








生物技术通报 ›› 2024, Vol. 40 ›› Issue (12): 20-33.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0392
周子莹1( ), 宋晓东2, 刘洋儿3,4, 吴一凡3, 朱龙佼3, 古东月5, 何国庆6, 李相阳1(
), 宋晓东2, 刘洋儿3,4, 吴一凡3, 朱龙佼3, 古东月5, 何国庆6, 李相阳1( ), 许文涛3(
), 许文涛3( )
)
收稿日期:2024-04-26
出版日期:2024-12-26
发布日期:2025-01-15
通讯作者:
李相阳,男,博士,副教授,研究方向:食品安全;E-mail: lxy2002cn@163.com;作者简介:周子莹,女,硕士研究生,研究方向:食品加工与安全;E-mail: 18210727086@163.com
基金资助:
ZHOU Zi-ying1( ), SONG Xiao-dong2, LIU Yang-er3,4, WU Yi-fan3, ZHU Long-jiao3, GU Dong-yue5, HE Guo-qing6, LI Xiang-yang1(
), SONG Xiao-dong2, LIU Yang-er3,4, WU Yi-fan3, ZHU Long-jiao3, GU Dong-yue5, HE Guo-qing6, LI Xiang-yang1( ), XU Wen-tao3(
), XU Wen-tao3( )
)
Received:2024-04-26
Published:2024-12-26
Online:2025-01-15
摘要:
随着合成生物学的兴起,基于转录因子的生物传感器逐渐从体内传感过渡到体外传感。这类传感器以其高安全性、强稳定性、快速响应等特点,在各个检测领域发挥着作用,特别是在食品安全领域。目前,关于变构转录因子(aTF)生物传感器的综述多侧重于体内构建全细胞生物传感器。本文在汲取前人研究基础上,专注于探讨体外构建aTF生物传感器,例如利用无细胞转录翻译体系和兼容性缓冲液体系作为反应载体。本文详细综述了基于aTF体外生物传感器的构建策略及在食品安全检测中的应用进展。首先,系统阐述了aTF生物传感器的构建,包括aTF分子识别机制,等温扩增与CRISPR-Cas两种信号放大策略、光学与电化学两种信号输出方式,以及兼容性缓冲液和无细胞两种传感体系的运用。其次,重点总结了aTF生物传感器在检测重金属离子、农兽药残留、食品添加剂以及食源性病原体等食品污染物方面的应用进展。最后,深入探讨了aTF生物传感器所面临的挑战,展望其未来发展趋势,以期进一步拓展其在新领域的应用潜力。
周子莹, 宋晓东, 刘洋儿, 吴一凡, 朱龙佼, 古东月, 何国庆, 李相阳, 许文涛. 变构转录因子生物传感器构建策略及在食品安全中的应用进展[J]. 生物技术通报, 2024, 40(12): 20-33.
ZHOU Zi-ying, SONG Xiao-dong, LIU Yang-er, WU Yi-fan, ZHU Long-jiao, GU Dong-yue, HE Guo-qing, LI Xiang-yang, XU Wen-tao. Construction Strategies of Allosteric Transcription Factor Biosensors and Their Application Advances in Food Safety[J]. Biotechnology Bulletin, 2024, 40(12): 20-33.
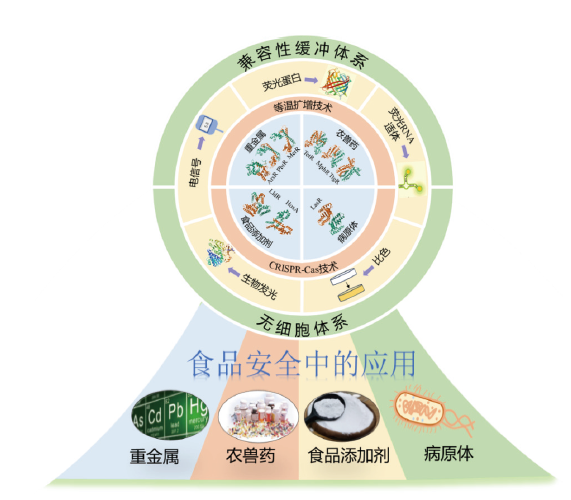
图1 aTF生物传感器构建策略及在食品安全中的应用 aTF生物传感器的典型系统包括分子识别元件aTF、信号放大策略、信号输出体系和传感体系。构建的aTF生物传感器可应用到检测食品中的重金属、农兽药残留、食品添加剂和病原体等食品安全领域中
Fig. 1 Construction strategy of aTF biosensor and its application in food safety A typical system of aTF biosensor includes a molecular recognition element-aTF, a signal amplification strategy, a signal output system and a sensing system. The constructed aTF biosensor can be applied to detect heavy metals, pesticide and veterinary drug residues, food additives and pathogens in food and other food safety fields
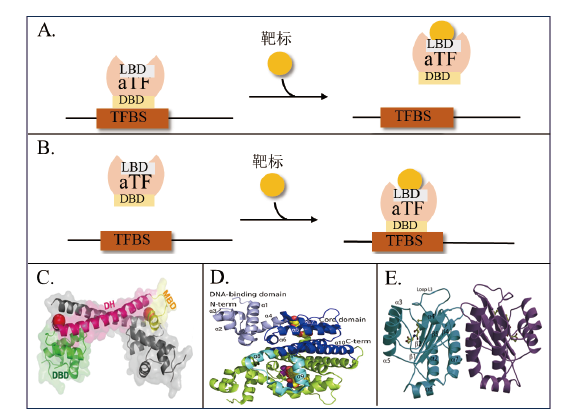
图2 aTF的变构调控机制示意图及aTF结构图 A:抑制型aTF调控机制; B:激活型aTF调控机制;C:汞(II)结合Tn501 MerR的整体结构立体图(PDB代码5CRL)[31]; D:TetR二聚体N82A与强力霉素-Mg2+复合物结合结构图(PDB代码6RGX)[25];E:LasR-N-3-氧十二烷基同型丝氨酸内酯二聚体的整体结构(PDB代码3Ⅸ3),其中一个单体为浅蓝色,另一个单体为紫色。同丝氨酸内酯以球棒表示,二级结构元素在浅蓝色单体上[26]
