








生物技术通报 ›› 2022, Vol. 38 ›› Issue (3): 234-245.doi: 10.13560/j.cnki.biotech.bull.1985.2021-0341
收稿日期:2021-03-18
出版日期:2022-03-26
发布日期:2022-04-06
作者简介:孙曼銮,女,博士,讲师,研究方向:微生物与分子酶学;E-mail: 基金资助:
SUN Man-luan1( ), GE Sai2, BU Jia1, ZHU Zhuang-yan1
), GE Sai2, BU Jia1, ZHU Zhuang-yan1
Received:2021-03-18
Published:2022-03-26
Online:2022-04-06
摘要:
核糖核酸酶参与体内多种RNA代谢反应,对细菌生理功能调节起重要作用。细菌需要进化出多种策略来对体内核糖核酸酶进行调节,以避免对RNA进行不必要地降解。目前对大肠杆菌核糖核酸酶调节机制的研究包括转录后调控、翻译后修饰、细胞定位及相关抑制剂等。本文系统性地阐述了大肠杆菌核糖核酸酶的分类、功能及其体内调控机制,总结了不同环境压力下大肠杆菌对自身核糖核酸酶进行适应性调节的响应机制及存在的问题。
孙曼銮, 葛赛, 卜佳, 朱壮彦. 大肠杆菌核糖核酸酶调控机制研究[J]. 生物技术通报, 2022, 38(3): 234-245.
SUN Man-luan, GE Sai, BU Jia, ZHU Zhuang-yan. Regulation Mechanism of Ribonucleases in Escherichia coli[J]. Biotechnology Bulletin, 2022, 38(3): 234-245.

图1 大肠杆菌体内RNase参与的RNA剪接与降解过程 A:核糖核酸内/外切酶共同参与的RNA剪接;B:sRNA介导的mRNA降解;C:核糖核酸内/外切酶共同参与的RNA降解
Fig.1 RNase-dependent RNA processing and degradation in Escherichia coli A:RNA processing mediated by endoribonucleas/exoribonuclease. B:sRNA-mediated mRNA degradation. C:RNA degradation mediated by endoribonuclease/exoribonuclease
| 名称 Name | 编码基因 Gene | 酶活特异性 Enzyme activity | 主要生理功能 Physiological functions | 已知主要调控方式 Regulation known | 主要参考文献 Reference | |
|---|---|---|---|---|---|---|
| 核 糖 核 酸 内 切 酶 | RNAse I | rna | 内切酶活性,Ca2+依赖的双链mRNA活性 | 尚不明确 | 可能与周质空间及胞内定位调节有关 | [ |
| RNAse III | rnc | 双链特异性内切酶活性 | rRNA的成熟 | 转录后自我调控;酶蛋白磷酸化修饰;反式作用因子调控 | [ | |
| RNase BN | rbn | 内切酶/外切酶活性 | sRNA的降解,mRNA降解 | 自身mRNA稳定性调节 | [ | |
| RNase E | rne | 内切酶活性 | mRNA的降解;rRNA及tRNA的成熟 | 转录后自我调控;酶蛋白磷酸化修饰;反式作用因子调控;sRNA;小分子结合及胞内定位 | [ | |
| RNase P | rnpA/rnpB | 内切酶活性 | 5' tRNA的成熟 | 未知 | [ | |
| RNase G | rng | 内切酶活性 | rRNA的成熟 | 未知 | [ | |
| RNase H家族 | rnhA/rnhB | RNA/DNA杂交体中RNA的切割与修饰 | 参与DNA复制过程中RNA的调整 | 未知 | [ | |
| YbeY | ybeY | 内切酶活性 | rRNA成熟与质控,16S rRNA3'末端的加工 | 未知 | [ | |
| RNase LS | rnlA/rnlB | 内切酶活性 | 降解T4噬菌体编码的mRNA | 未知 | [ | |
