








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 333-346.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0575
饶峻1( ), 赵晨1, 李端华1, 廖豪2, 黄加雨1, 王辂1(
), 赵晨1, 李端华1, 廖豪2, 黄加雨1, 王辂1( )
)
收稿日期:2024-06-14
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
王辂,男,博士,研究员,研究方向:微生物与生化药物产业化开发关键技术与应用推广;E-mail: wanglu@cdu.edu.cn作者简介:饶峻,男,硕士,研究方向:工程菌构建及发酵工艺优化;E-mail: 1274797431@qq.com
基金资助:
RAO Jun1( ), ZHAO Chen1, LI Duan-hua1, LIAO Hao2, HUANG Jia-yu1, WANG Lu1(
), ZHAO Chen1, LI Duan-hua1, LIAO Hao2, HUANG Jia-yu1, WANG Lu1( )
)
Received:2024-06-14
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】旨在挖掘细菌途径ERG合成潜力,为之后对细菌途径关键酶改造,提高ERG产量提供实验依据。【方法】在大肠杆菌中引入耻垢分枝杆菌ERG合成途径,创建阳性对照工程菌RE。通过摇瓶发酵比较常规诱导法与自诱导法ERG产量;随后对底盘细胞RE的His和Cys途径进行改造,增强其前体氨基酸内源合成能力,得到菌株RE-CH;使用RE-CH菌株在10 L罐上建立自诱导发酵工艺进行发酵放大研究,并对补料策略进行优化;最后,改变调控策略,增加菌体密度,在30 L罐上进行发酵,以期增加ERG产量。【结果】相比常规诱导法,自诱导法ERG产量提高2.8倍;发酵验证表明新工程菌(RE-CH)ERG合成能力得到增强;10 L罐优化补料策略后,ERG产量达到1.1 g/L;30 L罐调整调控策略后,发酵95.5 h,ERG的产量达到4.3 g/L。【结论】利用细菌ERG合成途径发酵ERG产量与真菌途径产量相当,且使用优化后的发酵工艺相比已报道的真菌途径发酵工艺发酵周期缩短约33%。
饶峻, 赵晨, 李端华, 廖豪, 黄加雨, 王辂. 自诱导策略在麦角硫因生物合成中的应用[J]. 生物技术通报, 2025, 41(1): 333-346.
RAO Jun, ZHAO Chen, LI Duan-hua, LIAO Hao, HUANG Jia-yu, WANG Lu. Application of Auto-induction Strategy in Ergothioneine Biosynthesis[J]. Biotechnology Bulletin, 2025, 41(1): 333-346.
| ERG合成途径来源 ERG synthesis pathway source | 关键酶* Key enzymes* | 工程菌 Engineering bacteria | 发酵工艺 Fermentation process | 诱导方式 Induction method | 发酵周期Fermentation period/h | 菌体密度 Bacterial density | ERG产量ERG production/(g·L-1) | ERG产率 ERG productivity/(g·L-1·h-1) | 参考文献 Reference |
|---|---|---|---|---|---|---|---|---|---|
| Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 73 | - | 0.6 | 8.2 | [ |
| Bacteria | W | Corynebacterium glutamicum | Fed-batch baffled flasks | - | 120 | OD600>80 | 0.1 | 0.8 | [ |
| Bacteria | W | E. coli | Baffled flasks | Manual addition of inducers | 192 | OD600=14.0 | 0.7 | 3.4 | [ |
| Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 216 | OD600≈55 | 1.3 | 6.1 | [ |
| Fungi | W | Yarrowia lipolytica | Fed-batch jar fermentation | - | 168 | OD600>150 | 7.3 | 43.5 | [ |
| Fungi | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 143 | OD600=130 | 4.3 | 30.3 | [ |
| Fungi | W | Yarrowia lipolytica | Fed-batch jar fermentation | - | 220 | 60.6 g/L CDW | 1.6 | 7.4 | [ |
| Fungi+Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 96 | OD600>75 | 2.6 | 27.4 | [ |
| Fungi+Bacteria | M | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 94 | OD600≈100 | 5.4 | 57.4 | [ |
| Fungi+Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 108 | OD600=45.7 | 0.7 | 6.6 | [ |
| Fungi+Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 108 | OD600=65 | 2 | 18.6 | [ |
表1 近几年微生物液体发酵法生物合成麦角硫因发酵水平
Table 1 Fermentation levels for biosynthesis of ergothioneine by microbial liquid fermentation in recent years
| ERG合成途径来源 ERG synthesis pathway source | 关键酶* Key enzymes* | 工程菌 Engineering bacteria | 发酵工艺 Fermentation process | 诱导方式 Induction method | 发酵周期Fermentation period/h | 菌体密度 Bacterial density | ERG产量ERG production/(g·L-1) | ERG产率 ERG productivity/(g·L-1·h-1) | 参考文献 Reference |
|---|---|---|---|---|---|---|---|---|---|
| Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 73 | - | 0.6 | 8.2 | [ |
| Bacteria | W | Corynebacterium glutamicum | Fed-batch baffled flasks | - | 120 | OD600>80 | 0.1 | 0.8 | [ |
| Bacteria | W | E. coli | Baffled flasks | Manual addition of inducers | 192 | OD600=14.0 | 0.7 | 3.4 | [ |
| Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 216 | OD600≈55 | 1.3 | 6.1 | [ |
| Fungi | W | Yarrowia lipolytica | Fed-batch jar fermentation | - | 168 | OD600>150 | 7.3 | 43.5 | [ |
| Fungi | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 143 | OD600=130 | 4.3 | 30.3 | [ |
| Fungi | W | Yarrowia lipolytica | Fed-batch jar fermentation | - | 220 | 60.6 g/L CDW | 1.6 | 7.4 | [ |
| Fungi+Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 96 | OD600>75 | 2.6 | 27.4 | [ |
| Fungi+Bacteria | M | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 94 | OD600≈100 | 5.4 | 57.4 | [ |
| Fungi+Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 108 | OD600=45.7 | 0.7 | 6.6 | [ |
| Fungi+Bacteria | W | E. coli | Fed-batch jar fermentation | Manual addition of inducers | 108 | OD600=65 | 2 | 18.6 | [ |

图1 麦角硫因及其前体氨基酸代谢途径 在大肠杆菌中导入耻垢分枝杆菌的ERG合成基因(egtABCDE)构建ERG工程菌。组氨酸、半胱氨酸、蛋氨酸和谷氨酸是ERG合成的前体氨基酸,其中谷氨酸可在合成过程中循环利用;HisG是合成组氨酸的关键酶,SerA和CysE是合成半胱氨酸的关键酶;γGC:γ-谷氨酰半胱氨酸;SAM:S-腺苷甲硫氨酸;SAH:S-腺苷-L-高半胱氨酸
Fig. 1 Metabolic pathways of ergothioneine and its precursor amino acids Introduction of the ERG synthesis gene(egtABCDE)from Mycobacterium smegmatis into E. coli results in ERG synthesis. Methionine, histidine, cysteine, and glutamate are the major precursors in this pathway, with glutamate being recycled and not physically consumed. HisG is the key enzyme in the synthesis of histidine whereas SerA and CysE are the key enzymes in the synthesis of cysteine. γGC: γ-glutamylcysteine; SAM: S-Adenosyl methionine; SAH: S-adenosyl-L-homocysteine
| 类型 Type | 名称 Name | 描述 Description | 来源 Source |
|---|---|---|---|
| Plasmid | pET-28a | Expression vector, Kanr | Lab stock |
| pET28a-egtABCDE | pET-28a containing egtABCDE | This work | |
| pET28a-egtA | pET-28a containing egtA | This work | |
| pET28a-egtB | pET-28a containing egtB | This work | |
| pET28a-egtC | pET-28a containing egtC | This work | |
| pET28a-egtD | pET-28a containing egtD | This work | |
| pET28a-egtE | pET-28a containing egtE | This work | |
| pACYC184 | Expression vector, Cmr, Tcr | General Biol | |
| pACYC184-cysE-serA | pACYC184 containing cysET167A, G203S, T234S, P252L, M256Q and serAT410stop | This work | |
| pACYC184-hisG | pACYC184 containing hisGS143F, ∆209-281 | This work | |
| pACYC184-hisG-cysE-serA | pACYC184 containing cysET167A, G203S, T234S, P252L, M256Q, serAT410stopand hisGS143F, ∆209-281 | This work | |
| Strain | E. coli Rosetta2(DE3) | Expression host | Lab stock |
| RE | E. coli Rosetta2(DE3)harboring pET28a-egtABCDE | This work | |
| REA | E. coli Rosetta2(DE3)harboring pET28a-egtA | This work | |
| REB | E. coli Rosetta2(DE3)harboring pET28a-egtB | This work | |
| REC | E. coli Rosetta2(DE3)harboring pET28a-egtC | This work | |
| RED | E. coli Rosetta2(DE3)harboring pET28a-egtD | This work | |
| REE | E. coli Rosetta2(DE3)harboring pET28a-egtE | This work | |
| RE-C | RE harboring pACYC184-cysE-serA | This work | |
| RE-H | RE harboring pACYC184-hisG | This work | |
| RE-CH | RE harboring pACYC184-hisG-cysE-serA | This work |
表2 菌株与质粒
Table 2 Plasmids and strains
| 类型 Type | 名称 Name | 描述 Description | 来源 Source |
|---|---|---|---|
| Plasmid | pET-28a | Expression vector, Kanr | Lab stock |
| pET28a-egtABCDE | pET-28a containing egtABCDE | This work | |
| pET28a-egtA | pET-28a containing egtA | This work | |
| pET28a-egtB | pET-28a containing egtB | This work | |
| pET28a-egtC | pET-28a containing egtC | This work | |
| pET28a-egtD | pET-28a containing egtD | This work | |