Fig. 2 Schematic diagram of the mechanism of metamorphic regulation of aTF and structure of aTF A: Repressed aTF regulatory mechanism. B: Activated aTF regulatory mechanism. C: Stereoview of the overall structure of mercury(II)-bound Tn501 MerR(PDB code 5CRL)[31]. D: Ribbon plot of the TetR dimer N82A in complex with doxycycline-Mg2+(PDB code 6RGX)[25]. E: Ribbon diagram showing the overall structure of the LasR-N-3-oxo-dodecanoyl homoserine lactone homodimer(PDB code 3Ⅸ3)with one monomer colored in light blue and the second colored in purple. The homoserine lactone is shown in a ball-and-stick representation and secondary structure elements are noted on the light-blue monomer[26]
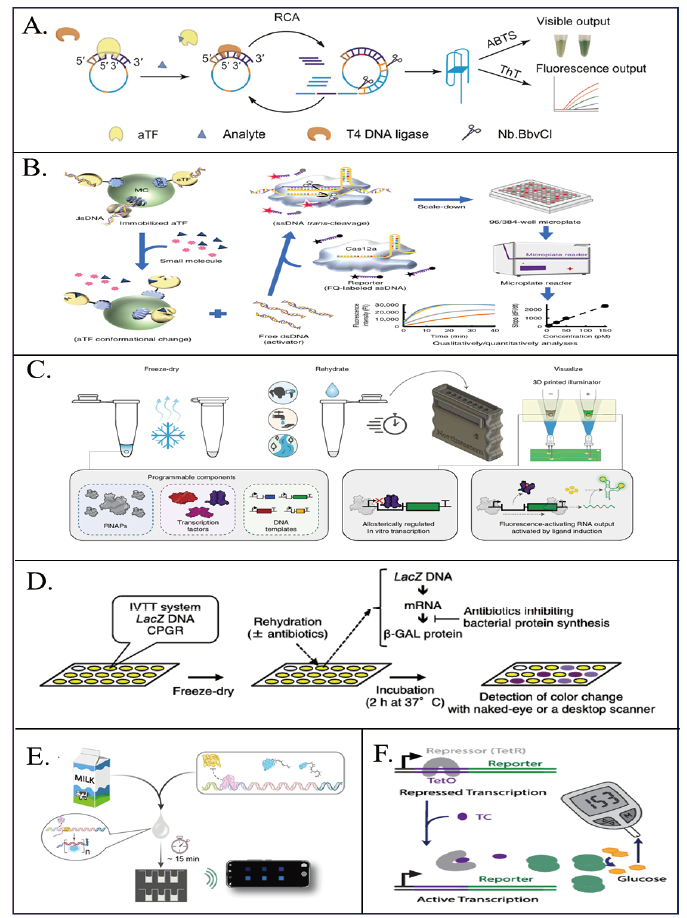
图3 基于aTF体外生物传感器在食品安全中的应用 A:基于aTF与RCA耦合的生物传感器检测小分子物质示意图[33]。B:基于aTF与CRISPR-Cas12a耦合的生物传感器检测小分子示意图[36]。C:在无细胞体外转录系统中,利用配体诱导激活的RNA输出传感器(ROSALIND)来检测水中的污染物示意图[39]。D:用于检测抑制细菌蛋白质合成的抗生素的比色纸基生物传感器示意图[43]。E:基于智能手机的aTF生物发光传感器定量牛奶中的四环素和红霉素示意图[11]。F:基于TetO/TetR的生物传感器利用血糖仪检测四环素示意图[47]
Fig. 3 Application of aTF-based in vivo biosensors in food safety A: Schematic of a biosensor based on aTF coupled with RCA for the detection of small molecules[33]. B: Schematic diagram of a biosensor based on aTF coupled to CRISPR-Cas12a for the detection of small molecules[36]. C: Schematic representation of the RNA Output Sensors Activated by Ligand Induction(ROSALIND)is used to detect contaminants in water in a cell-free in vivo transcription system[39]. D: Schematic of the colorimetric paper-based biosensor for the detection of antibiotics inhibiting bacterial protein synthesis[43]. E: Schematic representation of a smartphone-based aTF bioluminescent sensor for quantification of tetracycline and erythromycin in milk[11]. F: Schematic of a TetO/TetR-based biosensor to detect tetracycline using a glucometer[47]
| 物质类别 Material type | 靶标 Target | 识别元件 Recognition element | 信号输出 Signal output | 检测限 Detection limit | 检测时间 Detection time | 实际样品 Actual sample | 参考文献 Reference |
|---|---|---|---|---|---|---|---|
| 重金属离子 Heavy metal ion | Zn2+ | SmtB | 荧光Fluorescence | 2.5 µmol/L | — | 水样Water | [ |
| Cu2+ | CsoR | 荧光Fluorescence | 5 µmol/L | — | 水样Water | [ | |
| Pb2+ | CadC | 荧光Fluorescence | 259 μg/L | — | 水样Water | [ | |
| Cd2+ | CadC | 荧光Fluorescence | 141 μg/L | — | 水样Water | [ | |
| As3+ | ArsR | 比色Colorimetry | 0.5 mmol/L | 3 h | 水样Water | [ | |
| Hg2+ | MerR | 荧光、比色、生物发光Fluorescence,colorimetry, bioluminescence | 1 nmol/L | 30 min | —— | [ | |
| Hg2+ | MerR | 荧光Fluorescence | 0.5 nmol/L | 1 h | 河水、海水和废水River water, seawater, and wastewater | [ | |
| Pb2+ | PbrR | 荧光Fluorescence | 0.1 nmol/L | 1 h | 河水、海水和废水River water, seawater, and wastewater | [ | |
| Hg2+ | MerR | 荧光Fluorescence | 6 μg/L | 1 h | 水样Water | [ | |
| Hg2+ | MerR | 荧光Fluorescence | 1 μg/L | 1 h | 水样Water | [ | |
| As3+ | ArsR | 荧光Fluorescence | 3.65 μg/L | 2.5 h | 水样Water | [ | |