| 核 糖 核 酸 外 切 酶 | PNPase | pnp | 磷酸盐依赖的3'-5'外切酶活性 | mRNA的降解;rRNA及tRNA的成熟;sRNAs调控伴侣 | 自我调控;反式作用因子调控;sRNA的调控 | [ |
| RNAse II | rnb | 3'-5'外切酶活性 | mRNA的降解;rRNA及tRNA的成熟 | 酶蛋白的乙酰化修饰 | [ | |
| RNase R | rnr | 3'-5'外切酶活性 | mRNA的降解;rRNA的成熟与质控;tRNA的成熟 | 酶蛋白的乙酰化修饰 | [ | |
| RNase PH | rph | 磷酸盐依赖的3'-5'外切酶活性 | tRNA与rRNA的成熟;rRNA的降解 | 蛋白稳定性调节;RNase II调节(待发表资料) | [ | |
| RNase T | rnt | 3'-5'外切酶活性 | tRNA与rRNA的成熟 | 未知 | [ | |
| RNase D | rnd | 3'-5'外切酶活性 | tRNA的成熟 | 未知 | [ | |
| Oligo RNase | orn | 3'-5'外切酶活性 | 短链核苷酸的去除 | 未知 | [ | |
| RNase AM | trpH/yciV | 5'-3'外切酶活性 | 5S、23S 及16S rRNA的5'末端成熟 | 未知 | [ | |
表1 大肠杆菌RNase种类及其调控机制
Table 1 Properties and regulations of RNases in Escherichia coli
| 名称 Name | 编码基因 Gene | 酶活特异性 Enzyme activity | 主要生理功能 Physiological functions | 已知主要调控方式 Regulation known | 主要参考文献 Reference | |
|---|---|---|---|---|---|---|
| 核 糖 核 酸 内 切 酶 | RNAse I | rna | 内切酶活性,Ca2+依赖的双链mRNA活性 | 尚不明确 | 可能与周质空间及胞内定位调节有关 | [ |
| RNAse III | rnc | 双链特异性内切酶活性 | rRNA的成熟 | 转录后自我调控;酶蛋白磷酸化修饰;反式作用因子调控 | [ | |
| RNase BN | rbn | 内切酶/外切酶活性 | sRNA的降解,mRNA降解 | 自身mRNA稳定性调节 | [ | |
| RNase E | rne | 内切酶活性 | mRNA的降解;rRNA及tRNA的成熟 | 转录后自我调控;酶蛋白磷酸化修饰;反式作用因子调控;sRNA;小分子结合及胞内定位 | [ | |
| RNase P | rnpA/rnpB | 内切酶活性 | 5' tRNA的成熟 | 未知 | [ | |
| RNase G | rng | 内切酶活性 | rRNA的成熟 | 未知 | [ | |
| RNase H家族 | rnhA/rnhB | RNA/DNA杂交体中RNA的切割与修饰 | 参与DNA复制过程中RNA的调整 | 未知 | [ | |
| YbeY | ybeY | 内切酶活性 | rRNA成熟与质控,16S rRNA3'末端的加工 | 未知 | [ | |
| RNase LS | rnlA/rnlB | 内切酶活性 | 降解T4噬菌体编码的mRNA | 未知 | [ | |
| 核 糖 核 酸 外 切 酶 | PNPase | pnp | 磷酸盐依赖的3'-5'外切酶活性 | mRNA的降解;rRNA及tRNA的成熟;sRNAs调控伴侣 | 自我调控;反式作用因子调控;sRNA的调控 | [ |
| RNAse II | rnb | 3'-5'外切酶活性 | mRNA的降解;rRNA及tRNA的成熟 | 酶蛋白的乙酰化修饰 | [ | |
| RNase R | rnr | 3'-5'外切酶活性 | mRNA的降解;rRNA的成熟与质控;tRNA的成熟 | 酶蛋白的乙酰化修饰 | [ | |
| RNase PH | rph | 磷酸盐依赖的3'-5'外切酶活性 | tRNA与rRNA的成熟;rRNA的降解 | 蛋白稳定性调节;RNase II调节(待发表资料) | [ | |
