








• 研究报告 • 下一篇
收稿日期:2025-10-22
出版日期:2026-03-09
通讯作者:
高暝,女,研究员,研究方向 :天然香料树种良种选育;E-mail: 4862705@163.com作者简介:梁慧琪,女,硕士研究生,研究方向 :特色林木资源育种与培育;E-mail: liang13398769606@163.com
基金资助:
LIANG Hui-qi( ), CHEN Yi-cun, WANG Yang-dong, ZHAO Yun-xiao, GAO Ming(
), CHEN Yi-cun, WANG Yang-dong, ZHAO Yun-xiao, GAO Ming( )
)
Received:2025-10-22
Published:2026-03-09
摘要:
目的 探索山苍子(Litsea cubeba)PAT1亚家族(phytochrome A signal transduction 1 subfamily)在嫁接愈合中的作用,对其成员进行系统鉴定与表达分析,为后续的功能验证与分子改良提供理论基础。 方法 利用生物信息学方法对山苍子GRAS基因家族进行全基因组鉴定,共鉴定出45个成员,其中PAT1亚家族包含6个基因。通过分析其理化性质、基因结构、保守基序、顺式作用元件和共线性关系,结合转录组和RT-qPCR结果,探讨其在不同组织及嫁接愈合过程中的表达模式。同时利用加权基因共表达网络分析(WGCNA)挖掘核心调控基因及其潜在功能模块。 结果 6个LcPAT1基因不均匀分布于4条染色体上,其启动子富含光响应元件及ABA、MeJA、SA等激素响应元件。基于山苍子不同组织转录组数据的WGCNA结果显示,核心基因LcPAT1-3与346个基因显著共表达,这些基因主要富集于转录调控、激素信号转导及碳水化合物和次生代谢通路,提示其在嫁接愈合过程中的枢纽作用。不同组织的转录组和RT-qPCR结果均显示,6个LcPAT1基因在茎中表达较高。不同嫁接愈合时期的RT-qPCR结果显示,LcGRAS04与LcPAT1-2表达量在形成层连接期均升高,LcPAT1-3和LcPAT1-4在维管分化期持续高表达,呈现明显的时空特异性。 结论 山苍子PAT1亚家族成员在顺式元件和表达模式上存在分化。LcPAT1-3可能作为关键调控因子,协同其他成员在嫁接愈合的不同阶段调控形成层激活、维管再生及防御加固等过程。
梁慧琪, 陈益存, 汪阳东, 赵耘霄, 高暝. 山苍子PAT1基因亚家族鉴定及其在嫁接愈合过程中的表达分析[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1133.
LIANG Hui-qi, CHEN Yi-cun, WANG Yang-dong, ZHAO Yun-xiao, GAO Ming. Identification and Expression Analysis of the PAT1 Gene Subfamily in Litsea cubeba during Graft Union Healing[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1133.
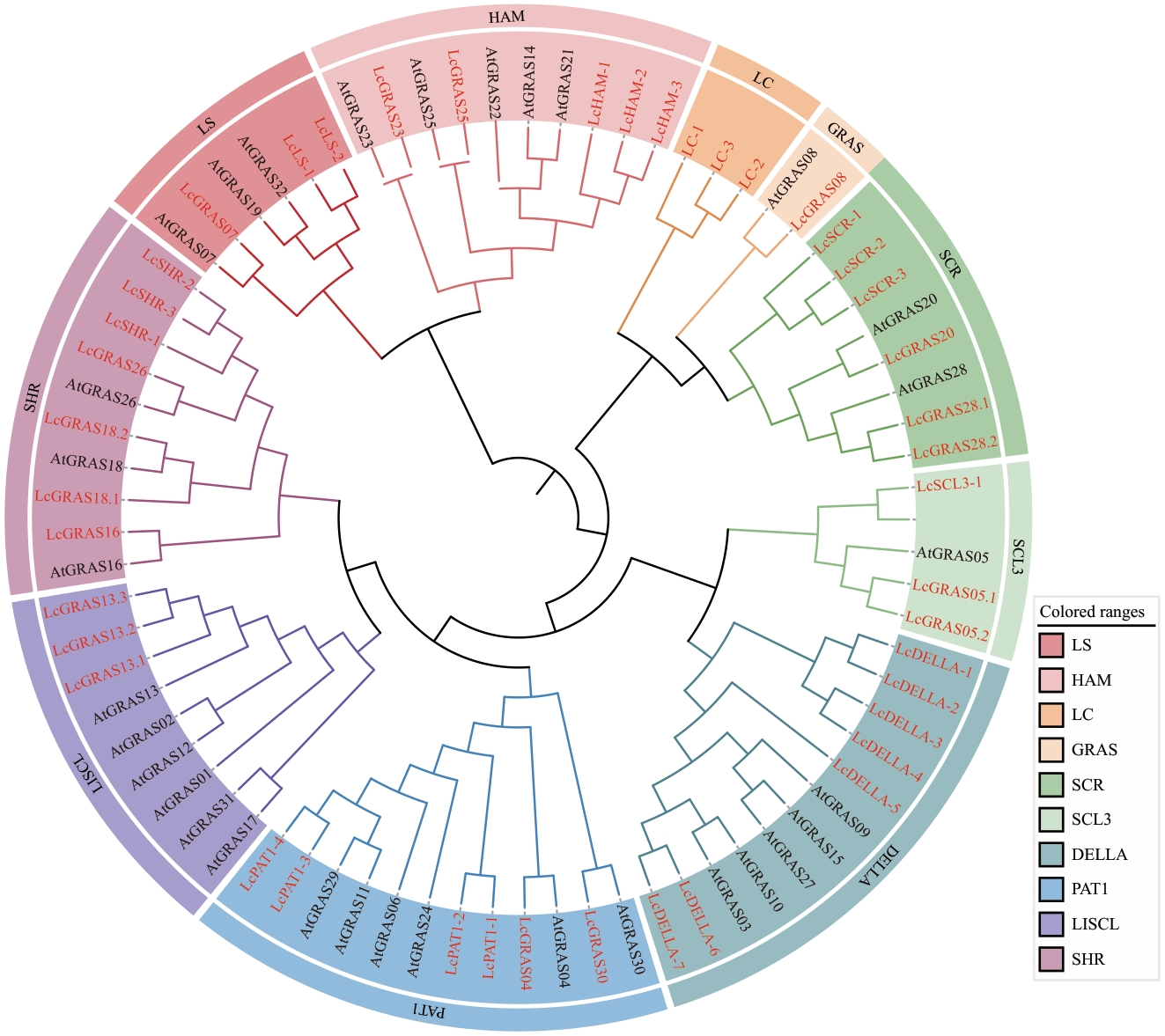
图1 山苍子及拟南芥GRAS基因系统发育进化树红色字体代表山苍子,黑色字体代表拟南芥
Fig. 1 Phylogenetic tree of GRAS genes from L. cubeba and A. thalianaThe red font indicates L. cubeba, the black font indicates A. thaliana
