








生物技术通报 ›› 2022, Vol. 38 ›› Issue (1): 15-32.doi: 10.13560/j.cnki.biotech.bull.1985.2021-1562
殷国良1,2( ), 孙文浩1,2, 庞效云1, 孙飞1,2(
), 孙文浩1,2, 庞效云1, 孙飞1,2( )
)
收稿日期:2021-12-16
出版日期:2022-01-26
发布日期:2022-02-22
作者简介:殷国良,男,硕士研究生,研究方向:生物大分子复合体的结构与功能;E-mail: 基金资助:
YIN Guo-liang1,2( ), SUN Wen-hao1,2, PANG Xiao-yun1, SUN Fei1,2(
), SUN Wen-hao1,2, PANG Xiao-yun1, SUN Fei1,2( )
)
Received:2021-12-16
Published:2022-01-26
Online:2022-02-22
摘要:
植物体的各项生理活动依赖于分子水平上多种植物蛋白质/蛋白质复合体的相互作用和动态变化,了解这些蛋白质/蛋白质复合体的结构和功能对于研究相关植物生理活动的分子机理至关重要。得益于最近的技术进步——包括直接电子探测器的发展和先进的图像处理算法,冷冻电镜技术已经逐步发展成为研究蛋白质/蛋白质复合体的重要技术手段,这也为深入理解植物生理活动分子机理提供了结构生物学研究利器。目前,冷冻电镜技术在分子植物学研究领域的应用仍处于早期,对于一些重要蛋白质复合体的结构功能研究还不够深入。本文在对冷冻电镜技术发展历史进行简要回顾的基础上,对近年来人们利用冷冻电镜技术在植物光合作用、胁迫响应等方面进行的分子机理研究进行了梳理,旨在为加强分子植物学和冷冻电镜技术两个研究领域的合作提供有益参考。
殷国良, 孙文浩, 庞效云, 孙飞. 冷冻电镜技术在分子植物学研究中的应用[J]. 生物技术通报, 2022, 38(1): 15-32.
YIN Guo-liang, SUN Wen-hao, PANG Xiao-yun, SUN Fei. Application of cryo-Electron Microscopy in Molecular Botany Research[J]. Biotechnology Bulletin, 2022, 38(1): 15-32.
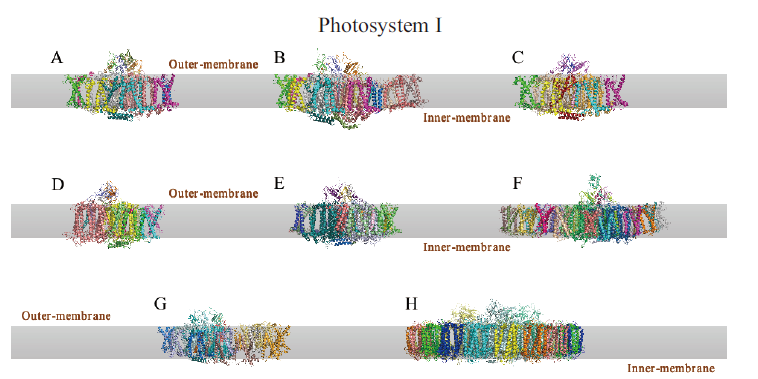
图2 光系统I的冷冻电镜结构 A:豌豆PSI-FD复合体;B:玉米PSI-LHCI-LHCII复合体;C:杜氏盐藻PSI复合体;D:红藻PSI复合体;E:莱茵衣藻PSI复合体;F:纤细角毛藻PSI-FCP复合体;G:小立碗藓PSI复合体;H:嗜热蓝藻PSI-isiA复合体
Fig. 2 cryo-EM structure of photosystem I A:PSI-FD complex of Pisum sativum;B:PSI-LHCI-LHCII complex of Zea mays;C:PSI complex of Dunaliella salina;D:PSI complex of Cyanidioschyzon merolae;E:PSI complex of Chlamydomonas reinhardtii;F:PSI-FCP complex of Chaeto-ceros gracilis;G:PSI complex of Physcomitrium patens;H:PSI-isiA complex of Thermosynechococcus vulcanus
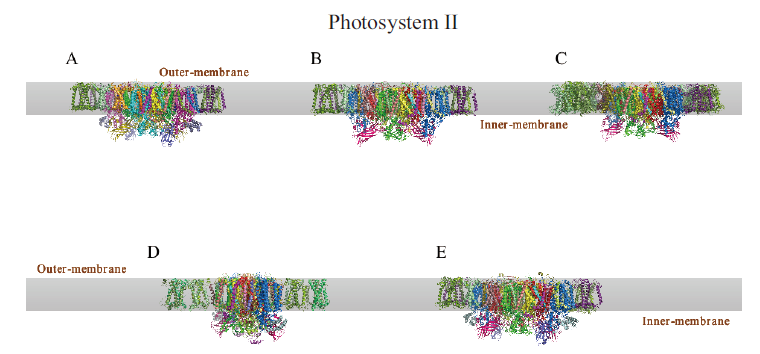
图3 光系统II的冷冻电镜结构 A:菠菜PSII-LHCII复合体;B:豌豆C2S2型PSII-LHCII复合体;C:拟南芥PSII复合体;D:纤细角毛藻C2S2型PSII-FCPII复合体;E:莱茵衣藻C2S2型PSII-LHCII复合体
Fig.3 Cryo-EM structure of photosystem II A:PSII-LHCII complex of Spinacia oleracea. B:PSII-LHCII complex of Pisum sativum. C:PSII complex of Arabidopsis thaliana. D:C2S2 type PSII-FCPII complex of Chaetoceros gracilis. E:C2S2 type PSII-LHCII complex of Chlamydomonas reinhardtii

