








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 186-197.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0606
李彩霞1( ), 李艺1, 穆宏秀1, 林俊轩1, 白龙强1,2, 孙美华1,2, 苗妍秀1,2(
), 李艺1, 穆宏秀1, 林俊轩1, 白龙强1,2, 孙美华1,2, 苗妍秀1,2( )
)
收稿日期:2024-06-25
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
苗妍秀,女,博士,副教授,研究方向:设施环境调控与园艺植物栽培生理;E-mail: miaoyanxiu@163.com作者简介:李彩霞,女,硕士研究生,研究方向:设施园艺工程与环境调控;E-mail: lcx6283@163.com
基金资助:
LI Cai-xia1( ), LI Yi1, MU Hong-xiu1, LIN Jun-xuan1, BAI Long-qiang1,2, SUN Mei-hua1,2, MIAO Yan-xiu1,2(
), LI Yi1, MU Hong-xiu1, LIN Jun-xuan1, BAI Long-qiang1,2, SUN Mei-hua1,2, MIAO Yan-xiu1,2( )
)
Received:2024-06-25
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】基于中国南瓜全基因组数据,鉴定中国南瓜bHLH转录因子家族成员并分析其理化性质和结构等,为中国南瓜bHLH转录因子家族的生物学功能研究提供理论支撑。【方法】利用NCBI和中国南瓜基因组数据库等对中国南瓜bHLH基因家族进行鉴定,利用ExPASy ProtParam tool、DNAMAN和MAGA5.1等软件进行基本理化性质、系统进化、保守基序、启动子作用元件和表达模式等生物信息学分析。【结果】在中国南瓜全基因组中共鉴定出153个CmobHLHs基因,不均等地分布在20条染色体上。CmobHLH蛋白质含有91-897个氨基酸,理论等电点介于4.77-10.33;亚细胞定位预测表明有139个CmobHLH蛋白位于细胞核;系统进化分析将CmobHLH蛋白分为8个亚家族;保守基序(motif)预测结果表明,中国南瓜共含10条motif,其中motif1和motif2普遍存在于153个bHLH蛋白中。CmobHLHs在中国南瓜的各个部位均有表达,CmobHLH41、CmobHLH5、CmobHLH36和CmobHLH88分别在果实、叶片、茎和根中表达量最高;盐胁迫下,100个CmobHLHs基因表达量上调,49个CmobHLHs基因表达量下调。RT-qPCR结果表明盐胁迫下CmobHLH52和CmobHLH148基因表达量显著上调,而CmobHLH8、CmobHLH83、CmobHLH93和CmobHLH115基因表达量显著下调,与转录组结果基本一致。【结论】鉴定出153个CmobHLHs基因,家族成员在不同部位和盐胁迫下的表达模式存在差异,为挖掘南瓜bHLH转录因子在抗盐等方面的功能提供理论基础。
李彩霞, 李艺, 穆宏秀, 林俊轩, 白龙强, 孙美华, 苗妍秀. 中国南瓜bHLH转录因子家族的鉴定与生物信息学分析[J]. 生物技术通报, 2025, 41(1): 186-197.
LI Cai-xia, LI Yi, MU Hong-xiu, LIN Jun-xuan, BAI Long-qiang, SUN Mei-hua, MIAO Yan-xiu. Identification and Bioinformatics Analysis of the bHLH Transcription Factor Family in Cucurbita moschata Duch.[J]. Biotechnology Bulletin, 2025, 41(1): 186-197.
