | LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development [J]. Biotechnology Bulletin, 2026, 42(3): 362-373. |
| [2] | LUO Wei, GONG Ao, ZHONG Yang, HU Di, ZHOU Hong-yuan, ZHANG Hong-xin, AI Ju, LUO You-wei, GAO Dong-li. Pleiotropic Effects of SEPALLATA2 Knock-out on Fruit and Wart Development in Cucumber [J]. Biotechnology Bulletin, 2026, 42(3): 283-293. |
| [3] | CHENG Yun-xia, ZHANG Jun-hong, YE Jie. Advances in the Genetic Regulation of Soluble Solid Accumulation in Tomato Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 145-155. |
| [4] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [5] | HU Qiu-ling, CHEN Ling, HUANG Jia-yi, ZHAO Zi-qiao, PAN Lu-yi, LIU Hui-li, LIU Tai-bo. Advances in the Regulation of Fruit Development by Polyamines [J]. Biotechnology Bulletin, 2026, 42(3): 203-212. |
| [6] | MA Shi-jie, LI Zheng, LI Wei, GUO Yang-dong, ZHANG Na. Research Progress in Light Signaling Regulation of Fruit Development in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 5-18. |
| [7] | CUI Zhi-han, WEI Qing-zhen, HU Na, BAO Chong-lai, WANG Hua-sen. Research Progress in IQD Genes in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 230-241. |
| [8] | ZHANG Gao-xiang, WU Yu-bi, GUO Ya-jing, JI Wei, YANG Zhong-yi. Identification and Expression Analysis of WD40 Gene Family in Grape [J]. Biotechnology Bulletin, 2026, 42(3): 324-337. |
| [9] | JIANG Zhe-hui, WANG Xiao-long, WANG Shou-chuang, ZHOU Ke. Advances in the Elucidation of Metabolic Pathways and Molecular Breeding for Tomato Flavor [J]. Biotechnology Bulletin, 2026, 42(3): 60-78. |
| [10] | YIN Yue, QIN Xiao-ya, MI Jia, AN Wei, HE Jun, ZHANG Feng-feng. Identification of FBN Gene Family and Its Relationship with Carotenoids Metabolism in Lyciumbarbarum [J]. Biotechnology Bulletin, 2026, 42(3): 338-348. |
| [11] | LIU Jia-li, SONG Jing-rong, ZHAO Wen-yu, ZHANG Xin-yuan, ZHAO Zi-yang, CAO Yi-bo, ZHANG Ling-yun. Identification of the R2R3-MYB Gene and Expression Analysis of Flavonoid Regulatory Genes in Blueberry [J]. Biotechnology Bulletin, 2025, 41(9): 124-138. |
| [12] | HOU Ying-xiang, FEI Si-tian, LI Ni, LI Lan, SONG Song-quan, WANG Wei-ping, ZHANG Chao. Research Progress in Response of Rice miRNAs to Biotic Stress [J]. Biotechnology Bulletin, 2025, 41(7): 69-80. |
| [13] | WANG Cong-huan, WU Guo-qiang, WEI Ming. Functional Mechanism of Plant CBL in Regulating the Responses to Abiotic and Biotic Stresses [J]. Biotechnology Bulletin, 2025, 41(7): 1-16. |
| [14] | ZHANG Xue-qiong, PAN Su-jun, LI Wei, DAI Liang-ying. Research Progress of Plant Phosphate Transporters in the Response to Stress [J]. Biotechnology Bulletin, 2025, 41(7): 28-36. |
| [15] | LUO Si-fang, ZHANG Zu-ming, XIE Li-fang, GUO Zi-jing, CHEN Zhao-xing, YANG Yue-hua, YAN Xiang, ZHANG Hong-ming. Genome-wide Identification of GATA Gene Family of Jindou Kumquat (Fortunella hindsii) and Their Expression Analysis in Fruit Development [J]. Biotechnology Bulletin, 2025, 41(5): 218-230. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||