| RNase T | rnt | 3'-5'外切酶活性 | tRNA与rRNA的成熟 | 未知 | [ | |
| RNase D | rnd | 3'-5'外切酶活性 | tRNA的成熟 | 未知 | [ | |
| Oligo RNase | orn | 3'-5'外切酶活性 | 短链核苷酸的去除 | 未知 | [ | |
| RNase AM | trpH/yciV | 5'-3'外切酶活性 | 5S、23S 及16S rRNA的5'末端成熟 | 未知 | [ | |

图2 大肠杆菌RNase E调控机制 A:自我调控;B:翻译后修饰(磷酸化)调节;C:sRNA-Hfq调节;D:调节蛋白调控;E:细胞膜定位调控;F:其它调控(小分子、酶及抗生素调节)
Fig. 2 Regulation mechanisms of RNase E in E. coli A:Autoregulation of RNase E. B:Regulation of RNase E by post-translational modification(phosphorylation). C:sRNA-Hfq regulation. D:RNase E regulated by regulator proteins. E:RNase E regulated by localization. F:Other regulations(by small molecules,enzymes,and antibiotics)
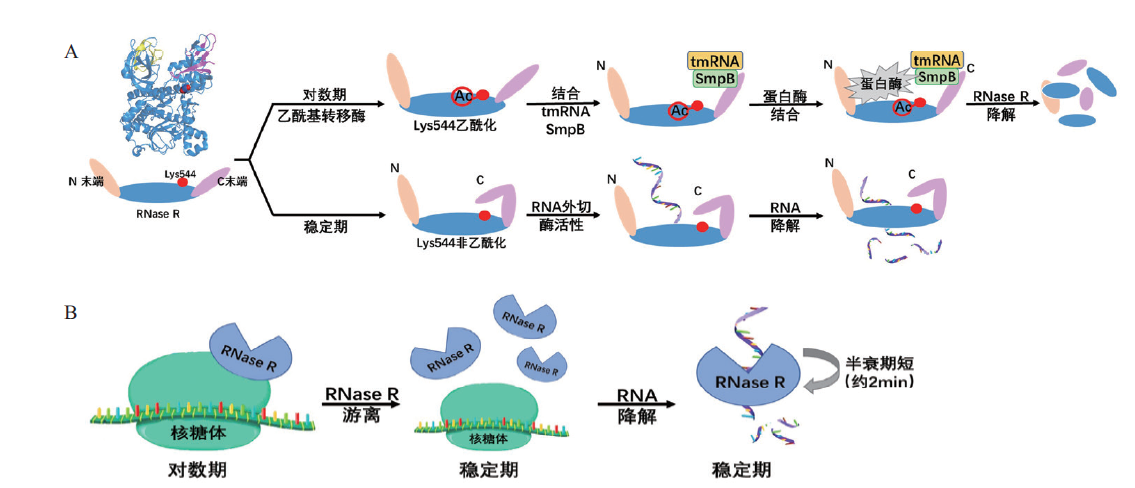
图4 大肠杆菌RNase R调控机制 A:翻译后修饰(乙酰化)调控RNase R蛋白稳定性;B:核糖体结合形式影响RNase R活性及稳定性
Fig. 4 Regulation mechanisms of RNase R in E. coli A:RNase R stability affected by post-translational modification(acetylation). B:Ribosome binding forms affect the activity and stability of RNase R
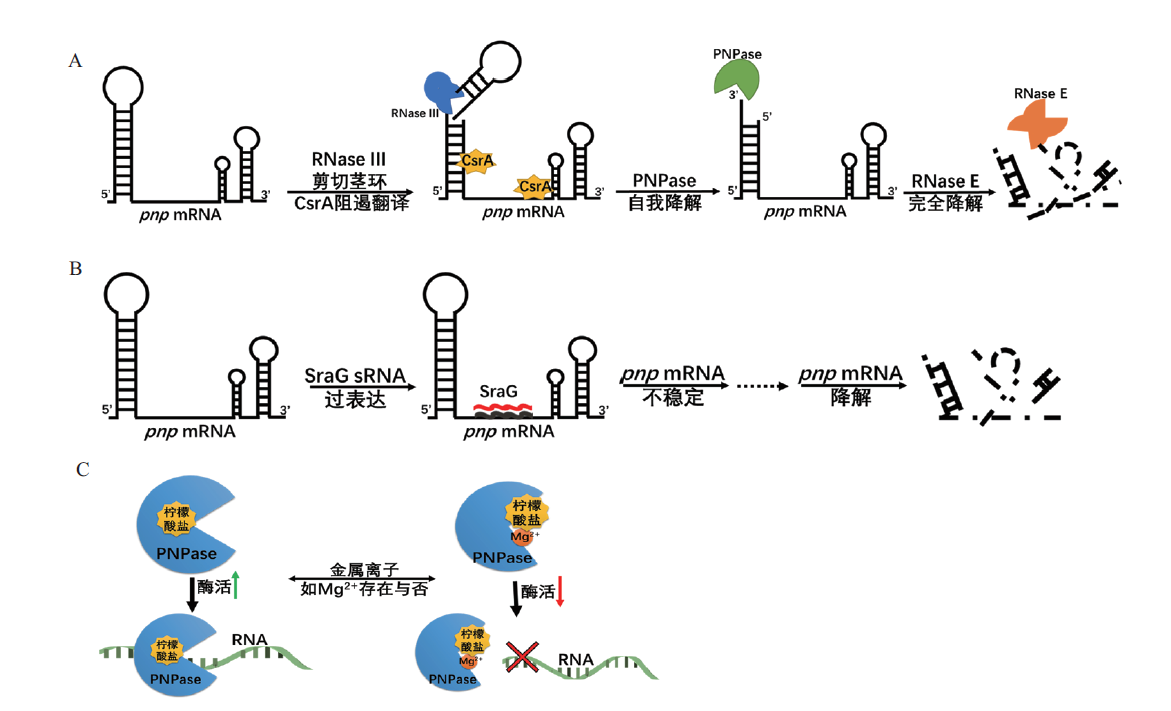
图5 大肠杆菌PNPase调控机制 A:PNPase的自我调控;B:sRNA SraG介导的PNPase调节;C:柠檬酸盐对PNPase活性的调节
Fig. 5 Regulation mechanisms of PNPase in E. coli A:Autoregulation of PNPase. B:sRNA SraG mediated regulation of PNPase. C:Activity of PNPase modulated by citrate
| [1] |
Deutscher MP. Twenty years of bacterial RNases and RNA processing:how we’ve matured[J]. RNA, 2015, 21(4):597-600.
doi: 10.1261/rna.049692.115 pmid: 25780155 |
| [2] | de Trinquier A, Durand S, Braun F, et al. Regulation of RNA processing and degradation in bacteria[J]. BBA Gene Regul Mech, 2020, 1863(5):194505. |
| [3] |
Canestrari E, Paroo Z. Ribonucleases as drug targets[J]. Trends Pharmacol Sci, 2018, 39(10):855-866.
doi: S0165-6147(18)30121-4 pmid: 30144949 |
| [4] |
Vargas-Blanco DA, Shell SS. Regulation of mRNA stability during bacterial stress responses[J]. Front Microbiol, 2020, 11:2111.
