








• 研究报告 • 下一篇
张欣1( ), 陈骋1, 霍元博1, 黄仁丽1, 张丽1, 张凯2, 童华荣1, 袁连玉1(
), 陈骋1, 霍元博1, 黄仁丽1, 张丽1, 张凯2, 童华荣1, 袁连玉1( )
)
收稿日期:2025-09-29
出版日期:2026-03-02
通讯作者:
袁连玉,女,博士,副教授,研究方向 :茶树遗传育种及抗逆性分子机理;E-mail: yuanlianyu88@163.com作者简介:张欣,女,硕士研究生,研究方向 :茶树抗逆性分子机理;E-mail: 15085472240@163.com
基金资助:
ZHANG Xin1( ), CHEN Cheng1, HUO Yuan-Bo1, HUANG Ren-Li1, ZHANG Li1, ZHANG Kai2, TONG Hua-rong1, YUAN Lian-Yu1(
), CHEN Cheng1, HUO Yuan-Bo1, HUANG Ren-Li1, ZHANG Li1, ZHANG Kai2, TONG Hua-rong1, YUAN Lian-Yu1( )
)
Received:2025-09-29
Published:2026-03-02
摘要:
目的 Mg-原卟啉Ⅸ单甲基酯环化酶(Mg-protoporphyrin Ⅸ monomethyl ester cyclase, MgPEC)是叶绿素生物合成途径中的关键酶,探究编码该酶的CsCRD1基因对茶树叶绿素生物合成的影响,有助于解析茶树叶色形成的分子机制,为选育叶色变异的茶树品种提供科学的理论依据。 方法 利用PCR技术克隆CsCRD1基因,对其进行生物信息学、系统发育、表达模式、亚细胞定位分析,并采用反义寡核苷酸沉默技术探究CsCRD1的功能。 结果 茶树CsCRD1的CDS长度为1 215 bp,编码404个氨基酸,蛋白分子量为46.87 kD,pI为8.56,定位于茶树第10号染色体;亚细胞定位结果显示CsCRD1蛋白定位于烟草叶肉细胞的叶绿体内。CsCRD1蛋白含高度保守的Rubrerythrin结构域,与番茄CRD1亲缘关系最近,保守基序种类和顺序与其他植物CRD1一致,进化上高度保守。启动子元件分析发现,CsCRD1含光响应、激素响应及逆境胁迫响应元件。组织特异性表达分析显示,CsCRD1在茶树嫩叶及成熟叶中的相对表达量最高,花中最低。不同叶色茶树中,绿色叶片中CsCRD1的表达量显著高于白化叶片,且白化叶片返绿过程中该基因表达量会提高。相关性分析表明,CsCRD1的表达水平与茶树叶片叶绿素含量呈正相关。反义寡核苷酸沉默试验结果发现,CsCRD1沉默后叶绿素a、b及总叶绿素含量显著下降,叶绿素合成相关基因CsCHLM、CsDVR、CsPORA表达下调。 结论 茶树CsCRD1定位于叶绿体,在绿色叶片中高表达,抑制CsCRD1表达会减少叶绿素的生物合成,从而证实其在叶绿素代谢及叶色形成过程中具有关键作用。
张欣, 陈骋, 霍元博, 黄仁丽, 张丽, 张凯, 童华荣, 袁连玉. 茶树CsCRD1基因的克隆及其对叶绿素生物合成的影响[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1048.
ZHANG Xin, CHEN Cheng, HUO Yuan-Bo, HUANG Ren-Li, ZHANG Li, ZHANG Kai, TONG Hua-rong, YUAN Lian-Yu. Cloning of the CsCRD1 in Tea Plants and Its Impact on Chlorophyll Biosynthesis[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1048.
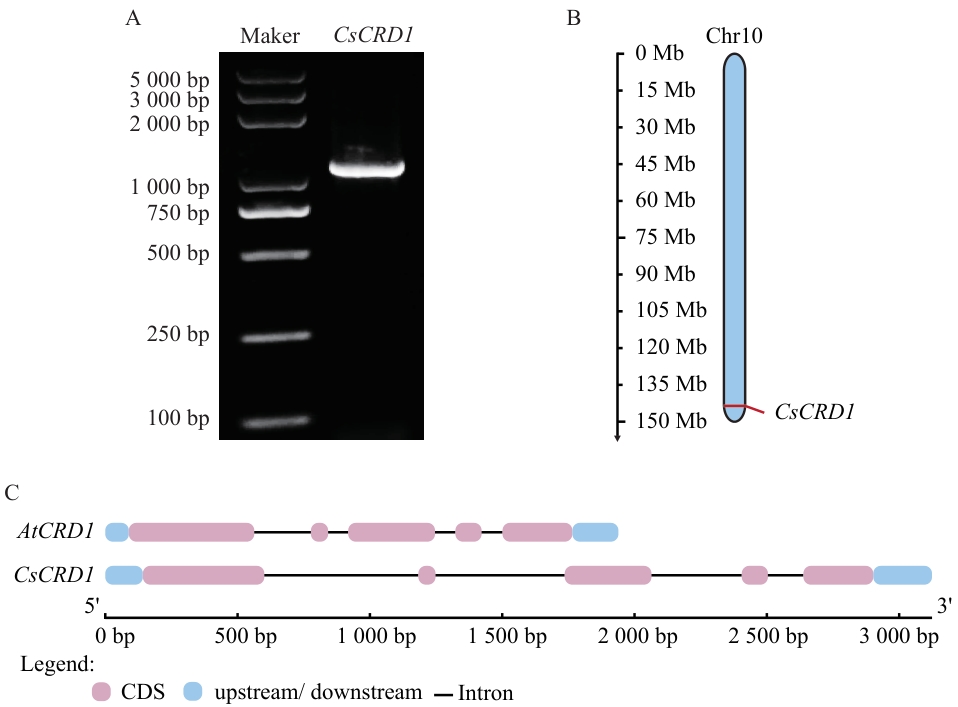
图1 茶树CsCRD1基因的克隆和特征分析A:CsCRD1基因的琼脂糖凝胶电泳图;B:CsCRD1基因染色体定位;C:CsCRD1基因结构分析
Fig. 1 Cloning and characterization analysis of CsCRD1 gene in tea plantA: Agarose gel electrophoresis image of the CsCRD1 gene. B: Chromosomal localization of CsCRD1 gene. C: Gene structural analysis of CsCRD1