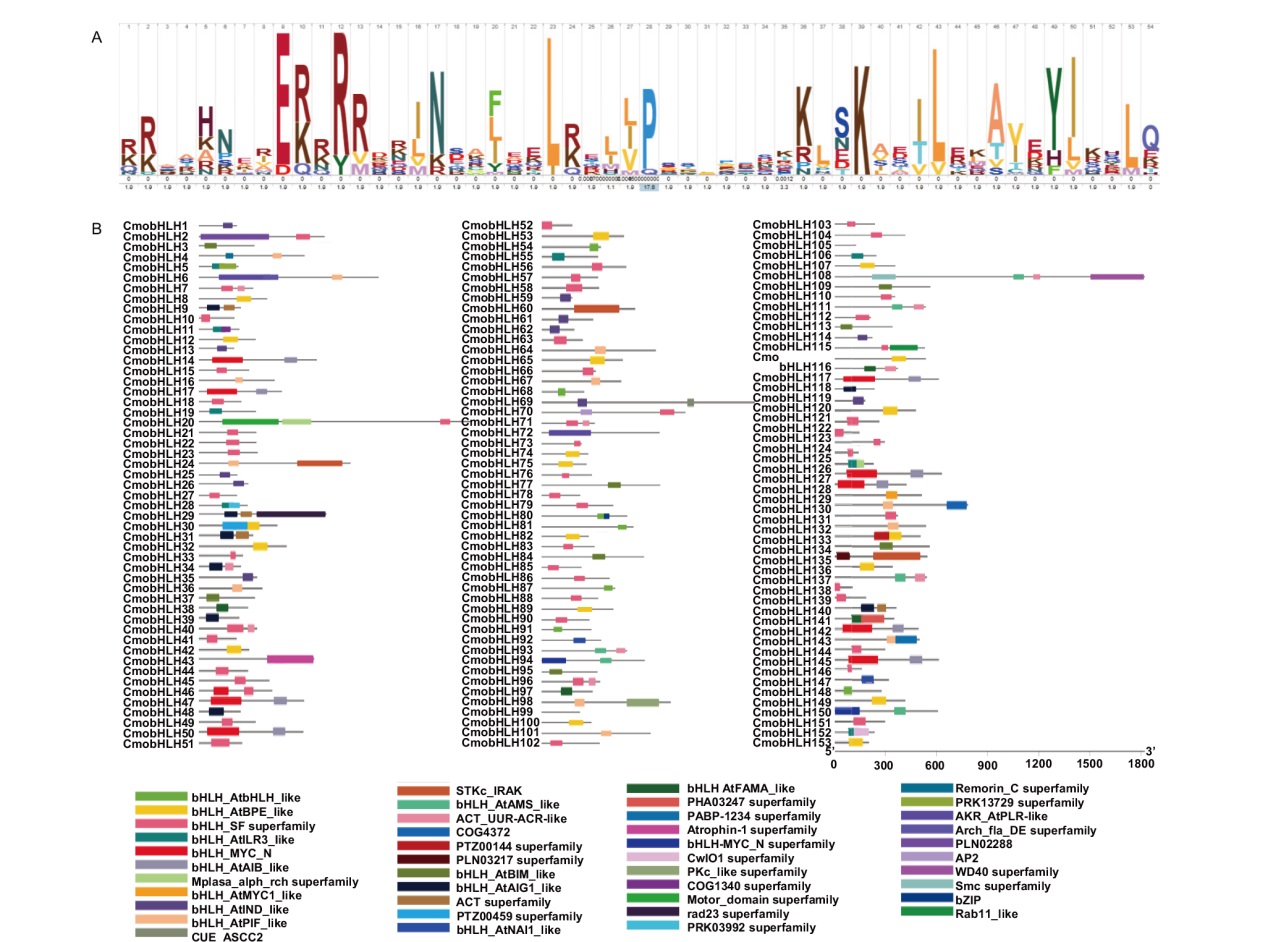
图2 中国南瓜bHLH家族的HLH保守结构域及分布 A:HLH保守结构域,不同位点上氨基酸字母的高度表示结构域保守性的高低;B:中国南瓜bHLH家族其他43类保守结构域的分布
Fig. 2 HLH conservative domain and distribution of bHLH family in C. moschata Duch. A: HLH conserved domain, where the height of amino acid letters at different sites indicates the level of domain conservation. B: Distribution of 43 other conserved domains of bHLH family in C. moschata Duch.

