








• 研究报告 • 下一篇
收稿日期:2025-10-08
出版日期:2026-03-09
通讯作者:
申琼,女,博士,副研究员,研究方向 :南瓜属分子遗传育种;E-mail: shenqiong08@163.com作者简介:孟洪玉,男,硕士研究生,研究方向 :园艺植物生物技术与遗传改良;E-mail: 13233572666@163.com
基金资助:
MENG Hong-yu( ), LIU Ya-nan, WU Jun-xin, SHEN Qiong(
), LIU Ya-nan, WU Jun-xin, SHEN Qiong( )
)
Received:2025-10-08
Published:2026-03-09
摘要:
目的 探究南瓜(Cucurbita moschata Duch.)NAC基因结构特征和表达模式,为阐明CmoNAC基因的生物学功能及进一步培育抗逆性强的砧用南瓜品种及裸仁南瓜品种提供参考。 方法 基于南瓜全基因组信息,使用生物信息学方法对NAC基因家族进行鉴定,系统分析该家族成员的理化性质、亚细胞定位、染色体定位、基因结构及保守基序等信息。结合转录组数据及实时荧光定量PCR(RT-qPCR)方法,分析CmoNAC的组织表达特征及对逆境胁迫应答和种皮发育的影响。 结果 CmoNAC基因家族有133个成员,其编码蛋白主要定位于细胞核,均含有典型的NAM结构域。这些成员不均匀分布于20条染色体上,4号染色体分布最多。系统进化分析将其划分为15个亚家族,其中NAM亚家族成员数量最多。保守基序和基因结构分析显示同一亚族内成员具有相似Motif组成和外显子‒内含子结构,Motif 2/3/4广泛存在,部分Motif具有亚家族特异性。顺式作用元件分析表明CmoNAC基因启动子区域包含光、激素、逆境响应及发育相关等多种调控元件。组织表达分析表明,CmoNAC家族成员在根中表达量最高。100个CmoNAC基因在盐胁迫下显著上调,64个CmoNAC基因在有种皮和裸仁南瓜种皮差异表达。RT-qPCR分析表明,15个CmoNAC基因在盐胁迫和不同类型种皮发育下的表达模式与转录组数据基本一致。 结论 在南瓜全基因组中共鉴定出133个NAC基因,可分为15个亚家族,其基因结构与保守基序在亚家族内高度保守。CmoNAC108、CmoNAC102和CmoNAC013分别与根部发育、茎部发育与逆境响应密切相关,CmoNAC026、CmoNAC062和CmoNAC121在裸仁种子中特异性上调,可能参与裸仁种皮的形成。
孟洪玉, 刘亚楠, 武峻新, 申琼. 南瓜NAC转录因子家族鉴定及生物信息学分析[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1069.
MENG Hong-yu, LIU Ya-nan, WU Jun-xin, SHEN Qiong. Identification and Analysis of the NAC Transcription Factors Family in Cucurbita moschata Duch.[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1069.

图3 南瓜NAC基因家族的结构分布模式和结构域预测A:蛋白保守基序(Motif)分析;B:基因结构分析;C:基因保守结构域分析
Fig. 3 Structure distribution patterns and domains prediction of the NAC gene family in C. moschata.A: Protein conserved motif analysis. B: Gene structure analysis. C: Gene conserved domain analysis
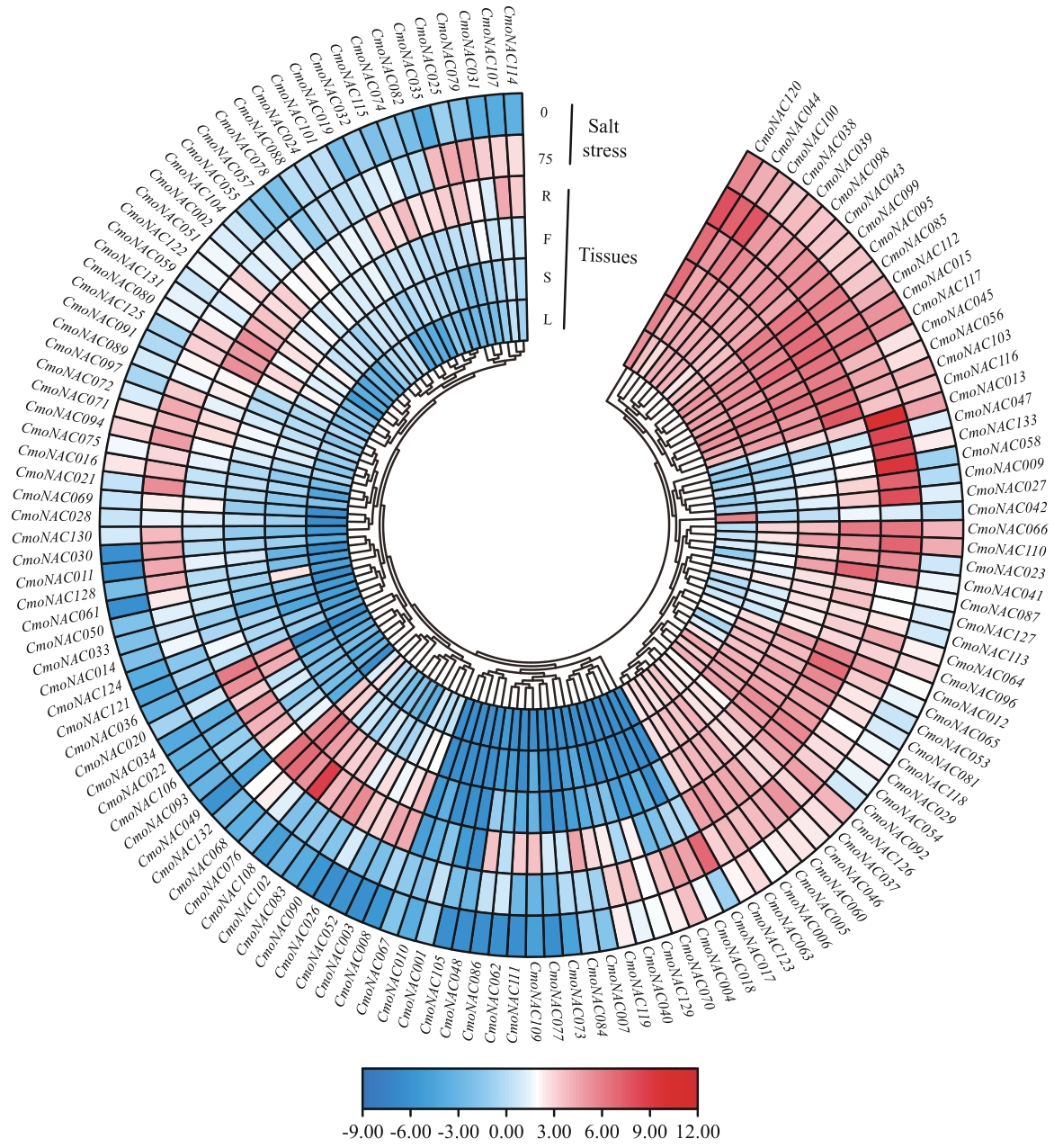
图6 南瓜NAC基因家族在各组织中的表达模式及响应盐胁迫的表达分析L、S、F、R分别代表叶、茎、果、根