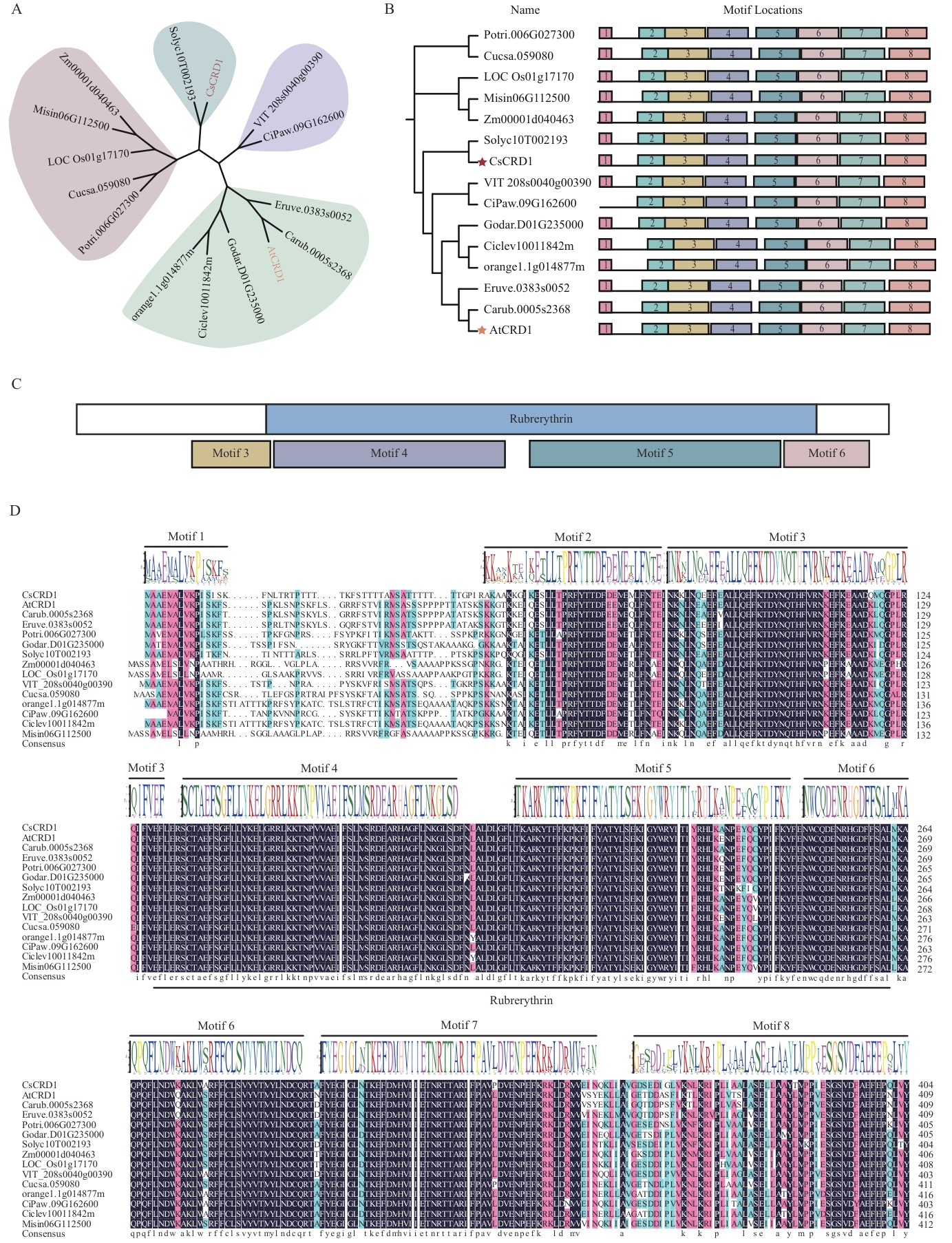
图2 植物CRD1蛋白的系统进化树和序列特征分析A:植物CRD1蛋白的系统进化树;构建系统进化树的CRD1同源蛋白来自茶树(Camellia sinensis, CsCRD1),拟南芥(Arabidopsis thaliana, AtCRD1),番茄(Solanum lycopersicum, Solyc10T002193),玉米(Zea mays, Zm00001d040463),水稻(Oryza sativa, LOC_Os02g46700),棉花(Gossypium darwinii, Godar.D01G235000),芥菜(Brassica juncea, Carub.0005s2368),芝麻菜(Eruca vesicaria, Eruve.0383s0052),毛果杨(Populus trichocarpa, Potri.006G027300),葡萄(Vitis vinifera, VIT_208s0040g00390),黄瓜(Cucumis sativus, Cucsa.059080),甜橙(Citrus sinensis, orange1.1g014877m),山核桃(Carya illinoinensis Pawnee, CiPaw.09G162600),柑橘(Citrus clementina, Ciclev10011842m),芒草(Miscanthus, Misin06G112500);B:植物CRD1蛋白的保守基序;C:植物CRD1蛋白的保守结构域示意图;D:植物CRD1蛋白的氨基酸多序列比对;蓝黑色:氨基酸一致性为100%;粉红色:氨基酸一致性为75%;蓝色:氨基酸一致性为50%。▬:植物CRD1蛋白的典型保守结构域Rubreythin
Fig. 2 Analysis of phylogenetic tree of CRD1 proteins and its sequence characteristics in plantA: Phylogenetic tree of CRD1 proteins in plant. The CRD1 homologous proteins used to construct the phylogenetic tree come from tea plant (Camellia sinensis, CsCRD1), Arabidopsis (Arabidopsis thaliana, AtCRD1), tomato (Solanum lycopersicum, Solyc10T002193), corn (Zea mays, Zm00001d040463), rice (Oryza sativa, LOC_Os02g46700), cotton (Gossypium darwinii, Godar.D01G235000), mustard (Brassica juncea, Carub.0005s2368), arugula (Eruca vesicaria, Eruve.0383s0052), poplar (Populus trichocarpa, Potri.006G027300), grape (Vitis vinifera, VIT_208s0040g00390), cucumber (Cucumis sativus, Cucsa.059080), sweet orange (Citrus sinensis, orange1.1g014877m), pecan (Carya illinoinensis Pawnee, CiPaw.09G162600), clementine (Citrus clementina, Ciclev10011842m), and Miscanthus (Miscanthus, Misin06G112500). B: Conserved motifs of CRD1 proteins in plant. C: The schematic representation for conserved structural domains of CRD1 