| 农兽药残留 Pesticide and veterinary drug residues | 土霉素Oxytetracycline | OtrR | 荧光Fluorescence | 0.03 nmol/L | — | — | [ |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 18.1 nmol/L | — | — | [ | |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 80 nmol/L | 32 min | — | [ | |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 12.5 nmol/L | 5 min | — | [ | |
| 无水四环素Anhydrotetracycline | TetR | 荧光Fluorescence | 0.5 μmol/L | 60 min | 牛奶Milk | [ | |
| 红霉素Erythromycin | MphR | 荧光Fluorescence | 0.1 μmol/L | 60 min | 牛奶Milk | [ | |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 25 nmol/L | 5-30 min | 水样Water | [ | |
| 红霉素Erythromycin | MphR | 荧光Fluorescence | 100 nmol/L | 5-30 min | 水样Water | [ | |
| 西维因Carbaryl | TtgR | qPCR | — | — | — | [ | |
| 阿特拉津Atrazine | AtzA | 荧光Fluorescence | 20 μmol/L | 1 h | 水样Water | [ | |
| 食品添加剂 Food additives | 4-HBA | HosA | RT-qPCR | 1.12 nmol/L | — | 河水River water | [ |
| 4-HBA | HosA | RPA | 0.005 nmol/L | — | 河水River water | [ | |
| 4-HBA | HosA | RCA | 1.73 nmol/L | — | 河水River water | [ | |
| 乳酸Lactate | STLldR | 荧光Fluorescence | 0.68 μmol/L | — | 细菌发酵样品、酵素和酸奶Bacterial fermentated samples,fermentand yogurt | [ | |
| 乳酸Lactate | LldR | 荧光Fluorescence | 2.34nmol/L | — | — | [ | |
| PHBA | HosA | 荧光Fluorescence | 1.8 nmol/L | 25 min | — | [ | |
| PHBA | HosA | 荧光Fluorescence | 1.35 nmol/L | — | — | [ | |
| PHBA | HosA | 比色Colorimetry | 2.55 nmol/L | — | — | [ | |
| 苯甲酸Benzoic acid | BenR | 荧光Fluorescence | — | ~1 h | 饮料Beverages | [ | |
| 苯甲酸Benzoic acid | BenR | 荧光Fluorescence | 21 702.843 nmol/L | — | 水样Water | [ | |
| 食源性病原体 Foodborne pathogen | 铜绿假单胞菌Pseudomonas aeruginosa | LasRV | 荧光Fluorescence | 4.9 nmol/L | — | 痰样Sputum | [ |
| 铜绿假单胞菌Pseudomonas aeruginosa | LasR | 荧光Fluorescence | 10 nmol/L | — | — | [ |
表1 aTF生物传感器在食品安全中的应用
Table 1 Applications of aTF biosensors in food safety
| 物质类别 Material type | 靶标 Target | 识别元件 Recognition element | 信号输出 Signal output | 检测限 Detection limit | 检测时间 Detection time | 实际样品 Actual sample | 参考文献 Reference |
|---|---|---|---|---|---|---|---|
| 重金属离子 Heavy metal ion | Zn2+ | SmtB | 荧光Fluorescence | 2.5 µmol/L | — | 水样Water | [ |
| Cu2+ | CsoR | 荧光Fluorescence | 5 µmol/L | — | 水样Water | [ | |
| Pb2+ | CadC | 荧光Fluorescence | 259 μg/L | — | 水样Water | [ | |
| Cd2+ | CadC | 荧光Fluorescence | 141 μg/L | — | 水样Water | [ | |
| As3+ | ArsR | 比色Colorimetry | 0.5 mmol/L | 3 h | 水样Water | [ | |
| Hg2+ | MerR | 荧光、比色、生物发光Fluorescence,colorimetry, bioluminescence | 1 nmol/L | 30 min | —— | [ | |
| Hg2+ | MerR | 荧光Fluorescence | 0.5 nmol/L | 1 h | 河水、海水和废水River water, seawater, and wastewater | [ | |