Fig. 6 Expression patterns in various tissues of the NAC gene family in C. moschata and its expression analysis in response to salt stressL, S, F, and R indicates leaves, stems, fruits and roots, respectively. 0: 0 mmol/L NaCl; 75: 75 mmol/L NaCl
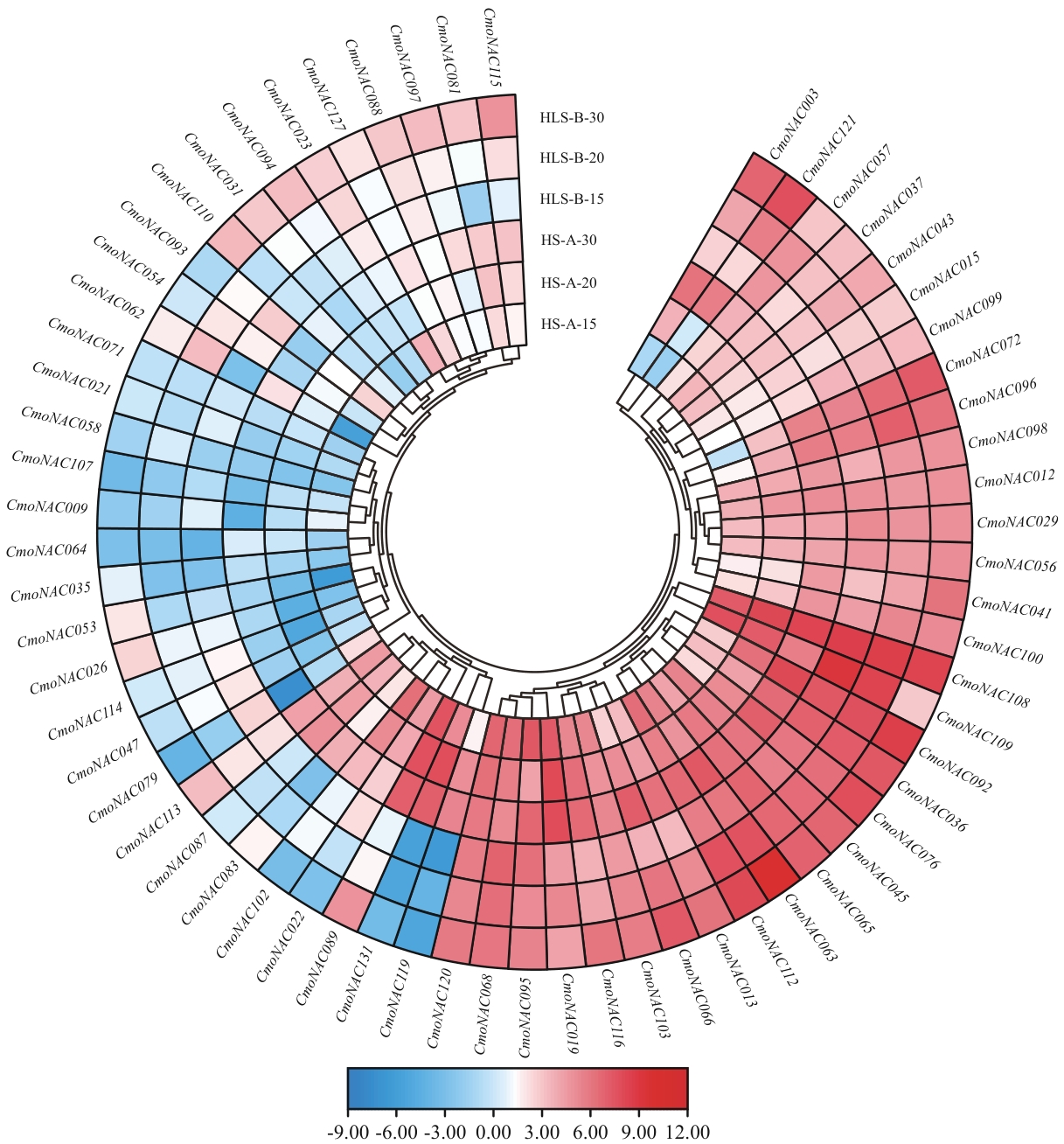
图7 NAC基因家族不同时期有种皮和裸仁南瓜种皮的表达分析HS-A-15、HS-A-20、HS-A-30:基因在有种皮南瓜HS-A的15、20和30 DPA种皮中的表达量;HLS-B-15、HLS-B-20、HLS-B-30:基因在裸仁南瓜HLS-B的15、20和30 DPA种皮中的表达量。下同
Fig. 7 Expression analysis of NAC gene family in seed coat of hull and hull-less seeded C. moschata at different developmental stagesHS-A-15, HS-A-20, HS-A-30: The expressions of the gene in the seed coat of seeded pumpkin HS-A at 15, 20 and 30 DPA. HLS-B-15, HLS-B-20, HLS-B-30: The expressions of the gene in the seed coat of hull-less pumpkin HLS-B at 15, 20 and 30 DPA. The same below
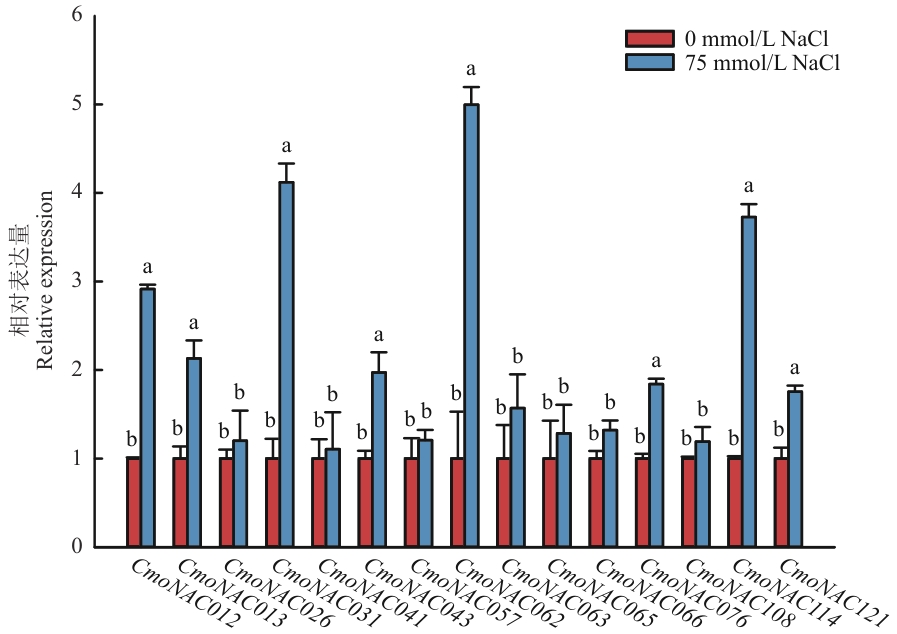
图8 15个CmoNACs在盐胁迫中的相对表达量不同小写字母表示在P<0.05水平差异显著,下同
Fig. 8 Relative expressions of 15 genes under salt stress of C. moschataDifferent lower letters indicate significant difference at P<0.05 level. The same below
| [1] | Kulczyński B, Gramza-Michałowska A. The profile of secondary metabolites and other bioactive compounds in Cucurbita pepo L. and Cucurbita moschata pumpkin cultivars [J]. Molecules, 2019, 24(16): 2945. |
| [2] | Young K, Woo K, Chang PS. Purification and characterization of a novel acid-tolerant and heterodimeric β-glucosidase from pumpkin (Cucurbita moschata) seed [J]. J Biosci Bioeng, 2021, 132(2): 125-131. |
| [3] | Medjakovic S, Hobiger S, Ardjomand-Woelkart K, et al. Pumpkin seed extract: Cell growth inhibition of hyperplastic and cancer cells, independent of steroid hormone receptors [J]. Fitoterapia, 2016, 110: 150-156. |
| [4] | Bergantin C, Maietti A, Tedeschi P, et al. HPLC-UV/Vis-APCI-MS/MS determination of major carotenoids and their bioaccessibility from “delica” (Cucurbita maxima) and “violina” (Cucurbita moschata) pumpkins as food traceability markers [J]. Molecules, 2018, 23(11): 2791. |