proteins in plant. D: Amino acid multiple sequence alignment of CRD1 proteins in plant. Blue-black: Amino acid identity is 100%. Pink: Amino acid identity is 75%; blue: amino acid identity is 50%. ▬ indicates predicted the typical conserved domain Rubreythin of CRD1 protein in plants
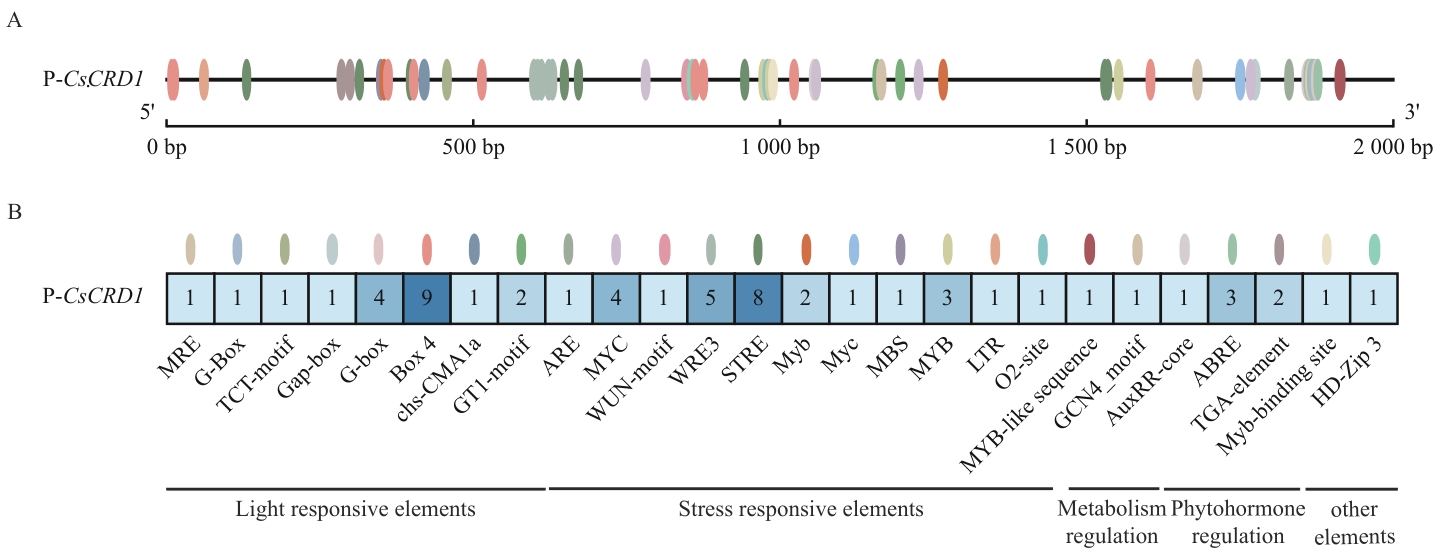
图3 茶树CsCRD1基因启动子顺式作用元件分析A:启动子顺式作用元件的分布;B:启动子顺式作用元件的数量
Fig. 3 Analysis of cis-acting elements in the promoters of gene CsCRD1 in tea plantA: Distribution of cis-acting elements in the promoter. B: Quantity of cis-acting elements in the promoter
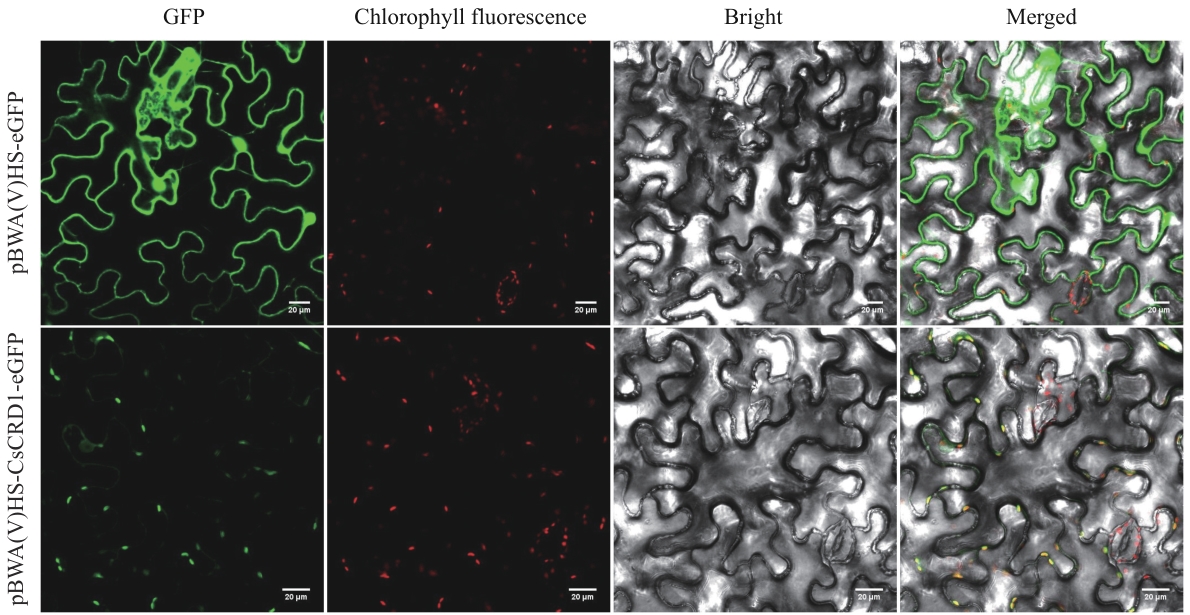
图4 茶树CsCRD1蛋白的亚细胞定位GFP:绿色荧光信号;Chlorophyll fluorescence:叶绿体自发荧光;Bright:明场;Merged:信号融合
Fig. 4 Subcellular localization of CsCRD1 protein in tea plantGFP: Green fluorescent signal. Chlorophyll fluorescence: chloroplast autofluorescence. Bright: Bright field. Merged: Signal merge
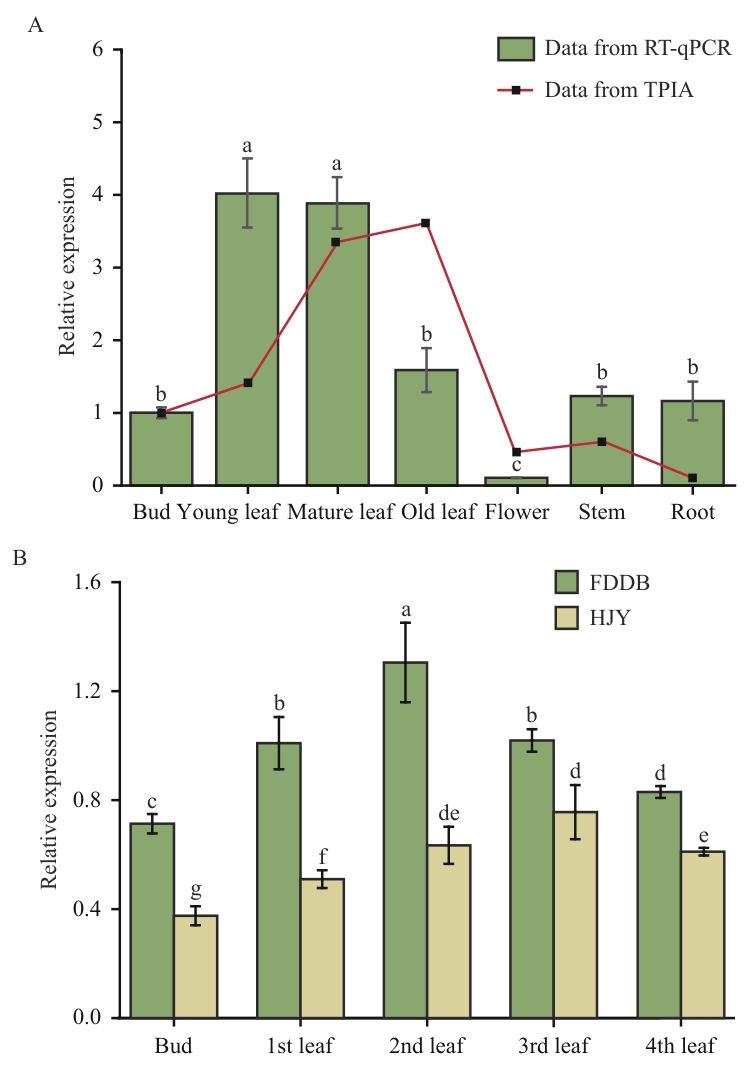
图5 茶树CsCRD1基因的组织特异性表达分析A:CsCRD1基因在茶树不同组织中的相对表达水平;B:CsCRD1基因在‘福鼎大白茶’和‘黄金叶’茶树不同叶发育时期的相对表达水平。不同小写字母表示在P<0.05水平差异显著(单因素方差分析结合Tukey事后检验)。下同
Fig. 5 Analysis of tissue-specific expression of CsCRD1 gene in tea plantA: Relative expression of the CsCRD1 gene in different tissues of the tea plant. B: Relative expression of the CsCRD1 gene in different leaf development stages of leaves from ‘Fudingdabaicha’ and ‘Huangjinye’. Different lowercase letters indicate significant differences at P<0.05 level (One-way ANOVA with Tukey’s post hoc test). The same below
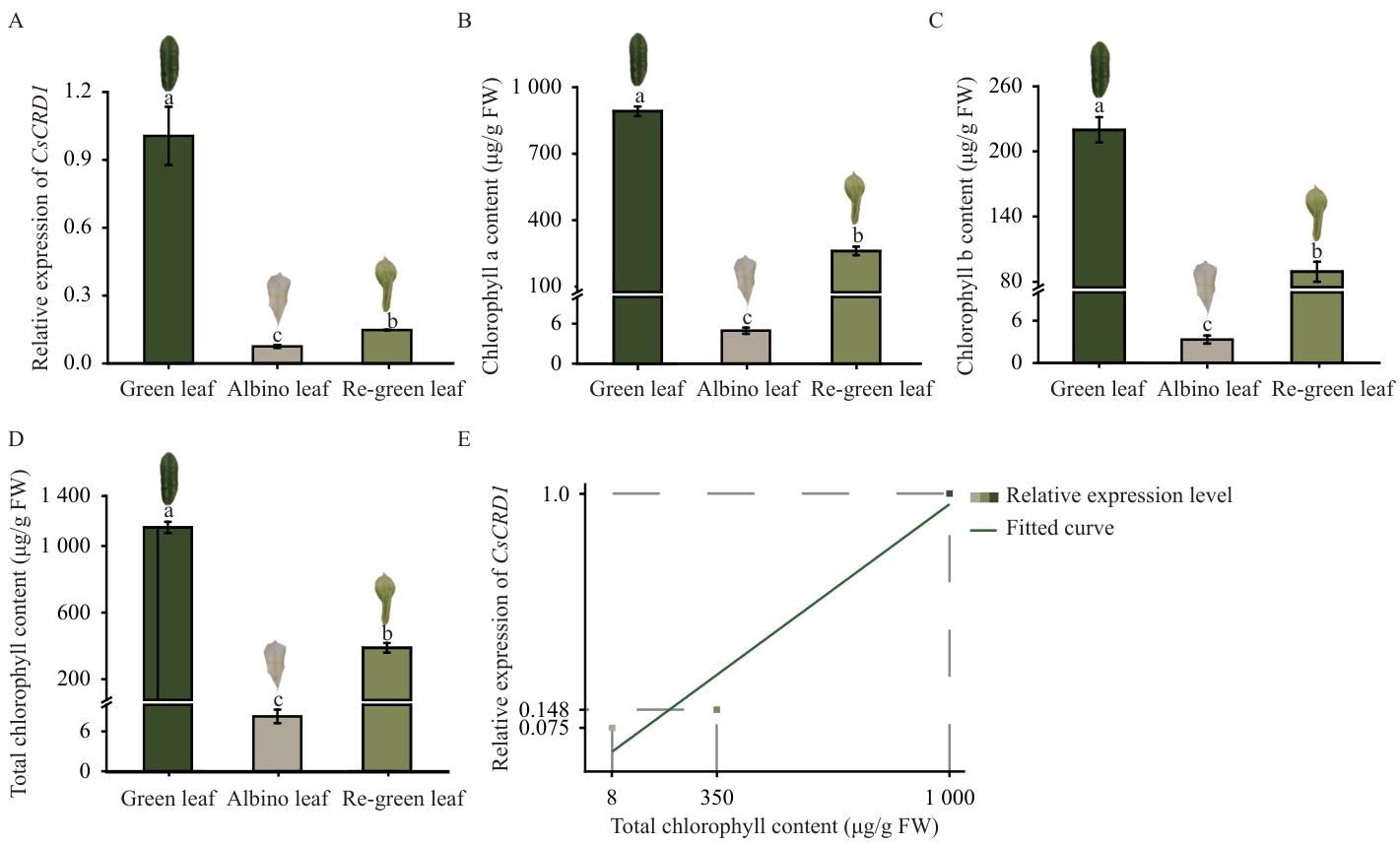
图6 茶树CsCRD1基因在叶色变化过程中的特异性表达分析A:CsCRD1基因的相对表达水平;B:叶绿素a含量;C:叶绿素b含量;D:叶绿素总量;E:叶绿素总量与CsCRD1基因相对表达水平的相关性分析
Fig. 6 Expression analysis of the CsCRD1 gene during the leaf color change process in tea plantsA: Relative expression of the CsCRD1 gene. B: Chlorophyll a content. C: Chlorophyll b content. D: Total chlorophyll content. E: Correlation analysis between total chlorophyll content and the relative expression of the CsCRD1 gene