| Pb2+ | PbrR | 荧光Fluorescence | 0.1 nmol/L | 1 h | 河水、海水和废水River water, seawater, and wastewater | [ | |
| Hg2+ | MerR | 荧光Fluorescence | 6 μg/L | 1 h | 水样Water | [ | |
| Hg2+ | MerR | 荧光Fluorescence | 1 μg/L | 1 h | 水样Water | [ | |
| As3+ | ArsR | 荧光Fluorescence | 3.65 μg/L | 2.5 h | 水样Water | [ | |
| 农兽药残留 Pesticide and veterinary drug residues | 土霉素Oxytetracycline | OtrR | 荧光Fluorescence | 0.03 nmol/L | — | — | [ |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 18.1 nmol/L | — | — | [ | |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 80 nmol/L | 32 min | — | [ | |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 12.5 nmol/L | 5 min | — | [ | |
| 无水四环素Anhydrotetracycline | TetR | 荧光Fluorescence | 0.5 μmol/L | 60 min | 牛奶Milk | [ | |
| 红霉素Erythromycin | MphR | 荧光Fluorescence | 0.1 μmol/L | 60 min | 牛奶Milk | [ | |
| 四环素Tetracycline | TetR | 荧光Fluorescence | 25 nmol/L | 5-30 min | 水样Water | [ | |
| 红霉素Erythromycin | MphR | 荧光Fluorescence | 100 nmol/L | 5-30 min | 水样Water | [ | |
| 西维因Carbaryl | TtgR | qPCR | — | — | — | [ | |
| 阿特拉津Atrazine | AtzA | 荧光Fluorescence | 20 μmol/L | 1 h | 水样Water | [ | |
| 食品添加剂 Food additives | 4-HBA | HosA | RT-qPCR | 1.12 nmol/L | — | 河水River water | [ |
| 4-HBA | HosA | RPA | 0.005 nmol/L | — | 河水River water | [ | |
| 4-HBA | HosA | RCA | 1.73 nmol/L | — | 河水River water | [ | |
| 乳酸Lactate | STLldR | 荧光Fluorescence | 0.68 μmol/L | — | 细菌发酵样品、酵素和酸奶Bacterial fermentated samples,fermentand yogurt | [ | |
| 乳酸Lactate | LldR | 荧光Fluorescence | 2.34nmol/L | — | — | [ | |
| PHBA | HosA | 荧光Fluorescence | 1.8 nmol/L | 25 min | — | [ | |
| PHBA | HosA | 荧光Fluorescence | 1.35 nmol/L | — | — | [ | |
| PHBA | HosA | 比色Colorimetry | 2.55 nmol/L | — | — | [ | |
| 苯甲酸Benzoic acid | BenR | 荧光Fluorescence | — | ~1 h | 饮料Beverages | [ | |
| 苯甲酸Benzoic acid | BenR | 荧光Fluorescence | 21 702.843 nmol/L | — | 水样Water | [ | |
| 食源性病原体 Foodborne pathogen | 铜绿假单胞菌Pseudomonas aeruginosa | LasRV | 荧光Fluorescence | 4.9 nmol/L | — | 痰样Sputum | [ |
| 铜绿假单胞菌Pseudomonas aeruginosa | LasR | 荧光Fluorescence | 10 nmol/L | — | — | [ |
| [1] | Wang PL, Xie LH, Joseph EA, et al. Metal-organic frameworks for food safety[J]. Chem Rev, 2019, 119(18): 10638-10690. |
| [2] | 张秋月. 土壤污染治理问题研究——以镉大米事件为视角[J]. 法制与社会, 2017(3): 77-78. |
| Zhang QY. Research on soil pollution control issues - taking the cadmium rice incident as a perspective[J]. Leg Syst Soc, 2017(3): 77-78. | |
| [3] |
Yamaguchi T, Okihashi M, Harada K, et al. Detection of antibiotics in chicken eggs obtained from supermarkets in Ho Chi Minh City, Vietnam[J]. J Environ Sci Health B, 2017, 52(6): 430-433.
