








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 173-185.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0479
王子傲1,2,3,4( ), 田瑞5, 崔永梅1,2,3,4, 白羿雄1,2,3,4, 姚晓华1,2,3,4, 安立昆1,2,3,4(
), 田瑞5, 崔永梅1,2,3,4, 白羿雄1,2,3,4, 姚晓华1,2,3,4, 安立昆1,2,3,4( ), 吴昆仑5(
), 吴昆仑5( )
)
收稿日期:2024-05-24
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
安立昆,男,副研究员,硕士研究生导师,研究方向:青稞遗传育种;E-mail: anlikun@163.com;作者简介:王子傲,男,硕士研究生,研究方向:青稞遗传育种;E-mail: 13043832262@163.com
基金资助:
WANG Zi-ao1,2,3,4( ), TIAN Rui5, CUI Yong-mei1,2,3,4, BAI Yi-xiong1,2,3,4, YAO Xiao-hua1,2,3,4, AN Li-kun1,2,3,4(
), TIAN Rui5, CUI Yong-mei1,2,3,4, BAI Yi-xiong1,2,3,4, YAO Xiao-hua1,2,3,4, AN Li-kun1,2,3,4( ), WU Kun-lun5(
), WU Kun-lun5( )
)
Received:2024-05-24
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】茉莉酸(JA)信号途径抑制因子JAZs家族是植物JA信号调控途径中的重要组成部分,探索青稞JA信号抑制因子HvnJAZ4基因和蛋白结构、分子动力学以及表达模式特点,为青稞HvnJAZs基因家族调控机制研究提供参考。【方法】采用多种生物信息学方法对HvnJAZ4的基因和蛋白结构,以及分子动力学特点进行分析,并采用qPCR和亚细胞定位对其表达模式进行研究。【结果】HvnJAZ4启动子区域有与JA、脱落酸(ABA)、赤霉素(GA)、水杨酸(SA)、低温响应相关的顺式作用元件。HvnJAZ4由418个氨基酸组成,其中,α螺旋、延长链、β折叠、无规则卷曲所占比例分别为23.50%、12.47%、2.64%和61.39%,HvnJAZ4与野生二粒小麦TdJAZ4亲缘关系最近。HvnJAZ4具有典型的NT、ZIM和Jas结构域,ZIM结构域是HvnJAZ4与其他HvnJAZs以及HvnNINJA1发生互作的关键结构域,Jas结构域是HvnJAZ4与HvnCOI1b发生互作的关键结构域。分子动力学模拟发现HvnJAZ4·Jas短肽与JA-Ile互作的关键氨基酸位点1个,与HvnCOI1b互作的关键氨基酸位点8个。相对于叶、根、茎和茎节,HvnJAZ4在籽粒、分蘖芽中表达量较高,并受低温、MeJA、ABA、GA诱导表达,亚细胞定位结果表明,HvnJAZ4定位在细胞核上。【结论】青稞HvnJAZ4可能在JA调控青稞生长发育和逆境适应以及与其他激素协同调控中扮演着关键角色。
王子傲, 田瑞, 崔永梅, 白羿雄, 姚晓华, 安立昆, 吴昆仑. 青稞HvnJAZ4的生物信息学和表达模式分析[J]. 生物技术通报, 2025, 41(1): 173-185.
WANG Zi-ao, TIAN Rui, CUI Yong-mei, BAI Yi-xiong, YAO Xiao-hua, AN Li-kun, WU Kun-lun. Bioinformatics and Expression Pattern Analysis of HvnJAZ4 Gene in Hulless Barley[J]. Biotechnology Bulletin, 2025, 41(1): 173-185.