doi: 10.3389/fmicb.2020.02111 pmid: 33013770 |
| [5] |
Barrangou R, Fremaux C, Deveau H, et al. CRISPR provides acquired resistance against viruses in prokaryotes[J]. Science, 2007, 315(5819):1709-1712.
doi: 10.1126/science.1138140 pmid: 17379808 |
| [6] | Roux C, Etienne TA, Hajnsdorf E, et al. The essential role of mRNA degradation in understanding and engineering E. coli metabolism[J]. Biotechnol Adv, 2021: 107805. |
| [7] | Stenum TS, Holmqvist E. CsrA enters Hfq's territory:Regulation of a base-pairing small RNA[J]. Mol Microbiol, 2021:mmi. 14785. |
| [8] |
Bechhofer DH, Deutscher MP. Bacterial ribonucleases and their roles in RNA metabolism[J]. Crit Rev Biochem Mol Biol, 2019, 54(3):242-300.
doi: 10.1080/10409238.2019.1651816 URL |
| [9] |
Zhu LQ, Gangopadhyay T, Padmanabha KP, et al. Escherichia coli RNA gene encoding RNase I:cloning, overexpression, subcellular distribution of the enzyme, and use of an RNA deletion to identify additional RNases[J]. J Bacteriol, 1990, 172(6):3146-3151.
pmid: 2188952 |
| [10] |
Altuvia Y, Bar A, Reiss N, et al. In vivo cleavage rules and target repertoire of RNase III in Escherichia coli[J]. Nucleic Acids Res, 2018, 46(19):10380-10394.
doi: 10.1093/nar/gky684 pmid: 30113670 |
| [11] |
Lee M, Joo M, Sim M, et al. The coordinated action of RNase III and RNase G controls enolase expression in response to oxygen availability in Escherichia coli[J]. Sci Rep, 2019, 9(1):17257.
doi: 10.1038/s41598-019-53883-y URL |
| [12] | Altuvia S, Storz G, Papenfort K. Cross-regulation between bacteria and phages at a posttranscriptional level[J]. Microbiol Spectr, 2018, 6(4):10. 1128. |
| [13] |
Chu LY, Hsieh TJ, Golzarroshan B, et al. Structural insights into RNA unwinding and degradation by RNase R[J]. Nucleic Acids Res, 2017, 45(20):12015-12024.
doi: 10.1093/nar/gkx880 URL |
| [14] |
Perwez T, Kushner SR. RNase Z in Escherichia coli plays a significant role in mRNA decay[J]. Mol Microbiol, 2006, 60(3):723-737.
pmid: 16629673 |
| [15] |
Chen H, Previero A, Deutscher MP. A novel mechanism of ribonuclease regulation:GcvB and Hfq stabilize the mRNA that encodes RNase BN/Z during exponential phase[J]. J Biol Chem, 2019, 294(52):19997-20008.
doi: 10.1074/jbc.RA119.011367 pmid: 31744883 |
| [16] |
Bandyra KJ, Luisi BF. RNase E and the high-fidelity orchestration of RNA metabolism[J]. Microbiol Spectr, 2018, 6(2). DOI: 10.1128/microbiolspec.rwr-0008-2017.
doi: 10.1128/microbiolspec.rwr-0008-2017 |
| [17] |
Ali N, Gowrishankar J. Cross-subunit catalysis and a new phenomenon of recessive resurrection in Escherichia coli RNase E[J]. Nucleic Acids Res, 2020, 48(2):847-861.
doi: 10.1093/nar/gkz1152 URL |
| [18] |
Hadjeras L, Poljak L, Bouvier M, et al. Detachment of the RNA degradosome from the inner membrane of Escherichia coli results in a global slowdown of mRNA degradation, proteolysis of RNase E and increased turnover of ribosome-free transcripts[J]. Mol Microbiol, 2019, 111(6):1715-1731.
doi: 10.1111/mmi.14248 pmid: 30903628 |
| [19] | Mardle CE, Goddard LR, Spelman BC, et al. Identification and analysis of novel small molecule inhibitors of RNase E:Implications for antibacterial targeting and regulation of RNase E[J]. Biochem Biophys Rep, 2020, 23:100773. |
| [20] |
Moore CJ, Go H, Shin E, et al. Substrate-dependent effects of quaternary structure on RNase E activity[J]. Genes Dev, 2021, 35(3/4):286-299.
doi: 10.1101/gad.335828.119 URL |
| [21] |
Hamouche L, Poljak L, Carpousis AJ. Ribosomal RNA degradation induced by the bacterial RNA polymerase inhibitor rifampicin[J]. bioRxiv, 2021. DOI: 10.1101/2021.04.24.441238.
doi: 10.1101/2021.04.24.441238 |
| [22] |
Mohanty BK, Agrawal A, Kushner SR. Generation of pre-tRNAs from polycistronic operons is the essential function of RNase P in Escherichia coli[J]. Nucleic Acids Res, 2020, 48(5):2564-2578.
doi: 10.1093/nar/gkz1188 pmid: 31993626 |
| [23] |
Kurata T, Nakanishi S, Hashimoto M, et al. Subunit composition of ribosome in the yqgF mutant Is deficient in pre-16S rRNA processing of Escherichia coli[J]. J Mol Microbiol Biotechnol, 2018, 28(4):179-182.
doi: 10.1159/000494494 URL |
| [24] |
Kouzminova EA, Kuzminov A. Ultraviolet-induced RNA:DNA hybrids interfere with chromosomal DNA synjournal[J]. Nucleic Acids Res, 2021, 49(7):3888-3906.
doi: 10.1093/nar/gkab147 pmid: 33693789 |
| [25] |
Prossliner T, Gerdes K, Sørensen MA, et al. Hibernation factors directly block ribonucleases from entering the ribosome in response to starvation[J]. Nucleic Acids Res, 2021, 49(4):2226-2239.
doi: 10.1093/nar/gkab017 pmid: 33503254 |
| [26] |
Garcia-Rodriguez G, Charlier D, Wilmaerts D, et al. Alternative dimerization is required for activity and inhibition of the HEPN ribonuclease RnlA[J]. Nucleic Acids Res, 2021, 49(12):7164-7178.
doi: 10.1093/nar/gkab513 pmid: 34139012 |
| [27] | Dendooven T, Sinha D, Roeselová A, et al. A cooperative PNPase-Hfq-RNA carrier complex facilitates bacterial riboregulation[J]. Mol Cell, 2021, 81(14):2901- 2913. e5. |
| [28] |
Dressaire C, Pobre V, Laguerre S, et al. PNPase is involved in the coordination of mRNA degradation and expression in stationary phase cells of Escherichia coli[J]. BMC Genomics, 2018, 19(1):848.
doi: 10.1186/s12864-018-5259-8 URL |
| [29] |
Park H, Yakhnin H, Connolly M, et al. CsrA participates in a PNPase autoregulatory mechanism by selectively repressing translation of pnp transcripts that have been previously processed by RNase III and PNPase[J]. J Bacteriol, 2015, 197(24):3751-3759.
doi: 10.1128/JB.00721-15 URL |
| [30] |
Fontaine F, Gasiorowski E, Gracia C, et al. The small RNA SraG participates in PNPase homeostasis[J]. RNA, 2016, 22(10):1560-1573.
doi: 10.1261/rna.055236.115 pmid: 27495318 |
| [31] |
Cairrão F, Chora A, Zilhão R, et al. RNase II levels change according to the growth conditions:characterization of gmr, a new Escherichia coli gene involved in the modulation of RNase II[J]. Mol Microbiol, 2001, 39(6):1550-1561.
pmid: 11260472 |
| [32] |
Lu F, Taghbalout A. Membrane association via an amino-terminal amphipathic helix is required for the cellular organization and function of RNase II[J]. J Biol Chem, 2013, 288(10):7241-7251.
doi: 10.1074/jbc.M112.408674 URL |
| [33] |
Song L, Wang G, Malhotra A, et al. Reversible acetylation on Lys501 regulates the activity of RNase II[J]. Nucleic Acids Res, 2016, 44(5):1979-1988.
doi: 10.1093/nar/gkw053 URL |
| [34] | dos Santos RF, Andrade JM, Pissarra J, et al. Hfq and RNase R mediate rRNA processing and degradation in a novel RNA quality control process[J]. mBio, 2020, 11(5):e02398-20. |
| [35] |
Liang W, Deutscher MP. Ribosomes regulate the stability and action of the exoribonuclease RNase R[J]. J Biol Chem, 2013, 288(48):34791-34798.
doi: 10.1074/jbc.M113.519553 URL |
| [36] |
Martínez VP, Dehò G, Simons RW, et al. Ribonuclease PH interacts with an acidic ribonuclease E site through a basic 80-amino acid domain[J]. FEMS Microbiol Lett, 2014, 355(1):51-60.
doi: 10.1111/fml.2014.355.issue-1 URL |
| [37] |
Sulthana S, Quesada E, Deutscher MP. RNase II regulates RNase PH and is essential for cell survival during starvation and stationary phase[J]. RNA, 2017, 23(9):1456-1464.
doi: 10.1261/rna.060558.116 pmid: 28625967 |
| [38] |
Zuo Y, Deutscher MP. The physiological role of RNase T can be explained by its unusual substrate specificity[J]. J Biol Chem, 2002, 277(33):29654-29661.
doi: 10.1074/jbc.M204252200 URL |
| [39] |
Jain C. RNase AM, a 5' to 3' exonuclease, matures the 5' end of all three ribosomal RNAs in E. coli[J]. Nucleic Acids Res, 2020, 48(10):5616-5623.
doi: 10.1093/nar/gkaa260 URL |
| [40] |
Mackie GA. RNase E:at the interface of bacterial RNA processing and decay[J]. Nat Rev Microbiol, 2013, 11(1):45-57.
doi: 10.1038/nrmicro2930 URL |
| [41] |
Mohanty BK, Petree JR, Kushner SR. Endonucleolytic cleavages by RNase E generate the mature 3' termini of the three proline tRNAs in Escherichia coli[J]. Nucleic Acids Res, 2016, 44(13):6350-6362.