| 基因名称Gene name | 蛋白长度Number of amino acids | 分子量Molecular weight (kD) | 等电点Theoretical pI | 不稳定系数Instability index | 脂肪系数Aliphatic index | 亲水性平均值Grand average of hydropathicity | 亚细胞定位Subcellular localization |
|---|---|---|---|---|---|---|---|
| LcGRAS04 | 582 | 78.07 | 5.05 | 50.50 | 80.43 | -0.24 | Nucleus |
| LcGRAS30 | 591 | 86.07 | 6.41 | 57.11 | 78.83 | -0.32 | Nucleus |
| LcPAT1-1 | 583 | 71.07 | 5.46 | 53.31 | 83.60 | -0.43 | Nucleus |
| LcPAT1-2 | 579 | 110.07 | 5.84 | 56.06 | 79.36 | -0.41 | Nucleus |
| LcPAT1-3 | 589 | 69.07 | 5.49 | 50.44 | 82.77 | -0.27 | Nucleus |
| LcPAT1-4 | 586 | 60.07 | 5.76 | 48.94 | 80.89 | -0.31 | Cytoplasm |
| 平均值 Average | 585 | 79.07 | 5.67 | 52.73 | 80.98 | -0.33 | - |
表1 LcPAT1蛋白理化性质与亚细胞定位
Table 1 Physicochemical properties and subcellular localization of LcPAT1 protein
| 基因名称Gene name | 蛋白长度Number of amino acids | 分子量Molecular weight (kD) | 等电点Theoretical pI | 不稳定系数Instability index | 脂肪系数Aliphatic index | 亲水性平均值Grand average of hydropathicity | 亚细胞定位Subcellular localization |
|---|---|---|---|---|---|---|---|
| LcGRAS04 | 582 | 78.07 | 5.05 | 50.50 | 80.43 | -0.24 | Nucleus |
| LcGRAS30 | 591 | 86.07 | 6.41 | 57.11 | 78.83 | -0.32 | Nucleus |
| LcPAT1-1 | 583 | 71.07 | 5.46 | 53.31 | 83.60 | -0.43 | Nucleus |
| LcPAT1-2 | 579 | 110.07 | 5.84 | 56.06 | 79.36 | -0.41 | Nucleus |
| LcPAT1-3 | 589 | 69.07 | 5.49 | 50.44 | 82.77 | -0.27 | Nucleus |
| LcPAT1-4 | 586 | 60.07 | 5.76 | 48.94 | 80.89 | -0.31 | Cytoplasm |
| 平均值 Average | 585 | 79.07 | 5.67 | 52.73 | 80.98 | -0.33 | - |
