








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 133-144.doi: 10.13560/j.cnki.biotech.bull.1985.2026-0164
Previous Articles Next Articles
PENG Chu1,2( ), SUN Juan-li1, ZHENG Bei-bei1, ZHANG Ruo-xi1, HAN Yue-peng1(
), SUN Juan-li1, ZHENG Bei-bei1, ZHANG Ruo-xi1, HAN Yue-peng1( ), ZHAO Yun1(
), ZHAO Yun1( )
)
Received:2026-02-02
Online:2026-03-26
Published:2026-04-23
Contact:
HAN Yue-peng, ZHAO Yun
E-mail:pengchu2026@163.com;yphan@wbgcas.cn;zhaoyun@wbgcas.cn
PENG Chu, SUN Juan-li, ZHENG Bei-bei, ZHANG Ruo-xi, HAN Yue-peng, ZHAO Yun. Research Progress in the Enzymatic Degradation Mechanism of Anthocyanins in Fruits[J]. Biotechnology Bulletin, 2026, 42(3): 133-144.
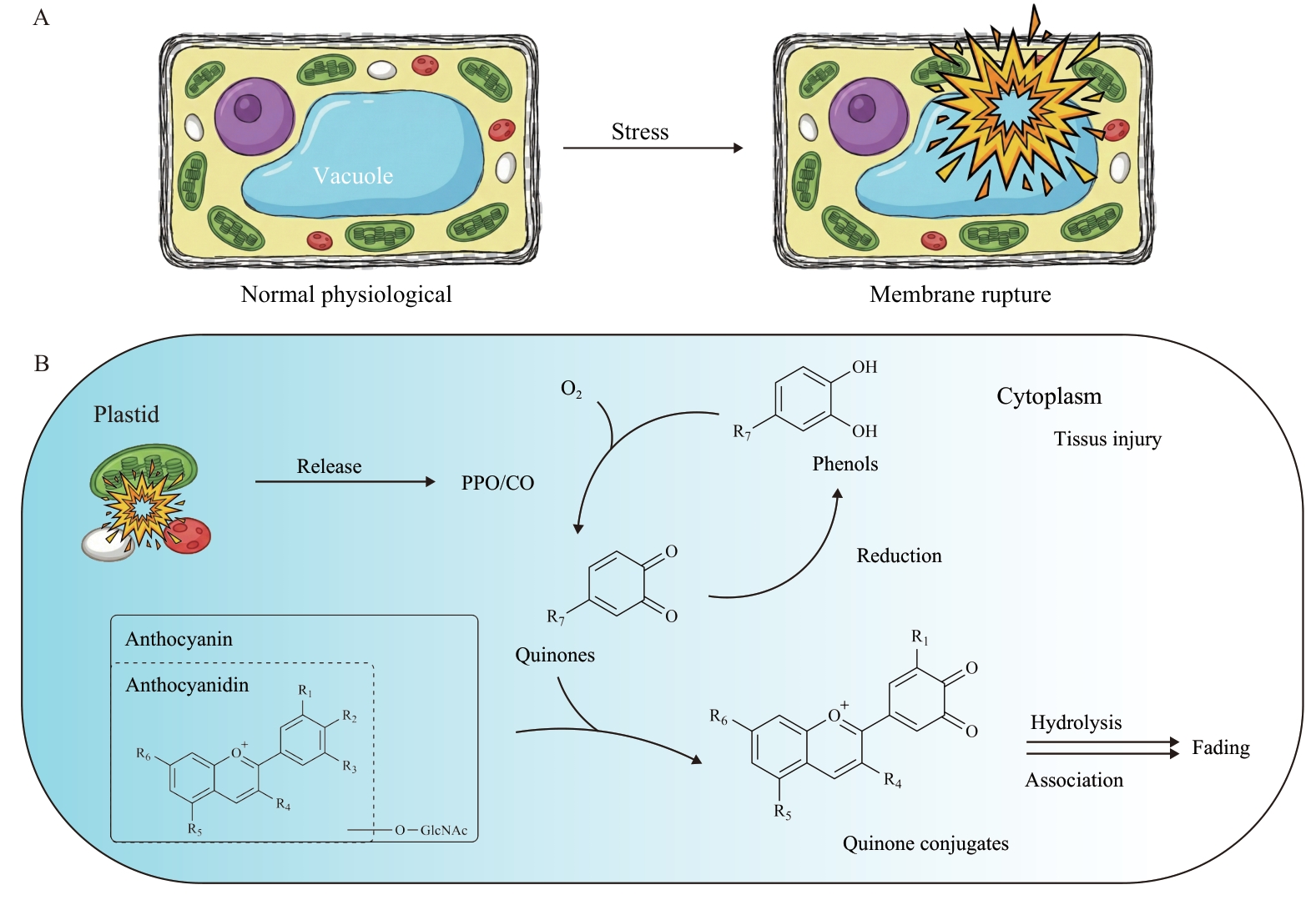
Fig. 1 Schematic representation of anthocyanin degradation pathway in the cytoplasm triggered by tissue injuryA: Under various stress conditions, the disruption of cellular membrane integrity leads to the breakdown of membrane systems, resulting in the release of plastid-localized PPO/CO into the cytoplasm. B: The released PPO/CO catalyzes the oxidation of free phenolic substrates to generate reactive quinone intermediates, which form conjugates with anthocyanins. These conjugates subsequently undergo hydrolysis and polymerization, ultimately causing tissue discoloration. Moreover, the quinone intermediates can be rapidly reduced back to phenolic substrates, establishing a redox cycle that facilitates efficient anthocyanin degradation in the cytoplasm. PPO/CO: Catechol oxidase belonging to the polyphenol oxidase family
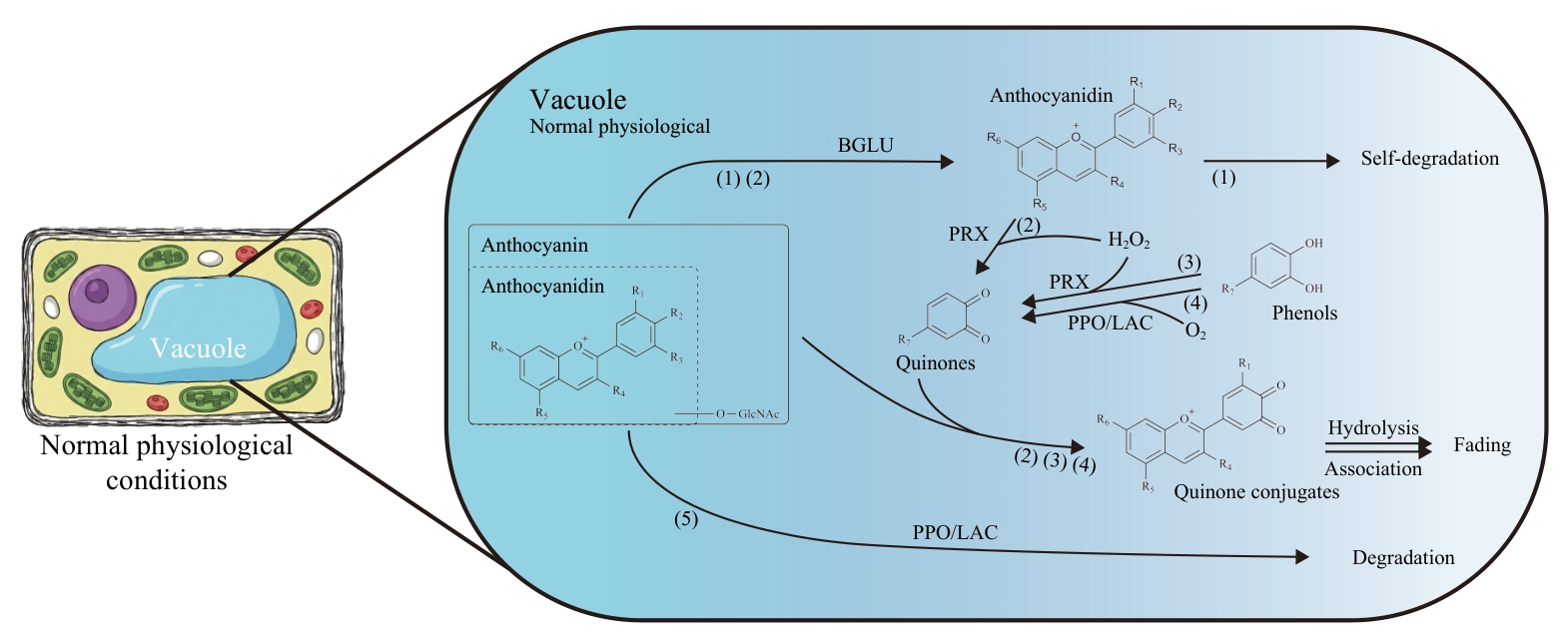
Fig. 2 Proposed schematic model illustrating enzymatic degradation pathways of anthocyanins in the vacuole under normal physiological conditionsUnder normal physiological conditions, anthocyanins sequestered in the vacuole undergo enzymatic degradation through multiple degradation routes. 1) Anthocyanins are hydrolyzed by BGLU to generate anthocyanidins, which are inherently unstable and prone to spontaneous decomposition. 2) Anthocyanidins generated by BGLU are further oxidized to quinones by PRX, which then couple with anthocyanins to form quinone conjugates, leading to anthocyanin degradation through hydrolysis or polymerization. 3) 4) Free phenol compounds present in the vacuole are oxidized to quinones by PRX and PPO/LAC, which then mediate anthocyanin degradation by a coupling reaction similar to athway 2). 5) Anthocyanins are directly oxidatively degraded by PPO/LAC. BGLU: β-glucosidase; PRX: Peroxidase; PPO/LAC: the laccase belonging to the polyphenol oxidase family
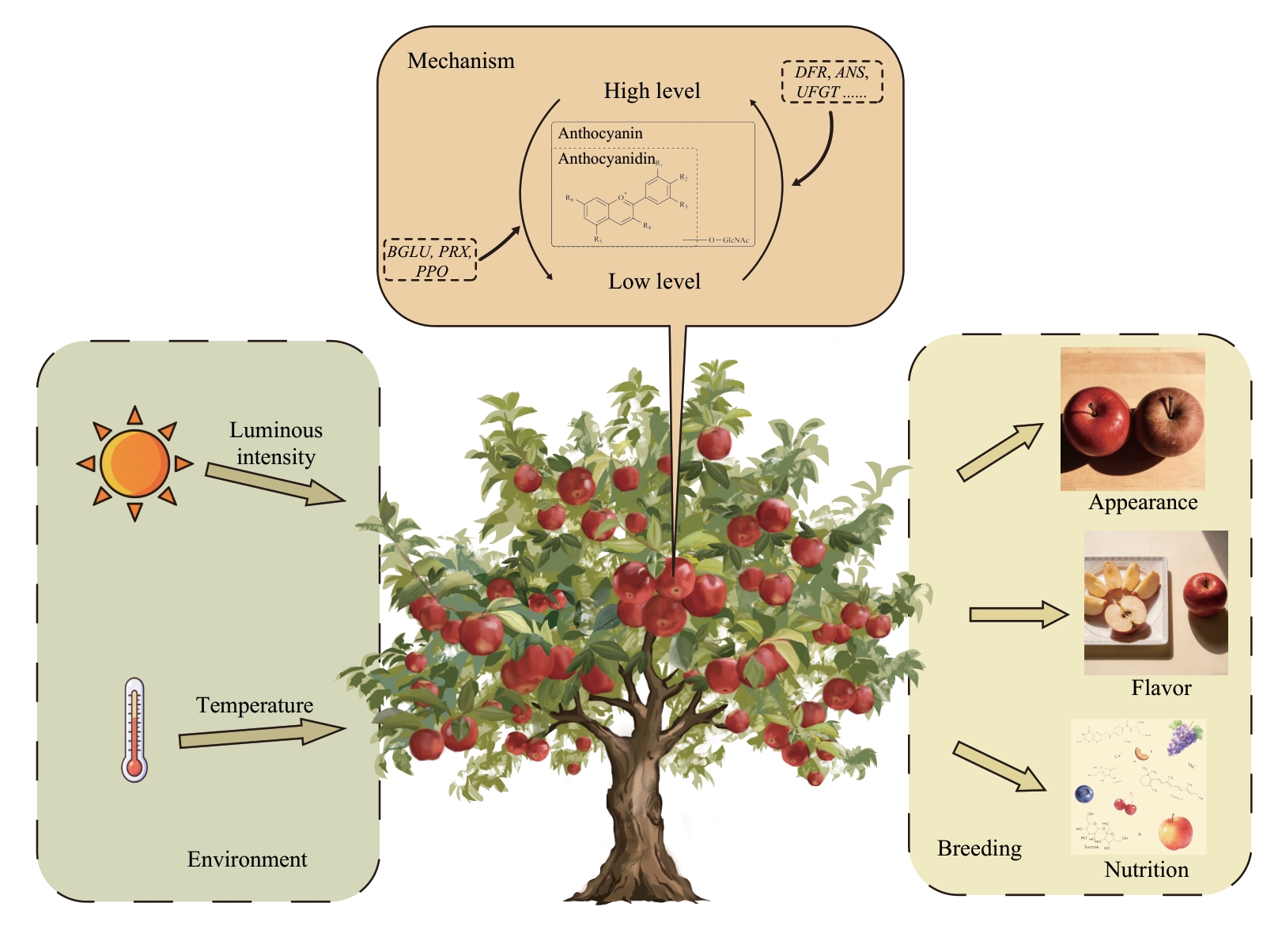
Fig. 3 Schematic diagram illustrating environmental regulation of anthocyanin metabolism in fruit and its applications in breeding programsEnvironmental factors such as light and temperature regulate anthocyanin accumulation in fruit by coordinately modulating the expressions of key genes involved in both biosynthesis and degradation. Enhanced expressions of key biosynthetic genes, including DFR, ANS and UFGT, promotes anthocyanin accumulation, whereas the upregulation of degradation-associated