| pET28a-egtE | pET-28a containing egtE | This work | |
| pACYC184 | Expression vector, Cmr, Tcr | General Biol | |
| pACYC184-cysE-serA | pACYC184 containing cysET167A, G203S, T234S, P252L, M256Q and serAT410stop | This work | |
| pACYC184-hisG | pACYC184 containing hisGS143F, ∆209-281 | This work | |
| pACYC184-hisG-cysE-serA | pACYC184 containing cysET167A, G203S, T234S, P252L, M256Q, serAT410stopand hisGS143F, ∆209-281 | This work | |
| Strain | E. coli Rosetta2(DE3) | Expression host | Lab stock |
| RE | E. coli Rosetta2(DE3)harboring pET28a-egtABCDE | This work | |
| REA | E. coli Rosetta2(DE3)harboring pET28a-egtA | This work | |
| REB | E. coli Rosetta2(DE3)harboring pET28a-egtB | This work | |
| REC | E. coli Rosetta2(DE3)harboring pET28a-egtC | This work | |
| RED | E. coli Rosetta2(DE3)harboring pET28a-egtD | This work | |
| REE | E. coli Rosetta2(DE3)harboring pET28a-egtE | This work | |
| RE-C | RE harboring pACYC184-cysE-serA | This work | |
| RE-H | RE harboring pACYC184-hisG | This work | |
| RE-CH | RE harboring pACYC184-hisG-cysE-serA | This work |
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') |
|---|---|
| EgtA-F | ATATACCATGGCCCTGCCGG |
| EgtA-R | CCCAAGCTTTCCTTCTTACAGTTCACCTTTTGCC |
| EgtB-F | CATGCCATGGAACTGATTGCACGTGAAACCCTGG |
| EgtB-R | CCCAAGCTTCTCCTTCTTACACATCCCATGCC |
| EgtC-F | CATGCCATGGAATGTCGCCATGTTGCCTGGCT |
| EgtC-R | CCCAAGCTTCTTCTTACAGCGGGGTAACAAC |
| EgtD-F | CATGCCATGGAAACCCTGAGCCTGGCCAATTATC |
| EgtD-R | CCCAAGCTTCCTTCTTAACGAACTGCCAGGC |
| EgtE-F | CATGCCATGGAAATGCTGGCACAGCAGTGGCGTGA |
| EgtE-R | CCCAAGCTTAGCTTTTACGGTGCTTCACGC |
| HisG-F | ATAAAATATTTCTAGTTTTTTTCATATGCCTGACGGAGTTCACAC |
| HisG-R | TGCACTGAAATCTAGTTAATTCTGTGCATGCAGAATACCCT |
表3 相关引物
Table 3 Related primers
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') |
|---|---|
| EgtA-F | ATATACCATGGCCCTGCCGG |
| EgtA-R | CCCAAGCTTTCCTTCTTACAGTTCACCTTTTGCC |
| EgtB-F | CATGCCATGGAACTGATTGCACGTGAAACCCTGG |
| EgtB-R | CCCAAGCTTCTCCTTCTTACACATCCCATGCC |
| EgtC-F | CATGCCATGGAATGTCGCCATGTTGCCTGGCT |
| EgtC-R | CCCAAGCTTCTTCTTACAGCGGGGTAACAAC |
| EgtD-F | CATGCCATGGAAACCCTGAGCCTGGCCAATTATC |
| EgtD-R | CCCAAGCTTCCTTCTTAACGAACTGCCAGGC |
| EgtE-F | CATGCCATGGAAATGCTGGCACAGCAGTGGCGTGA |
| EgtE-R | CCCAAGCTTAGCTTTTACGGTGCTTCACGC |
| HisG-F | ATAAAATATTTCTAGTTTTTTTCATATGCCTGACGGAGTTCACAC |
| HisG-R | TGCACTGAAATCTAGTTAATTCTGTGCATGCAGAATACCCT |
| 氨基酸 Amino acid | 终浓度Final concentration/(g·L-1) |
|---|---|
| Glu | 0.8 |
| 1.4 | |
| 2.0 | |
| His | 1.4 |
| 2.0 | |
| 2.6 | |
| Cys | 1.4 |
| 1.8 | |
| 2.0 | |
| Met | 0.25 |
| 0.5 | |
| 1.0 | |
| BLK | 0.0 |
表4 前体氨基酸添加浓度
Table 4 Added concentration of precursor amino acid
| 氨基酸 Amino acid | 终浓度Final concentration/(g·L-1) |
|---|---|
| Glu | 0.8 |
| 1.4 | |
| 2.0 | |
| His | 1.4 |
| 2.0 | |
| 2.6 | |
| Cys | 1.4 |
| 1.8 | |
| 2.0 | |
| Met | 0.25 |
| 0.5 | |
| 1.0 | |
| BLK | 0.0 |
| 水平 Level | 因素 Factors | ||
|---|---|---|---|
| A: Glu/(g·L-1) | B: His/(g·L-1) | C: Cys/(g·L-1) | |
| 1 | 1.1 | 1.7 | 1.6 |
| 2 | 1.4 | 2.0 | 1.8 |
| 3 | 1.7 | 2.3 | 2.0 |
表5 正交实验因素水平表
Table 5 Factors and levels of orthogonal experiments
| 水平 Level | 因素 Factors | ||
|---|---|---|---|
| A: Glu/(g·L-1) | B: His/(g·L-1) | C: Cys/(g·L-1) | |
| 1 | 1.1 | 1.7 | 1.6 |
| 2 | 1.4 | 2.0 | 1.8 |
| 3 | 1.7 | 2.3 | 2.0 |
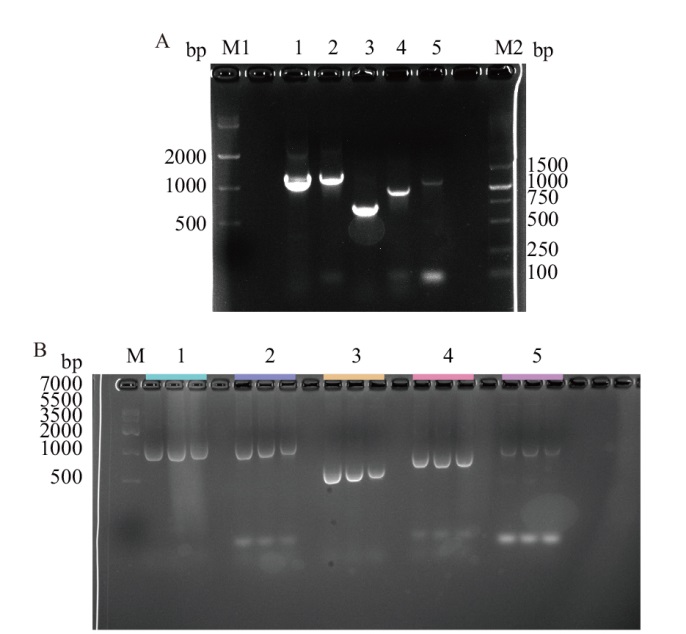
图2 阳性工程菌构建 A:单个egt基因扩增结果(M1:DNA marker IV;M2:DNA marker DL5000;1-5:egtA、egtB、egtC、egtD、egtE);B:PCR鉴定结果(M:DNA marker IV;1-5:egtA、egtB、egtC、egtD、egtE)