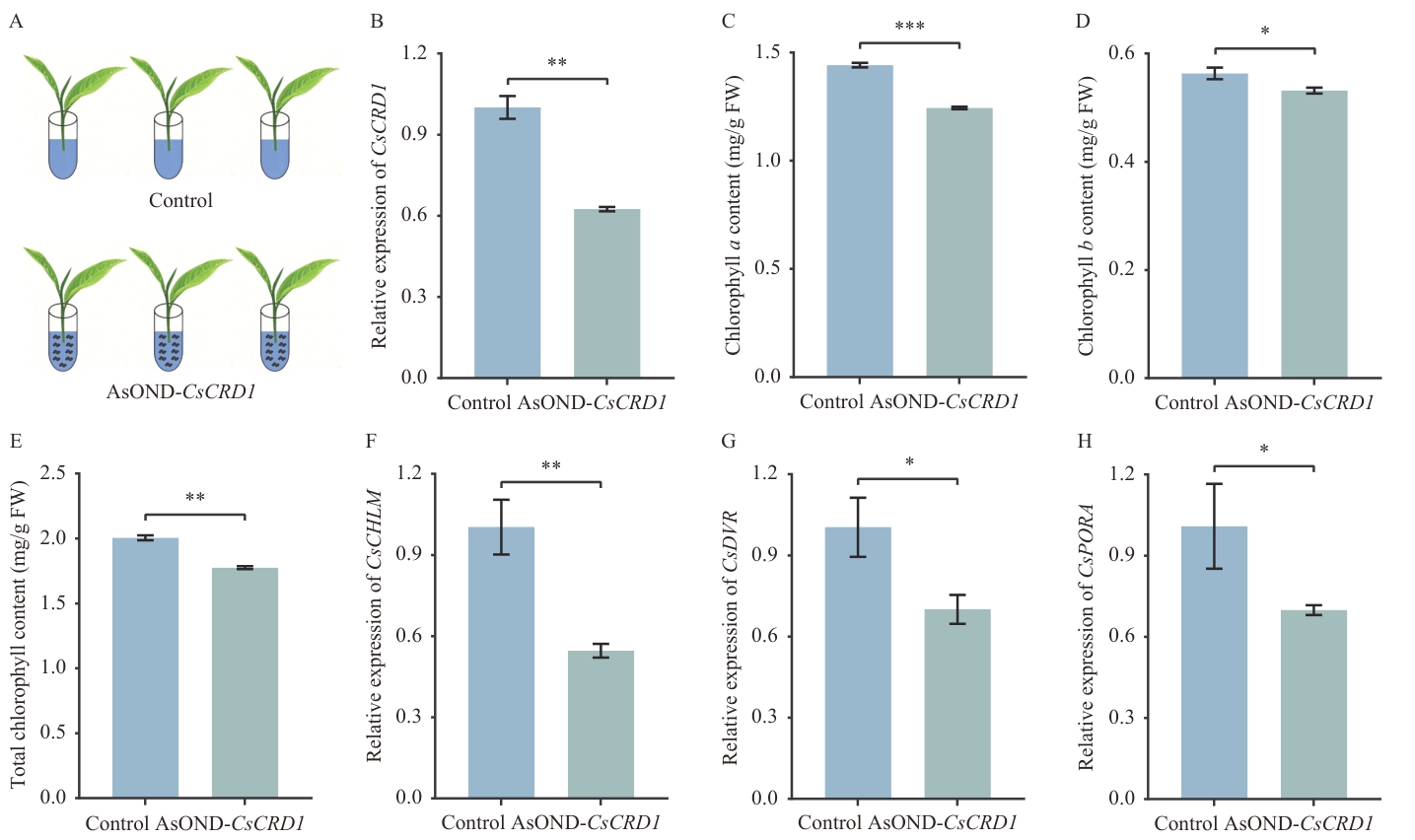
图8 茶树CsCRD1基因沉默对叶绿素合成响应的分析A:CsCRD1基因AsODN沉默示意图;B:AsODNs-CsCRD1沉默后CsCRD1基因的相对表达水平;C‒E:CsCRD1基因沉默后叶绿素含量的变化;F‒H:CsCRD1基因沉默后叶绿素生物合成途径相关基因的相对表达水平。*、**和***分别表示P<0.05、P<0.01和P<0.001的显著性水平(单因素方差分析结合Tukey事后检验)
Fig. 8 Analysis of the impact of CsCRD1 gene silencing on chlorophyll biosynthesis in tea plantsA: Schematic diagram of CsCRD1 gene AsODN silencing. B: Relative expression of the CsCRD1 gene after AsODNs-CsCRD1 silencing. C‒E: Changes in chlorophyll content after CsCRD1 gene silencing. F‒H: Relative expressions of chlorophyll biosynthesis-related genes after CsCRD1 gene silencing. *, ** and *** indicate the significances of P<0.05, P<0.01, and P<0.001, respectively (one-way ANOVA with Tukey’s post hoc test)
| [1] | 王开荣. 白化茶种质资源综合性状研究 [D]. 杭州: 浙江大学, 2006. |
| Wang KR. The research on the comprehensive characters of albino tea as germplasm resources [D]. Hangzhou: Zhejiang University, 2006. | |
| [2] | Cai WH, Zheng XQ, Liang YR. High-light-induced degradation of photosystem Ⅱ subunits’ involvement in the albino phenotype in tea plants [J]. Int J Mol Sci, 2022, 23(15): 8522. |
| [3] | Xiong JY, Wen GP, Song J, et al. Knockout of the chlorophyll a oxygenase gene OsCAO1 reduces chilling tolerance in rice seedlings [J]. Genes, 2024, 15(6): 721. |
| [4] | Ye JJ, Lin XY, Yang ZX, et al. The light-harvesting chlorophyll a/b-binding proteins of photosystem Ⅱ family members are responsible for temperature sensitivity and leaf color phenotype in albino tea plant [J]. J Adv Res, 2024, 66: 87-104. |
| [5] | Li CF, Ma JQ, Huang DJ, et al. Comprehensive dissection of metabolic changes in albino and green tea cultivars [J]. J Agric Food Chem, 2018, 66(8): 2040-2048. |
| [6] | Zhang CY, Wang MH, Gao XZ, et al. Multi-omics research in albino tea plants: Past, present, and future [J]. Sci Hortic, 2020, 261: 108943. |