doi: 10.1093/nar/gkw517 pmid: 27288443 |
| [42] |
Sulthana S, Basturea GN, Deutscher MP. Elucidation of pathways of ribosomal RNA degradation:an essential role for RNase E[J]. RNA, 2016, 22(8):1163-1171.
doi: 10.1261/rna.056275.116 pmid: 27298395 |
| [43] |
Ow MC, Kushner SR. Initiation of tRNA maturation by RNase E is essential for cell viability in E. coli[J]. Genes Dev, 2002, 16(9):1102-1115.
doi: 10.1101/gad.983502 URL |
| [44] |
Sousa S, Marchand I, Dreyfus M. Autoregulation allows Escherichia coli RNase E to adjust continuously its synjournal to that of its substrates[J]. Mol Microbiol, 2001, 42(3):867-878.
pmid: 11722748 |
| [45] |
Jain C, Deana A, Belasco JG. Consequences of RNase E scarcity in Escherichia coli[J]. Mol Microbiol, 2002, 43(4):1053-1064.
doi: 10.1046/j.1365-2958.2002.02808.x URL |
| [46] |
Mohanty BK, Kushner SR. Polyadenylation of Escherichia coli transcripts plays an integral role in regulating intracellular levels of polynucleotide phosphorylase and RNase E[J]. Mol Microbiol, 2002, 45(5):1315-1324.
pmid: 12207699 |
| [47] |
Jaso-Vera ME, Domínguez-Malfavón L, Curiel-Quesada E, et al. Dynamics of the canonical RNA degradosome components during glucose stress[J]. Biochimie, 2021, 187:67-74.
doi: 10.1016/j.biochi.2021.05.006 pmid: 34022290 |
| [48] |
McQuail J, Carpousis AJ, Wigneshweraraj S. The association between Hfq and RNase E in long-term nitrogen starved Escherichia coli[J]. bioRxiv, 2021. DOI: 10.1101/2021.04, 19. 440462.
doi: 10.1101/2021.04 |
| [49] |
Singh D, Murashko ON, Lin-Chao SE. Posttranscriptional regulation of tnaA by protein-RNA interaction mediated by ribosomal protein L4 in Escherichia coli[J]. J Bacteriol, 2020, 202(10):e00799-19. DOI: 10.1128/jb.00799-19.
doi: 10.1128/jb.00799-19 |
| [50] |
Lee K, Zhan X, Gao J, et al. RraA. a protein inhibitor of RNase E activity that globally modulates RNA abundance in E. coli[J]. Cell, 2003, 114(5):623-634.
doi: 10.1016/j.cell.2003.08.003 URL |
| [51] | Lim B, Sim M, Lee H, et al. Regulation of Escherichia coli RNase III activity[J]. J Microbiol Seoul Korea, 2015, 53(8):487-494. |
| [52] |
Sirdeshmukh R, Schlessinger D. Ordered processing of Escherichia coli 23S rRNA in vitro[J]. Nucleic Acids Res, 1985, 13(14):5041-5054.
pmid: 2991850 |
| [53] |
King TC, Sirdeshmukh R, Schlessinger D. RNase III cleavage is obligate for maturation but not for function of Escherichia coli pre-23S rRNA[J]. PNAS, 1984, 81(1):185-188.
pmid: 6364133 |
| [54] |
Stead MB, Marshburn S, Mohanty BK, et al. Analysis of Escherichia coli RNase E and RNase III activity in vivo using tiling microarrays[J]. Nucleic Acids Res, 2011, 39(8):3188-3203.
doi: 10.1093/nar/gkq1242 pmid: 21149258 |
| [55] |
Conrad C, Rauhut R. Ribonuclease III:new sense from nuisance[J]. Int J Biochem Cell Biol, 2002, 34(2):116-129.
doi: 10.1016/S1357-2725(01)00112-1 URL |
| [56] |
Deutscher MP. Regulation of bacterial ribonucleases[J]. Annu Rev Microbiol, 2021, 75(1). DOI: 10.1146/annurev-micro-020121-011201.
doi: 10.1146/annurev-micro-020121-011201 |
| [57] |
Lee J, Lee M, Lee K. Trans-acting regulators of ribonuclease activity[J]. J Microbiol, 2021, 59(4):341-359.
doi: 10.1007/s12275-021-0650-6 URL |
| [58] |
Paudyal S, Alfonso-Prieto M, Carnevale V, et al. Combined computational and experimental analysis of a complex of ribonuclease III and the regulatory macrodomain protein, YmdB[J]. Proteins, 2015, 83(3):459-472.
doi: 10.1002/prot.v83.3 URL |
| [59] |
Kavalchuk K, Madhusudan S, Schnetz K. RNase III initiates rapid degradation of proU mRNA upon hypo-osmotic stress in Escherichia coli[J]. RNA Biol, 2012, 9(1):98-109.
doi: 10.4161/rna.9.1.18228 URL |
| [60] |
Song W, Kim YH, Sim SH, et al. Antibiotic stress-induced modulation of the endoribonucleolytic activity of RNase III and RNase G confers resistance to aminoglycoside antibiotics in Escherichia coli[J]. Nucleic Acids Res, 2014, 42(7):4669-4681.