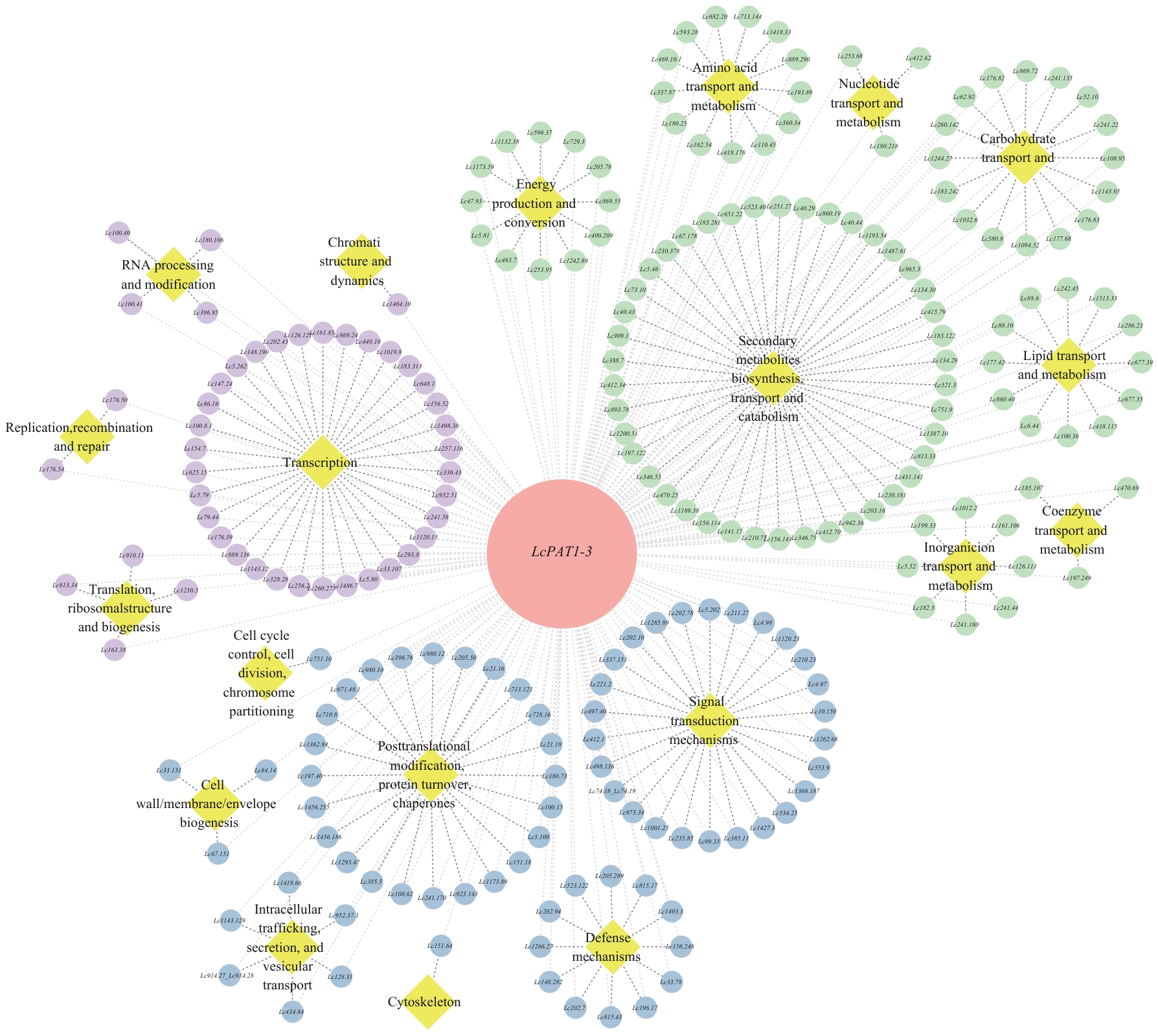
图6 LcPAT1-3共表达网络黄色菱形表示基于COG注释的功能簇,圆形表示基因。基因节点颜色对应其功能大类:紫色代表信息存储与处理,绿色代表代谢,蓝色代表细胞过程与信号转导
Fig. 6 Co-expression network diagram of LcPAT1-3Yellow diamonds indicate functional clusters based on COG annotations, while circles indicate genes. The color of each gene node corresponds to its functional category: Purple indicates information storage and processing, green indicates metabolism, and blue denotes cellular processes and signal transduction
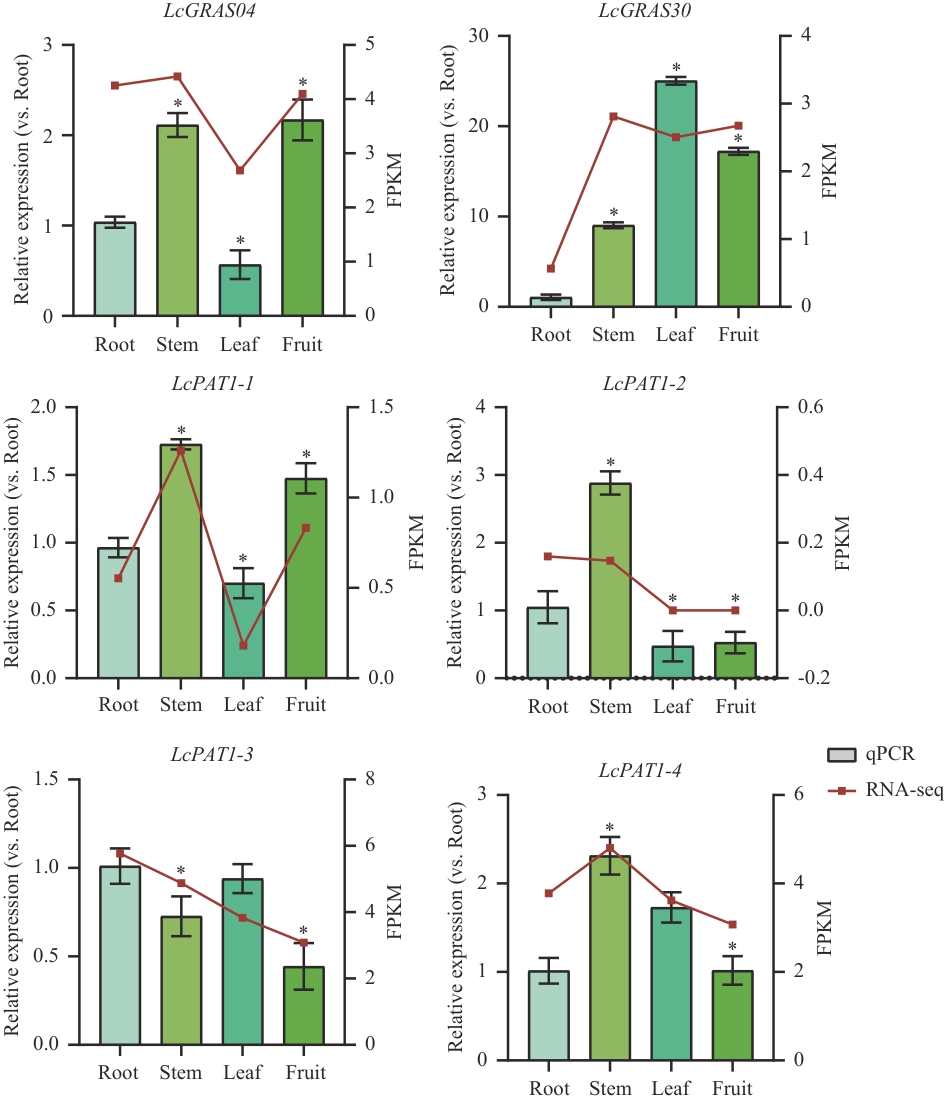
图8 山苍子不同组织中PAT1亚家族基因的相对表达量RT-qPCR结果(柱状图,左Y轴)与转录组log2(FPKM+1)值(折线图,右Y轴)展示于同一图中。样品采集自与转录组分析相同的植株。qPCR数据表示为均值±SD(n=3)。*P<0.05,下同
Fig. 8 Relative expressions of PAT1 subfamily genes in different tissues of L. cubebaThe RT-qPCR results (bar chart, left Y-axis) and transcriptome log₂(FPKM+1) values (line chart, right Y-axis) are presented in the same graph. The samples were collected from the same plants used for the transcriptome analysis. qPCR data are presented as mean ± SD (n=3). *P<0.05. The same below
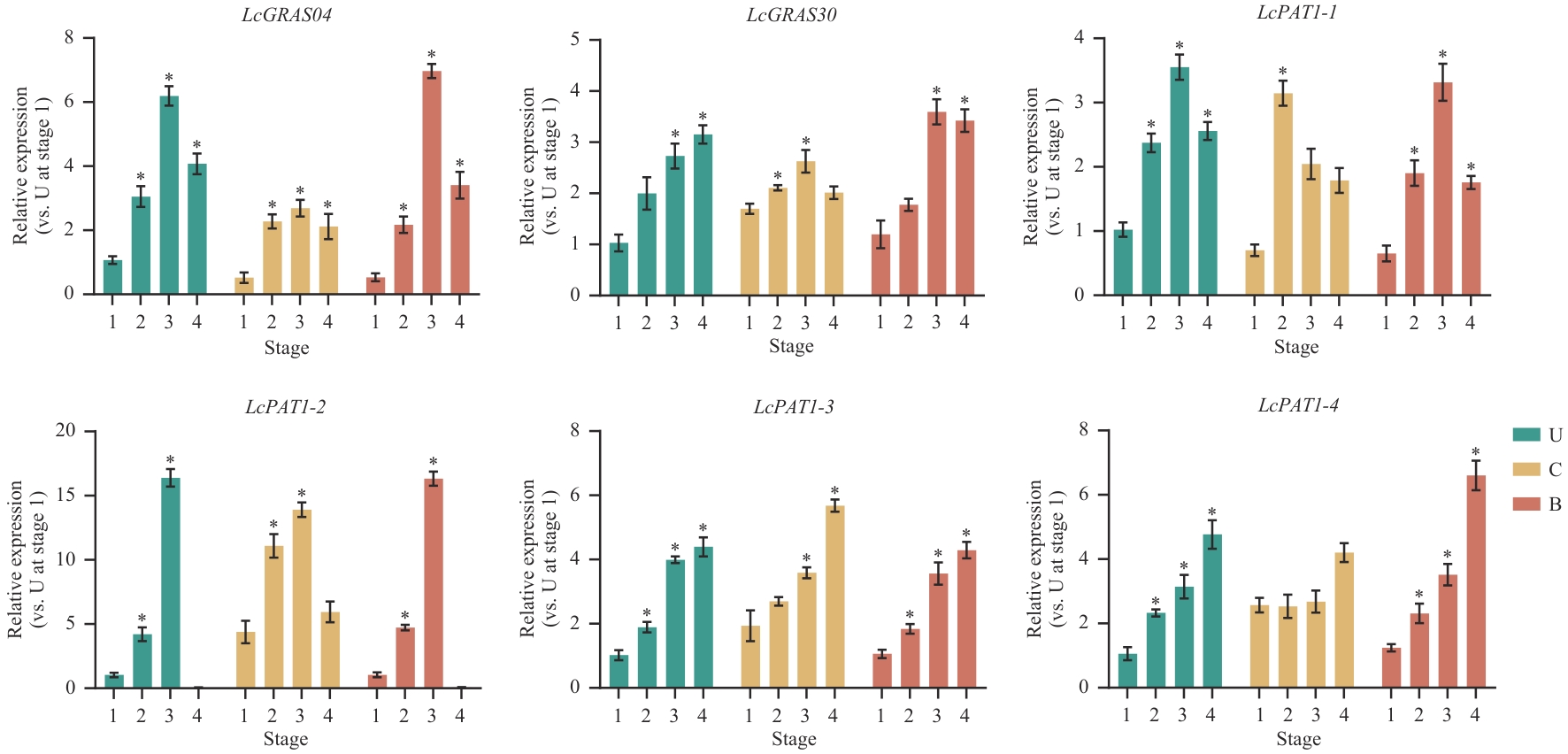
图9 山苍子嫁接愈合过程中PAT1亚家族基因的相对表达量U:结合处上部未嫁接砧木茎段;C:嫁接结合处砧穗茎段;B:结合处下部未嫁接砧木茎段;Stage 1:隔离层形成期;2:愈伤组织形成期;3:形成层连接期;4:维管组织分化期
Fig. 9 Relative expressions of PAT1 subfamily genes during graft union healing in L. cubebaU: Upper region of the junction without grafted rootstock stem segment. C: Stem segment of rootstock and scion at the junction of grafting. B: Basal region of the junction without grafted rootstock stem segment. Stage1: Formation period of the isolation layer. 2: Formation period of callus tissue. 3: Cambium connection. 4: Differentiation period of vascular tissue