genes such as BGLU, PRX and PPO accelerates anthocyanin turnover. Anthocyanin biosynthesis and degradation respond coordinately to changes in environmental cues at the transcriptional level, and their coordinated interplay maintains anthocyanin metabolic homeostasis in fruit. Deciphering the molecular mechanism by which environmental factors coordinately regulate anthocyanin biosynthesis and degradation will provide a foundation for coloration quality improvement strategies and genetic enhancement of fruit quality traits. DFR: Dihydroflavonol 4-reductase; ANS: Anthocyanidin synthase; UFGT: UDP-glucose: flavonoid 3-O-glucosyltransferase; BGLU: β-glucosidase; PRX: peroxidase; PPO: polyphenol oxidase
| [1] | Zhao L, Sun JL, Cai YM, et al. PpHYH is responsible for light-induced anthocyanin accumulation in fruit peel of Prunus persica [J]. Tree Physiol, 2022, 42(8): 1662-1677. |
| [2] | Li Z, Ahammed GJ. Plant stress response and adaptation via anthocyanins: a review [J]. Plant Stress, 2023, 10: 100230. |
| [3] | Grünig N, Horz JM, Pucker B. Diversity and ecological functions of anthocyanins [J]. BMC Plant Biol, 2026, 26: 146. |
| [4] | Moyer RA, Hummer KE, Finn CE, et al. Anthocyanins, phenolics, and antioxidant capacity in diverse small fruits: Vaccinium, Rubus, and Ribes [J]. J Agric Food Chem, 2002, 50(3): 519-525. |
| [5] | 常仁杰, 何勇, 朱祝军. 植物花色苷抗逆性作用研究进展 [J]. 北方园艺, 2013(9): 216-219. |
| Chang RJ, He Y, Zhu ZJ. Research progress of the resistance of plant anthocyanin to stress [J]. North Hortic, 2013(9): 216-219. | |
| [6] | Geng ZQ, Zhou LJ, Wang YG, et al. A U-box E3 ubiquitin ligase CmPUB15 targets CmMYB73 to regulate anthocyanin biosynthesis in response to low temperatures in Chrysanthemum [J]. New Phytol, 2025, 248(3): 1304-1320. |
| [7] | Zhu CL, Yang XH, Chen WW, et al. WD40 protein OsTTG1 promotes anthocyanin accumulation and CBF transcription factor-dependent pathways for rice cold tolerance [J]. Plant Physiol, 2024, 197: kiae604. |
| [8] | Hu YF, Zhao HY, Xue LY, et al. IbMYC2 contributes to salt and drought stress tolerance via modulating anthocyanin accumulation and ROS-scavenging system in sweet potato [J]. Int J Mol Sci, 2024, 25(4): 2096. |
| [9] | Wang Y, Yang Y, Chen MK, et al. The MpbZIP46-MpERF105 module responds to ABA and ethylene signalling to promote anthocyanin synthesis in Malus ‘profusion’ under rust stress [J]. Plant Cell Environ, 2026, 49(1): 193-208. |
| [10] | Duan XY, Wang KR, Tang RK, et al. Recent advances in biosynthesis and regulation of strawberry anthocyanins [J]. Hortic Res, 2025, 12(8): uhaf135. |
| [11] | Golovko TK. Plant anthocyanins: structure, biosynthesis regulation, functions, and ecology [J]. Russ J Plant Physiol, 2023, 70(7): 161. |
| [12] | Adnan, Bao TT, Zheng X, et al. Functional characterization of 5-O-glycosyltranferase transforming 3-O anthocyanins into 3,5-O anthocyanins in Freesia hybrida [J]. Int J Mol Sci, 2025, 26(10): 4542. |
| [13] | Aravena-Calvo J, Busck-Mellor S, Laursen T. Global organization of phenylpropanoid and anthocyanin pathways revealed by proximity labeling of trans-cinnamic acid 4-hydroxylase in Petunia inflata petal protoplasts [J]. Front Plant Sci, 2024, 15: 1295750. |