| [5] | Vinayashree S, Vasu P. Biochemical, nutritional and functional properties of protein isolate and fractions from pumpkin (Cucurbita moschata var. Kashi Harit) seeds [J]. Food Chem, 2021, 340: 128177. |
| [6] | Shen Q, Wu LY, Zhao XH, et al. Comparative analysis of electron microscopic structure of seed testa, lignin and biosynthesis-related enzyme activities in hulled and hull-less seeds of Cucurbita moschata [J]. Sci Hortic, 2019, 245: 137-143. |
| [7] | Singh S, Grover A, Nasim M. Biofuel potential of plants transformed genetically with NAC family genes [J]. Front Plant Sci, 2016, 7: 22. |
| [8] | Maugarny-Calès A, Gonçalves B, Jouannic S, et al. Apparition of the NAC transcription factors predates the emergence of land plants [J]. Mol Plant, 2016, 9(9): 1345-1348. |
| [9] | Shen Q, Weng YQ. Alternative splicing of NAC transcription factor gene CmNST1 is associated with naked seed mutation in pumpkin, Cucurbita moschata [J]. Genes, 2023, 14(5): 962. |
| [10] | Lyu XL, Shi L, Zhao M, et al. A natural mutation of the NST1 gene arrests secondary cell wall biosynthesis in the seed coat of a hull-less pumpkin accession [J]. Hortic Res, 2022, 9: uhac136. |
| [11] | Wang JL, Zhang XX, Yang HQ, et al. Eggplant NAC domain transcription factor SmNST1 as an activator promotes secondary cell wall thickening [J]. Plant Cell Environ, 2024, 47(11): 4293-4304. |
| [12] | Wang JF, Wang YP, Zhang J, et al. The NAC transcription factor ClNAC68 positively regulates sugar content and seed development in watermelon by repressing ClINV and ClGH3.6 [J]. Hortic Res, 2021, 8(1): 214. |
| [13] | Wang JF, Zhu Y, Li MY, et al. The NAC transcription factor ClNAC100 positively regulates plant height and fruit size in watermelon [J]. Plant J, 2025, 123(1): e70292. |
