








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 37-47.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1303
Previous Articles Next Articles
LIU Na1( ), ZENG Bao-zhen2, JIA Zhao-xing1, ZHU Ying-fang2(
), ZENG Bao-zhen2, JIA Zhao-xing1, ZHU Ying-fang2( )
)
Received:2025-11-29
Online:2026-03-26
Published:2026-04-23
Contact:
ZHU Ying-fang
E-mail:iliuna@163.com;zhuyf@henu.edu.cn
LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening[J]. Biotechnology Bulletin, 2026, 42(3): 37-47.
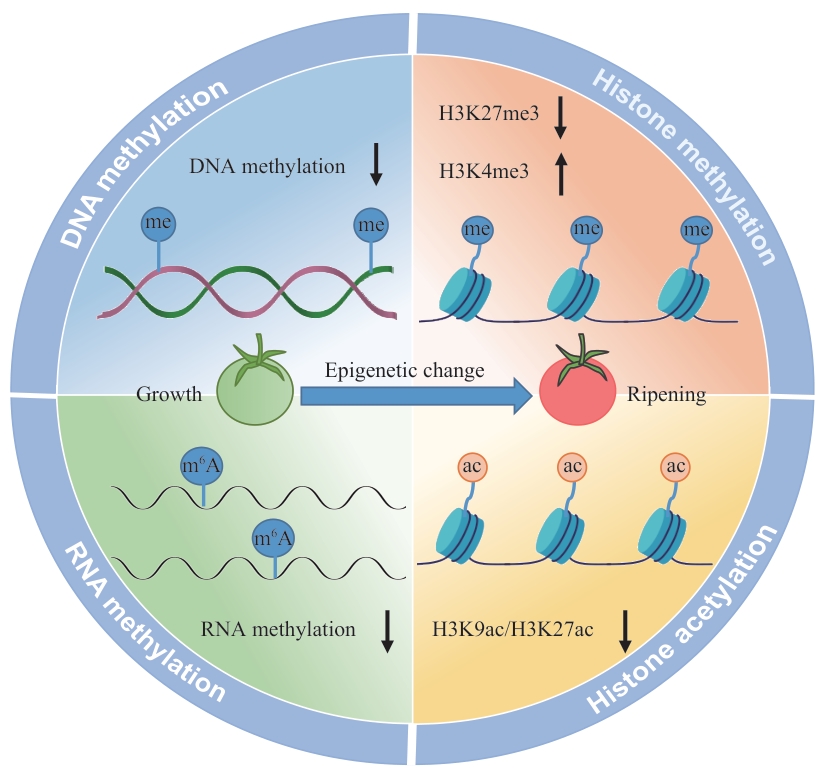
Fig. 1 Dynamic changes in epigenetic modification during fruit ripening of tomato ( Solanum lycopersicum)↑: Increase. ↓: Decrease. : DNA methylation. : m6A methylation. : Histone methylation. : Histone acetylation
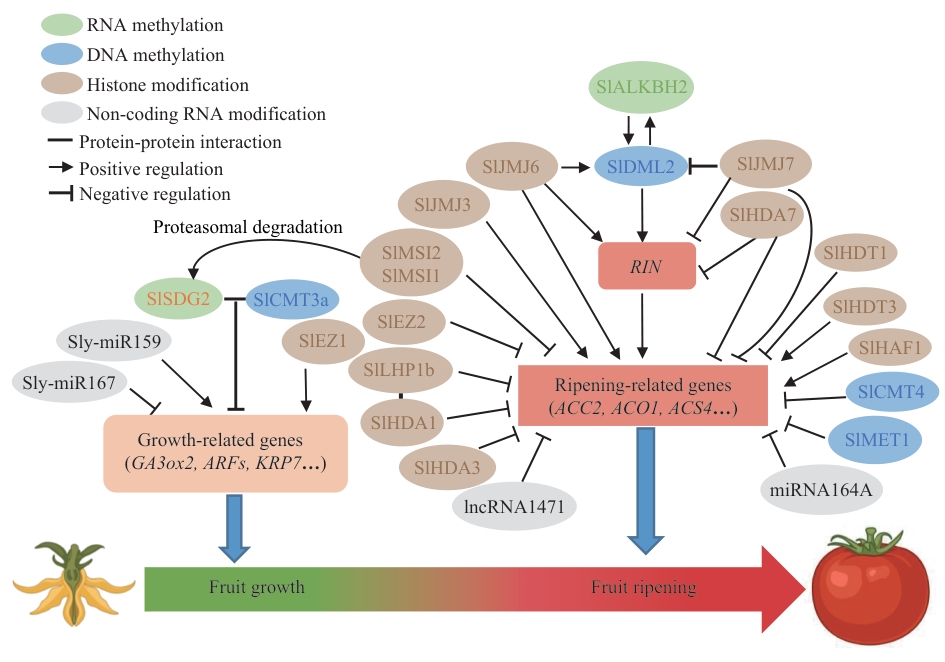
Fig. 2 Epigenetic regulatory networks of tomato fruit development and ripeningDuring tomato fruit development, SlSDG2 coordinates with SlCMT3a to negatively regulate cell proliferation, whereas SlSDG2 is ubiquitinated for degradation mediated by SlMSI2. microRNA is involved in regulating fruit size by targeting growth-related genes. During the fruit ripening stage, SlALKBH2 regulates the stability of SlDML2 transcripts through RNA m6A demethylation. Conversely, SlDML2 promotes the expressions of SlALKBH2 via DNA demethylation, forming a feedback loop that modulates fruit ripening. The histone demethylase SlJMJ6 promotes fruit ripening by removing the repressive H3K27me marks from SlDML2, RIN and other ripening-related genes, whereas SlJMJ7 suppresses fruit ripening by erasing the activating H3K4me marks from these loci. DNA methyltransferase SlMET1 and SlCMT4, together with histone deacetylases SlHDA1, SlHDA3 and SlHDT1, negatively regulate ripening-related genes to control fruit ripening, while histone acetyltransferase SlHAF1 and histone deacetylase SlHDT3 exert opposite regulatory effects. In addition, the components of PRC1 and PRC2 complexes deposit the H3K27me marks on ripening-related genes, thereby repressing fruit ripening. Non-coding RNAs, including miR164A and lncRNA1471, are also involved in the negative regulation of tomato fruit ripening
表观遗传修饰 Epigenetic modification | 表观遗传因子 Epigenetic factors | 生物功能 Biological function | 参考文献 Reference |
|---|---|---|---|
DNA甲基化 DNA methylation | SlDML2 | dml2突变,DNA甲基化升高,果实成熟延迟 | [ |
| SlCMT/DRM | 果实发育早期表达,随着果实成熟,表达量下降 | [ | |
| SlMETL | 成熟果实中表达丰度最高 | [ | |
| SlMET1 | met1突变,叶片组织的甲基化整体降低,RIN靶基因ACC2在叶片中异位表达 | [ | |
| SlCMT4 | 突变后,呈现叶小且厚、侧芽增加、花蕊异常、果实小、结实率低等生长发育缺陷 | [ | |
RNA甲基化 RNA methylation | SlALKBH2 | 敲除后,降低SlDML2表达,延迟果实成熟 | [ |
组蛋白修饰 Histone modification | SlHAF1 | 促进果实成熟 | [ |
| SlHAM1 | 种子和果实发育中表达 | [ | |