图3 中国南瓜bHLH家族的进化和结构分析 A:系统进化树;B:基因保守基序分析;C:内含子-外显子分布;内含子用黑线表示,外显子用绿色框表示;比例尺显示在底部
Fig. 3 Phylogenetic and structural analysis of the bHLH family in C. moschata Duch. A: Phylogenetic tree. B: Gene motif analysis. C: Intron and exon distribution. Introns are indicated by black lines, exons are indicated by green boxes. The scale bar is shown at the bottom

图5 中国南瓜bHLH基因家族组织表达分析 F、L、S、R分别代表果、叶、茎、根
Fig. 5 Tissue expression analysis of bHLH gene family in C. moschata Duch. F, L, S, and R indicate fruits, leaves, stems, and roots, respectively
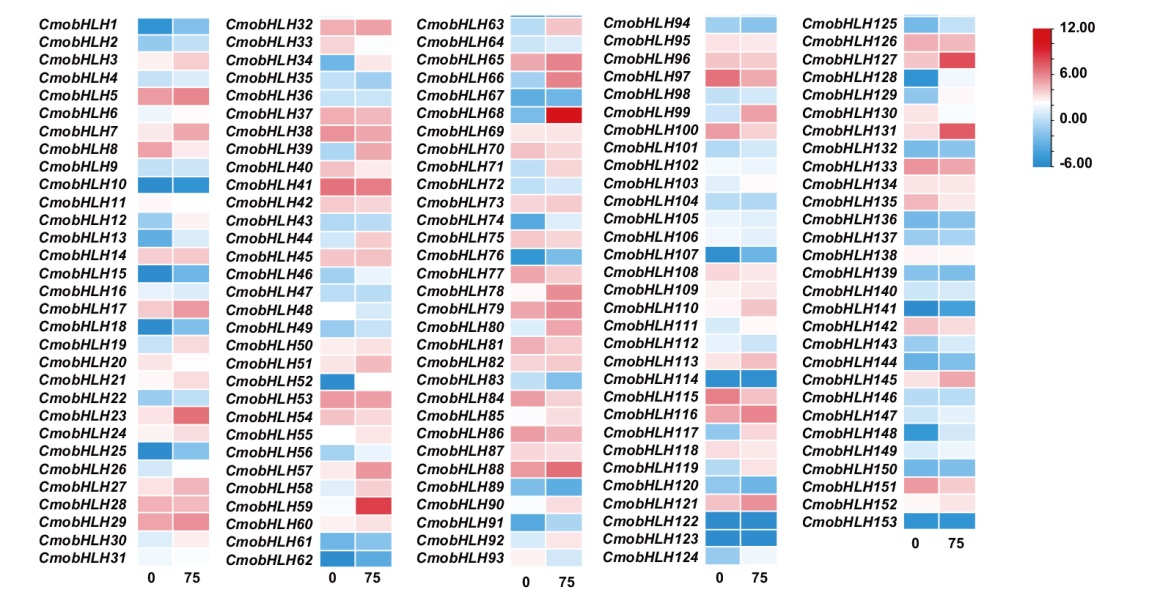
图6 中国南瓜bHLH基因家族响应盐胁迫的表达分析 “0”代表0 mmol/L NaCl;“75”代表75 mmol/L NaCl
Fig. 6 Expression analysis of bHLH gene family in response to salt stress in C. moschata Duch. “0” indicates 0 mmol/L NaCl; “75” indicates 75 mmol/L NaCl
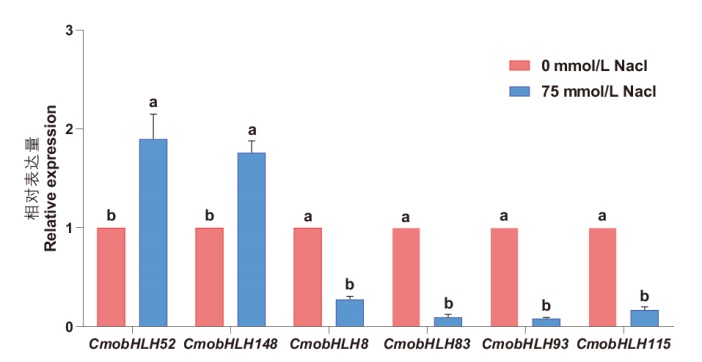
图7 中国南瓜bHLH基因家族响应盐胁迫的RT-qPCR验证 不同小写字母表示在P<0.05水平差异显著
Fig. 7 RT-qPCR validation of bHLH gene family in response to salt stress in C. moschata Duch. Different lower letters indicate significant difference at P<0.05 level
| [1] |
Robinson KA, Koepke JI, Kharodawala M, et al. A network of yeast basic helix-loop-helix interactions[J]. Nucleic Acids Res, 2000, 28(22): 4460-4466.