doi: 10.1080/03601234.2017.1293457 pmid: 28281880 |
| [4] | 李航, 刘议蔧, 余晓琴, 等. “穿透式” 监管下的酱油、食醋食品安全问题及对策研究[J]. 食品工业, 2024, 45(4): 323-326. |
| Li H, Liu YH, Yu XQ, et al. Study on food safety problems and countermeasures of soy sauce and vinegar from the “penetrating” supervision[J]. Food Ind, 2024, 45(4): 323-326. | |
| [5] | Griesche C, Baeumner AJ. Biosensors to support sustainable agriculture and food safety[J]. Trac Trends Anal Chem, 2020, 128: 115906. |
| [6] |
Ulrich LE, Koonin EV, Zhulin IB. One-component systems dominate signal transduction in prokaryotes[J]. Trends Microbiol, 2005, 13(2): 52-56.
pmid: 15680762 |
| [7] |
Libis V, Delépine B, Faulon JL. Sensing new chemicals with bacterial transcription factors[J]. Curr Opin Microbiol, 2016, 33: 105-112.
doi: S1369-5274(16)30094-7 pmid: 27472026 |
| [8] |
Baksh KA, Zamble DB. Allosteric control of metal-responsive transcriptional regulators in bacteria[J]. J Biol Chem, 2020, 295(6): 1673-1684.
doi: 10.1074/jbc.REV119.011444 pmid: 31857375 |
| [9] | Zhang YK, Zhao C, Bi HX, et al. A cell-free paper-based biosensor dependent on allosteric transcription factors(aTFs)for on-site detection of harmful metals Hg2+ and Pb2+ in water[J]. J Hazard Mater, 2022, 438: 129499. |
| [10] |
Zhang P, Feng HB, Yang JZ, et al. Detection of inorganic ions and organic molecules with cell-free biosensing systems[J]. J Biotechnol, 2019, 300: 78-86.
doi: S0168-1656(19)30175-0 pmid: 31141711 |
| [11] | Zhang R, Wang Y, Deng HF, et al. Fast and bioluminescent detection of antibiotic contaminants by on-demand transcription of RNA scaffold arrays[J]. Anal Chim Acta, 2023, 1273: 341538. |
| [12] | Li SS, Zhou L, Yao YP, et al. A platform for the development of novel biosensors by configuring allosteric transcription factor recognition with amplified luminescent proximity homogeneous assays[J]. Chem Commun, 2016, 53(1): 99-102. |
| [13] |
Silverman AD, Akova U, Alam KK, et al. Design and optimization of a cell-free atrazine biosensor[J]. ACS Synth Biol, 2020, 9(3): 671-677.
doi: 10.1021/acssynbio.9b00388 pmid: 32078765 |
| [14] | Yao YP, Li SS, Cao JQ, et al. Development of small molecule biosensors by coupling the recognition of the bacterial allosteric transcription factor with isothermal strand displacement amplification[J]. Chem Commun, 2018, 54(38): 4774-4777. |
| [15] | Xiao D, Hu CX, Xu XZ, et al. A d, l-lactate biosensor based on allosteric transcription factor LldR and amplified luminescent proximity homogeneous assay[J]. Biosens Bioelectron, 2022, 211: 114378. |
| [16] | Voyvodic PL, Pandi A, Koch M, et al. Plug-and-play metabolic transducers expand the chemical detection space of cell-free biosensors[J]. Nat Commun, 2019, 10(1): 1697. |
| [17] | Jong S. On a robust, sensitive cell-free method for Pseudomonas sensing and quantification in microfluidic templated hydrogels[J]. Micromachines, 2019, 10(8): 506. |
| [18] | Liu YE, Guo MZ, Du RX, et al. A gas reporting whole-cell microbial biosensor system for rapid on-site detection of mercury contamination in soils[J]. Biosens Bioelectron, 2020, 170: 112660. |
| [19] |
Li SS, Li ZL, Tan GY, et al. in vivo allosteric transcription factor-based biosensing[J]. Trends Biotechnol, 2023, 41(8): 1080-1095.