图7 植物NLR蛋白ZAR1由静息状态进入激活状态并寡聚化形成抗病小体的构象变化
Fig. 7 Conformational changes of plant NLR protein ZAR1 from inactive state to active state and then oligomerized to form resistsome

图9 OSCA1蛋白的冷冻电镜结构及其激活构象 AtOSCA1.1(A)、AtOSCA3.1(B)的冷冻电镜结构比对;(C)AtOSCA1.1构象变化以允许Ca2+进入
Fig. 9 cryo-EM structure and activated conformation of OSCA1 Cryo-EM structure of AtOSCA1.1(A),AtOSCA3.1(B);(C)Conformational changes of AtOSCA1.1 to allow Ca2 + entry
| Name | Organism(s) | Resolution | Year | PDB | Reference | |
|---|---|---|---|---|---|---|
| Photosynthesis | PSII-LHCII supercomplex | Spinacia oleracea | 3.20 Å | 2016 | 3JCU | [ |
| M-LHCII and CP24 complexes | Pisum sativum | 3.50 Å | 2017 | 5XNO | [ | |
| C2S2M2N2-type PSII-LHCII | Pisum sativum | 2.70 Å | 2017 | 5XNL | [ | |
| Phycobilisome | Griffithsia pacifica | 3.50 Å | 2017 | 5Y6P | [ | |
| PSI-LHCR | Cyanidioschyzon merolae strain 10D | 3.82 Å | 2018 | 5ZGH | [ | |
| Photosystem I supercomplex with light-harvesting complexes I and II | Zea mays,Zea mays subsp. mays | 3.30 Å | 2018 | 5ZJI | [ | |
| PSII-FCP supercomplex | Chaetoceros gracilis | 3.02 Å | 2019 | 6JLU | [ | |
| C2S2M2L2-type PSII-LHCII supercomplex | Chlamydomonas reinhardtii | 3.40 Å | 2019 | 6KAD | [ | |
| C2S2-type PSII-LHCII supercomplex | Chlamydomonas reinhardtii | 2.70 Å | 2019 | 6KAC | [ | |
| Photosystem I | Chlamydomonas reinhardtii | 3.30 Å | 2019 | 6IJO | [ | |
| Cytochrome b6f complex | Spinacia oleracea | 3.58 Å | 2019 | 6RQF | [ | |
| Phycobilisome | Porphyridium purpureum | 2.80 Å | 2019 | 6KGX | [ | |
| NDH | Thermosynechococcus vestitus BP-1 | 3.10 Å | 2019 | 6NBY | [ | |
| Photosystem I | Dunaliella salina | 3.20 Å | 2020 | 6RHZ | [ | |
| PSI-FCPI supercomplex | Chaetoceros gracilis | 2.40 Å | 2020 | 6L4U | [ | |
| Photosystem I complex | Synechocystis sp. PCC 6803 substr. Kazusa | 3.10 Å | 2020 | 6UZV | [ | |
| Fd-NDH-1L complex | Thermosynechococcus elongatus BP-1 | 3.20 Å | 2020 | 6L7O | [ | |