| [14] | Alabd A, Ahmad M, Zhang X, et al. Light-responsive transcription factor PpWRKY44 induces anthocyanin accumulation by regulating PpMYB10 expression in pear [J]. Hortic Res, 2022, 9: uhac199. |
| [15] | Alabd A, Cheng HY, Ahmad M, et al. Abre-binding factor3-wrky dna-binding protein44 module promotes salinity-induced malate accumulation in pear [J]. Plant Physiol, 2023, 192(3): 1982-1996. |
| [16] | Hu DG, Sun CH, Ma QJ, et al. MdMYB1 regulates anthocyanin and malate accumulation by directly facilitating their transport into vacuoles in apples [J]. Plant Physiol, 2016, 170(3): 1315-1330. |
| [17] | Xie XB, Li S, Zhang RF, et al. The bHLH transcription factor MdbHLH3 promotes anthocyanin accumulation and fruit colouration in response to low temperature in apples [J]. Plant Cell Environ, 2012, 35(11): 1884-1897. |
| [18] | Yu JQ, Gu KD, Sun CH, et al. The apple bHLH transcription factor MdbHLH3 functions in determining the fruit carbohydrates and malate [J]. Plant Biotechnol J, 2021, 19(2): 285-299. |
| [19] | Cao XM, Su YK, Zhao T, et al. Multi-omics analysis unravels chemical roadmap and genetic basis for peach fruit aroma improvement [J]. Cell Rep, 2024, 43(8): 114623. |
| [20] | Oren-Shamir M. Does anthocyanin degradation play a significant role in determining pigment concentration in plants? [J]. Plant Sci, 2009, 177(4): 310-316. |
| [21] | Li LQ, Liu HY, Liu TY, et al. β-glucosidase: progress from basic mechanism to frontier application [J]. Fermentation, 2025, 11(10): 588. |
| [22] | Wang H, Zhang YJ, Feng XF, et al. Analysis of the β-glucosidase family reveals genes involved in the lignification of stone cells in Chinese white pear (Pyrus bretschneideri Rehd.) [J]. Front Plant Sci, 2022, 13: 852001. |
| [23] | Kongdin M, Mahong BC, Lee SK, et al. Action of multiple rice β-glucosidases on abscisic acid glucose ester [J]. Int J Mol Sci, 2021, 22(14): 7593. |
| [24] | Ma LJ, Liu X, Guo LW, et al. Discovery of plant chemical defence mediated by a two-component system involving β-glucosidase in Panax species [J]. Nat Commun, 2024, 15: 602. |
| [25] | Godse R, Fernandes JM, Kulkarni R. Characterization of β-glucosidase activity of a lactiplantibacillus plantarum 6-phospho-β-glucosidase [J]. Appl Microbiol Biotechnol, 2025, 109: 86. |
| [26] | Yuan KL, Wu GK, Li XS, et al. Anthocyanins degradation mediated by β-glycosidase contributes to the color loss during alcoholic fermentation in a structure-dependent manner [J]. Food Res Int, 2024, 175: 113732. |
| [27] | Yuan KL, Li XS, Zeng YY, et al. Chemical stability of carboxylpyranocyanidin-3-O-glucoside under β-glucosidase treatment and description of their interaction [J]. Food Chem, 2024, 447: 138840. |
| [28] | Sakamura S, Obata Y. Anthocyanase and anthocyanins occurring in eggplant, Solanum melongena L.(I) [J]. Agric Biol Chem, 1961, 25(10): 750-756. |
| [29] | Nadarajan S, Kumar V, Ovadia R, et al. Active anthocyanin degradation in Solanum macranthum flowers involves both peroxidase and β-glucosidase enzymes [J]. Physiol Plant, 2025, 177(6): e70603. |