Fig. 2 Construction of positive engineering strain A: Amplified results of a single egt gene(M1:DNA marker IV;M2:DNA marker DL5000;1-5:egtA, egtB, egtC, egtD, and egtE)). B: Results of PCR identification(M:DNA marker IV;1-5:egtA,egtB,egtC, egtD, and egtE)
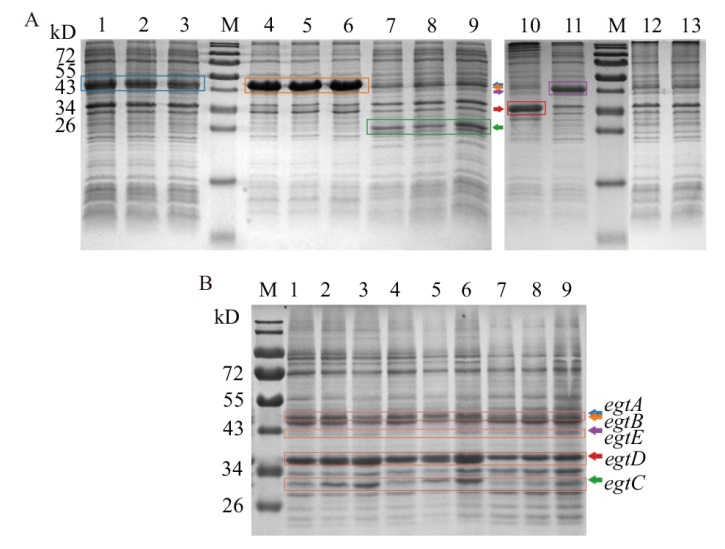
图3 工程菌表达鉴定结果 A:单基因表达SDS-PAGE结果(M:蛋白marker;1-3:egtA的表达结果;4-6:egtB的表达结果;7-9:egtC的表达结果;10-12:分别为egtD、egtE和NC(E. coli Rosetta2(DE3)/pET28a)的表达结果);B:RE菌株表达SDS-PAGE结果(M:蛋白marker;1-3,4-6:RE菌株在诱导后2、4和20 h的表达结果;7-9:NC的表达结果)
Fig. 3 Identification results of engineering bacteria expression A: SDS-PAGE results of single gene expression(M: Protein marker; 1-3: expression results for egtA; 4-6: expression results for egtB; 7-9: expression results for egtC; 10-12: expression results of egtD, egtE and NC(E. coli Rosetta2(DE3)/pET28a)respectively). B: RE strain expression SDS-PAGE results(M: Protein marker; 1-3, 4-6: expression results of RE strain at 2, 4, and 20 h after induction, respectively; 7-9: expression results of NC)

图4 发酵液中麦角硫因检测 A:HPLC定量曲线;B:LC-MS检测结果,i:发酵液上清液的高效液相色谱结果显示(使用氨基柱),目标化合物的保留时间为17.98 min;ii:保留时间在(17.985 1±0.058 5)min范围内检出峰的m/z结果图;C:发酵产物与ERG标准品HPLC结果对比
Fig. 4 Determination of ergothioneine content in liquid fermentation broth A: HPLC quantitation profile. B: LC-MS detection outcomes. i: The HPLC results of the supernatant of the fermentation broth(using an amino column)show that the retention time of the target compounds is 17.98 min; ii: m/z result plots for peaks with retention times in the range of(17.985 1±0.058 5)min. C: Comparison of HPLC results of fermentation products and ERG standards
| 诱导剂 Inducers | 诱导温度 Induction temperature/℃ | 麦角硫因含量 ERG content/(mg·L-1) | 上清中蛋白含量 Protein content of supernatant | 沉淀中蛋白含量 Protein content of precipitation |
|---|---|---|---|---|
| 0.4 mmol/L IPTG | 37 | 10.7 | + | +++ |
| 28 | 17.3 | ++ | ++ | |
| 16 | 9.8 | +++ | + | |
| 0.8% lactose | 37 | 5.4 | + | ++ |
| 28 | 10.2 | + | + | |
| 16 | 3.3 | + | + |
表6 表达条件优化
Table 6 Optimization of expression conditions
| 诱导剂 Inducers | 诱导温度 Induction temperature/℃ | 麦角硫因含量 ERG content/(mg·L-1) | 上清中蛋白含量 Protein content of supernatant | 沉淀中蛋白含量 Protein content of precipitation |
|---|---|---|---|---|
| 0.4 mmol/L IPTG | 37 | 10.7 | + | +++ |
| 28 | 17.3 | ++ | ++ | |
| 16 | 9.8 | +++ | + | |
| 0.8% lactose | 37 | 5.4 | + | ++ |
| 28 | 10.2 | + | + | |
| 16 | 3.3 | + | + |

图5 前体氨基酸对ERG产量的影响 A:外源添加前体氨基酸单因素实验结果;B:底盘细胞改造摇瓶验证;*P≤0.05,**P≤0.01
Fig. 5 Effects of precursor amino acids on ERG production A: Results of one-factor experiments with exogenously added precursor amino acids. B: Chassis cell modification shake flask validation; *P≤0.05, **P≤0.01
| 实验号 Experiment No. | 列号Column number | ERG产量 ERG yield/(mg·L-1) | ||
|---|---|---|---|---|
| A | B | C | ||
| 1 | 1 | 1 | 1 | 77.4±4.2 |
| 2 | 1 | 2 | 3 | 81.8±1.4 |
| 3 | 1 | 3 | 2 | 81.4±2.0 |
| 4 | 2 | 1 | 3 | 82.8±1.9 |
| 5 | 2 | 2 | 2 | 89.7±2.0 |
| 6 | 2 | 3 | 1 | 82.5±1.2 |
| 7 | 3 | 1 | 2 | 80.8±2.2 |
| 8 | 3 | 2 | 1 | 82.7±3.5 |
| 9 | 3 | 3 | 3 | 91.2±6.0 |
| Ij | 240.7 | 241.0 | 242.7 | T=750.5 |
| IIj | 255.1 | 254.3 | 251.9 | |
| IIIj | 254.8 | 255.2 | 255.9 | |
| Rj | 14.4 | 14.2 | 13.2 | |
表7 正交实验结果
Table 7 Results of orthogonal experiments
| 实验号 Experiment No. | 列号Column number | ERG产量 ERG yield/(mg·L-1) | ||
|---|---|---|---|---|
| A | B | C | ||
| 1 | 1 | 1 | 1 | 77.4±4.2 |
| 2 | 1 | 2 | 3 | 81.8±1.4 |
| 3 | 1 | 3 | 2 | 81.4±2.0 |
| 4 | 2 | 1 | 3 | 82.8±1.9 |
| 5 | 2 | 2 | 2 | 89.7±2.0 |
| 6 | 2 | 3 | 1 | 82.5±1.2 |
| 7 | 3 | 1 | 2 | 80.8±2.2 |
| 8 | 3 | 2 | 1 | 82.7±3.5 |
| 9 | 3 | 3 | 3 | 91.2±6.0 |
| Ij | 240.7 | 241.0 | 242.7 | T=750.5 |
| IIj | 255.1 | 254.3 | 251.9 | |
| IIIj | 254.8 | 255.2 | 255.9 | |
| Rj | 14.4 | 14.2 | 13.2 | |
| 工艺 Process | 菌株 Strain | 补料 Feeding | 发酵规模 Scale/L | 周期 Period/h | 湿重Wet weight/(g·L-1) | 产量Yield/(g·L-1) | 产率Productivity/(mg·L-1·h-1) | 单位湿重菌体产量Yield per wet weight/(mg·g-1) |
|---|---|---|---|---|---|---|---|---|
| 常规诱导Routine induction | RE | Met | 10 | 142.0 | 12.3 | 0.2 | 1.2 | 13.7 |
| 自诱导 Self-induced | RE | Met | 10 | 135.5 | 123.2 | 0.4 | 3.1 | 3.4 |
| RE-CH | Met | 10 | 114.6 | 84.8 | 0.5 | 4.3 | 5.8 | |
| 自诱导优化Self-induced optimization | RE-CH | Met | 10 | 118.0 | 91.5 | 1.1 | 9.7 | 12.0 |
| 自诱导放大Self-induced scale-up | RE-CH | Met, His, Cys | 30 | 95.5 | 142.6 | 4.3 | 45.2 | 30.2 |
表8 10 L和30 L罐发酵结果
Table 8 Results of fermentation experiments in 10 L and 30 L bioreactor
| 工艺 Process | 菌株 Strain | 补料 Feeding | 发酵规模 Scale/L | 周期 Period/h | 湿重Wet weight/(g·L-1) | 产量Yield/(g·L-1) | 产率Productivity/(mg·L-1·h-1) | 单位湿重菌体产量Yield per wet weight/(mg·g-1) |
|---|---|---|---|---|---|---|---|---|
| 常规诱导Routine induction | RE | Met | 10 | 142.0 | 12.3 | 0.2 | 1.2 | 13.7 |
| 自诱导 Self-induced | RE | Met | 10 | 135.5 | 123.2 | 0.4 | 3.1 | 3.4 |
| RE-CH | Met | 10 | 114.6 | 84.8 | 0.5 | 4.3 | 5.8 | |
| 自诱导优化Self-induced optimization | RE-CH | Met | 10 | 118.0 | 91.5 | 1.1 | 9.7 | 12.0 |
| 自诱导放大Self-induced scale-up | RE-CH | Met, His, Cys | 30 | 95.5 | 142.6 | 4.3 | 45.2 | 30.2 |
| 发酵周期 Period/h | 上清Supernatant | 菌体Microorganism | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Cys | Met | His | γGC | Cys | Met | His | γGC | ||
| 15.9 | - | + | - | - | - | + | - | - | |
| 20.7 | - | + | - | - | - | + | - | - | |
| 38.4 | - | - | - | - | - | + | + | + | |
| 43.8 | - | - | - | - | - | - | - | + | |
| 46.3 | - | - | - | - | - | + | + | + | |
| 62.2 | - | + | - | - | - | + | - | + | |
| 86.4 | - | + | - | - | - | + | - | + | |
| 95.0 | - | + | - | - | - | + | + | + | |
| 111.0 | - | + | - | - | - | + | + | + | |
| 114.6 | - | + | - | - | - | + | - | + | |
表9 发酵液中主要前体和重要中间体检测结果
Table 9 Determination of major precursors and important intermediates in fermentation broths
| 发酵周期 Period/h | 上清Supernatant | 菌体Microorganism | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Cys | Met | His | γGC | Cys | Met | His | γGC | ||
| 15.9 | - | + | - | - | - | + | - | - | |
| 20.7 | - | + | - | - | - | + | - | - | |
| 38.4 | - | - | - | - | - | + | + | + | |
| 43.8 | - | - | - | - | - | - | - | + | |
| 46.3 | - | - | - | - | - | + | + | + | |
| 62.2 | - | + | - | - | - | + | - | + | |
| 86.4 | - | + | - | - | - | + | - | + | |
| 95.0 | - | + | - | - | - | + | + | + | |
| 111.0 | - | + | - | - | - | + | + | + | |
| 114.6 | - | + | - | - | - | + | - | + | |
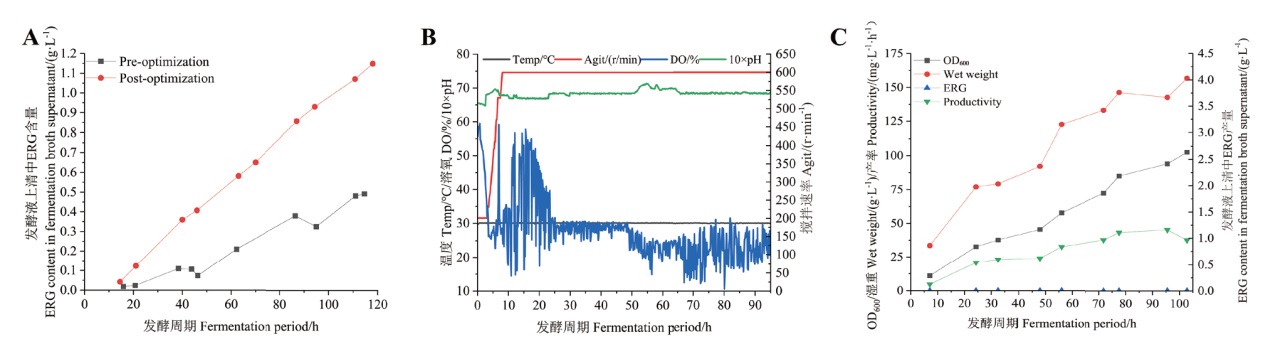
图6 发酵曲线及产量曲线 A:10 L罐发酵工艺优化前后产量比较;B:30 L罐扩大发酵发酵曲线;C:30 L罐发酵产量曲线
Fig. 6 Fermentation and yield curves A: Comparison of yield before and after optimization of 10 L-bioreactor process. B: Fermentation curves for enlarged fermentation in 30 L-bioreactor. C: Yield curve for fermentation in 30 L-bioreactor
| [1] | Melville DB. Ergothioneine[M]// Vitamins & Hormones. Amsterdam: Elsevier, 1959: 155-204. |
| [2] |
Hunter G. A new test for ergothioneine upon which is based a method for its estimation in simple solution and in blood-filtrates[J]. Biochem J, 1928, 22(1): 4-10.
pmid: 16744012 |
| [3] |
Chen ZH, He YZ, Wu XY, et al. Toward more efficient ergothioneine production using the fungal ergothioneine biosynthetic pathway[J]. Microb Cell Fact, 2022, 21(1): 76.
doi: 10.1186/s12934-022-01807-3 pmid: 35525939 |
| [4] |
Chen L, Zhang LP, Ye XJ, et al. Ergothioneine and its congeners: anti-ageing mechanisms and pharmacophore biosynthesis[J]. Protein Cell, 2024, 15(3): 191-206.
doi: 10.1093/procel/pwad048 |
| [5] | Duan R, Pan HT, Li DC, et al. Ergothioneine improves myocardial remodeling and heart function after acute myocardial infarction via S-glutathionylation through the NF-ĸB dependent Wnt5a-sFlt-1 pathway[J]. Eur J Pharmacol, 2023, 950: 175759. |
| [6] | Mayayo-Vallverdú C, López de Heredia M, Prat E, et al. The antioxidant L-Ergothioneine prevents cystine lithiasis in the Slc7a9-/- mouse model of cystinuria[J]. Redox Biol, 2023, 64: 102801. |
| [7] | Wang Z, Ma JW, Miao ZM, et al. Ergothioneine inhibits the progression of osteoarthritis via the Sirt6/NF-κB axis both in vitro and in vivo[J]. Int Immunopharmacol, 2023, 119: 110211. |
| [8] | Chen Q, Zhou RR, Yang C, et al. Ergothioneine attenuates varicocele-induced testicular damage by upregulating HSP90AA1 in rats[J]. J Biochem Mol Toxicol, 2023, 37(4): e23301. |
| [9] | Leow DMK, Cheah IKM, Fong ZWJ, et al. Protective effect of ergothioneine against 7-ketocholesterol-induced mitochondrial damage in hCMEC/D3 human brain endothelial cells[J]. Int J Mol Sci, 2023, 24(6): 5498. |
| [10] | Hartmann L, Seebeck FP, Schmalz HG, et al. Isotope-labeled ergothioneine clarifies the mechanism of reaction with singlet oxygen[J]. Free Radic Biol Med, 2023, 198: 12-26. |
| [11] | Iqbal S, Jabeen F, Aslam N, et al. Anti-EMT properties of ergothioneine attenuate lipopolysaccharide-induced oxidative stress-mediated acute lung injury via modulating TGF-β/smad/snail signaling pathway[J]. Hum Exp Toxicol, 2023, 42: 9603271231178015. |
| [12] | Bernardo VS, Torres FF, de Paula CP, et al. Potential cytoprotective and regulatory effects of ergothioneine on gene expression of proteins involved in erythroid adaptation mechanisms and redox pathways in K562 cells[J]. Genes, 2022, 13(12): 2368. |
| [13] |
Nakamichi N, Tsuzuku S, Shibagaki F. Ergothioneine and central nervous system diseases[J]. Neurochem Res, 2022, 47(9): 2513-2521.
doi: 10.1007/s11064-022-03665-2 pmid: 35788879 |
| [14] |
Alamgir KM, Masuda S, Fujitani Y, et al. Production of ergothioneine by Methylobacterium species[J]. Front Microbiol, 2015, 6: 1185.
doi: 10.3389/fmicb.2015.01185 pmid: 26579093 |
| [15] | Pluskal T, Ueno M, Yanagida M. Genetic and metabolomic dissection of the ergothioneine and selenoneine biosynthetic pathway in the fission yeast, S. pombe, and construction of an overproduction system[J]. PLoS One, 2014, 9(5): e97774. |
| [16] |
Genghof DS, Van Damme O. Biosynthesis of ergothioneine from endogenous hercynine in Mycobacterium smegmatis[J]. J Bacteriol, 1968, 95(2): 340-344.
doi: 10.1128/jb.95.2.340-344.1968 pmid: 5644441 |
| [17] |
Genghof DS, Inamine E, Kovalenko V, et al. Ergothioneine in microorganisms[J]. J Biol Chem, 1956, 223(1): 9-17.