doi: 10.1093/nar/gku093 URL |
| [61] |
Li de la Sierra-Gallay I, Pellegrini O, Condon C. Structural basis for substrate binding, cleavage and allostery in the tRNA maturase RNase Z[J]. Nature, 2005, 433(7026):657-661.
doi: 10.1038/nature03284 URL |
| [62] |
Chen H, Dutta T, Deutscher MP. Growth phase-dependent variation of RNase BN/Z affects small RNAs[J]. J Biol Chem, 2016, 291(51):26435-26442.
pmid: 27875308 |
| [63] |
Gupta AK, Siddiqui N, Dutta T. A novel mechanism of RyeA/SraC induction under acid stress[J]. Biochem Biophys Res Commun, 2020, 525(2):298-302.
doi: 10.1016/j.bbrc.2020.02.085 URL |
| [64] |
Grünberg S, Coxam B, Chen TH, et al. E. coli RNase I exhibits a strong Ca2+-dependent inherent double-stranded RNase activity[J]. Nucleic Acids Res, 2021, 49(9):5265-5277.
doi: 10.1093/nar/gkab284 URL |
| [65] |
Fontaine BM, Martin KS, Garcia-Rodriguez JM, et al. RNase I regulates Escherichia coli 2', 3'-cyclic nucleotide monophosphate levels and biofilm formation[J]. Biochem J, 2018, 475(8):1491-1506.
doi: 10.1042/BCJ20170906 URL |
| [66] |
Hossain ST, Malhotra A, Deutscher MP. How RNase R degrades structured rna:role of the helicase activity and the S1 domain[J]. J Biol Chem, 2016, 291(15):7877-7887.
doi: 10.1074/jbc.M116.717991 URL |
| [67] |
Cheng ZF, Deutscher MP. Purification and characterization of the Escherichia coli exoribonuclease RNase R. Comparison with RNase II[J]. J Biol Chem, 2002, 277(24):21624-21629.
doi: 10.1074/jbc.M202942200 URL |
| [68] |
Chen C, Deutscher MP. Elevation of RNase R in response to multiple stress conditions[J]. J Biol Chem, 2005, 280(41):34393-34396.
doi: 10.1074/jbc.C500333200 URL |
| [69] |
Liang WX, Malhotra A, Deutscher MP. Acetylation regulates the stability of a bacterial protein:growth stage-dependent modification of RNase R[J]. Mol Cell, 2011, 44(1):160-166.
doi: 10.1016/j.molcel.2011.06.037 URL |
| [70] |
Mohanty BK, Kushner SR. Polynucleotide phosphorylase functions both as a 3' right-arrow 5' exonuclease and a poly(A)polymerase in Escherichiacoli[J]. PNAS, 2000, 97(22):11966-11971.
pmid: 11035800 |
| [71] |
Briani F, Carzaniga T, Dehò G. Regulation and functions of bacterial PNPase[J]. Wiley Interdiscip Rev RNA, 2016, 7(2):241-258.
doi: 10.1002/wrna.2016.7.issue-2 URL |
| [72] |
Cameron TA, de Lay NR. The phosphorolytic exoribonucleases polynucleotide phosphorylase and RNase PH stabilize sRNAs and facilitate regulation of their mRNA targets[J]. J Bacteriol, 2016, 198(24):3309-3317.
pmid: 27698082 |
| [73] |
Saramago M, Bárria C, dos Santos RF, et al. The role of RNases in the regulation of small RNAs[J]. Curr Opin Microbiol, 2014, 18:105-115.
doi: 10.1016/j.mib.2014.02.009 URL |
| [74] |
Pobre V, Arraiano CM. Next generation sequencing analysis reveals that the ribonucleases RNase II, RNase R and PNPase affect bacterial motility and biofilm formation in E. coli[J]. BMC Genomics, 2015, 16:72.
doi: 10.1186/s12864-015-1237-6 URL |
| [75] |
Wu J, Jiang Z, Liu M, et al. Polynucleotide phosphorylase protects Escherichia coli against oxidative stress[J]. Biochemistry, 2009, 48(9):2012-2020.
doi: 10.1021/bi801752p URL |
| [76] |
Hör J, Matera G, Vogel J, et al. Trans-acting small RNAs and their effects on gene expression in Escherichia coli and Salmonella enterica[J]. Ecosal Plus, 2020,9:ecosalplus. ESP-30. DOI: 10.1128/ecosalplus.esp-0030-2019.
doi: 10.1128/ecosalplus.esp-0030-2019 |
| [77] |
Nurmohamed S, Vincent HA, Titman CM, et al. Polynucleotide phosphorylase activity may be modulated by metabolites in Escherichia coli[J]. J Biol Chem, 2011, 286(16):14315-14323.