| [30] | Barbagallo RN, Palmeri R, Fabiano S, et al. Characteristic of β-glucosidase from Sicilian blood oranges in relation to anthocyanin degradation [J]. Enzyme Microb Technol, 2007, 41(5): 570-575. |
| [31] | 庞学群, 黄雪梅, 杨晓棠, 等. 多酚氧化酶在荔枝果皮花色素苷降解中的作用 [J]. 中国农业科学, 2008, 41(2): 540-545. |
| Pang XQ, Huang XM, Yang XT, et al. Role of polyphenol oxidase in anthocyanin degradation of lychee pericarp [J]. Sci Agric Sin, 2008, 41(2): 540-545. | |
| [32] | Veitch NC. Horseradish peroxidase: a modern view of a classic enzyme [J]. Phytochemistry, 2004, 65(3): 249-259. |
| [33] | Freitas CDT, Costa JH, Germano TA, et al. Class III plant peroxidases: From classification to physiological functions [J]. Int J Biol Macromol, 2024, 263: 130306. |
| [34] | Mathé C, Barre A, Jourda C, et al. Evolution and expression of class III peroxidases [J]. Arch Biochem Biophys, 2010, 500(1): 58-65. |
| [35] | Passardi F, Penel C, Dunand C. Performing the paradoxical: how plant peroxidases modify the cell wall [J]. Trends Plant Sci, 2004, 9(11): 534-540. |
| [36] | Hiraga S, Sasaki K, Ito H, et al. A large family of class III plant peroxidases [J]. Plant Cell Physiol, 2001, 42(5): 462-468. |
| [37] | Quiroga M, Guerrero C, Botella MA, et al. A tomato peroxidase involved in the synthesis of lignin and suberin [J]. Plant Physiol, 2000, 122(4): 1119-1128. |
| [38] | Zhang ZQ, Pang XQ, Duan XW, et al. Role of peroxidase in anthocyanin degradation in litchi fruit pericarp [J]. Food Chem, 2005, 90(1/2): 47-52. |
| [39] | Sun TT, Wang MZ, Ren HF, et al. Comprehensive analysis of the physiological, metabolome, and transcriptome provided insights into anthocyanin biosynthesis and degradation of Malus crabapple [J]. Plant Physiol Biochem, 2025, 223: 109821. |
| [40] | Zipor G, Duarte P, Carqueijeiro I, et al. In planta anthocyanin degradation by a vacuolar class III peroxidase in Brunfelsia calycina flowers [J]. New Phytol, 2015, 205(2): 653-665. |
| [41] | Liu J, Wang YX, Zhang MH, et al. Color fading in lotus (Nelumbo nucifera) petals is manipulated both by anthocyanin biosynthesis reduction and active degradation [J]. Plant Physiol Biochem, 2022, 179: 100-107. |
| [42] | Yamada Y, Nakayama M, Shibata H, et al. Anthocyanin production and enzymatic degradation during the development of dark purple and lilac paprika fruit [J]. J Amer Soc Hort Sci, 2019, 144(5): 329-338. |
| [43] | Movahed N, Pastore C, Cellini A, et al. The grapevine VviPrx31 peroxidase as a candidate gene involved in anthocyanin degradation in ripening berries under high temperature [J]. J Plant Res, 2016, 129(3): 513-526. |
| [44] | Xu JH, Xiong L, Yao JL, et al. Hypermethylation in the promoter regions of flavonoid pathway genes is associated with skin color fading during ‘Daihong’ apple fruit development [J]. Hortic Res, 2024, 11(3): uhae031. |
| [45] | Zhao G, Li YN, Peng JL, et al. Anthocyanin degradation drives heat-induced petal fading in Chrysanthemum morifolium at full bloom: a multi-omics analysis [J]. Agriculture, 2025, 15(9): 950. |
| [46] | Zou H, Xiao Q, Li GC, et al. Revisiting the advancements in plant polyphenol oxidases research [J]. Sci Hortic, 2025, 341: 113960. |
| [47] | Zhou F, Lin HY, Luo Y, et al. Advances and application of polyphenol oxidase immobilization technology in plants [J]. Plants, 2025, 14(15): 2335. |
| [48] | Araji S, Grammer TA, Gertzen R, et al. Novel roles for the polyphenol oxidase enzyme in secondary metabolism and the regulation of cell death in walnut [J]. Plant Physiol, 2014, 164(3): 1191-1203. |
| [49] | Zhang J, Sun XL. Recent advances in polyphenol oxidase-mediated plant stress responses [J]. Phytochemistry, 2021, 181: 112588. |
| [50] | Zhang S. Recent advances of polyphenol oxidases in plants [J]. Molecules, 2023, 28(5): 2158. |
| [51] | Fronk P, Hartmann H, Bauer M, et al. Polyphenoloxidase from Riesling and Dornfelder wine grapes (Vitis vinifera) is a tyrosinase [J]. Food Chem, 2015, 183: 49-57. |
| [52] | Abbasi E. Melanin biosynthesis and functional roles in insects: insights into immunological defense, physiological regulation, and environmental adaptation [J]. Ann Med Surg, 2025, 87(12): 8643-8653. |
| [53] | Sullivan ML. Beyond brown: polyphenol oxidases as enzymes of plant specialized metabolism [J]. Front Plant Sci, 2015, 5: 783. |
| [54] | Prexler SM, Frassek M, Moerschbacher BM, et al. Catechol oxidase versus tyrosinase classification revisited by site-directed mutagenesis studies [J]. Angew Chem Int Ed, 2019, 58(26): 8757-8761. |
| [55] | Mayer AM, Staples RC. Laccase: new functions for an old enzyme [J]. Phytochemistry, 2002, 60(6): 551-565. |
| [56] | Munk L, Sitarz AK, Kalyani DC, et al. Can laccases catalyze bond cleavage in lignin? [J]. Biotechnol Adv, 2015, 33(1): 13-24. |
| [57] | Turlapati PV, Kim KW, Davin LB, et al. The laccase multigene family in Arabidopsis thaliana: towards addressing the mystery of their gene function(s) [J]. Planta, 2011, 233(3): 439-470. |
| [58] | Wang H, Zhong HX, Zhang FC, et al. Identification of grape laccase genes and their potential role in secondary metabolite synthesis [J]. Int J Mol Sci, 2024, 25(19): 10574. |
| [59] | Balarynová J, Klčová B, Sekaninová J, et al. The loss of polyphenol oxidase function is associated with hilum pigmentation and has been selected during pea domestication [J]. New Phytol, 2022, 235(5): 1807-1821. |
| [60] | Fan XT. Chemical inhibition of polyphenol oxidase and cut surface browning of fresh-cut apples [J]. Crit Rev Food Sci Nutr, 2023, 63(27): 8737-8751. |
| [61] | Li X, Wu XY, Bi JF, et al. Polyphenols accumulation effects on surface color variation in apple slices hot air drying process [J]. LWT, 2019, 108: 421-428. |
| [62] | Burke AE. Quantifying flesh browning, polyphenoloxidase, total phenolic content and vitamin C in select apple varieties and progeny [D]. Ithaca: Cornell University, 2010. |
| [63] | Taranto F, Pasqualone A, Mangini G, et al. Polyphenol oxidases in crops: biochemical, physiological and genetic aspects [J]. Int J Mol Sci, 2017, 18(2): 377. |
| [64] | Bisswanger H. Enzyme assays [J]. Perspect Sci, 2014, 1(1/2/3/4/5/6): 41-55. |
| [65] | Shrestha L, Kulig B, Moscetti R, et al. Optimisation of physical and chemical treatments to control browning development and enzymatic activity on fresh-cut apple slices [J]. Foods, 2020, 9(1): 76. |