| 引物Primer | 引物序列Primer sequence(5'-3') |
|---|---|
| HvnJAZ4 | AGCCGGCTGGATGGAGAGGGA |
| TATTATATCGGATCAGATGT | |
| HvnJAZ4 promotor | CATGCTAACTGCTCCTAG |
| CCAGCCGGCTCGGGAA | |
| HvnJAZ4 qPCR | TCATCCTGTGACACTAGCCTCC |
| GACTTGCTGTTGTTGTCGTT | |
| HvnJAZ1 | ATGGATCTGCTGGA |
| TTACTGGGCCTT | |
| HvnNINJA1 | ATGGAGGATGGCCTTGA |
| TTAGTTTTGGGCCGAGGC | |
| HvnCOI1b | AGAAAGGGTGGGAGGGAGGAGGA |
| CCCAAGCGACGAGGGGCAATAAG | |
| 18SrRNA | ACGAGTCAGCCTTCGTCGT |
| GCGATCTCTGTGCATGATG | |
| HvnJAZ4-GFP | GCTCTAGAAGAAAGGGTGGGAGGGAGGAGGA |
| CGGGTACCCTCAGATGTGTAGTTTTGT |
表1 引物名称及序列
Table 1 Primers’ names and sequences
| 引物Primer | 引物序列Primer sequence(5'-3') |
|---|---|
| HvnJAZ4 | AGCCGGCTGGATGGAGAGGGA |
| TATTATATCGGATCAGATGT | |
| HvnJAZ4 promotor | CATGCTAACTGCTCCTAG |
| CCAGCCGGCTCGGGAA | |
| HvnJAZ4 qPCR | TCATCCTGTGACACTAGCCTCC |
| GACTTGCTGTTGTTGTCGTT | |
| HvnJAZ1 | ATGGATCTGCTGGA |
| TTACTGGGCCTT | |
| HvnNINJA1 | ATGGAGGATGGCCTTGA |
| TTAGTTTTGGGCCGAGGC | |
| HvnCOI1b | AGAAAGGGTGGGAGGGAGGAGGA |
| CCCAAGCGACGAGGGGCAATAAG | |
| 18SrRNA | ACGAGTCAGCCTTCGTCGT |
| GCGATCTCTGTGCATGATG | |
| HvnJAZ4-GFP | GCTCTAGAAGAAAGGGTGGGAGGGAGGAGGA |
| CGGGTACCCTCAGATGTGTAGTTTTGT |
| 元件 Site name | 序列 Sequence | 功能 Function | 数量 Amouny |
|---|---|---|---|
| A-box | CCGTCC | 顺式作用调节元件Cis-acting regulatory element | 1 |
| ABRE | ACGTG | 参与脱落酸反应的顺式作用元件 Cis-acting element involved in the abscisic acid responsiveness | 5 |
| ARE | AAACCA | 对厌氧诱导调节顺式作用元件 Cis-acting regulatory element essential for the anaerobic induction | 3 |
| AuxRR-core | GGTCCAT | 参与生长素反应的顺式作用调节元件 Cis-acting regulatory element involved in auxin responsiveness | 1 |
| Box 4 | ATTAAT | 参与光响应的保守DNA模块的一部分 Part of a conserved DNA module involved in light responsiveness | 1 |
| CAAT-box | CCAAT; CAAAT | 启动子和增强子区域中常见的顺式作用元件 Common Cis-acting element in promoter and enhancer regions | 10 |
| CAT-box | GCCACT | 与分生组织表达相关的顺式作用调控元件 Cis-acting regulatory element related to meristem expression | 3 |
| CGTCA-motif | CGTCA | 参与MeJA反应性的顺式作用调节元件 Cis-acting regulatory element involved in the MeJA-responsiveness | 2 |
| G-Box | CACGTC | 参与光响应的顺式作用调节元件 Cis-acting regulatory element involved in light responsiveness | 2 |
| G-box | CACGTC; CACGAC; GCCACGTGGA | 参与光响应的顺式作用调节元件 Cis-acting regulatory element involved in light responsiveness | 5 |
| GC-motif | CCCCCG | 参与缺氧特异性诱导的增强子样元件 Enhancer-like element involved in anoxic specific inducibility | 2 |
| I-box | CCATATCCAAT; CGATAAGGCG | 光响应元件Part of a light responsive element | 2 |
| LTR | CCGAAA | 参与低温响应的顺式作用元件 Cis-acting element involved in low-temperature responsiveness | 1 |
| MBS | CAACTG | MYB结合位点参与干旱诱导MYB binding site involved in drought-inducibility | 1 |
| O2-site | GATGATGTGG | 参与醇溶蛋白代谢调节的顺式作用调节元件 Cis-acting regulatory element involved in zein metabolism regulation | 1 |
| TATA-box | TATATA; ATATAT; TATA; ACAAAA; TATAA; TATATTTATATTT | 启动子核心元件Core promoter element around -30 of transcription start | 46 |
| TATC-box | TATCCCA | 参与赤霉素反应的顺式作用元件 Cis-acting element involved in gibberellin-responsiveness | 1 |
| TC-rich repeats | GTTTTCTTAC | 参与防御和压力反应的顺式作用元件 Cis-acting element involved in defense and stress responsiveness | 1 |
| TCA-element | CCATCTTTTT; TCAGAAGAGG | 参与水杨酸反应性的顺式作用元件 Cis-acting element involved in salicylic acid responsiveness | 2 |
| TCCC-motif | TCTCCCT | 光响应元件Part of a light responsive element | 3 |
| TGACG-motif | TGACG | 参与MeJA反应性的顺式作用调节元件 Cis-acting regulatory element involved in the MeJA-responsiveness | 2 |
| Unnamed_5 | TGTAATAATATATTTATATT | SEF1因子结合位点SEF1 factor binding site | 1 |
表2 青稞HvnJAZ4启动子区域顺式元件分析
Table 2 Prediction of elements in the promoter region of HvnJAZ4 gene in hulless barely
| 元件 Site name | 序列 Sequence | 功能 Function | 数量 Amouny |
|---|---|---|---|
| A-box | CCGTCC | 顺式作用调节元件Cis-acting regulatory element | 1 |
| ABRE | ACGTG | 参与脱落酸反应的顺式作用元件 Cis-acting element involved in the abscisic acid responsiveness | 5 |
| ARE | AAACCA | 对厌氧诱导调节顺式作用元件 Cis-acting regulatory element essential for the anaerobic induction | 3 |
| AuxRR-core | GGTCCAT | 参与生长素反应的顺式作用调节元件 Cis-acting regulatory element involved in auxin responsiveness | 1 |
| Box 4 | ATTAAT | 参与光响应的保守DNA模块的一部分 Part of a conserved DNA module involved in light responsiveness | 1 |
| CAAT-box | CCAAT; CAAAT | 启动子和增强子区域中常见的顺式作用元件 Common Cis-acting element in promoter and enhancer regions | 10 |
| CAT-box | GCCACT | 与分生组织表达相关的顺式作用调控元件 Cis-acting regulatory element related to meristem expression | 3 |
| CGTCA-motif | CGTCA | 参与MeJA反应性的顺式作用调节元件 Cis-acting regulatory element involved in the MeJA-responsiveness | 2 |
| G-Box | CACGTC | 参与光响应的顺式作用调节元件 Cis-acting regulatory element involved in light responsiveness | 2 |
| G-box | CACGTC; CACGAC; GCCACGTGGA | 参与光响应的顺式作用调节元件 Cis-acting regulatory element involved in light responsiveness | 5 |
| GC-motif | CCCCCG | 参与缺氧特异性诱导的增强子样元件 Enhancer-like element involved in anoxic specific inducibility | 2 |
| I-box | CCATATCCAAT; CGATAAGGCG | 光响应元件Part of a light responsive element | 2 |
| LTR | CCGAAA | 参与低温响应的顺式作用元件 Cis-acting element involved in low-temperature responsiveness | 1 |
| MBS | CAACTG | MYB结合位点参与干旱诱导MYB binding site involved in drought-inducibility | 1 |
| O2-site | GATGATGTGG | 参与醇溶蛋白代谢调节的顺式作用调节元件 Cis-acting regulatory element involved in zein metabolism regulation | 1 |
| TATA-box | TATATA; ATATAT; TATA; ACAAAA; TATAA; TATATTTATATTT | 启动子核心元件Core promoter element around -30 of transcription start | 46 |
| TATC-box | TATCCCA | 参与赤霉素反应的顺式作用元件 Cis-acting element involved in gibberellin-responsiveness | 1 |
| TC-rich repeats | GTTTTCTTAC | 参与防御和压力反应的顺式作用元件 Cis-acting element involved in defense and stress responsiveness | 1 |
| TCA-element | CCATCTTTTT; TCAGAAGAGG | 参与水杨酸反应性的顺式作用元件 Cis-acting element involved in salicylic acid responsiveness | 2 |
| TCCC-motif | TCTCCCT | 光响应元件Part of a light responsive element | 3 |
| TGACG-motif | TGACG | 参与MeJA反应性的顺式作用调节元件 Cis-acting regulatory element involved in the MeJA-responsiveness | 2 |
| Unnamed_5 | TGTAATAATATATTTATATT | SEF1因子结合位点SEF1 factor binding site | 1 |
| 理化性质Physicochemical property | HvnJAZ4 | 理化性质Physicochemical property | HvnJAZ4 | |
|---|---|---|---|---|
| 分子重量Molecular weight/Da | 44 055.39 | α 螺旋比例Proportions of α helix/% | 23.50 | |
| 总原子数Total number of atoms | 6 124 | 延伸链比例Proportions of extended strand/% | 12.47 | |
| 分子式Formula | C1908H3030N568O603S12 | β-折叠比例Proportions of β turn/% | 2.64 | |
| 亲水系数GRAVY | -0.497 | 无规则卷曲比例Proportions of random coil/% | 61.39 | |
| 理论等电点Theoretical pI | 10.0 | 亚细胞定位预测Prediction of subcellular localization | 细胞核Nucleus | |
| 不稳定指数Instability index(II) | 71.38 | 信号肽Signal peptide | 无No | |
| 跨膜结构Transmembrane structures | 无No | 磷酸化位点Phosphorylation site | 丝氨酸43个 | |
| 糖基化位点Glycosylation site 脂溶指数Aliphatic index | 无No 57.03 | 苏氨酸11个 酪氨酸2个 |
表3 青稞HvnJAZ4蛋白质理化性质分析
Table 3 Physical and chemical properties of HvnJAZ4 protein in hulless barely
| 理化性质Physicochemical property | HvnJAZ4 | 理化性质Physicochemical property | HvnJAZ4 | |
|---|---|---|---|---|
| 分子重量Molecular weight/Da | 44 055.39 | α 螺旋比例Proportions of α helix/% | 23.50 | |
| 总原子数Total number of atoms | 6 124 | 延伸链比例Proportions of extended strand/% | 12.47 | |
| 分子式Formula | C1908H3030N568O603S12 | β-折叠比例Proportions of β turn/% | 2.64 | |
| 亲水系数GRAVY | -0.497 | 无规则卷曲比例Proportions of random coil/% | 61.39 | |
| 理论等电点Theoretical pI | 10.0 | 亚细胞定位预测Prediction of subcellular localization | 细胞核Nucleus | |
| 不稳定指数Instability index(II) | 71.38 | 信号肽Signal peptide | 无No | |
| 跨膜结构Transmembrane structures | 无No | 磷酸化位点Phosphorylation site | 丝氨酸43个 | |
| 糖基化位点Glycosylation site 脂溶指数Aliphatic index | 无No 57.03 | 苏氨酸11个 酪氨酸2个 |

图1 青稞HvnJAZ4蛋白序列分析 *:HvnJAZ4·jas与HvnCOI1b由疏水作用形成互作的氨基酸位点;●:HvnJAZ4·jas与HvnCOI1b由氢键形成互作的氨基酸位点
Fig. 1 Protein sequence analysis of HvnJAZ4 in hulless barely *: Key amino acid sites for the hydrophobic interaction between HvnJAZ4·jas and HvnCOI1b. ●: Key amino acid sites for the hydrogen bond interaction between HvnJAZ4·jas and HvnCOI1b
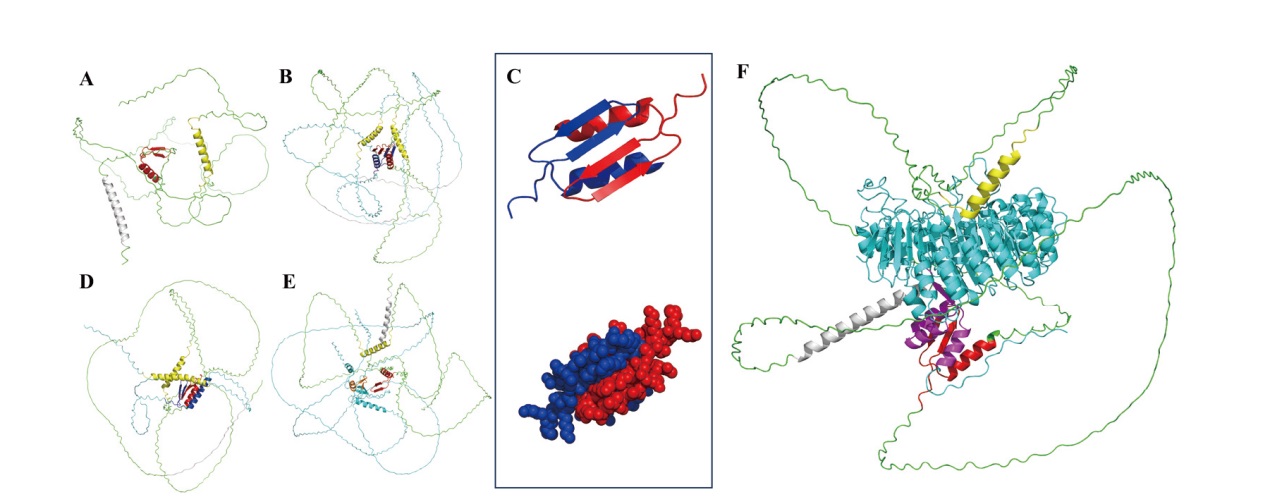
图3 HvnJAZ4与HvnJAZ4、HvnJAZ1、HvnNINJA1、HvnCOI1b互作模型预测 A:HvnJAZ4结构模型;B:HvnJAZ4和HvnJAZ4互作模型;C:HvnJAZ4和HvnJAZ4互作模型中ZIM结构域;D:HvnJAZ4和HvnJAZ1互作模型;E:HvnJAZ4和HvnNINJA1互作模型;F:HvnJAZ4和HvnCOI1b互作模型;Jas结构域(黄色),ZIM结构域(红色,蓝色),NT结构域(灰色),HvnJAZ4分子(绿色),HvnJAZ1分子(青色),HvnNINJA1分子(青色),TDBD结构域(橙色),HvnCOI1b分子(青色),HvnCOI1b的F-box结构域(紫色)
Fig. 3 Interaction model prediction of HvnJAZ4 with HvnJAZ4, HvnJAZ1, HvnNINJA1, and HvnCOI1b A: HvnJAZ4. B: Interaction model of HvnJAZ4 and HvnJAZ4. C: Interaction model of HvnJAZ4 ZIM domain. D: Interaction model of HvnJAZ4 and HvnJAZ1. E: Interaction model of HvnJAZ4 and HvnNINJA1. F: Interaction model of HvnJAZ4 and HvnCOI1b, Jas domain(yellow), ZIM domain(red, bule), NT domain(gray), HvnJAZ4(green), HvnJAZ1(cyans)HvnNINJA1(cyans), HvnNINJA1 TDBD domain(orange), HvnCOI1b(cyans), HvnCOI1b F-box domain(purple)

图4 在30 ns模拟中HvnCOI1b-HvnJAZ4·jas复合物和游离蛋白HvnCOI1b的RMSD和RMSF分析 A:RMSD分析;B:RMSF分析
Fig. 4 RMSD and RMSF analysis of the HvnCOI1b-HvnJAZ4·jas complex and free protein HvnCOI1b during the 30 ns simulation A: RMSD analysis; B: RMSF analysis

图5 在30 ns模拟中蛋白HvnCOI1b和HvnJAZ4·jas关键结合氨基酸能量分解分析 A:HvnCOI1b关键结合氨基酸能量分解分析;B:HvnJAZ4·jas关键结合氨基酸能量分解分析
Fig. 5 Decomposition of the binding energy on a per-residue basis for HvnCOI1b and HvnJAZ4·jas during the 30 ns simulation A: Decomposition of the binding energy on a per-residue basis for HvnCOI1b in the HvnCOI1b-HvnJAZ4 complex. B: Decomposition of the binding energy on a per-residue basis for HvnJAZ4·jas in the HvnCOI1b-HvnJAZ9 complex

图6 HvnCOI1b-HvnJAZ4·jas复合物结合模式图 A:JA-Ile、HvnCOI1b和HvnJAZ4·jas对接模型,JA-Ile(黄色),HvnCOI1b(绿色),HvnJAZ4·Jas(玫红色);B:JA-Ile、HvnCOI1b和HvnJAZ4·jas短肽互作关键氨基酸位点分析
Fig. 6 Binding model of HvnCOI1b-HvnJAZ4·jas complex A: Interaction model of JA-Ile, HvnCOI1b and HvnJAZ4·Jas, JA-Ile(yellow), HvnCOI1b(green), and HvnJAZ4·Jas(rose). B: Key amino acid sites analysis of the interaction of JA-Ile, HvnCOI1b, and HvnJAZ4·jas interaction

图7 青稞HvnJAZ4表达模式分析 A:不同组织中HvnJAZ4的表达;B:低温胁迫下HvnJAZ4的表达:C-H:MeJA、ABA、SA、GA、NAA、6-BA处理下HvnJAZ4的表达。不同小写字母表示差异显著(P<0.05)
Fig. 7 Expression pattern of HvnJAZ4 in hulless barley A: Expression of HvnJAZ4 in different tissues. B: Expression of HvnJAZ4 under low temperatures treatments. C-H: Expression of HvnJAZ4 under MeJA, ABA, SA, GA, NAA and 6-BA treatments. Different lowercase letters indicate significant differences(P<0.05)

图8 青稞HvnJAZ4蛋白亚细胞定位 A:35:HvnJAZ4-GFP荧光场;B:35:HvnJAZ4-GFP叶绿体自发荧光场;C:35:HvnJAZ4-GFP明场;D:35:HvnJAZ4-GFP叠加场;E:35:GFP空载体明场;F:35:GFP叶绿体自发荧光场;G:35:GFP荧光场;H:35:GFP叠加场
Fig. 8 Subcellular localization of HvnJAZ4 protein in hulless barley A: The fluorescence field of 35:HvnJAZ4-GFP. B: The chlorophyll autofluorescence field of 35:HvnJAZ4-GFP. C: The bright field of 35:HvnJAZ4-GFP. D: The merged field of 35:HvnJAZ4-GFP. E: The fluorescence field of 35:GFP. F: The chlorophyll autofluorescence field of 35:GFP. G: The bright field of 35:GFP. H: The merged field of 35:GFP
| [1] | 安立昆, 姚有华, 姚晓华, 等. 青稞耐低氮相关类甜蛋白基因HvTOND1克隆和亚细胞定位研究[J]. 西北农业学报, 2021, 30(8): 1157-1166. |
| An LK, Yao YH, Yao XH, et al. Cloning and subcellular localization of related thaumatin-like protein gene HvTOND1 tolerant to low nitrogen in hulless barley[J]. Acta Agric Boreali Occidentalis Sin, 2021, 30(8): 1157-1166. | |