| [1] | Tian CG, Wan P, Sun SH, et al. Genome-wide analysis of the GRAS gene family in rice and Arabidopsis [J]. Plant Mol Biol, 2004, 54(4): 519-532. |
| [2] | Pysh LD, Wysocka-Diller JW, Camilleri C, et al. The GRAS gene family in Arabidopsis: sequence characterization and basic expression analysis of the SCARECROW-LIKE genes [J]. Plant J, 1999, 18(1): 111-119. |
| [3] | Sun XL, Jones WT, Rikkerink EHA. GRAS proteins: the versatile roles of intrinsically disordered proteins in plant signalling [J]. Biochem J, 2012, 442(1): 1-12. |
| [4] | Bolle C, Koncz C, Chua NH. PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction [J]. Genes Dev, 2000, 14(10): 1269-1278. |
| [5] | Muntha ST, Zhang LL, Zhou YF, et al. Phytochrome A signal transduction 1 and CONSTANS-LIKE 13 coordinately orchestrate shoot branching and flowering in leafy Brassica juncea [J]. Plant Biotechnol J, 2019, 17(7): 1333-1343. |
| [6] | Ren J, Zhang QW, Xu YF, et al. Decreased expression of a phosphoribosylanthranilate transferase-encoding gene, OsPAT1, causes lesion mimics in rice [J]. Int J Mol Sci, 2025, 26(19): 9428. |
| [7] | Feng M, Zhang A, Nguyen V, et al. A conserved graft formation process in Norway spruce and Arabidopsis identifies the PAT gene family as central regulators of wound healing [J]. Nat Plants, 2024, 10(1): 53-65. |
| [8] | Bisht A, Eekhout T, Canher B, et al. PAT1-type GRAS-domain proteins control regeneration by activating DOF3.4 to drive cell proliferation in Arabidopsis roots [J]. Plant Cell, 2023, 35(5): 1513-1531. |