| [66] | Fan J, Du W, Chen QL, et al. Comparative transcriptomic analyses provide insights into the enzymatic browning mechanism of fresh-cut sand pear fruit [J]. Horticulturae, 2021, 7(11): 502. |
| [67] | Wei JB, Zhang X, Zhong RH, et al. Laccase-mediated flavonoid polymerization leads to the pericarp browning of litchi fruit [J]. J Agric Food Chem, 2021, 69(50): 15218-15230. |
| [68] | Li CL, Qiu XX, Hou XM, et al. Polymerization of proanthocyanidins under the catalysis of miR397a-regulated laccases in Salvia miltiorrhiza and Populus trichocarpa [J]. Nat Commun, 2025, 16: 1513. |
| [69] | Tilley A, McHenry MP, McHenry JA, et al. Enzymatic browning: The role of substrates in polyphenol oxidase mediated browning [J]. Curr Res Food Sci, 2023, 7: 100623. |
| [70] | Kader F, Haluk JP, Nicolas JP, et al. Degradation of cyanidin 3-glucoside by blueberry polyphenol oxidase: kinetic studies and mechanisms [J]. J Agric Food Chem, 1998, 46(8): 3060-3065. |
| [71] | Shen YW, Wang S, Wan R, et al. Integrated transcriptomic and physiological analysis highlights the candidate genes regulating enzymatic browning, antioxidant capacity, and lipid metabolism in pomegranate aril browning [J]. Postharvest Biol Technol, 2026, 231: 113889. |
| [72] | Fang F, Zhang XL, Luo HH, et al. An intracellular laccase is responsible for the epicatechin mediated anthocyanin degradation in litchi fruit pericarp [J]. Plant Physiol, 2015: pp.00359.2015. |
| [73] | Zhao GP, Xiang FX, Zhang SC, et al. PbLAC4-like, activated by PbMYB26, related to the degradation of anthocyanin during color fading in pear [J]. BMC Plant Biol, 2021, 21: 469. |
| [74] | Giménez P, Just-Borràs A, Gombau J, et al. Effects of laccase from Botrytis cinerea on the oxidative degradation of anthocyanins [J]. OENO One, 2023, 57(3): 243-253. |
| [75] | Dong BY, Luo HH, Liu B, et al. BcXyl, a β-xylosidase isolated from Brunfelsia calycina flowers with anthocyanin-β-glycosidase activity [J]. Int J Mol Sci, 2019, 20(6): 1423. |
| [76] | Ren J, Liu ZY, Chen WS, et al. Anthocyanin degrading and chlorophyll accumulation lead to the formation of bicolor leaf in ornamental kale [J]. Int J Mol Sci, 2019, 20(3): 603. |
| [77] | Xue HK, Zhao JD, Wang Y, et al. Factors affecting the stability of anthocyanins and strategies for improving their stability: a review [J]. Food Chem X, 2024, 24: 101883. |
| [78] | Tkaczyńska A, Sendra E, Jiménez-Redondo N, et al. Studying the stability of anthocyanin pigments isolated from juices of colored-fleshed potatoes [J]. Int J Mol Sci, 2024, 25(20): 11116. |
| [79] | 孙建霞, 张燕, 胡小松, 等. 花色苷的结构稳定性与降解机制研究进展 [J]. 中国农业科学, 2009, 42(3): 996-1008. |
| Sun JX, Zhang Y, Hu XS, et al. Structural stability and degradation mechanisms of anthocyanins [J]. Sci Agric Sin, 2009, 42(3): 996-1008. | |
| [80] | 白新祥. 菊花花色形成的表型分析 [D]. 北京: 北京林业大学, 2007. |
| Bai XX. Phenotype analysis of flower coloration of Chrysanthemum×morifolium Ramat. [D]. Beijing: Beijing Forestry University, 2007. | |
| [81] | 赵昶灵, 陈俊愉, 刘雪兰, 等. 理化因素对梅花'南京红须'花色色素颜色呈现的效应 [J]. 南京林业大学学报: 自然科学版, 2004, 28(2): 27-32. |