pmid: 13376573 |
| [18] | 李亚欢. 杏鲍菇中麦角硫因的提取纯化和抗氧化活性研究[D]. 广州: 华南农业大学, 2016. |
| Li YH. Study on extraction, purification and antioxidant activity of ergothionine from Pleurotus eryngii and its antioxidant activity[D]. Guangzhou: South China Agricultural University, 2016. | |
| [19] | Xu J, Yadan JC. ChemInform Abstract: synthesis of L-(+)-ergothioneine[J]. ChemInform, 1996, 27(9): no. |
| [20] | Seebeck FP. In vitro reconstitution of mycobacterial ergothioneine biosynthesis[J]. J Am Chem Soc, 2010, 132(19): 6632-6633. |
| [21] | Qiu YB, Chen ZL, Su EZ, et al. Recent strategies for the biosynthesis of ergothioneine[J]. J Agric Food Chem, 2021, 69(46): 13682-13690. |
| [22] | 刘琦, 毛雨丰, 廖小平, 等. 麦角硫因生物合成研究的新进展[J]. 生物工程学报, 2022, 38(4): 1408-1420. |
| Liu Q, Mao YF, Liao XP, et al. Recent progress in ergothioneine biosynthesis: a review[J]. Chin J Biotechnol, 2022, 38(4): 1408-1420. | |
| [23] | Tanaka N, Kawano Y, Satoh Y, et al. Gram-scale fermentative production of ergothioneine driven by overproduction of cysteine in Escherichia coli[J]. Sci Rep, 2019, 9(1): 1895. |
| [24] | Kamide T, Takusagawa S, Tanaka N, et al. High Production of ergothioneine in Escherichia coli using the sulfoxide synthase from Methylobacterium strains[J]. J Agric Food Chem, 2020, 68(23): 6390-6394. |
| [25] | Zhang HF, Zhang YF, Zhao M, et al. Fermentative production of ergothioneine by exploring novel biosynthetic pathway and remodulating precursor synthesis pathways[J]. J Agric Food Chem, 2024, 72(25): 14264-14273. |
| [26] | Hirasawa T, Shimoyamada Y, Tachikawa Y, et al. Ergothioneine production by Corynebacterium glutamicum harboring heterologous biosynthesis pathways[J]. J Biosci Bioeng, 2023, 135(1): 25-33. |
| [27] | Liu MS, Wu JJ, Yue MY, et al. YaliCMulti and YaliHMulti: stable, efficient multi-copy integration tools for engineering Yarrowia lipolytica[J]. Metab Eng, 2024, 82: 29-40. |
| [28] | van der Hoek SA, Rusnák M, Jacobsen IH, et al. Engineering ergothioneine production in Yarrowia lipolytica[J]. FEBS Lett, 2022, 596(10): 1356-1364. |
| [29] | 陈佳敏. 大肠杆菌麦角硫因合成体系的构建与优化[D]. 无锡: 江南大学, 2023. |
| Chen JM. Construction and optimization of ergothionine synthesis system for Escherichia coli[D]. Wuxi: Jiangnan University, 2023. | |
| [30] | Zhang LW, Tang JW, Feng MQ, et al. Engineering methyltransferase and sulfoxide synthase for high-yield production of ergothioneine[J]. J Agric Food Chem, 2023, 71(1): 671-679. |
| [31] | 王丽, 王阳, 李江华, 等. 产麦角硫因大肠杆菌工程菌株的构建与优化[J]. 生物工程学报, 2022, 38(2): 796-806. |
| Wang L, Wang Y, Li JH, et al. Construction and optimization of ergothioneine-producing Escherichia coli[J]. Chin J Biotechnol, 2022, 38(2): 796-806. | |
| [32] | 陈佳敏, 王阳, 堵国成, 等. 优化前体供给与细胞膜通透性强化大肠杆菌合成麦角硫因[J]. 食品与生物技术学报, 2022, 41(8): 43-52. |
| Chen JM, Wang Y, Du GC, et al. Enhancement of ergothioneine synthesis in Escherichia coli via optimization of precursor supply and cell membrane permeability[J]. J Food Sci Biotechnol, 2022, 41(8): 43-52. | |
| [33] |
Jones GW, Doyle S, Fitzpatrick DA. The evolutionary history of the genes involved in the biosynthesis of the antioxidant ergothioneine[J]. Gene, 2014, 549(1): 161-170.
doi: 10.1016/j.gene.2014.07.065 pmid: 25068406 |
| [34] | Genghof DS, Vandamme O. Biosynthesis of ergothioneine and hercynine by mycobacteria[J]. J Bacteriol, 1964, 87(4): 852-862. |
| [35] | Xiong LB, Xie ZY, Ke J, et al. Engineering Mycolicibacterium neoaurum for the production of antioxidant ergothioneine[J]. Food Bioeng, 2022, 1(1): 26-36. |
| [36] | 刘琦, 张维亚, 姜文侠. 麦角硫因的合成与降解代谢[J]. 天然产物研究与开发, 2015, 27(6): 1112-1117, 1002. |
|
Liu Q, Zhang WY, Jiang WX. Biosynthesis and catabolism of L-ergothioneine[J]. Nat Prod Res Dev, 2015, 27(6): 1112-1117, 1002.
doi: 10.16333/j.1001-6880.2015.06.031 |
|
| [37] | Studier FW. Protein production by auto-induction in high density shaking cultures[J]. Protein Expr Purif, 2005, 41(1): 207-234. |
| [38] | Tahara N, Tachibana I, Takeo K, et al. Boosting auto-induction of recombinant proteins in Escherichia coli with glucose and lactose additives[J]. Protein Pept Lett, 2021, 28(10): 1180-1190. |
| [39] | Crowley EL, Rafferty SP. Review of lactose-driven auto-induction expression of isotope-labelled proteins[J]. Protein Expr Purif, 2019, 157: 70-85. |
| [40] | Fathi-Roudsari M, Maghsoudi N, Maghsoudi A, et al. Auto-induction for high level production of biologically active reteplase in Escherichia coli[J]. Protein Expr Purif, 2018, 151: 18-22. |
| [41] |
EL-Baky NA, Linjawi MH, Redwan EM. Auto-induction expression of human consensus interferon-alpha in Escherichia coli[J]. BMC Biotechnol, 2015, 15: 14.