doi: 10.1093/nar/28.22.4460 pmid: 11071933 |
| [2] |
Murre C, McCaw PS, Baltimore D. A new DNA binding and dimerization motif in immunoglobulin enhancer binding, daughterless, MyoD, and myc proteins[J]. Cell, 1989, 56(5): 777-783.
doi: 10.1016/0092-8674(89)90682-x pmid: 2493990 |
| [3] |
Ledent V, Vervoort M. The basic helix-loop-helix protein family: comparative genomics and phylogenetic analysis[J]. Genome Res, 2001, 11(5): 754-770.
doi: 10.1101/gr.177001 pmid: 11337472 |
| [4] |
Jones S. An overview of the basic helix-loop-helix proteins[J]. Genome Biol, 2004, 5(6): 226.
doi: 10.1186/gb-2004-5-6-226 pmid: 15186484 |
| [5] |
Heim MA, Jakoby M, Werber M, et al. The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity[J]. Mol Biol Evol, 2003, 20(5): 735-747.
doi: 10.1093/molbev/msg088 pmid: 12679534 |
| [6] | Li XX, Duan XP, Jiang HX, et al. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis[J]. Plant Physiol, 2006, 141(4): 1167-1184. |
| [7] | 唐文武, 吴秀兰, 钟佩桥. 白菜bHLH转录因子家族的全基因鉴定及表达特征分析[J]. 江西农业学报, 2020, 32(6): 1-5. |
| Tang WW, Wu XL, Zhong PQ. Genome-wide identification and expression analysis of bHLH transcription factor family in B. rapa[J]. Acta Agric Jiangxi, 2020, 32(6): 1-5. | |
| [8] | 郭鹏宇. 番茄bHLH转录因子SlPRE3的克隆与功能研究[D]. 重庆: 重庆大学, 2021. |
| Guo PY. Cloning and functional study of tomato bHLH transcription factor SlPRE3[D]. Chongqing: Chongqing University, 2021. | |
| [9] | 薛宝平. 辣椒bHLH基因家族的鉴定、表达分析及CabHLH94在应答青枯菌侵染中的作用[D]. 延安: 延安大学, 2019. |
| Xue BP. Identification and expression analysis of bHLH gene family in pepper and the role of CabHLH94 in responding to Ralstonia solanacearum infection[D]. Yan'an: Yan'an University, 2019. | |
| [10] | 张鲲, 史后蕊, 马婧, 等. 花生bHLH基因家族的鉴定与表达分析[J/OL]. 分子植物育种, 2022. http://kns.cnki.net/kcms/detail/46.1068.S.20221121.0836.002.html |
| Zhang K, Shi HR, Ma J, et al. Identification and expression analysis of bHLH gene family in peanut[J/OL]. Mol Plant Breed, 2022. http://kns.cnki.net/kcms/detail/46.1068.S.20221121.0836.002.html | |
| [11] | 何开平, 吴楚. bHLH转录因子对植物形态发生的影响[J]. 安徽农业科学, 2010, 38(35): 19957-19959. |
| He KP, Wu C. The effects of bHLH transcription factors on plant morphogenesis[J]. J Anhui Agric Sci, 2010, 38(35): 19957-19959. | |
| [12] | 唐琪琦, 李金帅, 邱帅, 等. 绣球花型全基因组关联分析[J]. 植物遗传资源学报, 2023, 24(4): 1174-1185. |
| Tang QQ, Li JS, Qiu S, et al. Genome-wide association study of Hydrangea macrophylla inflorescence type[J]. J Plant Genet Resour, 2023, 24(4): 1174-1185. | |
| [13] | Xiang LL, Liu XF, Shi YN, et al. Comparative transcriptome analysis revealed two alternative splicing bHLHs account for flower color alteration in Chrysanthemum[J]. Int J Mol Sci, 2021, 22(23): 12769. |
| [14] |
Zhang F, Gonzalez A, Zhao MZ, et al. A network of redundant bHLH proteins functions in all TTG1-dependent pathways of Arabidopsis[J]. Development, 2003, 130(20): 4859-4869.