| Zhao CL, Chen JY, Liu XL, et al. Effects of physical and chemical factors on the color expression of the flower color pigment of Prunus mume sieb.et zucc. ‘Nanjing hongxu’ (Nanjing red-bearded) [J]. J Nanjing For Univ, 2004, 28(2): 27-32. | |
| [82] | Zhu HF, Li XF, Zhai W, et al. Effects of low light on photosynthetic properties, antioxidant enzyme activity, and anthocyanin accumulation in purple pak-choi (Brassica campestris ssp. Chinensis Makino) [J]. PLoS One, 2017, 12(6): e0179305. |
| [83] | Liu YK, Zhang XW, Liu X, et al. Phytochrome interacting factor MdPIF7 modulates anthocyanin biosynthesis and hypocotyl growth in apple [J]. Plant Physiol, 2022, 188(4): 2342-2363. |
| [84] | Bu HD, Gu GJ, Hu YH, et al. Research advances in the synthesis and regulation of apple anthocyanins [J]. Biology, 2025, 14(10): 1322. |
| [85] | Li SR, Ou CQ, Wang F, et al. Ppbbx24-del mutant positively regulates light-induced anthocyanin accumulation in the ‘Red Zaosu’ pear (Pyrus pyrifolia White Pear Group) [J]. J Integr Agric, 2025, 24(7): 2619-2639. |
| [86] | Cao HW, Qu YY, Guo L, et al. Anthocyanin accumulation differences in European pears caused by phytochrome-interacting factor 3 (PcPIF3) promoter mutations under UV-B [J]. J Adv Res, 2026, 80: 125-136. |
| [87] | Du MN, Wang YN, Zhang XZ, et al. Integrated metabolomic and transcriptomic analyses elucidate anthocyanin-mediated flesh coloration mechanisms in red-fleshed pear [J]. Front Plant Sci, 2025, 16: 1670229. |
| [88] | 孙萍, 朱文灿, 林贤锐, 等. 基于代谢组和转录组解析多雨寡照对桃果皮着色和类黄酮积累的影响 [J]. 中国农业科学, 2025, 58(6): 1173-1194. |
| Sun P, Zhu WC, Lin XR, et al. Effects of rainy and low light conditions on coloration and flavonoid accumulation in peach peel based on metabolomic and transcriptomic analyses [J]. Sci Agric Sin, 2025, 58(6): 1173-1194. | |
| [89] | Chen QQ, Man YP, Lu XM, et al. High temperature affects synthesis and degradation of anthocyanin in red-fleshed kiwifruit (Actinidia chinensis): pathway and kinetics analysis [J]. Plant Physiol Biochem, 2026, 230: 110936. |
| [90] | Zhou M, Chen QQ, Bi JF, et al. Degradation kinetics of cyanidin 3-O-glucoside and cyanidin 3-O-rutinoside during hot air and vacuum drying in mulberry (Morus alba L.) fruit: a comparative study based on solid food system [J]. Food Chem, 2017, 229: 574-579. |
| [91] | Gouot JC, Smith JP, Holzapfel BP, et al. Grape berry flavonoids: a review of their biochemical responses to high and extreme high temperatures [J]. J Exp Bot, 2019, 70(2): 397-423. |
| [92] | Kui LW, Micheletti D, Palmer J, et al. High temperature reduces apple fruit colour via modulation of the anthocyanin regulatory complex [J]. Plant Cell Environ, 2011, 34(7): 1176-1190. |
| [93] | Zhang LP, Wang L, Zeng XG, et al. Comparative transcriptome analysis reveals fruit discoloration mechanisms in postharvest strawberries in response to high ambient temperature [J]. Food Chem X, 2019, 2: 100025. |
| [94] | Niu JP, Zhang GJ, Zhang WT, et al. Anthocyanin concentration depends on the counterbalance between its synthesis and degradation in plum fruit at high temperature [J]. Sci Rep, 2017, 7: 7684. |