| SlHDA1 | 果实成熟负调控因子;还能够与SlERF.F12,TPL蛋白形成三元复合物,抑制关键成熟相关基因,延缓果实成熟 | [ | |
| SlHDA3 | 突变后乙烯和番茄红素合成积累,加速果实成熟 | [ | |
| SlHDA7 | 通过调控成熟相关基因表达,促进果实成熟 | [ | |
| SlHDT1 | 沉默后,果实成熟加快 | [ | |
| SlHDT3 | 沉默后,延缓果实成熟 | [ | |
| SlSDG33/SlSDG8 | 参与果实成熟 | [ | |
| SlEZ1/SlEZ2 | SlEZ1调控果实发育,SlEZ2调控成熟 | [ | |
| SlSDG2 | 与DNA甲基转移酶SlCMT3a协同调控果实大小,又能被SlCUL4泛素化降解 | [ | |
| SlLHP1b | 一方面与SlMSI1相互作用,调控果实成熟;另一方面与SlHDA1互作,招募转录因子SlPLT6,抑制果实成熟 | [ | |
| SlJMJ3 | 过表达促进果实成熟 | [ | |
| SlJMJ6 | 通过去除H3K27甲基化,并上调SlDML2表达,促进果实成熟 | [ | |
| SlJMJ7 | 果实成熟负调控因子,功能缺失后,激活成熟相关基因和DML2的表达,促进果实成熟 | [ | |
| 非编码RNA | microRNA164A | 过表达延缓番茄成熟 | [ |
| Non-coding RNA | Sly-miR159 | 靶向赤霉素相关基因,影响果实形状和大小 | [ |
| Sly-miR167 | 抑制生长素响应基因,参与腔室和胎座发育 | [ | |
| lncRNA1471 | 结合转录因子ASR,参与果实成熟 | [ |
Table 1 Overview of epigenetic regulations involved in tomato fruit development and ripening
表观遗传修饰 Epigenetic modification | 表观遗传因子 Epigenetic factors | 生物功能 Biological function | 参考文献 Reference |
|---|---|---|---|
DNA甲基化 DNA methylation | SlDML2 | dml2突变,DNA甲基化升高,果实成熟延迟 | [ |
| SlCMT/DRM | 果实发育早期表达,随着果实成熟,表达量下降 | [ | |
| SlMETL | 成熟果实中表达丰度最高 | [ | |
| SlMET1 | met1突变,叶片组织的甲基化整体降低,RIN靶基因ACC2在叶片中异位表达 | [ | |
| SlCMT4 | 突变后,呈现叶小且厚、侧芽增加、花蕊异常、果实小、结实率低等生长发育缺陷 | [ | |
RNA甲基化 RNA methylation | SlALKBH2 | 敲除后,降低SlDML2表达,延迟果实成熟 | [ |
组蛋白修饰 Histone modification | SlHAF1 | 促进果实成熟 | [ |
| SlHAM1 | 种子和果实发育中表达 | [ | |
| SlHDA1 | 果实成熟负调控因子;还能够与SlERF.F12,TPL蛋白形成三元复合物,抑制关键成熟相关基因,延缓果实成熟 | [ | |
| SlHDA3 | 突变后乙烯和番茄红素合成积累,加速果实成熟 | [ | |
| SlHDA7 | 通过调控成熟相关基因表达,促进果实成熟 | [ | |
| SlHDT1 | 沉默后,果实成熟加快 | [ | |
| SlHDT3 | 沉默后,延缓果实成熟 | [ | |
| SlSDG33/SlSDG8 | 参与果实成熟 | [ | |
| SlEZ1/SlEZ2 | SlEZ1调控果实发育,SlEZ2调控成熟 | [ | |
| SlSDG2 | 与DNA甲基转移酶SlCMT3a协同调控果实大小,又能被SlCUL4泛素化降解 | [ | |
| SlLHP1b | 一方面与SlMSI1相互作用,调控果实成熟;另一方面与SlHDA1互作,招募转录因子SlPLT6,抑制果实成熟 | [ | |
| SlJMJ3 | 过表达促进果实成熟 | [ | |
| SlJMJ6 | 通过去除H3K27甲基化,并上调SlDML2表达,促进果实成熟 | [ | |
| SlJMJ7 | 果实成熟负调控因子,功能缺失后,激活成熟相关基因和DML2的表达,促进果实成熟 | [ | |
| 非编码RNA | microRNA164A | 过表达延缓番茄成熟 | [ |
| Non-coding RNA | Sly-miR159 | 靶向赤霉素相关基因,影响果实形状和大小 | [ |
| Sly-miR167 | 抑制生长素响应基因,参与腔室和胎座发育 | [ | |
| lncRNA1471 | 结合转录因子ASR,参与果实成熟 | [ |
| [1] | Seymour GB, Østergaard L, Chapman NH, et al. Fruit development and ripening [J]. Annu Rev Plant Biol, 2013, 64: 219-241. |
| [2] | Prasanna V, Prabha TN, Tharanathan RN. Fruit ripening phenomena—an overview [J]. Crit Rev Food Sci Nutr, 2007, 47(1): 1-19. |
| [3] | Li S, Chen KS, Grierson D. Molecular and hormonal mechanisms regulating fleshy fruit ripening [J]. Cells, 2021, 10(5): 1136. |
| [4] | Cammarano D, Jamshidi S, Hoogenboom G, et al. Processing tomato production is expected to decrease by 2050 due to the projected increase in temperature [J]. Nat Food, 2022, 3(6): 437-444. |
| [5] | The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution [J]. Nature, 2012, 485(7400): 635-641. |
| [6] | Liu ZY, Wu XT, Liu HW, et al. DNA methylation in tomato fruit ripening [J]. Physiol Plant, 2022, 174(1): e13627. |
| [7] | Aprilyanto V, Wang XW, Wang RF, et al. A comprehensive model of tomato fruit ripening regulation by the transcription factors NOR-like1, NAC-NOR, and MADS-RIN [J]. Plant Physiol, 2025, 198(3): kiaf291. |
| [8] | Li S, Chen KS, Grierson D. A critical evaluation of the role of ethylene and MADS transcription factors in the network controlling fleshy fruit ripening [J]. New Phytol, 2019, 221(4): 1724-1741. |
| [9] | Vrebalov J, Pan IL, Arroyo AJM, et al. Fleshy fruit expansion and ripening are regulated by the tomato SHATTERPROOF gene TAGL1 [J]. Plant Cell, 2009, 21(10): 3041-3062. |
| [10] | Fujisawa M, Shima Y, Nakagawa H, et al. Transcriptional regulation of fruit ripening by tomato FRUITFULL homologs and associated MADS box proteins [J]. Plant Cell, 2014, 26(1): 89-101. |
| [11] | Chang YN, Zhu C, Jiang J, et al. Epigenetic regulation in plant abiotic stress responses [J]. J Integr Plant Biol, 2020, 62(5): 563-580. |
| [12] | Zhu JP, Cao XF, Deng X. Epigenetic and transcription factors synergistically promote the high temperature response in plants [J]. Trends Biochem Sci, 2023, 48(9): 788-800. |
| [13] | Zhao HH, Shin D, Zhu YF, et al. Bridging the knowledge gap: utilization of mediator subunits for crop improvement [J]. Plant Cell Environ, 2025, 48(1): 213-225. |
| [14] | Wu SQ, Hou YW, Ma Y, et al. Epigenetic regulation and its applications in plants [J]. Epigenetics Insights, 2025, 18: e010. |
| [15] | Du MM, Sun CL, Deng L, et al. Molecular breeding of tomato: Advances and challenges [J]. J Integr Plant Biol, 2025, 67(3): 669-721. |
| [16] | Yang XD, Sohail H, Noor I, et al. Epigenetic crop improvement: Integrating ENCODE strategies into horticultural breeding [J]. Hortic Res, 2025, 12(11): uhaf213. |
| [17] | An CT, Liu ZS, Pan XJ, et al. Effect of histone modifications on fruit ripening [J]. Physiol Plant, 2024, 176(6): e14639. |
| [18] | Ming YC, Jiang LB, Ji DC. Epigenetic regulation in tomato fruit ripening [J]. Front Plant Sci, 2023, 14: 1269090. |
| [19] | Wang DD, Seymour GB. Molecular and biochemical basis of softening in tomato [J]. Mol Hortic, 2022, 2(1): 5. |
| [20] | Slotkin RK, Martienssen R. Transposable elements and the epigenetic regulation of the genome [J]. Nat Rev Genet, 2007, 8(4): 272-285. |
| [21] | Gwee J, Tian WW, Qian SM, et al. DNA methylation dynamics: patterns, regulation, and function [J]. Curr Opin Plant Biol, 2025, 88: 102787. |