| [9] | Chen YC, Wang YD, Han XJ, et al. Biology and chemistry of Litsea cubeba, a promising industrial tree in China [J]. J Essent Oil Res, 2013, 25(2): 103-111. |
| [10] | Melnyk CW, Gabel A, Hardcastle TJ, et al. Transcriptome dynamics at Arabidopsis graft junctions reveal an intertissue recognition mechanism that activates vascular regeneration [J]. Proc Natl Acad Sci USA, 2018, 115(10): E2447-E2456. |
| [11] | Chen YC, Li Z, Zhao YX, et al. The Litsea genome and the evolution of the laurel family [J]. Nat Commun, 2020, 11(1): 1675. |
| [12] | Ni FF, Zhao YX, Gao M, et al. LcMYB43 enhances monoterpene biosynthesis by activating 1-deoxy-D-xylulose-5-phosphate synthase gene expression in Litsea cubeba [J]. Int J Biol Macromol, 2025, 304(Pt 1): 140860. |
| [13] | Wang SS, Duan Z, Yan Q, et al. Genome-wide identification of the GRAS family genes in Melilotus albus and expression analysis under various tissues and abiotic stresses [J]. Int J Mol Sci, 2022, 23(13): 7403. |
| [14] | Wang N, Wang KY, Li SK, et al. Transcriptome-wide identification, evolutionary analysis, and GA stress response of the GRAS gene family in Panax ginseng C. A. Meyer [J]. Plants, 2020, 9(2): 190. |
| [15] | Guo YY, Wu HY, Li X, et al. Identification and expression of GRAS family genes in maize (Zea mays L.) [J]. PLoS One, 2017, 12(9): e0185418. |
| [16] | Yang Y, Agassin RH, Ji KS. Transcriptome-wide identification of the GRAS transcription factor family in Pinus massoniana and its role in regulating development and stress response [J]. Int J Mol Sci, 2023, 24(13): 10690. |
| [17] | He ZH, Tian ZZ, Zhang Q, et al. Genome-wide identification, expression and salt stress tolerance analysis of the GRAS transcription factor family in Betula platyphylla [J]. Front Plant Sci, 2022, 13: 1022076. |
| [18] | Hou SJ, Zhang QQ, Chen J, et al. Genome-wide identification and analysis of the GRAS transcription factor gene family in Theobroma cacao [J]. Genes, 2022, 14(1): 57. |
| [19] | 李晨, 赵雪惠, 王庆杰, 等. 桃GRAS家族全基因组鉴定与响应UV-B的表达模式分析 [J]. 中国农业科学, 2019, 52(24): 4567-4581. |
| Li C, Zhao XH, Wang QJ, et al. Genome identification of pp GRAS family and expression pattern analysis of responding to UV-B in peach [J]. Sci Agric Sin, 2019, 52(24): 4567-4581. | |
| [20] | Cenci A, Rouard M. Evolutionary analyses of GRAS transcription factors in angiosperms [J]. Front Plant Sci, 2017, 8: 273. |
| [21] | Cannon SB, Mitra A, Baumgarten A, et al. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana [J]. BMC Plant Biol, 2004, 4: 10. |
| [22] | Serivichyaswat PT, Kareem A, Feng M, et al. Auxin signaling in the cambium promotes tissue adhesion and vascular formation during Arabidopsis graft healing [J]. Plant Physiol, 2024, 196(2): 754-762. |
| [23] | Robert A Creelman MLT. Jasmonic acid/methyl jasmonate accumulate in wounded soybean hypocotyls and modulate wound gene expression [J]. Proc Natl Acad Sci U S A, 1992, 89(11): 4938-4941. |
| [24] | Ruan JJ, Zhou YX, Zhou ML, et al. Jasmonic acid signaling pathway in plants [J]. Int J Mol Sci, 2019, 20(10): 2479. |
| [25] | Li C, Xu MX, Cai X, et al. Jasmonate signaling pathway modulates plant defense, growth, and their trade-offs [J]. Int J Mol Sci, 2022, 23(7): 3945. |
| [26] | Bharath P, Gahir S, Raghavendra AS. Abscisic acid-induced stomatal closure: an important component of plant defense against abiotic and biotic stress [J]. Front Plant Sci, 2021, 12: 615114. |
| [27] | Wang LX, Liao YM, Liu JM, et al. Advances in understanding the graft healing mechanism: a review of factors and regulatory pathways [J]. Hortic Res, 2024, 11(8): uhae175. |
| [28] | Wang QH, Liu N, Yang RR, et al. Essential role of ABA signaling and related transcription factors in phenolic acid and lignin synthesis during muskmelon wound healing [J]. Front Plant Sci, 2024, 15: 1404477. |
| [29] | Jeyasri R, Muthuramalingam P, Karthick K, et al. Methyl jasmonate and salicylic acid as powerful elicitors for enhancing the production of secondary metabolites in medicinal plants: an updated review [J]. Plant Cell Tissue Organ Cult, 2023, 153(3): 447-458. |
| [30] | Feng M, Augstein F, Kareem A, et al. Plant grafting: Molecular mechanisms and applications [J]. Mol Plant, 2024, 17(1): 75-91. |
| [31] | Laura Ragni KN. Mobile gibberellin directly stimulates Arabidopsis hypocotyl xylem expansion [J]. Plant Cell, 2011, 23(4): 1322-1336. |
| [32] | Fan L, Wang RM, García-Caparrós P, et al. Genome-wide identification and functional analysis of the GRAS gene family in Medicago lupulina L [J]. BMC Genomics, 2025, 26(1): 682. |
| [33] | Tang GP, Ni TT, Pedro GC, et al. Identification and characterization of nine PAT1 genes subfamily in Medicago edgeworthii [J]. Plant Signal Behav, 2025, 20(1): 2527380. |
| [1] | 农韦优, 赵昌祖, 钱禛锋, 丁倩, 王誉洁, 陈疏影, 何丽莲, 李富生. 蔗茅EfBBX基因家族鉴定及冷胁迫表达模式分析[J]. 生物技术通报, 2026, 42(2): 1-11. |
| [2] | 杨娟, 冯慧, 吉乃喆, 孙丽萍, 王赟, 张佳楠, 赵世伟. 月季AP2/ERF转录因子RcERF4和RcRAP2-12的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 150-160. |
| [3] | 吴翠翠, 陈登科, 兰刚, 夏芝, 李朋波. 花生转录因子AhHDZ70的生物信息学分析及耐盐耐旱性研究[J]. 生物技术通报, 2026, 42(1): 198-207. |
| [4] | 龙林茜, 曾银萍, 王茜, 邓玉萍, 葛敏茜, 陈彦灼, 李鑫娟, 杨军, 邹建. 向日葵GH3基因家族鉴定及其在花发育中的功能分析[J]. 生物技术通报, 2026, 42(1): 125-138. |
| [5] | 张超超, 韩开元, 王彤, 陈仲. 毛白杨PtoYABBY2和PtoYABBY12的克隆及功能分析[J]. 生物技术通报, 2025, 41(9): 256-264. |
| [6] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [7] | 史发超, 姜永华, 刘海伦, 文英杰, 严倩. 荔枝LcTFL1基因的克隆与功能分析[J]. 生物技术通报, 2025, 41(9): 159-167. |
| [8] | 李玉珍, 李梦丹, 张蔚, 彭婷. 基于月季扩展蛋白基因家族鉴定的野蔷薇RmEXPB2基因功能研究[J]. 生物技术通报, 2025, 41(9): 182-194. |
| [9] | 康琴, 汪霞, 谌明洋, 徐静天, 陈诗兰, 廖平杨, 许文志, 吴卫, 徐东北. 薄荷UV-B受体基因McUVR8的克隆与表达分析[J]. 生物技术通报, 2025, 41(8): 255-266. |
| [10] | 黄诗宇, 田姗姗, 杨天为, 高曼熔, 张尚文. 赤苍藤WRI1基因家族的全基因组鉴定及表达模式分析[J]. 生物技术通报, 2025, 41(8): 242-254. |
| [11] | 朱丽娟, 张锴, 温晓蕾, 褚佳豪, 史凤玉, 王艳丽. 基于WGCNA挖掘野生大豆耐镉关键基因[J]. 生物技术通报, 2025, 41(8): 124-136. |
| [12] | 王芳, 乔帅, 宋伟, 崔鹏娟, 廖安忠, 谭文芳, 杨松涛. 甘薯IbNRT2基因家族全基因组鉴定和表达分析[J]. 生物技术通报, 2025, 41(7): 193-204. |
| [13] | 龚钰涵, 陈兰, 尚方慧子, 郝灵颖, 刘硕谦. 茶树TRB基因家族鉴定及表达模式分析[J]. 生物技术通报, 2025, 41(7): 214-225. |
| [14] | 许慧珍, SHANTWANA Ghimire, RAJU Kharel, 岳云, 司怀军, 唐勋. 马铃薯SUMO E3连接酶基因家族分析及StSIZ1基因的克隆与表达模式分析[J]. 生物技术通报, 2025, 41(6): 119-129. |
| [15] | 王天禧, 杨炳松, 潘荣君, 盖文贤, 梁美霞. 苹果PLATZ基因家族鉴定及MdPLATZ9基因功能研究[J]. 生物技术通报, 2025, 41(4): 176-187. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||