| NDH-1LdelV complex | Thermosynechococcus elongatus BP-1 | 3.60 Å | 2020 | 6L7P | [ | |
| PSI-NDH supercomplex | Hordeum vulgare subsp. spontaneum | 4.50 Å | 2021 | 7F9O | [ | |
| Chloroplast NDH complex | Hordeum vulgare subsp. spontaneum | 3.70 Å | 2021 | 7EU3 | [ | |
| PSI-LHCI-Lhca5 supercomplex | Hordeum vulgare subsp. spontaneum | 3.40 Å | 2021 | 7EW6 | [ | |
| PSI-LHCI-Lhca6 supercomplex | Hordeum vulgare subsp. spontaneum | 3.88 Å | 2021 | 7EWK | [ | |
| Immunity | NLR complex | Arabidopsis thaliana | 3.70 Å | 2019 | 6J5W | [ |
| NLR RPP1 LRR-ID domain in complex with ATR1 | Hyaloperonospora arabidopsidis Emoy2,Arabidopsis thaliana | 3.16 Å | 2020 | 7CRB | [ | |
| Activated Roq1 resistosome | Nicotiana benthamiana,Xanthomonas euvesicatoria | 3.80 Å | 2020 | 7JLU | [ | |
| Transportation | atOSCA3.1 channel | Arabidopsis thaliana | 4.80 Å | 2018 | 5Z1F | [ |
| atOSCA1.1 channel | Arabidopsis thaliana | 3.52 Å | 2018 | 6JPF | [ | |
| Activated ion channel OSCA1.2 | Arabidopsis thaliana | 3.10 Å | 2018 | 6MGV | [ | |
| Cation channel | Arabidopsis thaliana | 3.68 Å | 2018 | 6IJZ | [ | |
| Ion channel OSCA1.2 | Oryza sativa | 4.90 Å | 2019 | 6OCE | [ | |
| MSL1 | Arabidopsis thaliana | 3.06 Å | 2020 | 6VXM | [ |
表1 冷冻电镜技术解析获得的植物相关重要生物大分子结构
Table 1 Structure of important plant-related biological macromolecules solved by cryo-EM
| Name | Organism(s) | Resolution | Year | PDB | Reference | |
|---|---|---|---|---|---|---|
| Photosynthesis | PSII-LHCII supercomplex | Spinacia oleracea | 3.20 Å | 2016 | 3JCU | [ |
| M-LHCII and CP24 complexes | Pisum sativum | 3.50 Å | 2017 | 5XNO | [ | |
| C2S2M2N2-type PSII-LHCII | Pisum sativum | 2.70 Å | 2017 | 5XNL | [ | |
| Phycobilisome | Griffithsia pacifica | 3.50 Å | 2017 | 5Y6P | [ | |
| PSI-LHCR | Cyanidioschyzon merolae strain 10D | 3.82 Å | 2018 | 5ZGH | [ | |
| Photosystem I supercomplex with light-harvesting complexes I and II | