








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 255-262.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1242
Previous Articles Next Articles
LI Ya-ni1,2( ), HAN Hong-yu3, GENG Meng-shuang1, MI Ruo-lan1, Wang Wei-qi3, Yu Wen-jing3, MENG Xian-wen1,2(
), HAN Hong-yu3, GENG Meng-shuang1, MI Ruo-lan1, Wang Wei-qi3, Yu Wen-jing3, MENG Xian-wen1,2( ), LI Chuan-you1,2,3(
), LI Chuan-you1,2,3( )
)
Received:2025-11-16
Online:2026-03-26
Published:2026-04-23
Contact:
MENG Xian-wen, LI Chuan-you
E-mail:17793655826@163.com;xwmeng@sdau.edu.cn;chuanyouli@sdau.edu.cn
LI Ya-ni, HAN Hong-yu, GENG Meng-shuang, MI Ruo-lan, Wang Wei-qi, Yu Wen-jing, MENG Xian-wen, LI Chuan-you. Mechanistic Study on ChiC-mediated Regulation Mechanism of Tomato Resistance to Botrytis cinerea[J]. Biotechnology Bulletin, 2026, 42(3): 255-262.
引物名称 Primer name | 引物序列 Primer sequence (5′‒3′) |
|---|---|
| ACTIN2-qRT-F | TTGCTGACCGTATGAGCAAG |
| ACTIN2-qRT-R | GGACAATGGATGGACCAGAC |
| ChiC-qRT-F | TGGACTTGATCTTGATTGGGAA |
| ChiC-qRT-R | CCGGATAGTTCAAACCATTGAC |
| ERF.C3-qRT-F | GAATTCGTCGTCTAGCTATGGA |
| ERF.C3-qRT-R | GAACCTCTCATAGCAAATGCAG |
| PR-STH2c-qRT-F | AATCCTTCTGTCTACGCTTGAA |
| PR-STH2c-qRT-R | TTAACATCACAACACGTTCACG |
| PR-STH2d-qRT-F | AATAAGAATCAGGTCCACACGT |
| PR-STH2d-qRT-R | CACACAACTCAACAAAAAGCAC |
Table 1 Primers for RT-qPCR used in this study
引物名称 Primer name | 引物序列 Primer sequence (5′‒3′) |
|---|---|
| ACTIN2-qRT-F | TTGCTGACCGTATGAGCAAG |
| ACTIN2-qRT-R | GGACAATGGATGGACCAGAC |
| ChiC-qRT-F | TGGACTTGATCTTGATTGGGAA |
| ChiC-qRT-R | CCGGATAGTTCAAACCATTGAC |
| ERF.C3-qRT-F | GAATTCGTCGTCTAGCTATGGA |
| ERF.C3-qRT-R | GAACCTCTCATAGCAAATGCAG |
| PR-STH2c-qRT-F | AATCCTTCTGTCTACGCTTGAA |
| PR-STH2c-qRT-R | TTAACATCACAACACGTTCACG |
| PR-STH2d-qRT-F | AATAAGAATCAGGTCCACACGT |
| PR-STH2d-qRT-R | CACACAACTCAACAAAAAGCAC |
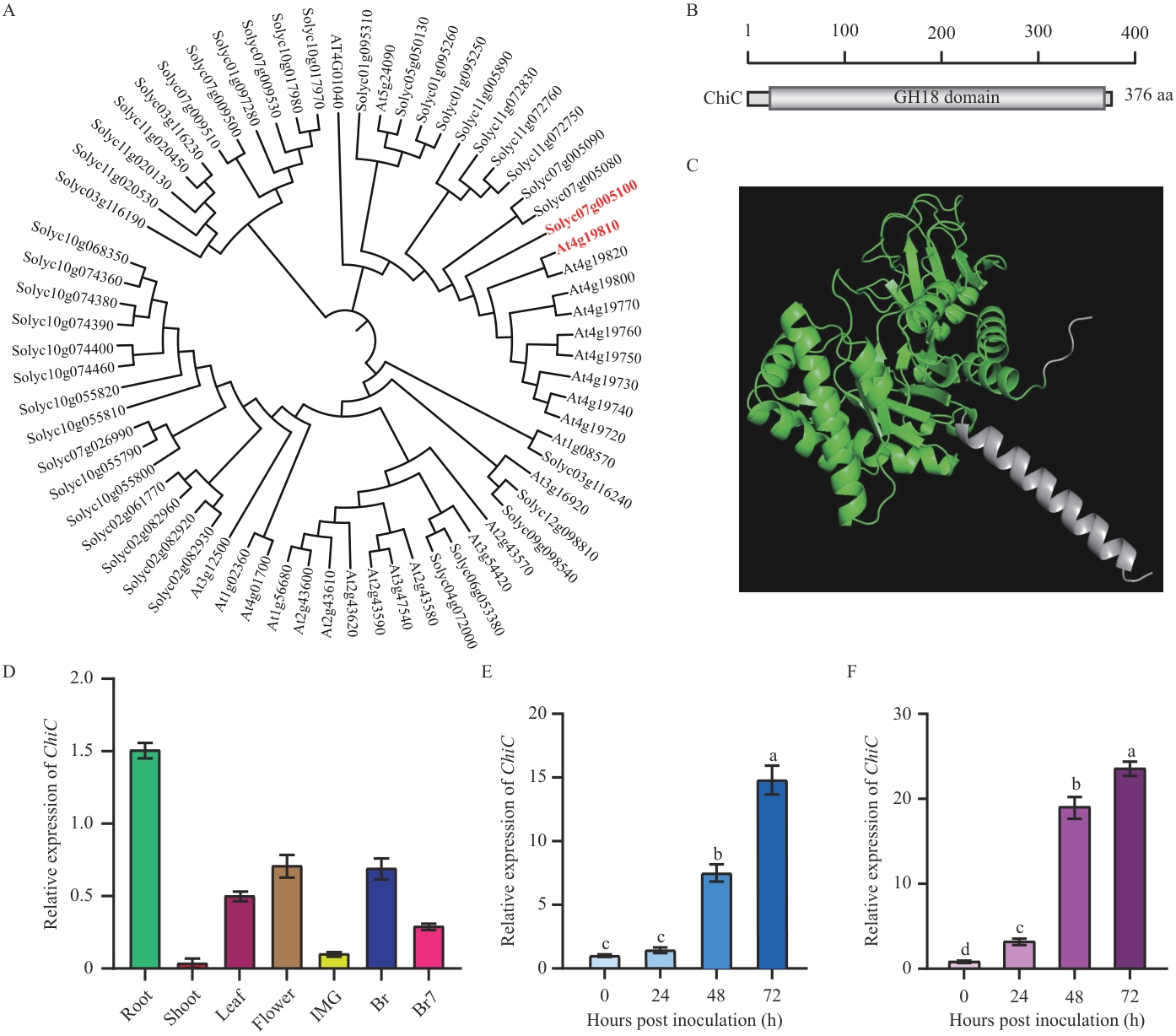
Fig. 1 Structure of tomato ChiC and its expression patternsA: Phylogenetic tree of chitinase genes from tomato and Arabidopsis. B: ChiC protein domains. C: Three-dimensional structure of ChiC. D: Relative expression of the ChiC gene in different tissues (IMG: immature green; Br: breaker; Br7: 7 d after breaker stage). E: Expression patterns of ChiC before and after B. cinerea infection in the fruits. F: Expression patterns of ChiC before and after B. cinerea infection in leaves. The values in the bar charts are presented as mean ± standard error (n=3). Different lowercase letters indicate significant differences at P<0.05 level. The same below
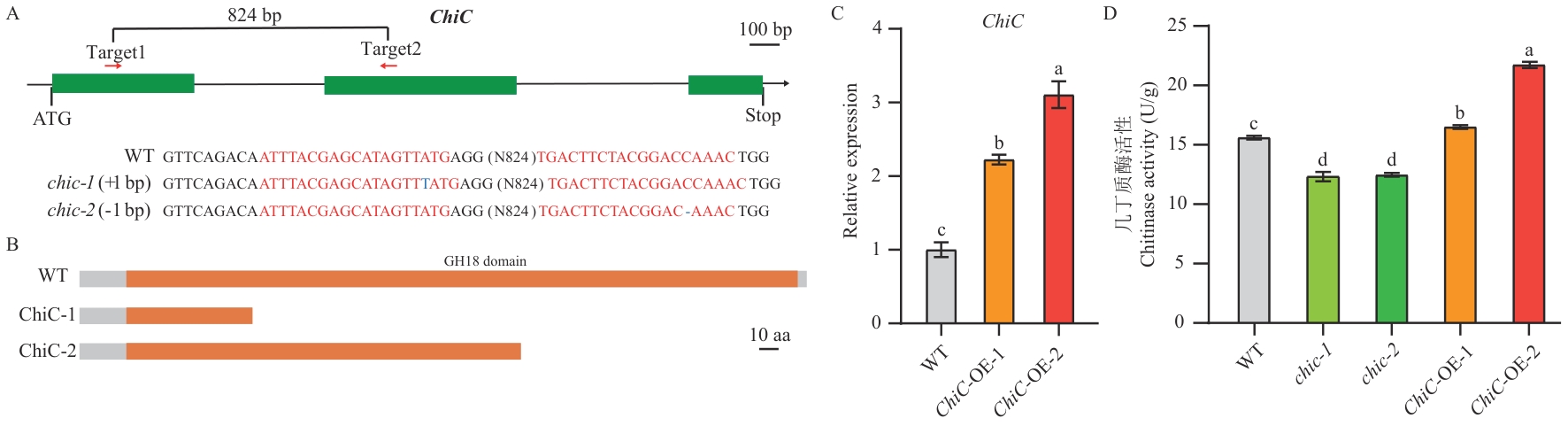
Fig. 2 Generation of ChiC genetic materials and assays of chitinase activityA: Positions of the gene-editing targets and mutation types in the chic-1 and chic-2 lines. B: ChiC protein domain structure in the chic-1 and chic-2 lines. C: Relative expression of ChiC in overexpression lines. D: Detection of chitinase activity in transgenic materials
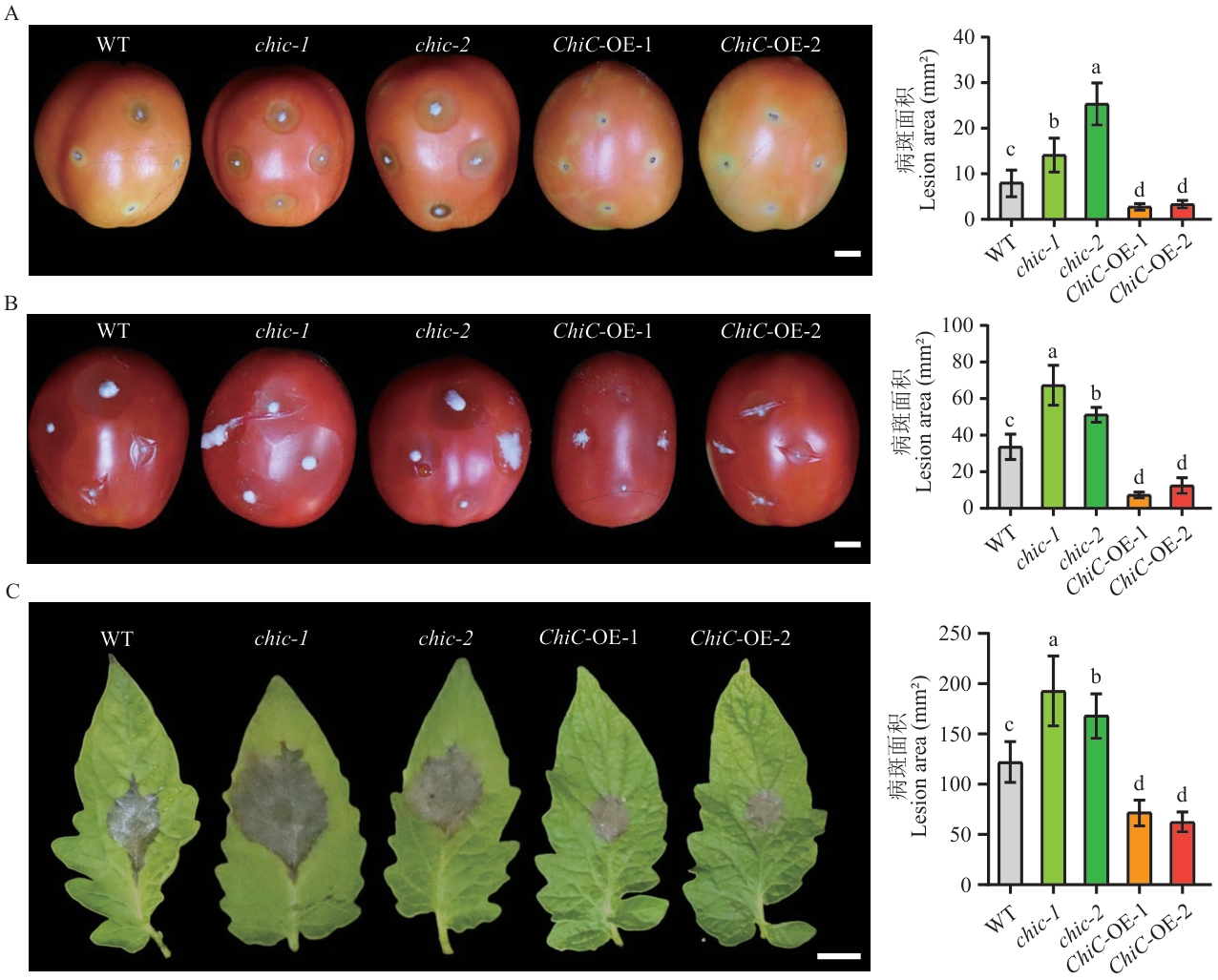
Fig. 3 ChiC positively regulates the resistance to B. cinerea in tomatoA: Disease symptoms and quantification of lesion areas in breaker-stage fruits at 3 d after B. cinerea infection. B: Disease symptoms and quantification of lesion areas in red-ripe fruits at 3 d after B. cinerea infection. C: Disease symptoms and quantification of lesion areas in leaves at 3 d after B. cinerea infection. Bar=1 cm
| [1] | Wang SQ, Zhang XY, Zhong S, et al. BcatrB mediates pyrimethanil resistance in Botrytis cinerea revealed by transcriptomics analysis [J]. Sci Rep, 2025, 15(1): 11478. |
| [2] | Williamson B, Tudzynski B, Tudzynski P, et al. Botrytis cinerea: the cause of grey mould disease [J]. Mol Plant Pathol, 2007, 8(5): 561-580. |
| [3] | Mobasher Amini M, Mirzaei S, Heidari A. A growing threat: Investigating the high incidence of benzimidazole fungicides resistance in Iranian Botrytis cinerea isolates [J]. PLoS One, 2023, 18(11): e0294530. |
| [4] | Shawky A, Hatawsh A, Al-Saadi N, et al. Revolutionizing tomato cultivation: CRISPR/Cas9 mediated biotic stress resistance [J]. Plants, 2024, 13(16): 2269. |
| [5] | Arie T, Takahashi H, Kodama M, et al. Tomato as a model plant for plant-pathogen interactions [J]. Plant Biotechnol, 2007, 24(1): 135-147. |
| [6] | Huang H, Zhao WC, Li CH, et al. SlVQ15 interacts with jasmonate-ZIM domain proteins and SlWRKY31 to regulate defense response in tomato [J]. Plant Physiol, 2022, 190(1): 828-842. |
| [7] | 李杰, 梁郅林, 孙燕, 等. SlSnRK1.2调控番茄抗灰霉病功能分析 [J]. 中国农业科学, 2024(21): 4238-4247. |
| Li J, Liang ZL, Sun Y, et al. Functional analysis of SlSnRK1.2 in regulating tomato resistance to grey mould [J]. Sci Agric Sin, 2024(21): 4238-4247. | |
| [8] | Ding ST, Feng SX, Zhou SB, et al. A novel LRR receptor-like kinase BRAK reciprocally phosphorylates PSKR1 to enhance growth and defense in tomato [J]. EMBO J, 2024, 43(23): 6104-6123. |
| [9] | Li S, Huang YL, Zhao Y, et al. Ripening-induced defence signalling in Botrytis cinerea-infected tomato fruits involves activation of ERF.F4 by a MYC2-NOR/RIN protein complex [J]. Plant Biotechnol J, 2025, 23(9): 4126-4139. |
| [10] | Wang XW, Gao M, Li HX, et al. SCFSlRAE1 regulates tomato resistance to Botrytis cinerea by modulating SlWRKY1 stability [J]. J Integr Plant Biol, 2025, 67(8): 2167-2183. |
| [11] | Zhu KY, Merzendorfer H, Zhang WQ, et al. Biosynthesis, turnover, and functions of chitin in insects [J]. Annu Rev Entomol, 2016, 61: 177-196. |
| [12] | Grover A. Plant chitinases: genetic diversity and physiological roles [J]. Crit Rev Plant Sci, 2012, 31(1): 57-73. |
| [13] | Khan RS, Iqbal A, Bibi A, et al. Plant chitinases: Types, structural classification, antifungal potential and transgenic expression in plants for enhanced disease resistance [J]. Plant Cell Tissue Organ Cult, 2024, 156(3): 75. |
| [14] | 郭林霞, 董旋, 赵德刚. 转杜仲几丁质酶基因EuCHIT1番茄提高对灰霉病的抗性 [J]. 植物生理学报, 2016, 52(5): 703-714. |
| Guo LX, Dong X, Zhao DG. Transgenic tomato plants expressing a Eucommia ulmoides chitinase gene EuCHIT1 and their resistance to Botrytis cinerea [J]. Plant Physiol J, 2016, 52(5): 703-714. | |
| [15] | Tabei Y, Kitade S, Nishizawa Y, et al. Transgenic cucumber plants harboring a rice chitinase gene exhibit enhanced resistance to gray mold (Botrytis cinerea) [J]. Plant Cell Rep, 1998, 17(3): 159-164. |
| [16] | Liu ZH, Yang CP, Qi XT, et al. Cloning, heterologous expression, and functional characterization of a chitinase gene, Lbchi32, from Limonium bicolor [J]. Biochem Genet, 2010, 48(7/8): 669-679. |
| [17] | Liu JJ, Wen JX, Li JF, et al. Nepenthes chitinase NkChit2b-1 confers broad-spectrum resistance to chitin-containing pathogens and insects in plants [J]. Adv Biotechnol, 2025, 3(2): 12. |
| [18] | Cao J, Tan XN. Comprehensive analysis of the chitinase family genes in tomato (Solanum lycopersicum) [J]. Plants, 2019, 8(3): 52. |
| [19] | Ohnuma T, Numata T, Osawa T, et al. A class V chitinase from Arabidopsis thaliana: gene responses, enzymatic properties, and crystallographic analysis [J]. Planta, 2011, 234(1): 123-137. |
| [20] | Abramson J, Adler J, Dunger J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3 [J]. Nature, 2024, 630(8016): 493-500. |
| [21] | Lian JJ, Han HY, Zhao JH, et al. In-vitro and in-planta Botrytis cinerea inoculation assays for tomato [J]. Bio Protoc, 2018, 8(8): e2810. |
| [22] | Du MM, Zhao JH, Tzeng DT, et al. MYC2 orchestrates a hierarchical transcriptional cascade that regulates jasmonate-mediated plant immunity in tomato [J]. Plant Cell, 2017, 29(8): 1883-1906. |
| [23] | Wang T, Zhang HY, Zhu HL. CRISPR technology is revolutionizing the improvement of tomato and other fruit crops [J]. Hortic Res, 2019, 6: 77. |
| [24] | Zhang JL, Dong DH, Jia CY, et al. Fine-tuning of MYC2-mediated Botrytis defense response by the LBD40/42-CRL3BPM4 module in tomato [J]. Plant Cell, 2025, 37(11): koaf258. |
| [25] | Vaghela B, Vashi R, Rajput K, et al. Plant chitinases and their role in plant defense: a comprehensive review [J]. Enzyme Microb Technol, 2022, 159: 110055. |
| [26] | Jabeen N, Chaudhary Z, Gulfraz M, et al. Expression of rice chitinase gene in genetically engineered tomato confers enhanced resistance to Fusarium wilt and early blight [J]. Plant Pathol J, 2015, 31(3): 252-258. |
| [27] | Li QQ, Yang YY, Bai X, et al. Systematic analysis and functional characterization of the chitinase gene family in Fagopyrum tataricum under salt stress [J]. BMC Plant Biol, 2024, 24(1): 1222. |
| [28] | Hamid R, Khan M, Ahmad M, et al. Chitinases: an update [J]. J Pharm Bioall Sci, 2013, 5(1): 21-29. |
| [29] | He TN, Fan JS, Jiao GZ, et al. Bioinformatics and expression analysis of the chitinase genes in strawberry (Fragaria vesca) and functional study of FvChi-14 [J]. Plants, 2023, 12(7): 1543. |
| [30] | Ali M, Li QH, Zou T, et al. Chitinase gene positively regulates hypersensitive and defense responses of pepper to Colletotrichum acutatum infection [J]. Int J Mol Sci, 2020, 21(18): 6624. |
| [31] | Xiao YH, Li XB, Yang XY, et al. Cloning and characterization of a balsam pear class I chitinase gene (Mcchit1) and its ectopic expression enhances fungal resistance in transgenic plants [J]. Biosci Biotechnol Biochem, 2007, 71(5): 1211-1219. |
| [32] | Lombardo L, Zelasco S. Biotech approaches to overcome the limitations of using transgenic plants in organic farming [J]. Sustainability, 2016, 8(5): 497. |
| [1] | LIU Miao, LIN Tao, JIA Le-song, HU Feng, LI Tao, LI Zhi-wan, LIU Mei-fang, ZHENG Fang-yan, CUI Long. From Wild to Cultivated: Evolution and Regulatory Mechanisms of Tomato Fruit Color [J]. Biotechnology Bulletin, 2026, 42(3): 187-202. |
| [2] | CHENG Yun-xia, ZHANG Jun-hong, YE Jie. Advances in the Genetic Regulation of Soluble Solid Accumulation in Tomato Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 145-155. |
| [3] | YAN Chen-lin, LI Fan, YAN Chun-ting, CHENG Jiao-wen, HU Kai-lin, YE Zhi-biao, SONG Jian-wen. Advances in Genes Related to Tomato Fruit Morphogenesis [J]. Biotechnology Bulletin, 2026, 42(3): 172-186. |
| [4] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [5] | DU Dan, GUO Xiang, HU Xin, PAN Yu. Advances in the Regulatory Mechanisms of Plastid Development on Fruit Ripening and Quality [J]. Biotechnology Bulletin, 2026, 42(3): 48-59. |
| [6] | JIANG Zhe-hui, WANG Xiao-long, WANG Shou-chuang, ZHOU Ke. Advances in the Elucidation of Metabolic Pathways and Molecular Breeding for Tomato Flavor [J]. Biotechnology Bulletin, 2026, 42(3): 60-78. |
| [7] | WANG Xiao-yi, LI Jin-yan, XING Xing, ZHU Hong-liang. Screening and Functional Analysis of Ethylene-responsive Genes Regulating Tomato Fruit Ripening and Respiration [J]. Biotechnology Bulletin, 2026, 42(3): 275-282. |
| [8] | LI Ying-hui, WANG Yang-bo-han, ZHOU Hao-bo, LU Xin-ru, ZHANG Ke-xin, YU Yang, LI Chuan-you, SUN Chuan-long. Identification of VPE Gene Family and Their Functional Analysis under Abiotic Stress in Tomato [J]. Biotechnology Bulletin, 2026, 42(3): 263-274. |
| [9] | WANG Ting-ting, HE Meng-ya, SHENG Jia-shun, GAO Chen, CAI Han-fang, FU Tong, SUN Yu, GAO Teng-yun, ZHANG Tian-liu. Preparation of NCAPG Knockout Bovine Fibroblast Cell Lines Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2026, 42(2): 317-324. |
| [10] | ZENG Ting, ZHANG Lan, LUO Rui. Functional Analysis of the Transcription Factor MpR2R3-MYB17 in Regulating Gemma Development in Marchantia polymorpha L. [J]. Biotechnology Bulletin, 2026, 42(1): 208-217. |
| [11] | SU Xiu-min, HAN Wen-qing, WANG Jiao, LI Peng, WANG Qiu-lan, LI Wan-xing, CAO Jin-jun. Isolation, Identification, Biological Characteristics and Biocontrol Effects of Trichoderma harzianum M408 against Tomato Early Blight [J]. Biotechnology Bulletin, 2025, 41(9): 277-288. |
| [12] | LI Shan, MA Deng-hui, MA Hong-yi, YAO Wen-kong, YIN Xiao. Identification and Expression Analysis of SKP1 Gene Family in Grapevine (Vitis vinifera L.) [J]. Biotechnology Bulletin, 2025, 41(9): 147-158. |
| [13] | YU Yong-xia, DU Zai-hui, ZHU Long-jiao, XU Wen-tao. Application and Research Progress of Gene Editing Technology in Bovine [J]. Biotechnology Bulletin, 2025, 41(8): 34-41. |
| [14] | QU Shan, ZHAO Yue, LI Ya-hua, ZHENG Gui-ling, XIAN Hong-quan. A Study on the Interaction between Transcriptional Factor and Protein of Tachi2 Chitinase Gene in Trichoderma asperellum [J]. Biotechnology Bulletin, 2025, 41(5): 310-319. |
| [15] | LU Yong-jie, XIA Hai-qian, LI Yong-ling, ZHANG Wen-jian, YU Jing, ZHAO Hui-na, WANG Bing, XU Ben-bo, LEI Bo. Cloning and Expression Analysis of AP2/ERF Transcription Factor NtESR2 in Nicotiana tabacum [J]. Biotechnology Bulletin, 2025, 41(4): 266-277. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||