doi: 10.1242/dev.00681 pmid: 12917293 |
| [15] | Liu WW, Tai HH, Li SS, et al. bHLH122 is important for drought and osmotic stress resistance in Arabidopsis and in the repression of ABA catabolism[J]. New Phytol, 2014, 201(4): 1192-1204. |
| [16] | Wang PF, Su L, Gao HH, et al. Genome-wide characterization of bHLH genes in grape and analysis of their potential relevance to abiotic stress tolerance and secondary metabolite biosynthesis[J]. Front Plant Sci, 2018, 9: 64. |
| [17] | Shen TJ, Wen XP, Wen Z, et al. Genome-wide identification and expression analysis of bHLH transcription factor family in response to cold stress in sweet cherry(Prunus avium L.)[J]. Sci Hortic, 2021, 279: 109905. |
| [18] | Dong Y, Wang CP, Han X, et al. A novel bHLH transcription factor PebHLH35 from Populus euphratica confers drought tolerance through regulating stomatal development, photosynthesis and growth in Arabidopsis[J]. Biochem Biophys Res Commun, 2014, 450(1): 453-458. |
| [19] | 徐颖超, 张思程, 薛舒丹, 等. 南瓜叶黄素基因紧密连锁的InDel分子标记开发及应用[J]. 江苏农业学报, 2024, 40(2): 348-358. |
| Xu YC, Zhang SC, Xue SD, et al. Development and application of closely linked InDel molecular markers of lutein gene in Cucurbita moschata Duch[J]. Jiangsu J Agric Sci, 2024, 40(2): 348-358. | |
| [20] | 杨建坤, 殷缘, 车易达, 等. 胁迫条件下山新杨PdPapMYC2基因的组织表达模式[J]. 西北农林科技大学学报: 自然科学版, 2023, 51(6): 18-27. |
| Yang JK, Yin Y, Che YD, et al. Tissue-specific expression profiles of PdPapMYC2 in Shanxin poplar under various stresses[J]. J Northwest A F Univ Nat Sci Ed, 2023, 51(6): 18-27. | |
| [21] |
隋心意, 赵小刚, 毛欣, 等. 生菜bHLH转录因子家族的鉴定与生物信息学分析[J]. 中国农学通报, 2023, 39(3): 104-110.
doi: 10.11924/j.issn.1000-6850.casb2022-0214 |
| Sui XY, Zhao XG, Mao X, et al. Identification and bioinformatics analysis of bHLH transcription factor family in Lactuca sativaL[J]. Chin Agric Sci Bull, 2023, 39(3): 104-110. | |
| [22] | Eailey TL, Elkan C. Proceedings of the second international conference on intelligent systems for molecular biology[C]. Menlo Park, California: AAAI Press, 1994: 28-36. |
| [23] | 李君霞, 樊永强, 代书桃, 等. bHLH转录因子在植物抗非生物胁迫基因工程中的应用进展[J]. 江苏农业科学, 2022, 50(12): 1-9. |
| Li JX, Fan YQ, Dai ST, et al. Application progress of bHLH transcription factor in plant abiotic stress resistance genetic engineering[J]. Jiangsu Agric Sci, 2022, 50(12): 1-9. | |
| [24] | 高珂. 基于与BcAS1启动子互作及表达同步性筛选调控柴胡皂苷合成转录因子[D]. 北京: 北京协和医学院, 2016. |
| Gao K. Screening transcription factors regulating saikosaponin synthesis based on interaction with BcAS1 promoter and synchronization of expression[D]. Beijing: Peking Union Medical College, 2016. | |
| [25] |
Bailey PC, Martin C, Toledo-Ortiz G, et al. Update on the basic helix-loop-helix transcription factor gene family in Arabidopsis thaliana[J]. Plant Cell, 2003, 15(11): 2497-2502.
pmid: 14600211 |
| [26] | Tan C, Qiao HL, Ma M, et al. Genome-wide identification and characterization of melon bHLH transcription factors in regulation of fruit development[J]. Plants, 2021, 10(12): 2721. |
| [27] | 丁冬, 李嘉欣, 魏玉磊, 等. 拟南芥和玉米中膜结合bHLH转录因子的鉴定与分析[J]. 植物生理学报, 2020, 56(4): 700-710. |
| Ding D, Li JX, Wei YL, et al. Identification and analysis of membrane-bound bHLH transcription factors in Arabidopsis thaliana and maize[J]. Plant Physiol J, 2020, 56(4): 700-710. | |
| [28] | 李晓星, 蒋海雄, 黄青云, 等. 水稻bHLH基因家族成员表达模式分析[J]. 厦门大学学报: 自然科学版, 2006, S1: 73-77. |
| Li XX, Jiang HX, Huang QY, et al. Expression pattern analysis of rice bHLH gene family members[J]. J Xiamen Univ Nat Sci, 2006, S1: 73-77. | |
| [29] | 王艳敏, 白卉, 曹焱. bHLH转录因子研究进展及其在植物抗逆中的应用[J]. 安徽农业科学, 2015, 43(21): 34-35, 50. |
| Wang YM, Bai H, Cao Y. Research progress of bHLH transcription factor and application in plant abiotic stress tolerance[J]. J Anhui Agric Sci, 2015, 43(21): 34-35, 50. | |
| [30] |
范子培, 李龙, 史雨刚, 等. 小麦TabHLH112-2B基因克隆及每穗小穗数相关功能标记开发[J]. 作物学报, 2024, 50(2): 403-413.
doi: 10.3724/SP.J.1006.2024.31016 |
| Fan ZP, Li L, Shi YG, et al. Cloning of TabHLH112-2B gene and development of its functional marker associated with the number of spikelet per spike in wheat[J]. Acta Agron Sin, 2024, 50(2): 403-413. | |
| [31] | 刘选明, 彭玉冲, 杨远柱, 等. 水稻 bHLH转录因子 Os11g39000的功能研究[J]. 湖南大学学报: 自然科学版, 2016, 43(6): 109-116. |
| Liu XM, Peng YC, Yang YZ, et al. Functional analysis of a bHLH transcription factor Os11g39000 in rice[J]. J Hunan Univ Nat Sci, 2016, 43(6): 109-116. | |
| [32] | Zhou J, Li F, Wang JL, et al. Basic helix-loop-helix transcription factor from wild rice(OrbHLH2)improves tolerance to salt- and osmotic stress in Arabidopsis[J]. J Plant Physiol, 2009, 166(12): 1296-1306. |
| [33] | 吕宝莲, 杨宇昕, 崔立操, 等. 小麦bHLH家族转录因子的鉴定及其在盐胁迫条件下的表达分析[J]. 作物杂志, 2024(1): 65-72. |