| [2] | 任晴雯, 安立昆, 姚有华, 等. 青稞HvnPHO1;2基因克隆、亚细胞定位和表达模式分析[J]. 西北农业学报, 2021, 30(10): 1461-1472. |
| Ren QW, An LK, Yao YH, et al. Cloning,subcellular localization and expression analysis of phosphate transporter gene HvPHO1;2 in hulless barely[J]. Acta Agriculturae Boreali - occidentalis Sinica, 2021, 30(10): 1461-1472. | |
| [3] | 颜昌兰. 青稞品种稳定性、适应性及主要农艺性状的评价与分析[D]. 杨凌: 西北农林科技大学, 2015. |
| Yan CL. Evaluation and analysis of stability, adaptability and main agronomic characters of highland barley varieties[D]. Yangling: Northwest A&F University, 2015. | |
| [4] | 弓开元. 青藏高原青稞产量和光温生产潜力对气候变化的响应[D]. 杨凌: 西北农林科技大学, 2020. |
| Gong KY. Response of highland barley yield and light and temperature production potential to climate change in Qinghai-Tibet Plateau[D]. Yangling: Northwest A&F University, 2020. | |
| [5] | 张银乐, 张文静, 杨洋, 等. 低温对青稞种子萌发及幼苗生长的影响[J]. 安徽农学通报, 2018, 24(16): 28-31. |
| Zhang YL, Zhang WJ, Yang Y, et al. Effects of low temperature on seed germination and seedling growth of highland barley[J]. Anhui Agric Sci Bull, 2018, 24(16): 28-31. | |
| [6] | 王玉林, 徐齐君, 原红军, 等. 西藏青稞品种喜马拉雅8号低温处理的SSH-cDNA文库构建及分析[J]. 麦类作物学报, 2017, 37(8): 1025-1030. |
| Wang YL, Xu QJ, Yuan HJ, et al. Construction and analyses of SSH-cDNA library of Tibetan hulless barley(Hordeum vulgare l. var. nudum HK.f.)ximalaya 8 under low temperature treatment[J]. J Triticeae Crops, 2017, 37(8): 1025-1030. | |
| [7] | Yan YX, Christensen S, Isakeit T, et al. Disruption of OPR7 and OPR8 reveals the versatile functions of jasmonic acid in maize development and defense[J]. Plant Cell, 2012, 24(4): 1420-1436. |
| [8] | An LK, Ahmad RM, Ren H, et al. Jasmonate signal receptor gene family ZmCOIs restore male fertility and defense response of Arabidopsis mutant coi1-1[J]. Plant Growth Regul, 2019, 38(2): 479-493. |
| [9] | Chini A, Fonseca S, Fernández G, et al. The JAZ family of repressors is the missing link in jasmonate signalling[J]. Nature, 2007, 448(7154): 666-671. |
| [10] | Chung HS, Cooke TF, Depew CL, et al. Alternative splicing expands the repertoire of dominant JAZ repressors of jasmonate signaling[J]. Plant J, 2010, 63(4): 613-622. |
| [11] | 孙程, 周晓今, 陈茹梅, 等. 植物JAZ蛋白的功能概述[J]. 生物技术通报, 2014(6): 1-8. |
| Sun C, Zhou XJ, Chen RM, et al. Comprehensive overview of JAZ proteins in plants[J]. Biotechnol Bull, 2014(6): 1-8. | |
| [12] | Thines B, Katsir L, Melotto M, et al. JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling[J]. Nature, 2007, 448: 661-665. |
| [13] |
Oblessuc PR, Obulareddy N, DeMott L, et al. JAZ4 is involved in plant defense, growth, and development in Arabidopsis[J]. Plant J, 2020, 101(2): 371-383.
doi: 10.1111/tpj.14548 |
| [14] |
晏胜伟, 孙程, 周晓今, 等. 玉米JAZ家族基因ZmJAZ4的克隆及功能分析[J]. 生物技术通报, 2015(3): 96-101.