| [7] | Kumari M, Clarke HJ, Small I, et al. Albinism in plants: a major bottleneck in wide hybridization, androgenesis and doubled haploid culture [J]. Crit Rev Plant Sci, 2009, 28(6): 393-409. |
| [8] | 史琳, 陈彦. 植物白化基因的作用机制与研究进展 [J]. 安徽农业科学, 2017, 45(12): 132-135. |
| Shi L, Chen Y. Function mechanism and research progress of plant albino gene [J]. J Anhui Agric Sci, 2017, 45(12): 132-135. | |
| [9] | Fan YG, Zhao XX, Wang HY, et al. Effects of light intensity on metabolism of light-harvesting pigment and photosynthetic system in Camellia sinensis L. cultivar ‘Huangjinya’ [J]. Environ Exp Bot, 2019, 166: 103796. |
| [10] | Tian YY, Wang HY, Zhang ZQ, et al. An rna-seq analysis reveals differential transcriptional responses to different light qualities in leaf color of Camellia sinensis cv. huangjinya [J]. J Plant Growth Regul, 2022, 41(2): 612-627. |
| [11] | Ma CL, Chen L, Wang XC, et al. Differential expression analysis of different albescent stages of ‘Anji Baicha’ (Camellia sinensis (L.) O. Kuntze) using cDNA microarray [J]. Sci Hortic, 2012, 148: 246-254. |
| [12] | Chen XF, Li JJ, Yu Y, et al. STAY-GREEN and light-harvesting complex Ⅱ chlorophyll a/b binding protein are involved in albinism of a novel albino tea germplasm ‘Huabai 1’ [J]. Sci Hortic, 2022, 293: 110653. |
| [13] | Wu ZJ, Liu KY, Zhang X, et al. CsNYC1a mediates chlorophyll degradation and albino trait formation in the Arbor-type tea plant Camellia nanchuanica [J]. J Agric Food Chem, 2024, 72(24): 14087-14098. |
| [14] | Liu WM, Liu SY, Zhang KY, et al. Chlorophyllase is transcriptionally regulated by CsMYB308/CsDOF3 in young leaves of tea plant [J]. Hortic Plant J, 2023, 9(6): 1162-1176. |
| [15] | Liu XY, Cheng X, Cao JJ, et al. GOLDEN 2-LIKE transcription factors regulate chlorophyll biosynthesis and flavonoid accumulation in response to UV-B in tea plants [J]. Hortic Plant J, 2023, 9(5): 1055-1066. |