| Lü BL, Yang YX, Cui LC, et al. Identification of bHLH family transcription factors of wheat and expression analysis under salt stress[J]. Crops, 2024(1): 65-72. | |
| [34] | 朱涛, 李芳菲, 杨海涵, 等. 山药bHLH基因家族鉴定及表达分析[J]. 信阳师范学院学报: 自然科学版, 2022, 35(3): 393-399. |
| Zhu T, Li FF, Yang HH, et al. Identification and expression ananlysis of bHLH gene family in Chinese yam(Dioscoreae Rhizoma)[J]. J Xinyang Norm Univ Nat Sci Ed, 2022, 35(3): 393-399. |
| [1] | 李雨晴, 吴楠, 罗建让. 卵叶牡丹花色苷合成相关基因bHLH的克隆与功能分析[J]. 生物技术通报, 2024, 40(8): 174-185. |
| [2] | 周麟, 黄顺满, 苏文坤, 姚响, 屈燕. 滇山茶bHLH基因家族鉴定及花色形成相关基因筛选[J]. 生物技术通报, 2024, 40(8): 142-151. |
| [3] | 王健, 杨莎, 孙庆文, 陈宏宇, 杨涛, 黄园. 金钗石斛bHLH转录因子家族全基因组鉴定及表达分析[J]. 生物技术通报, 2024, 40(6): 203-218. |
| [4] | 刘保财, 张武君, 赵云青, 黄颖桢, 陈菁瑛. 石仙桃中4种转录因子鉴定及其组织表达的研究[J]. 生物技术通报, 2024, 40(5): 141-152. |
| [5] | 刘佳宁, 李梦, 杨新森, 吴伟, 裴新梧, 袁潜华. 不同水分管理栽培方式对山栏稻根际土壤细菌群落的影响[J]. 生物技术通报, 2024, 40(3): 242-250. |
| [6] | 王斌, 袁晓, 蒋园园, 王玉昆, 肖艳辉, 何金明. bHLH96的克隆及其在薄荷萜烯生物合成调控中的功能[J]. 生物技术通报, 2024, 40(1): 281-293. |
| [7] | 林红妍, 郭晓蕊, 刘迪, 李慧, 陆海. 转录组分析转录因子AtbHLH68调控细胞壁发育的分子机制[J]. 生物技术通报, 2023, 39(9): 105-116. |
| [8] | 吕秋谕, 孙培媛, 冉彬, 王佳蕊, 陈庆富, 李洪有. 苦荞转录因子基因FtbHLH3的克隆、亚细胞定位及表达分析[J]. 生物技术通报, 2023, 39(8): 194-203. |
| [9] | 李博, 刘合霞, 陈宇玲, 周兴文, 朱宇林. 金花茶CnbHLH79转录因子的克隆、亚细胞定位及表达分析[J]. 生物技术通报, 2023, 39(8): 241-250. |
| [10] | 李宇, 李素贞, 陈茹梅, 卢海强. 植物bHLH转录因子调控铁稳态的研究进展[J]. 生物技术通报, 2023, 39(7): 26-36. |
| [11] | 鄢梦雨, 韦晓薇, 曹婧, 兰海燕. 异子蓬SabHLH169基因的克隆及抗旱功能分析[J]. 生物技术通报, 2023, 39(11): 328-339. |
| [12] | 安昌, 陆琳, 沈梦千, 陈盛圳, 叶康卓, 秦源, 郑平. 植物bHLH基因家族研究进展及在药用植物中的应用前景[J]. 生物技术通报, 2023, 39(10): 1-16. |
| [13] | 赵林艳, 官会林, 王克书, 卢燕磊, 向萍, 魏富刚, 杨绍周, 徐武美. 土壤含水量对三七连作土壤微生物群落的影响[J]. 生物技术通报, 2022, 38(7): 215-223. |
| [14] | 赵忠娟, 杨凯, 扈进冬, 魏艳丽, 李玲, 徐维生, 李纪顺. 盐胁迫条件下哈茨木霉ST02对椒样薄荷生长及根区土壤理化性质的影响[J]. 生物技术通报, 2022, 38(7): 224-235. |
| [15] | 冯建英, 李立芹, 鲁黎明. 马铃薯bHLH转录因子家族全基因组鉴定与表达分析[J]. 生物技术通报, 2022, 38(2): 21-33. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||