| [14] | Thirumalaikumar VP, Devkar V, Mehterov N, et al. NAC transcription factor JUNGBRUNNEN1 enhances drought tolerance in tomato [J]. Plant Biotechnol J, 2018, 16(2): 354-366. |
| [15] | Wang T, Ma XM, Chen Y, et al. SlNAC3 suppresses cold tolerance in tomatoes by enhancing ethylene biosynthesis [J]. Plant Cell Environ, 2024, 47(8): 3132-3146. |
| [16] | Zhang HF, Pei YP, Zhu FL, et al. CaSnRK2.4-mediated phosphorylation of CaNAC035 regulates abscisic acid synthesis in pepper (Capsicum annuum L.) responding to cold stress [J]. Plant J, 2024, 117(5): 1377-1391. |
| [17] | Chen SQ, Zhang WX, Zhang Q, et al. SlNAC12, a novel NAC-type transcription factor, confers salt stress tolerance in tomato [J]. Plant Cell Rep, 2024, 44(1): 5. |
| [18] | Duan AQ, Tao JP, Jia LL, et al. AgNAC1, a celery transcription factor, related to regulation on lignin biosynthesis and salt tolerance [J]. Genomics, 2020, 112(6): 5254-5264. |
| [19] | Peng YQ, Cao HS, Cui LJ, et al. CmoNAC1 in pumpkin rootstocks improves salt tolerance of grafted cucumbers by binding to the promoters of CmoRBOHD1, CmoNCED6, CmoAKT1;2 and CmoHKT1;1 to regulate H2O2, ABA signaling and K+/Na+ homeostasis [J]. Hortic Res, 2023, 10(9): uhad157. |
| [20] | Gao Y, Fan ZQ, Zhang Q, et al. A tomato NAC transcription factor, SlNAM1, positively regulates ethylene biosynthesis and the onset of tomato fruit ripening [J]. Plant J, 2021, 108(5): 1317-1331. |
| [21] | Wang JF, Tian SW, Yu YT, et al. Natural variation in the NAC transcription factor NONRIPENING contributes to melon fruit ripening [J]. J Integr Plant Biol, 2022, 64(7): 1448-1461. |
| [22] | Zhang XM, Yu HJ, Sun C, et al. Genome-wide characterization and expression profiling of the NAC genes under abiotic stresses in Cucumis sativus [J]. Plant Physiol Biochem, 2017, 113: 98-109. |