doi: 10.1016/j.tibtech.2023.03.001 pmid: 36967257 |
| [20] | Wang GH, Wang F, Huang Q, et al. Understanding transcription factor regulation by integrating gene expression and DNase I hypersensitive sites[J]. Biomed Res Int, 2015, 2015: 757530. |
| [21] | Ding NN, Zhou SH, Deng Y. Transcription-factor-based biosensor engineering for applications in synthetic biology[J]. ACS Synth Biol, 2021, 10(5): 911-922. |
| [22] |
Tulin G, Figueroa NR, Checa SK, et al. The multifarious MerR family of transcriptional regulators[J]. Mol Microbiol, 2024, 121(2): 230-242.
doi: 10.1111/mmi.15212 pmid: 38105009 |
| [23] | Bhukya H, Anand R. TetR regulators: a structural and functional perspective[J]. J Indian Inst Sci, 2017, 97(2): 245-259. |
| [24] | Deochand DK, Grove A. MarR family transcription factors: dynamic variations on a common scaffold[J]. Crit Rev Biochem Mol Biol, 2017, 52(6): 595-613. |
| [25] | Palm GJ, Buchholz I, Werten S, et al. Thermodynamics, cooperativity and stability of the tetracycline repressor(TetR)upon tetracycline binding[J]. Biochim Biophys Acta Proteins Proteom, 2020, 1868(6): 140404. |
| [26] | Zou YZ, Nair SK. Molecular basis for the recognition of structurally distinct autoinducer mimics by the Pseudomonas aeruginosa LasR quorum-sensing signaling receptor[J]. Chem Biol, 2009, 16(9): 961-970. |
| [27] |
Kasey CM, Zerrad M, Li YW, et al. Development of transcription factor-based designer macrolide biosensors for metabolic engineering and synthetic biology[J]. ACS Synth Biol, 2018, 7(1): 227-239.
doi: 10.1021/acssynbio.7b00287 pmid: 28950701 |
| [28] |
Machado LFM, Currin A, Dixon N. Directed evolution of the PcaV allosteric transcription factor to generate a biosensor for aromatic aldehydes[J]. J Biol Eng, 2019, 13: 91.
doi: 10.1186/s13036-019-0214-z pmid: 31798685 |
| [29] |
Bai DY, Ding DQ, Li JL, et al. Pinpointing the L-phenylalanine binding sites of TyrR using biosensors and computer-aided simulation[J]. Biotechnol Lett, 2019, 41(3): 401-408.
doi: 10.1007/s10529-019-02645-x pmid: 30680497 |
| [30] | Liang YY, Luo J, Yang CH, et al. Directed evolution of the PobR allosteric transcription factor to generate a biosensor for 4-hydroxymandelic acid[J]. World J Microbiol Biotechnol, 2022, 38(6): 104. |
| [31] |
Wang D, Huang SQ, Liu PY, et al. Structural analysis of the Hg(II)-regulatory protein Tn501 MerR from Pseudomonas aeruginosa[J]. Sci Rep, 2016, 6: 33391.
doi: 10.1038/srep33391 pmid: 27641146 |
| [32] | Liu RN, Liu X, Yang H, et al. A cell-free biosensor based on strand displacement amplification and hybridization chain reaction for fluorescence detection of tetracycline[J]. Microchem J, 2023, 185: 108239. |
| [33] | Cao JQ, Yao YP, Fan KQ, et al. Harnessing a previously unidentified capability of bacterial allosteric transcription factors for sensing diverse small molecules in vivo[J]. Sci Adv, 2018, 4(11): eaau4602. |
| [34] | Zhong JL, Zhao XH. Isothermal amplification technologies for the detection of foodborne pathogens[J]. Food Anal Meth, 2018, 11(6): 1543-1560. |
| [35] | Iwasaki RS, Batey RT. SPRINT: a Cas13a-based platform for detection of small molecules[J]. Nucleic Acids Res, 2020, 48(17): e101. |
| [36] | Liang MD, Li ZL, Wang WS, et al. A CRISPR-Cas12a-derived biosensing platform for the highly sensitive detection of diverse small molecules[J]. Nat Commun, 2019, 10(1): 3672. |
| [37] |
Özyurt C, Üstükarcı H, Evran S, et al. MerR-fluorescent protein chimera biosensor for fast and sensitive detection of Hg2+ in drinking water[J]. Biotechnol Appl Biochem, 2019, 66(5): 731-737.
doi: 10.1002/bab.1805 pmid: 31411759 |
| [38] | Gräwe A, Dreyer A, Vornholt T, et al. A paper-based, cell-free biosensor system for the detection of heavy metals and date rape drugs[J]. PLoS One, 2019, 14(3): e0210940. |
| [39] | Jung JK, Alam KK, Verosloff MS, et al. Cell-free biosensors for rapid detection of water contaminants[J]. Nat Biotechnol, 2020, 38(12): 1451-1459. |
| [40] | Nguyen TT, Chern M, Baer RC, et al. A Förster resonance energy transfer-based ratiometric sensor with the allosteric transcription factor TetR[J]. Small, 2020, 16(17): e1907522. |
| [41] | Chen MF, Nguyen TT, Varongchayakul N, et al. Surface immobilized nucleic acid-transcription factor quantum dots for biosensing[J]. Adv Healthc Mater, 2020, 9(17): e2000403. |
| [42] |
Lin XM, Li YT, Li ZX, et al. Portable environment-signal detection biosensors with cell-free synthetic biosystems[J]. RSC Adv, 2020, 10(64): 39261-39265.