| [16] | Zhang CY, Liu HR, Wang JY, et al. A key mutation in magnesium chelatase I subunit leads to a chlorophyll-deficient mutant of tea (Camellia sinensis) [J]. J Exp Bot, 2024, 75(3): 935-946. |
| [17] | Zhang XN, Tong JW, Wen BB, et al. The phytochrome-interacting factor CsPIF3a positively regulates chlorophyll biosynthesis in tea plants (Camellia sinensis) [J]. Plant Physiol Biochem, 2025, 229: 110419. |
| [18] | Liu XY, Cheng X, Cao JJ, et al. UV-B regulates seasonal greening of albino leaves by modulating CsHY5-inhibiting chlorophyll biosynthesis in Camellia sinensis cv. Huangkui [J]. Plant Sci, 2023, 328: 111569. |
| [19] | Stenbaek A, Jensen PE. Redox regulation of chlorophyll biosynthesis [J]. Phytochemistry, 2010, 71(8/9): 853-859. |
| [20] | Xue YJ, Dong HX, Huang HR, et al. Mutation in Mg-protoporphyrin Ⅸ monomethyl ester (oxidative) cyclase gene ZmCRD1 causes chlorophyll-deficiency in maize [J]. Front Plant Sci, 2022, 13: 912215. |
| [21] | Kong WY, Yu XW, Chen HY, et al. The catalytic subunit of magnesium-protoporphyrin Ⅸ monomethyl ester cyclase forms a chloroplast complex to regulate chlorophyll biosynthesis in rice [J]. Plant Mol Biol, 2016, 92(1/2): 177-191. |
| [22] | Wang XX, Huang RF, Quan RD. Mutation in Mg-protoporphyrin Ⅸ monomethyl ester cyclase decreases photosynthesis capacity in rice [J]. PLoS One, 2017, 12(1): e0171118. |
| [23] | Li CY, Hu SY, Yang WT, et al. Conversion obstacle from Mg-protoporphyrin Ⅸ to protochlorophyllide might be responsible for chlorophyll-deficient phenotype of the Huangjinya’s albino offspring [J]. Plant Physiol Biochem, 2024, 212: 108778. |
| [24] | Wu ZJ, Tian C, Jiang Q, et al. Selection of suitable reference genes for qRT-PCR normalization during leaf development and hormonal stimuli in tea plant (Camellia sinensis) [J]. Sci Rep, 2016, 6: 19748. |