| [22] | Cokus SJ, Feng SH, Zhang XY, et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning [J]. Nature, 2008, 452(7184): 215-219. |
| [23] | Zhang HM, Lang ZB, Zhu JK. Dynamics and function of DNA methylation in plants [J]. Nat Rev Mol Cell Biol, 2018, 19(8): 489-506. |
| [24] | Zhu JK. Active DNA demethylation mediated by DNA glycosylases [J]. Annu Rev Genet, 2009, 43: 143-166. |
| [25] | Zhong SL, Fei ZJ, Chen YR, et al. Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening [J]. Nat Biotechnol, 2013, 31(2): 154-159. |
| [26] | Lü PT, Yu S, Zhu N, et al. Genome encode analyses reveal the basis of convergent evolution of fleshy fruit ripening [J]. Nat Plants, 2018, 4(10): 784-791. |
| [27] | Liu GZ, Li CX, Yu HY, et al. GREEN STRIPE, encoding methylated TOMATO AGAMOUS-LIKE 1, regulates chloroplast development and Chl synthesis in fruit [J]. New Phytol, 2020, 228(1): 302-317. |
| [28] | Manning K, Tör M, Poole M, et al. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening [J]. Nat Genet, 2006, 38(8): 948-952. |
| [29] | Huang H, Liu RE, Niu QF, et al. Global increase in DNA methylation during orange fruit development and ripening [J]. Proc Natl Acad Sci USA, 2019, 116(4): 1430-1436. |
| [30] | Guo XH, Xie Q, Li BY, et al. Molecular characterization and transcription analysis of DNA methyltransferase genes in tomato (Solanum lycopersicum) [J]. Genet Mol Biol, 2020, 43(1): e20180295. |
| [31] | Yang Y, Tang K, Datsenka TU, et al. Critical function of DNA methyltransferase 1 in tomato development and regulation of the DNA methylome and transcriptome [J]. J Integr Plant Biol, 2019, 61(12): 1224-1242. |
| [32] | Guo XH, Zhao JG, Chen ZW, et al. CRISPR/Cas9-targeted mutagenesis of SlCMT4 causes changes in plant architecture and reproductive organs in tomato [J]. Hortic Res, 2022, 9: uhac081. |
| [33] | Liu RE, How-Kit A, Stammitti L, et al. A DEMETER-like DNA demethylase governs tomato fruit ripening [J]. Proc Natl Acad Sci USA, 2015, 112(34): 10804-10809. |
| [34] | Lang ZB, Wang YH, Tang K, et al. Critical roles of DNA demethylation in the activation of ripening-induced genes and inhibition of ripening-repressed genes in tomato fruit [J]. Proc Natl Acad Sci USA, 2017, 114(22): E4511-E4519. |
| [35] | Niu QF, Xu YP, Huang H, et al. Two transcription factors play critical roles in mediating epigenetic regulation of fruit ripening in tomato [J]. Proc Natl Acad Sci USA, 2025, 122(15): e2422798122. |
| [36] | Zhang ZX, Zhang J, Wang Y, et al. Light regulates tomato fruit metabolome via SlDML2-mediated global DNA demethylation [J]. J Integr Plant Biol, 2025, 68(2): 383-405. |
| [37] | Hu JZ, Cai J, Umme A, et al. Unique features of mRNA m6A methylomes during expansion of tomato (Solanum lycopersicum) fruits [J]. Plant Physiol, 2022, 188(4): 2215-2227. |
| [38] | Bian HX, Song PZ, Gao Y, et al. The m6A reader SlYTH2 negatively regulates tomato fruit aroma by impeding the translation process [J]. Proc Natl Acad Sci USA, 2024, 121(28): e2405100121. |
| [39] | Zhou LL, Tian SP, Qin GZ. RNA methylomes reveal the m6A-mediated regulation of DNA demethylase gene SlDML2 in tomato fruit ripening [J]. Genome Biol, 2019, 20(1): 156. |
| [40] | Aiese Cigliano R, Sanseverino W, Cremona G, et al. Genome-wide analysis of histone modifiers in tomato: Gaining an insight into their developmental roles [J]. BMC Genomics, 2013, 14: 57. |
| [41] | Xu JD, Xu HD, Liu YL, et al. Genome-wide identification of sweet orange (Citrus sinensis) histone modification gene families and their expression analysis during the fruit development and fruit-blue mold infection process [J]. Front Plant Sci, 2015, 6: 607. |
| [42] | Han YC, Kuang JF, Chen JY, et al. Banana transcription factor MaERF11 recruits histone deacetylase MaHDA1 and represses the expression of MaACO1 and expansins during fruit ripening [J]. Plant Physiol, 2016, 171(2): 1070-1084. |
| [43] | Guo JE, Hu ZL, Zhu MK, et al. The tomato histone deacetylase SlHDA1 contributes to the repression of fruit ripening and carotenoid accumulation [J]. Sci Rep, 2017, 7(1): 7930. |
| [44] | Guo JE, Hu ZL, Yu XH, et al. A histone deacetylase gene, SlHDA3, acts as a negative regulator of fruit ripening and carotenoid accumulation [J]. Plant Cell Rep, 2018, 37(1): 125-135. |
| [45] | Guo JE. Histone deacetylase gene SlHDT1 regulates tomato fruit ripening by affecting carotenoid accumulation and ethylene biosynthesis [J]. Plant Sci, 2022, 318: 111235. |
| [46] | Zhou YJ, Li ZW, Su XG, et al. Histone deacetylase SlHDA7 impacts fruit ripening and shelf life in tomato [J]. Hortic Res, 2024, 11(11): uhae234. |
| [47] | Guo JE, Hu ZL, Li FF, et al. Silencing of histone deacetylase SlHDT3 delays fruit ripening and suppresses carotenoid accumulation in tomato [J]. Plant Sci, 2017, 265: 29-38. |
| [48] | Deng H, Chen Y, Liu ZY, et al. SlERF.F12 modulates the transition to ripening in tomato fruit by recruiting the co-repressor TOPLESS and histone deacetylases to repress key ripening genes [J]. Plant Cell, 2022, 34(4): 1250-1272. |
| [49] | Shi L, Cui XY, Shen Y. The roles of histone methylation in the regulation of abiotic stress responses in plants [J]. Plant Stress, 2024, 11: 100303. |
| [50] | Teperino R, Schoonjans K, Auwerx J. Histone methyl transferases and demethylases; can they link metabolism and transcription [J]. Cell Metab, 2010, 12(4): 321-327. |
| [51] | Cazzonelli CI, Cuttriss AJ, Cossetto SB, et al. Regulation of carotenoid composition and shoot branching in Arabidopsis by a chromatin modifying histone methyltransferase, SDG8 [J]. Plant Cell, 2009, 21(1): 39-53. |
| [52] | How Kit A, Boureau L, Stammitti-Bert L, et al. Functional analysis of SlEZ1 a tomato enhancer of zeste (E(z)) gene demonstrates a role in flower development [J]. Plant Mol Biol, 2010, 74(3): 201-213. |
| [53] | Boureau L, How-Kit A, Teyssier E, et al. A CURLY LEAF homologue controls both vegetative and reproductive development of tomato plants [J]. Plant Mol Biol, 2016, 90(4/5): 485-501. |
| [54] | Liu DD, Zhou LJ, Fang MJ, et al. Polycomb-group protein SlMSI1 represses the expression of fruit-ripening genes to prolong shelf life in tomato [J]. Sci Rep, 2016, 6: 31806. |
| [55] | Miao M, Tang XF, Lin YZ, et al. Cullin4-Ring ligase-mediated degradation of an H3K9 methyltransferase compromises cell proliferation and fruit size in tomato [J]. New Phytol, 2026, 249(3): 1303-1324. |
| [56] | Liang Q, Deng H, Li YX, et al. Like Heterochromatin Protein 1b represses fruit ripening via regulating the H3K27me3 levels in ripening-related genes in tomato [J]. New Phytol, 2020, 227(2): 485-497. |
| [57] | He XQ, Wu Y, Shu P, et al. SlPLT6 controls ripening initiation and quality traits through modulation of histone acetylation and methylation in tomato [J]. Proc Natl Acad Sci USA, 2025, 122(24): e2503732122. |
| [58] | Li ZW, Zeng J, Zhou YJ, et al. Histone H3K27 demethylase SlJMJ3 modulates fruit ripening in tomato [J]. Plant Physiol, 2024, 195(4): 2727-2742. |
| [59] | Li ZW, Jiang GX, Liu XC, et al. Histone demethylase SlJMJ6 promotes fruit ripening by removing H3K27 methylation of ripening-related genes in tomato [J]. New Phytol, 2020, 227(4): 1138-1156. |
| [60] | Ding XC, Liu XC, Jiang GX, et al. SlJMJ7 orchestrates tomato fruit ripening via crosstalk between H3K4me3 and DML2-mediated DNA demethylation [J]. New Phytol, 2022, 233(3): 1202-1219. |
| [61] | Cheng YZ, Sun YD, Pei MS, et al. Transcription factor VviAGL6a regulates fruit ripening by directly activating grape VviJMJ21 [J]. Sci Hortic, 2024, 336: 113396. |
| [62] | Haas AL, Warms JV, Hershko A, et al. Ubiquitin-activating enzyme. Mechanism and role in protein-ubiquitin conjugation [J]. J Biol Chem, 1982, 257(5): 2543-2548. |
| [63] | Miricescu A, Goslin K, Graciet E. Ubiquitylation in plants: signaling hub for the integration of environmental signals [J]. J Exp Bot, 2018, 69(19): 4511-4527. |
| [64] | Le H, Simmons CH, Zhong XH. Functions and mechanisms of histone modifications in plants [J]. Annu Rev Plant Biol, 2025, 76(1): 551-578. |
| [65] | Wang YY, Wang WH, Cai JH, et al. Tomato nuclear proteome reveals the involvement of specific E2 ubiquitin-conjugating enzymes in fruit ripening [J]. Genome Biol, 2014, 15(12): 548. |
| [66] | Wang YY, Kong LX, Wang WH, et al. Global ubiquitinome analysis reveals the role of E3 ubiquitin ligase FaBRIZ in strawberry fruit ripening [J]. J Exp Bot, 2023, 74(1): 214-232. |
| [67] | Li T, Liu L, Yang GX, et al. Ethylene-activated E3 ubiquitin ligase MdEAEL1 promotes apple fruit softening by facilitating the dissociation of transcriptional repressor complexes [J]. Adv Sci, 2025, 12(22): e2417393. |
| [68] | Waititu JK, Zhang CY, Liu J, et al. Plant non-coding RNAs: origin, biogenesis, mode of action and their roles in abiotic stress [J]. Int J Mol Sci, 2020, 21(21): 8401. |
| [69] | Lin DB, Zhu XE, Qi BL, et al. SlMIR164A regulates fruit ripening and quality by controlling SlNAM2 and SlNAM3 in tomato [J]. Plant Biotechnol J, 2022, 20(8): 1456-1469. |
| [70] | Zhao PP, Wang FP, Deng YJ, et al. Sly-miR159 regulates fruit morphology by modulating GA biosynthesis in tomato [J]. Plant Biotechnol J, 2022, 20(5): 833-845. |
| [71] | Hua B, Wu JQ, Han XQ, et al. Auxin homeostasis is maintained by sly-miR167-SlARF8A/B-SlGH3.4 feedback module in the development of locular and placental tissues of tomato fruits [J]. New Phytol, 2024, 241(3): 1177-1192. |
| [72] | Zhang LL, Zhu GN, Ma LQ, et al. lncRNA1471 mediates tomato-ripening initiation by binding to the ASR transcription factor [J]. Plant J, 2025, 121(5): e70050. |
| [73] | Zuo JH, Grierson D, Courtney LT, et al. Relationships between genome methylation, levels of non-coding RNAs, mRNAs and metabolites in ripening tomato fruit [J]. Plant J, 2020, 103(3): 980-994. |
| [74] | Wang JC, Li XM, Li JJ, et al. Manipulating the light systemic signal HY5 greatly improve fruit quality in tomato [J]. Adv Sci, 2025, 12(23): 2500110. |
| [75] | Quinet M, Angosto T, Yuste-Lisbona FJ, et al. Tomato fruit development and metabolism [J]. Front Plant Sci, 2019, 10: 1554. |
| [76] | Yang CP, Ying SY, Tang BB, et al. The mechanistic insights into fruit ripening: integrating phytohormones, transcription factors, and epigenetic modification [J]. J Genet Genom, 2025, 52(12): 1475-1489. |
| [1] | YIN Yue, QIN Xiao-ya, MI Jia, AN Wei, HE Jun, ZHANG Feng-feng. Identification of FBN Gene Family and Its Relationship with Carotenoids Metabolism in Lyciumbarbarum [J]. Biotechnology Bulletin, 2026, 42(3): 338-348. |
| [2] | LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development [J]. Biotechnology Bulletin, 2026, 42(3): 362-373. |
| [3] | LUO Wei, GONG Ao, ZHONG Yang, HU Di, ZHOU Hong-yuan, ZHANG Hong-xin, AI Ju, LUO You-wei, GAO Dong-li. Pleiotropic Effects of SEPALLATA2 Knock-out on Fruit and Wart Development in Cucumber [J]. Biotechnology Bulletin, 2026, 42(3): 283-293. |