doi: 10.1186/s12896-015-0128-x pmid: 25886839 |
| [42] | Studier FW. Stable expression clones and auto-induction for protein production in E. coli[J]. Methods Mol Biol, 2014, 1091: 17-32. |
| [43] | Yu YH, Pan HY, Guo LQ, et al. Successful biosynthesis of natural antioxidant ergothioneine in Saccharomyces cerevisiae required only two genes from Grifola frondosa[J]. Microb Cell Fact, 2020, 19(1): 164. |
| [44] |
Kulis-Horn RK, Persicke M, Kalinowski J. Corynebacterium glutamicum ATP-phosphoribosyl transferases suitable for L-histidine production—Strategies for the elimination of feedback inhibition[J]. J Biotechnol, 2015, 206: 26-37.
doi: 10.1016/j.jbiotec.2015.04.001 pmid: 25892668 |
| [45] |
Nakatani T, Ohtsu I, Nonaka G, et al. Enhancement of thioredoxin/glutaredoxin-mediated L-cysteine synthesis from S-sulfocysteine increases L-cysteine production in Escherichia coli[J]. Microb Cell Fact, 2012, 11: 62.
doi: 10.1186/1475-2859-11-62 pmid: 22607201 |
| [46] |
Ohtsu I, Wiriyathanawudhiwong N, Morigasaki S, et al. The L-cysteine/L-cystine shuttle system provides reducing equivalents to the periplasm in Escherichia coli[J]. J Biol Chem, 2010, 285(23): 17479-17487.
doi: 10.1074/jbc.M109.081356 pmid: 20351115 |
| [47] |
Konstantinov K, Kishimoto M, Seki T, et al. A balanced DO-stat and its application to the control of acetic acid excretion by recombinant Escherichia coli[J]. Biotechnol Bioeng, 1990, 36(7): 750-758.
pmid: 18597268 |
| [1] | 张静安, 胡孝龙, 曹蓓蓓, 廖敏, 束长龙, 张杰, 王奎, 操海群. 苏云金芽胞杆菌可视化快速表达载体的构建与特性分析[J]. 生物技术通报, 2025, 41(1): 95-102. |
| [2] | 沈真辉, 曹瑶, 杨林雷, 罗祥英, 子灵山, 陆青青, 李荣春. 金耳和毛韧革菌麦角硫因生物合成基因的克隆及生物信息学分析[J]. 生物技术通报, 2024, 40(7): 259-272. |
| [3] | 何玙冰, 付振浩, 李仁瀚, 刘秀霞, 刘春立, 杨艳坤, 李业, 白仲虎. 利用代谢工程在酿酒酵母中高效合成2-萘乙醇[J]. 生物技术通报, 2024, 40(7): 99-107. |
| [4] | 张美玉, 赵玉斌, 王灵云, 宋元达, 赵新河, 任晓洁. 微藻破囊壶菌产功能性脂肪酸DHA研究进展[J]. 生物技术通报, 2024, 40(6): 81-94. |
| [5] | 王周, 余杰, 王金华, 王永泽, 赵筱. 厌氧表达乳酸脱氢酶以提高大肠杆菌产D-乳酸光学纯度[J]. 生物技术通报, 2024, 40(5): 290-299. |
| [6] | 庄棵, 梁至轩, 何英婷, 谢秋玲. 大肠杆菌DH5α通过外膜囊泡传递抗生素抗性基因AmpR[J]. 生物技术通报, 2024, 40(12): 275-281. |
| [7] | 何思成, 张紫瑗, 韩雨晴, 苗琳, 张翠英, 于爱群. 解脂耶氏酵母细胞工厂生产多不饱和脂肪酸的研究进展[J]. 生物技术通报, 2024, 40(1): 72-85. |
| [8] | 李亮, 徐姗姗, 姜艳军. 生物合成法生产麦角硫因的研究进展[J]. 生物技术通报, 2024, 40(1): 86-99. |
| [9] | 杨红艳, 韩筱, 杨建军. pDNA质粒在一次性生物反应器中的放大生产研究[J]. 生物技术通报, 2024, 40(1): 168-175. |
| [10] | 薛宁, 王瑾, 李世新, 刘叶, 程海娇, 张玥, 毛雨丰, 王猛. 多基因同步调控结合高通量筛选构建高产L-苯丙氨酸的谷氨酸棒杆菌工程菌株[J]. 生物技术通报, 2023, 39(9): 268-280. |
| [11] | 程亚楠, 张文聪, 周圆, 孙雪, 李玉, 李庆刚. 乳酸乳球菌生产2'-岩藻糖基乳糖的途径构建及发酵培养基优化[J]. 生物技术通报, 2023, 39(9): 84-96. |
| [12] | 赵思佳, 王晓璐, 孙纪录, 田健, 张杰. 代谢工程改造毕赤酵母生产赤藓糖醇[J]. 生物技术通报, 2023, 39(8): 137-147. |
| [13] | 李雨真, 梅天秀, 李治文, 王淇, 李俊, 邹岳, 赵心清. 红酵母基因组和代谢工程改造研究进展[J]. 生物技术通报, 2023, 39(7): 67-79. |
| [14] | 陈彩萍, 任昊, 龙腾飞, 何冰, 鲁兆祥, 孙坚. 大肠杆菌Nissle 1917对炎症性肠病治疗作用的研究进展[J]. 生物技术通报, 2023, 39(6): 109-118. |
| [15] | 郁慧丽, 李爱涛. 细胞色素P450酶在香精香料绿色生物合成中的应用[J]. 生物技术通报, 2023, 39(4): 24-37. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||