| [25] | Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR [J]. Nucleic Acids Res, 2001, 29(9): e45. |
| [26] | Chazaux M, Schiphorst C, Lazzari G, et al. Precise estimation of chlorophyll a, b and carotenoid content by deconvolution of the absorption spectrum and new simultaneous equations for Chl determination [J]. Plant J, 2022, 109(6): 1630-1648. |
| [27] | Tang QH, Liu KY, Yue C, et al. CsXDH1 gene promotes caffeine catabolism induced by continuous strong light in tea plant [J]. Hortic Res, 2023, 10(6): uhad090. |
| [28] | Yamatani H, Ito T, Nishimura K, et al. Genetic analysis of chlorophyll synthesis and degradation regulated by BALANCE of CHLOROPHYLL METABOLISM [J]. Plant Physiol, 2022, 189(1): 419-432. |
| [29] | Eckhardt U, Grimm B, Hörtensteiner S. Recent advances in chlorophyll biosynthesis and breakdown in higher plants [J]. Plant Mol Biol, 2004, 56(1): 1-14. |
| [30] | Tripathy BC, Pattanayak GK. Chlorophyll biosynthesis in higher plants [M]//Photosynthesis. Dordrecht: Springer Netherlands, 2011: 63-94. |
| [31] | Gadjieva R, Axelsson E, Olsson U, et al. Analysis of Gun phenotype in barley magnesium chelatase and Mg-protoporphyrin Ⅸ monomethyl ester cyclase mutants [J]. Plant Physiol Biochem, 2005, 43(10/11): 901-908. |
| [32] | Yamanashi K, Minamizaki K, Fujita Y. Identification of the chlE gene encoding oxygen-independent Mg-protoporphyrin Ⅸ monomethyl ester cyclase in cyanobacteria [J]. Biochem Biophys Res Commun, 2015, 463(4): 1328-1333. |
| [33] | Luo J, Wang H, Chen SJ, et al. CmNAC73 mediates the formation of green color in Chrysanthemum flowers by directly activating the expression of chlorophyll biosynthesis genes HEMA1 and CRD1 [J]. Genes, 2021, 12(5): 704. |