doi: 10.1039/d0ra05293k pmid: 35518409 |
| [43] |
Duyen TTM, Matsuura H, Ujiie K, et al. Paper-based colorimetric biosensor for antibiotics inhibiting bacterial protein synthesis[J]. J Biosci Bioeng, 2017, 123(1): 96-100.
doi: S1389-1723(16)30186-4 pmid: 27514909 |
| [44] | Brodl E, Winkler A, Macheroux P. Molecular mechanisms of bacterial bioluminescence[J]. Comput Struct Biotechnol J, 2018, 16: 551-564. |
| [45] |
Pellinen T, Huovinen T, Karp M. A cell-free biosensor for the detection of transcriptional inducers using firefly luciferase as a reporter[J]. Anal Biochem, 2004, 330(1): 52-57.
pmid: 15183761 |
| [46] |
Lopreside A, Wan XY, Michelini E, et al. Comprehensive profiling of diverse genetic reporters with application to whole-cell and cell-free biosensors[J]. Anal Chem, 2019, 91(23): 15284-15292.
doi: 10.1021/acs.analchem.9b04444 pmid: 31690077 |
| [47] | Amalfitano E, Karlikow M, Norouzi M, et al. A glucose meter interface for point-of-care gene circuit-based diagnostics[J]. Nat Commun, 2021, 12(1): 724. |
| [48] | Sankar K, Baer R, Grazon C, et al. An allosteric transcription factor DNA-binding electrochemical biosensor for progesterone[J]. ACS Sens, 2022, 7(4): 1132-1137. |
| [49] |
Yao YP, Li SS, Cao JQ, et al. A novel signal transduction system for development of uric acid biosensors[J]. Appl Microbiol Biotechnol, 2018, 102(17): 7489-7497.
doi: 10.1007/s00253-018-9056-8 pmid: 29961098 |
| [50] | Wang T, Lu Y. Advances, challenges and future trends of cell-free transcription-translation biosensors[J]. Biosensors, 2022, 12(5): 318. |
| [51] | Bi HX, Zhao C, Zhang YK, et al. IVT cell-free biosensors for tetracycline and macrolide detection based on allosteric transcription factors(aTFs)[J]. Anal Methods, 2022, 14(44): 4545-4554. |
| [52] | Kudo A, Fujikawa Y, Miyahara S, et al. Lessons from Minamata mercury pollution, Japan - after a continuous 22 years of observation[J]. Water Sci Technol, 1998, 38(7): 187-193. |
| [53] |
Gupta S, Sarkar S, Katranidis A, et al. Development of a cell-free optical biosensor for detection of a broad range of mercury contaminants in water: a plasmid DNA-based approach[J]. ACS Omega, 2019, 4(5): 9480-9487.
doi: 10.1021/acsomega.9b00205 pmid: 31460039 |
| [54] | Wang XY, Zhu KL, Chen DD, et al. Monitoring arsenic using genetically encoded biosensors in vivo: the role of evolved regulatory genes[J]. Ecotoxicol Environ Saf, 2021, 207: 111273. |
| [55] | Wang JS, Davidson JL, Kaur S, et al. Paper-based biosensors for the detection of nucleic acids from pathogens[J]. Biosensors, 2022, 12(12): 1094. |
| [56] | Jia M, Zhongbo E, Zhai F, et al. Rapid multi-residue detection methods for pesticides and veterinary drugs[J]. Molecules, 2020, 25(16): 3590. |
| [57] | Chern M, Garden PM, Baer RC, et al. Transcription factor based small-molecule sensing with a rapid cell phone enabled fluorescent bead assay[J]. Angew Chem Int Ed Engl, 2020, 59(48): 21597-21602. |
| [58] |
Rodríguez-Serrano AF, Hsing IM. Allosteric regulation of DNA circuits enables minimal and rapid biosensors of small molecules[J]. ACS Synth Biol, 2021, 10(2): 371-378.
doi: 10.1021/acssynbio.0c00545 pmid: 33481567 |
| [59] |
Mahas A, Wang QC, Marsic T, et al. Development of Cas12a-based cell-free small-molecule biosensors via allosteric regulation of CRISPR array expression[J]. Anal Chem, 2022, 94(11): 4617-4626.
doi: 10.1021/acs.analchem.1c04332 pmid: 35266687 |
| [60] |
Chen W, Zhang XX, Xiong DD, et al. Engineering the effector specificity of regulatory proteins for the in vivo detection of biomarkers and pesticide residues[J]. Appl Microbiol Biotechnol, 2019, 103(7): 3205-3213.
doi: 10.1007/s00253-019-09679-1 pmid: 30770965 |
| [61] | Warner JO. Artificial food additives: hazardous to long-term health[J]. Arch Dis Child, 2024: archdischild-archdisc2023-326565. |
| [62] | Marques C, Wojeicchowski JP, Cardoso T, et al. Lactobionic acid as a suitable food preservative for yacon juice[J]. Innov Food Sci Emerg Technol, 2020, 64: 102400. |
| [63] | Xu XZ, Xu R, Hou S, et al. A selective fluorescent l-lactate biosensor based on an l-lactate-specific transcription regulator and Förster resonance energy transfer[J]. Biosensors, 2022, 12(12): 1111. |
| [64] | Saravanan A, Kumar PS, Hemavathy RV, et al. Methods of detection of food-borne pathogens: a review[J]. Environ Chem Lett, 2021, 19(1): 189-207. |
| [65] | Verbeke F, De Craemer S, Debunne N, et al. Peptides as quorum sensing molecules: measurement techniques and obtained levels in vivo and In vivo[J]. Front Neurosci, 2017, 11: 183. |
| [66] | Wen KY, Cameron L, Chappell J, et al. A cell-free biosensor for detecting quorum sensing molecules in P. aeruginosa-infected respiratory samples[J]. ACS Synth Biol, 2017, 6(12): 2293-2301. |
| [1] | 陈墨岩, 祝诚. 基于CRISPR/Cas12a的生物传感平台的机制研究及应用[J]. 生物技术通报, 2024, 40(7): 90-98. |
| [2] | 王梦雅, 刘家齐, 姜海霖, 李菁华, 赵春燕, 黄红兰. 肠侵袭性大肠埃希菌噬菌体DK-13的生物学特性及应用[J]. 生物技术通报, 2024, 40(3): 296-304. |
| [3] | 余永霞, 祝宁, 刘光敏, 朱龙佼, 许文涛. 肺炎支原体核酸分子诊断技术研究进展[J]. 生物技术通报, 2024, 40(12): 72-83. |
| [4] | 薛宁, 王瑾, 李世新, 刘叶, 程海娇, 张玥, 毛雨丰, 王猛. 多基因同步调控结合高通量筛选构建高产L-苯丙氨酸的谷氨酸棒杆菌工程菌株[J]. 生物技术通报, 2023, 39(9): 268-280. |
| [5] | 李仁瀚, 张乐乐, 刘春立, 刘秀霞, 白仲虎, 杨艳坤, 李业. 基于紫色杆菌素生物合成途径的L-色氨酸生物传感器的构建[J]. 生物技术通报, 2023, 39(10): 80-92. |
| [6] | 陈晓琳, 刘洋儿, 许文涛, 郭明璋, 刘慧琳. 合成生物学细胞传感技术在食品安全快速检测中的应用[J]. 生物技术通报, 2023, 39(1): 137-149. |
| [7] | 张雅涵, 朱丽霞, 胡静, 朱亚静, 张雪婧, 曹叶中. 草甘膦在我国生物育种产业化应用中的机遇与挑战[J]. 生物技术通报, 2022, 38(11): 1-9. |
| [8] | 王鹏飞, 杨敏, 朱龙佼, 许文涛. 基于铂纳米团簇的生物传感研究进展[J]. 生物技术通报, 2021, 37(12): 235-242. |
| [9] | 赵颖, 王楠, 陆安祥, 冯晓元, 郭晓军, 栾云霞. 核酸适配体侧流层析分析技术在真菌毒素检测中的应用[J]. 生物技术通报, 2020, 36(8): 217-227. |
| [10] | 方顺燕, 宋丹, 刘艳萍, 徐文娟, 刘佳瑶, 韩向峙, 龙峰. 用于Escherichia coli O157∶H7直接快速检测的倏逝波荧光核酸适配体传感器研究[J]. 生物技术通报, 2020, 36(7): 228-234. |
| [11] | 叶健文, 陈江楠, 张旭, 吴赴清, 陈国强. 动态调控:一种高效的细胞工厂工程化代谢改造策略[J]. 生物技术通报, 2020, 36(6): 1-12. |
| [12] | 杨敏, 李舒婷, 杨文平, 李相阳, 许文涛. DNA/银纳米簇介导的功能核酸生物传感器研究进展[J]. 生物技术通报, 2020, 36(6): 245-254. |
| [13] | 柳苏月, 田晶晶, 田洪涛, 许文涛. 铽(III)离子及其复合物:从发光特性到传感应用[J]. 生物技术通报, 2020, 36(4): 192-207. |
| [14] | 孙雨阁, 李宸葳, 杜再慧, 许文涛. FEN1酶介导的功能核酸生物传感器的研究进展[J]. 生物技术通报, 2020, 36(4): 208-224. |
| [15] | 王琦, 颜春蕾, 高洪伟, 吴薇, 杨庆利. 基于核酸适配体传感器检测食品致病菌的研究进展[J]. 生物技术通报, 2020, 36(11): 245-258. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||