| [4] | LIU Lin-ya, LIU Huan-yan, LIANG Xin-yu, SONG Shu-yi, HE Bin, WANG Xu-ying, HUANG Ya-cheng. Genome-wide Identification and Expression Analysis of BGAL Gene Family in Actinidiachinensis var. Hongyang [J]. Biotechnology Bulletin, 2026, 42(3): 312-323. |
| [5] | LIU Miao, LIN Tao, JIA Le-song, HU Feng, LI Tao, LI Zhi-wan, LIU Mei-fang, ZHENG Fang-yan, CUI Long. From Wild to Cultivated: Evolution and Regulatory Mechanisms of Tomato Fruit Color [J]. Biotechnology Bulletin, 2026, 42(3): 187-202. |
| [6] | LI Ya-ni, HAN Hong-yu, GENG Meng-shuang, MI Ruo-lan, Wang Wei-qi, Yu Wen-jing, MENG Xian-wen, LI Chuan-you. Mechanistic Study on ChiC-mediated Regulation Mechanism of Tomato Resistance to Botrytis cinerea [J]. Biotechnology Bulletin, 2026, 42(3): 255-262. |
| [7] | WANG He-yao, SUN Hong-mei. Research Progress in the Function and Formation Mechanism of Trichomes in Horticultural Plants [J]. Biotechnology Bulletin, 2026, 42(3): 242-254. |
| [8] | CHENG Yun-xia, ZHANG Jun-hong, YE Jie. Advances in the Genetic Regulation of Soluble Solid Accumulation in Tomato Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 145-155. |
| [9] | YAN Chen-lin, LI Fan, YAN Chun-ting, CHENG Jiao-wen, HU Kai-lin, YE Zhi-biao, SONG Jian-wen. Advances in Genes Related to Tomato Fruit Morphogenesis [J]. Biotechnology Bulletin, 2026, 42(3): 172-186. |
| [10] | HU Qiu-ling, CHEN Ling, HUANG Jia-yi, ZHAO Zi-qiao, PAN Lu-yi, LIU Hui-li, LIU Tai-bo. Advances in the Regulation of Fruit Development by Polyamines [J]. Biotechnology Bulletin, 2026, 42(3): 203-212. |
| [11] | MA Shi-jie, LI Zheng, LI Wei, GUO Yang-dong, ZHANG Na. Research Progress in Light Signaling Regulation of Fruit Development in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 5-18. |
| [12] | ZHANG Gao-xiang, WU Yu-bi, GUO Ya-jing, JI Wei, YANG Zhong-yi. Identification and Expression Analysis of WD40 Gene Family in Grape [J]. Biotechnology Bulletin, 2026, 42(3): 324-337. |
| [13] | YIN Shi-qing, TIAN Tai, HUANG Feng-ting, FENG Long-qiang, WANG Hao, ZHANG Jing, HE Wen, CHEN Qing, WANG Xiao-rong, WANG Yan. Advances in Regulatory Mechanism of Fruit Firmness in Fruit Crops [J]. Biotechnology Bulletin, 2026, 42(3): 213-229. |
| [14] | DU Dan, GUO Xiang, HU Xin, PAN Yu. Advances in the Regulatory Mechanisms of Plastid Development on Fruit Ripening and Quality [J]. Biotechnology Bulletin, 2026, 42(3): 48-59. |
| [15] | JIANG Zhe-hui, WANG Xiao-long, WANG Shou-chuang, ZHOU Ke. Advances in the Elucidation of Metabolic Pathways and Molecular Breeding for Tomato Flavor [J]. Biotechnology Bulletin, 2026, 42(3): 60-78. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||