| [34] | Tottey S, Block MA, Allen M, et al. Arabidopsis CHL27, located in both envelope and thylakoid membranes, is required for the synthesis of protochlorophyllide [J]. Proc Natl Acad Sci USA, 2003, 100(26): 16119-16124. |
| [35] | Li C, Ma FR, Jiao RJ, et al. Mutation in Mg-protoporphyrin Ⅸ monomethyl ester cyclase causes yellow and spotted leaf phenotype in rice [J]. Plant Mol Biol Report, 2019, 37(4): 253-264. |
| [1] | 苏燕竹, 李达, 张爱爱, 刘永光, 张秀荣, 薛其勤. 大豆CAD基因家族的鉴定及表达分析[J]. 生物技术通报, 2026, 42(4): 101-113. |
| [2] | 刘青媛, 吴洪启, 陈秀娥, 陈剑, 姜远泽, 何燕子, 喻奇伟, 刘仁祥. 转录因子NtMYB96a调控烟草耐旱性的功能研究[J]. 生物技术通报, 2026, 42(4): 239-250. |
| [3] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [4] | 曾厅, 张兰, 罗睿. 转录因子MpR2R3-MYB17调控地钱胞芽发育的功能研究[J]. 生物技术通报, 2026, 42(1): 208-217. |
| [5] | 陈静欢, 房国楠, 朱文豪, 叶广继, 苏旺, 贺苗苗, 杨生龙, 周云. 马铃薯种质资源淀粉表征及相关基因表达分析[J]. 生物技术通报, 2026, 42(1): 170-183. |
| [6] | 倪莹, 李雷, 汪进萱, 马波, 孟昕, 冷平生, 吴静, 胡增辉. 紫丁香So4CL的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 139-149. |
| [7] | 任云儿, 伍国强, 成斌, 魏明. 甜菜BvATGs基因家族全基因组鉴定及盐胁迫下表达模式分析[J]. 生物技术通报, 2026, 42(1): 184-197. |
| [8] | 杨跃琴, 邢英, 仲子荷, 田维军, 杨雪清, 王建旭. 甲基汞胁迫下水稻OsMATE34的表达及功能分析[J]. 生物技术通报, 2026, 42(1): 86-94. |
| [9] | 杨娟, 冯慧, 吉乃喆, 孙丽萍, 王赟, 张佳楠, 赵世伟. 月季AP2/ERF转录因子RcERF4和RcRAP2-12的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 150-160. |
| [10] | 董向向, 缪百灵, 许贺娟, 陈娟娟, 李亮杰, 龚守富, 朱庆松. 森林草莓FveBBX20基因的生物信息学分析及开花调控功能[J]. 生物技术通报, 2025, 41(9): 115-123. |
| [11] | 张永艳, 郭思健, 李晶, 郝思怡, 李瑞得, 刘嘉鹏, 程春振. 蓝莓花青素相关VcGSTF19基因的克隆及功能研究[J]. 生物技术通报, 2025, 41(9): 139-146. |
| [12] | 李珊, 马登辉, 马红义, 姚文孔, 尹晓. 葡萄SKP1基因家族鉴定与表达分析[J]. 生物技术通报, 2025, 41(9): 147-158. |
| [13] | 巩慧玲, 邢玉洁, 马俊贤, 蔡霞, 冯再平. 马铃薯LAC基因家族的鉴定及盐胁迫下表达分析[J]. 生物技术通报, 2025, 41(9): 82-93. |
| [14] | 徐小萍, 杨成龙, 和兴, 郭文杰, 吴健, 方少忠. 百合LoAPS1克隆及其在休眠解除过程的功能分析[J]. 生物技术通报, 2025, 41(9): 195-206. |
| [15] | 赖诗雨, 梁巧兰, 魏列新, 牛二波, 陈应娥, 周鑫, 杨思正, 王博. NbJAZ3在苜蓿花叶病毒侵染本氏烟过程中的作用[J]. 生物技术通报, 2025, 41(8): 186-196. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||