| [1] | YIN Yue, QIN Xiao-ya, MI Jia, AN Wei, HE Jun, ZHANG Feng-feng. Identification of FBN Gene Family and Its Relationship with Carotenoids Metabolism in Lyciumbarbarum [J]. Biotechnology Bulletin, 2026, 42(3): 338-348. |
| [2] | LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development [J]. Biotechnology Bulletin, 2026, 42(3): 362-373. |
| [3] | XU Ze, ZHOU Chen-ping, KUANG Rui-bin, WU Xia-ming, YANG Min, LIU Chuan-he, HE Han, WEI Yue-rong. Identification of PG Gene Family and Their Roles in Papaya Fruit Softening [J]. Biotechnology Bulletin, 2026, 42(3): 349-361. |
| [4] | LUO Wei, GONG Ao, ZHONG Yang, HU Di, ZHOU Hong-yuan, ZHANG Hong-xin, AI Ju, LUO You-wei, GAO Dong-li. Pleiotropic Effects of SEPALLATA2 Knock-out on Fruit and Wart Development in Cucumber [J]. Biotechnology Bulletin, 2026, 42(3): 283-293. |
| [5] | FU Wan-xiang, WANG Tao, TAN Shu, YU Si-yuan, XIONG Huan, ZOU Feng. Effects of Exogenous Sugar on the Fruit Quality, Amylase Activity and Gene Expression of Castanea henryi [J]. Biotechnology Bulletin, 2026, 42(3): 374-382. |
| [6] | LIU Lin-ya, LIU Huan-yan, LIANG Xin-yu, SONG Shu-yi, HE Bin, WANG Xu-ying, HUANG Ya-cheng. Genome-wide Identification and Expression Analysis of BGAL Gene Family in Actinidiachinensis var. Hongyang [J]. Biotechnology Bulletin, 2026, 42(3): 312-323. |
| [7] | LIU Miao, LIN Tao, JIA Le-song, HU Feng, LI Tao, LI Zhi-wan, LIU Mei-fang, ZHENG Fang-yan, CUI Long. From Wild to Cultivated: Evolution and Regulatory Mechanisms of Tomato Fruit Color [J]. Biotechnology Bulletin, 2026, 42(3): 187-202. |
| [8] | WANG He-yao, SUN Hong-mei. Research Progress in the Function and Formation Mechanism of Trichomes in Horticultural Plants [J]. Biotechnology Bulletin, 2026, 42(3): 242-254. |
| [9] | YAN Chen-lin, LI Fan, YAN Chun-ting, CHENG Jiao-wen, HU Kai-lin, YE Zhi-biao, SONG Jian-wen. Advances in Genes Related to Tomato Fruit Morphogenesis [J]. Biotechnology Bulletin, 2026, 42(3): 172-186. |
| [10] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [11] | HU Qiu-ling, CHEN Ling, HUANG Jia-yi, ZHAO Zi-qiao, PAN Lu-yi, LIU Hui-li, LIU Tai-bo. Advances in the Regulation of Fruit Development by Polyamines [J]. Biotechnology Bulletin, 2026, 42(3): 203-212. |
| [12] | MA Shi-jie, LI Zheng, LI Wei, GUO Yang-dong, ZHANG Na. Research Progress in Light Signaling Regulation of Fruit Development in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 5-18. |
| [13] | CUI Zhi-han, WEI Qing-zhen, HU Na, BAO Chong-lai, WANG Hua-sen. Research Progress in IQD Genes in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 230-241. |
| [14] | ZHANG Gao-xiang, WU Yu-bi, GUO Ya-jing, JI Wei, YANG Zhong-yi. Identification and Expression Analysis of WD40 Gene Family in Grape [J]. Biotechnology Bulletin, 2026, 42(3): 324-337. |
| [15] | LUO Long-xin, LI Zhi, LI Tong, FENG Zi-quan, ZHAI Xin-yue, LIANG Cheng-lin, ZHANG Ya-li, WU Shang, LI Yuan-yuan, JIANG Han. Molecular Basis of Sugar Accumulation in Apple Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 156-171. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||