doi: 10.13560/j.cnki.biotech.bull.1985.2015.04.013 |
| Yan SW, Sun C, Zhou XJ, et al. Cloning and characterization analysis of ZmJAZ4, a JAZ family gene in maize[J]. Biotechnol Bull, 2015,(3): 96-101. | |
| [15] | 王世通. 基于PhJAZ4转基因的矮牵牛新种质创制与评价[D]. 武汉: 华中农业大学, 2023. |
| Wang ST. Creation and evaluation of new petunia germplasm based on PhJAZ4 transgenic[D]. Wuhan: Huazhong Agricultural University, 2023. | |
| [16] | 杨娆. 丹参SmJAZ4调控酚酸类物质合成的分子机制[D]. 西安: 陕西师范大学, 2022. |
| Yang R. Molecular mechanism of the regulation of phenolic acid synthesis by Salvia miltiorrhiza SmJAZ4[D]. Xi'an: Shaanxi Normal University, 2022. | |
| [17] | Miccono MLA, Yang HW, DeMott L, et al. Review: Losing JAZ4 for growth and defense[J]. Plant Sci, 2023, 335: 111816. |
| [18] | Zhang MF, Luo X, He W, et al. OsJAZ4 fine-tunes rice blast resistance and yield traits[J]. Plants, 2024, 13(3): 348. |
| [19] | DeMott L, Oblessuc PR, Pierce A, et al. Spatiotemporal regulation of JAZ4 expression and splicing contribute to ethylene- and auxin-mediated responses in Arabidopsis roots[J]. Plant J, 2021, 108(5): 1266-1282. |
| [20] | Sheard LB, Tan X, Mao HB, et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor[J]. Nature, 2010, 468(7322): 400-405. |
| [21] | Abramson J, Adler J, Dunger J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3[J]. Nature, 2024, 630(8016): 493-500. |
| [22] |
Sanner MF. Python: a programming language for software integration and development[J]. J Mol Graph Model, 1999, 17(1): 57-61.
pmid: 10660911 |
| [23] |
Morris GM, Huey R, Lindstrom W, et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility[J]. J Comput Chem, 2009, 30(16): 2785-2791.
doi: 10.1002/jcc.21256 pmid: 19399780 |
| [24] |
Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading[J]. J Comput Chem, 2010, 31(2): 455-461.
doi: 10.1002/jcc.21334 pmid: 19499576 |
| [25] |
Pierce LCT, Salomon-Ferrer R, Augusto F de Oliveira C, et al. Routine access to millisecond time scale events with accelerated molecular dynamics[J]. J Chem Theory Comput, 2012, 8(9): 2997-3002.
pmid: 22984356 |
| [26] |
Götz AW, Williamson MJ, Xu D, et al. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. generalized born[J]. J Chem Theory Comput, 2012, 8(5): 1542-1555.
pmid: 22582031 |
| [27] |
Salomon-Ferrer R, Götz AW, Poole D, et al. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. explicit solvent particle mesh Ewald[J]. J Chem Theory Comput, 2013, 9(9): 3878-3888.
doi: 10.1021/ct400314y pmid: 26592383 |
| [28] |
Wang JM, Wolf RM, Caldwell JW, et al. Development and testing of a general amber force field[J]. J Comput Chem, 2004, 25(9): 1157-1174.
doi: 10.1002/jcc.20035 pmid: 15116359 |
| [29] |
Wang JM, Wang W, Kollman PA, et al. Automatic atom type and bond type perception in molecular mechanical calculations[J]. J Mol Graph Model, 2006, 25(2): 247-260.
pmid: 16458552 |
| [30] |
Sousa da Silva AW, Vranken WF. ACPYPE-AnteChamber PYthon parser interface[J]. BMC Res Notes, 2012, 5: 367.
doi: 10.1186/1756-0500-5-367 pmid: 22824207 |
| [31] | 魏昕, 刘雨恒, 刘宇阳, 等. 植物JAZ蛋白家族研究进展[J]. 植物生理学报, 2021, 57(5): 1039-1046. |
| Wei X, Liu YH, Liu YY, et al. Advances of JAZ family in plants[J]. China Ind Econ, 2021, 57(5): 1039-1046. | |
| [32] | Fu J, Wu H, Ma SQ, et al. OsJAZ1 attenuates drought resistance by regulating JA and ABA signaling in rice[J]. Front Plant Sci, 2017, 8: 2108. |
| [33] | 黄文峰, 王立丰, 田维敏. 茉莉酸反应基因转录抑制因子JAZ蛋白家族研究进展[J]. 热带作物学报, 2009, 30(9): 1383-1387. |
| Huang WF, Wang LF, Tian WM. Advances in JAZ protein family mediating JA signal pathway[J]. Chin J Trop Crops, 2009, 30(9): 1383-1387. | |
| [34] | 闫会转. JAZ2和JAZ7调控茉莉酸介导的转录与代谢重编程的机理研究[D]. 杭州: 浙江大学, 2014. |
| Yan HZ. Mechanism of JAZ2 and JAZ7 in regulating jasmonic acid mediated transcription and metabolic reprogramming[D]. Hangzhou: Zhejiang University, 2014. | |
| [35] | 安立昆. 玉米茉莉酸信号受体基因家族ZmCOIs功能研究[D]. 南京: 南京农业大学, 2018. |
| An LK. Study on the functional of maize jasmonic acid signal receptor gene family ZmCOIs[D]. Nanjing: Nanjing Agricultural University, 2018. | |
| [36] |
Saito R, Hayashi K, Nomoto H, et al. Extended JAZ degron sequence for plant hormone binding in jasmonate co receptor of tomato S1COI1 S1JAZ[J]. Sci Rep, 2021, 11(1): 13612.