| [23] | Li WQ, Ping F, Jiang HX, et al. Genome-wide identification of the NAC gene family in Brassica rapa (L.) and expression pattern analysis of BrNAC2s [J]. Plants, 2025, 14(6): 834. |
| [24] | Diao WP, Snyder JC, Wang SB, et al. Genome-wide analyses of the NAC transcription factor gene family in pepper (Capsicum annuum L.): chromosome location, phylogeny, structure, expression patterns, cis-elements in the promoter, and interaction network [J]. Int J Mol Sci, 2018, 19(4): 1028. |
| [25] | Sun HH, Wu S, Zhang GY, et al. Karyotype stability and unbiased fractionation in the paleo-allotetraploid Cucurbita genomes [J]. Mol Plant, 2017, 10(10): 1293-1306. |
| [26] | Ooka H, Satoh K, Doi K, et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana [J]. DNA Res, 2003, 10(6): 239-247. |
| [27] | Wang HZ, Dixon RA. On-off switches for secondary cell wall biosynthesis [J]. Mol Plant, 2012, 5(2): 297-303. |
| [28] | Han KJ, Zhao Y, Sun YH, et al. NACs, generalist in plant life [J]. Plant Biotechnol J, 2023, 21(12): 2433-2457. |
| [29] | 黄畋柳, 张锐, 贺迎骁, 等. 辣椒NAC家族成员鉴定及其编码基因在NaCl胁迫下的表达分析 [J]. 植物资源与环境学报, 2023, 32(4): 12-24. |
| Huang TL, Zhang R, He YX, et al. Identification of NAC family members of Capsicum annuum and analysis on expressions of their coding genes under NaCl stress [J]. J Plant Resour Environ, 2023, 32(4): 12-24. | |
| [30] | 王会文, 范军强, 路晓明, 等. 白菜型冬油菜NAC基因家族鉴定及表达分析 [J]. 2022(5): 1315-1329. |
| Wang HW, Fan JQ, Lu XM, et al. Identification and expression analysis of NAC gene family in Brassica rapa L [J]. Jiangsu J. Agric. Sci, 2022(5): 1315-1329. | |
| [31] | 荣欢, 任师杰, 汪梓坪, 等. 植物NAC转录因子的结构及功能研究进展 [J]. 江苏农业科学, 2020(18): 44-53. |
| Rong H, Ren SJ, Wang ZP, et al. Research progress on the structure and function of plant NAC transcription factors [J]. Jiangsu Agric. Sci., 2020(18): 44-53. | |