doi: 10.1074/jbc.M110.200741 pmid: 21324911 |
| [78] | Arraiano CM, Mauxion F, Viegas SC, et al. Intracellular ribonucleases involved in transcript processing and decay:Precision tools for RNA[J]. Biochim et Biophys Acta BBA Gene Regul Mech, 2013, 1829(6/7):491-513. |
| [79] |
Mohanty BK, Maples VF, Kushner SR. Polyadenylation helps regulate functional tRNA levels in Escherichia coli[J]. Nucleic Acids Res, 2012, 40(10):4589-4603.
doi: 10.1093/nar/gks006 pmid: 22287637 |
| [80] |
Garza-Sánchez F, Shoji S, Fredrick K, et al. RNase II is important for A-site mRNA cleavage during ribosome pausing[J]. Mol Microbiol, 2009, 73(5):882-897.
doi: 10.1111/j.1365-2958.2009.06813.x pmid: 19627501 |
| [81] |
Pobre V, Barahona S, Dobrzanski T, et al. Defining the impact of exoribonucleases in the shift between exponential and stationary phases[J]. Sci Rep, 2019, 9(1):16271.
doi: 10.1038/s41598-019-52453-6 URL |
| [82] |
Gutgsell NS, Jain C. Role of precursor sequences in the ordered maturation of E. coli 23S ribosomal RNA[J]. RNA, 2012, 18(2):345-353.
doi: 10.1261/rna.027854.111 pmid: 22190745 |
| [83] |
Li Z, Pandit S, Deutscher MP. 3' exoribonucleolytic trimming is a common feature of the maturation of small, stable RNAs in Escherichia coli[J]. PNAS, 1998, 95(6):2856-2861.
pmid: 9501180 |
| [1] | 陈彩萍, 任昊, 龙腾飞, 何冰, 鲁兆祥, 孙坚. 大肠杆菌Nissle 1917对炎症性肠病治疗作用的研究进展[J]. 生物技术通报, 2023, 39(6): 109-118. |
| [2] | 唐瑞琪, 赵心清, 朱笃, 汪涯. 大肠杆菌对木质纤维素水解液抑制物的胁迫耐受性[J]. 生物技术通报, 2023, 39(11): 205-216. |
| [3] | 李仁瀚, 张乐乐, 刘春立, 刘秀霞, 白仲虎, 杨艳坤, 李业. 基于紫色杆菌素生物合成途径的L-色氨酸生物传感器的构建[J]. 生物技术通报, 2023, 39(10): 80-92. |
| [4] | 位欣欣, 兰海燕. 植物MYB转录因子调控次生代谢及逆境响应的研究进展[J]. 生物技术通报, 2022, 38(8): 12-23. |
| [5] | 张婵, 吴友根, 于靖, 杨东梅, 姚广龙, 杨华庚, 张军锋, 陈萍. 光与茉莉酸信号介导的萜类化合物合成分子机制[J]. 生物技术通报, 2022, 38(8): 32-40. |
| [6] | 高伟欣, 黄火清, 赵晶, 张鑫, 杨宁, 杨浩萌. 应用于基因编辑的核糖核蛋白复合体的构建与活性验证[J]. 生物技术通报, 2022, 38(8): 60-68. |
| [7] | 李晓芳, 刘慧燕, 潘琳, 艾治宇, 李一鸣, 张恒, 方海田. 常温常压等离子体诱变选育高产L-异亮氨酸大肠杆菌[J]. 生物技术通报, 2022, 38(1): 150-156. |
| [8] | 张凤, 陈伟. 代谢组学在植物逆境生物学中的研究进展[J]. 生物技术通报, 2021, 37(8): 1-11. |
| [9] | 张婵, 姚广龙, 张军锋, 于靖, 杨东梅, 陈萍, 吴友根. 广藿香百秋李醇分子调控及合成生物学研究进展[J]. 生物技术通报, 2021, 37(8): 55-64. |
| [10] | 钱虹萍, 陈博, 林金星, 崔亚宁. RNA聚合酶II动态调控及其成像技术的研究进展[J]. 生物技术通报, 2021, 37(4): 293-302. |
| [11] | 吴蓉, 曹佳睿, 曹君, 刘飞翔, 杨猛, 苏二正. 南极假丝酵母脂肪酶B基因在大肠杆菌中的表达和发酵优化[J]. 生物技术通报, 2021, 37(2): 138-148. |
| [12] | 王凯凯, 王晓璐, 苏小运, 张杰. 大肠杆菌双质粒CRISPR-Cas9系统的优化及应用[J]. 生物技术通报, 2021, 37(12): 252-264. |
| [13] | 陈桥, 吴海英, 王宗寿, 谢雨康, 李宜青, 孙俊松. 聚羟基丁酸酯合成引发的高密度生长大肠杆菌的多位点突变分析[J]. 生物技术通报, 2020, 36(7): 112-118. |
| [14] | 邹坤, 路丽丽, Collins Asiamah Amponsah, 薛缘, 张少伟, 苏瑛, 赵志辉. 家禽卵泡闭锁机制的研究进展[J]. 生物技术通报, 2020, 36(4): 185-191. |
| [15] | 张春晨, 胡双艳, 阮海华. 人源溶菌酶在大肠杆菌中的表达与复性研究[J]. 生物技术通报, 2020, 36(3): 153-161. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||