doi: 10.1038/s41598-021-93067-1 pmid: 34193940 |
| [1] | 孔青洋, 张晓龙, 李娜, 张晨洁, 张雪云, 于超, 张启翔, 罗乐. 单叶蔷薇GRAS转录因子家族鉴定及表达分析[J]. 生物技术通报, 2025, 41(1): 210-220. |
| [2] | 宋兵芳, 柳宁, 程新艳, 徐晓斌, 田文茂, 高悦, 毕阳, 王毅. 马铃薯G6PDH基因家族鉴定及其在损伤块茎的表达分析[J]. 生物技术通报, 2024, 40(9): 104-112. |
| [3] | 吴娟, 武小娟, 王沛捷, 谢锐, 聂虎帅, 李楠, 马艳红. 彩色马铃薯花青素合成相关ERF基因筛选及表达分析[J]. 生物技术通报, 2024, 40(9): 82-91. |
| [4] | 武帅, 辛燕妮, 买春海, 穆晓娅, 王敏, 岳爱琴, 赵晋忠, 吴慎杰, 杜维俊, 王利祥. 大豆GS基因家族全基因组鉴定及胁迫响应分析[J]. 生物技术通报, 2024, 40(8): 63-73. |
| [5] | 王梦帆, 赵子玉, 王春光, 刘廷玉, 陈曦, 张铁. 基于药效团模型筛选CTX-M-14型超广谱β-内酰胺酶抑制剂[J]. 生物技术通报, 2024, 40(6): 319-329. |
| [6] | 刘蓉, 田闵玉, 李光泽, 谭成方, 阮颖, 刘春林. 甘蓝型油菜REVEILLE家族鉴定及诱导表达分析[J]. 生物技术通报, 2024, 40(6): 161-171. |
| [7] | 李嘉欣, 李鸿燕, 刘丽娥, 张恬, 周武. 沙棘NRAMP基因家族鉴定及铅胁迫下表达分析[J]. 生物技术通报, 2024, 40(5): 191-202. |
| [8] | 钟匀, 林春, 刘正杰, 董陈文华, 毛自朝, 李兴玉. 芦笋皂苷合成相关糖基转移酶基因克隆及原核表达分析[J]. 生物技术通报, 2024, 40(4): 255-263. |
| [9] | 郝楠, 耿珊, 赵雨薇, 侯智涵, 赵斌, 刘颖超. 拟轮枝镰孢丙氨酸转氨酶FvALT的克隆与表达分析[J]. 生物技术通报, 2024, 40(12): 256-263. |
| [10] | 杨冲, 程莎莎, 艾长丰, 赵璇, 刘孟军. 枣ABF/AREB基因家族鉴定及其在果实发育中的表达分析[J]. 生物技术通报, 2024, 40(11): 184-191. |
| [11] | 史京辉, 陈文慧, 陆坤, 郑婷婷, 任志远, 鲍国庆, 王敏, 骆健美. 定点饱和突变提高赭曲霉11α羟化酶的催化性能[J]. 生物技术通报, 2024, 40(1): 322-331. |
| [12] | 王艺清, 王涛, 韦朝领, 戴浩民, 曹士先, 孙威江, 曾雯. 茶树SMAS基因家族的鉴定及互作分析[J]. 生物技术通报, 2023, 39(4): 246-258. |
| [13] | 平怀磊, 郭雪, 余潇, 宋静, 杜春, 王娟, 张怀璧. 滇牡丹PdANS的克隆、表达及与花青素含量的相关性[J]. 生物技术通报, 2023, 39(3): 206-217. |
| [14] | 郭志浩, 金泽鑫, 刘琦, 高利. 小麦矮腥黑粉菌效应蛋白g11335的生物信息学分析、亚细胞定位及毒性验证[J]. 生物技术通报, 2022, 38(8): 110-117. |
| [15] | 于秋琳, 马婧怡, 赵盼, 孙鹏芳, 何玉美, 刘世彪, 郭惠红. 绞股蓝GpMIR156a和GpMIR166b的克隆与功能分析[J]. 生物技术通报, 2022, 38(7): 186-193. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||