| [32] | He XJ, Mu RL, Cao WH, et al. AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development [J]. Plant J, 2005, 44(6): 903-916. |
| [33] | Mitsuda N, Iwase A, Yamamoto H, et al. NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis [J]. Plant Cell, 2007, 19(1): 270-280. |
| [34] | Wu YR, Deng ZY, Lai JB, et al. Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses [J]. Cell Res, 2009, 19(11): 1279-1290. |
| [1] | 龙林茜, 曾银萍, 王茜, 邓玉萍, 葛敏茜, 陈彦灼, 李鑫娟, 杨军, 邹建. 向日葵GH3基因家族鉴定及其在花发育中的功能分析[J]. 生物技术通报, 2026, 42(1): 125-138. |
| [2] | 李玉珍, 李梦丹, 张蔚, 彭婷. 基于月季扩展蛋白基因家族鉴定的野蔷薇RmEXPB2基因功能研究[J]. 生物技术通报, 2025, 41(9): 182-194. |
| [3] | 龚钰涵, 陈兰, 尚方慧子, 郝灵颖, 刘硕谦. 茶树TRB基因家族鉴定及表达模式分析[J]. 生物技术通报, 2025, 41(7): 214-225. |
| [4] | 秦文俊, 熊言杰, 赵冉, 马萧然, 叶霄萌, 宋江华. 甘蓝ARF基因家族的鉴定及其在非生物胁迫下的表达分析[J]. 生物技术通报, 2025, 41(10): 253-263. |
| [5] | 李彩霞, 李艺, 穆宏秀, 林俊轩, 白龙强, 孙美华, 苗妍秀. 中国南瓜bHLH转录因子家族的鉴定与生物信息学分析[J]. 生物技术通报, 2025, 41(1): 186-197. |
| [6] | 田栩瑞, 霍信屹, 郭云涵, 向林, 产祝龙, 王艳平. 百合LoSAUR10基因的表达特征及功能分析[J]. 生物技术通报, 2025, 41(1): 221-229. |
| [7] | 申鹏, 高雅彬, 丁红. 马铃薯SAT基因家族的鉴定和表达分析[J]. 生物技术通报, 2024, 40(9): 64-73. |
| [8] | 胡永波, 雷雨田, 杨永森, 陈馨, 林黄昉, 林碧英, 刘爽, 毕格, 申宝营. 黄瓜和南瓜Bcl-2相关抗凋亡家族全基因组鉴定与表达模式分析[J]. 生物技术通报, 2024, 40(6): 219-237. |
| [9] | 沈天虹, 齐孝博, 赵瑞丰, 马欣荣. 微藻盐胁迫响应分子机制研究进展[J]. 生物技术通报, 2024, 40(3): 89-99. |
| [10] | 李明坤, 毕美营, 张天航, 吴翔宇, 杨培儒, 应明. UgRNA/Cas9多基因编辑法恢复根际细菌农用功能的研究[J]. 生物技术通报, 2024, 40(10): 275-287. |
| [11] | 张洁萍, 关跃峰. 基于启动子编辑的作物育种[J]. 生物技术通报, 2024, 40(10): 53-61. |
| [12] | 徐红云, 张恩辉, 于存. 柽柳ThWRKY4转录因子结合ARR1AT元件调控基因表达[J]. 生物技术通报, 2021, 37(3): 18-26. |
| [13] | 张桐, 李智强, 伍国强. WRKY转录因子在植物逆境响应中的作用[J]. 生物技术通报, 2021, 37(10): 203-215. |
| [14] | 王博, 王莲萍, 杨春雨, 国会艳, 魏继承. 白桦E-box元件的载体构建及与BplMYB46转录因子的互作分析[J]. 生物技术通报, 2018, 34(12): 110-115. |
| [15] | 肖晶,查星琴,成文敏,霍金龙,潘伟荣,王淑燕,王配,刘海京,张秀琼,曾养志. 版纳微型猪近交系CATSPER3基因克隆、序列及发育进化分析[J]. 生物技术通报, 2016, 32(12): 103-107. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||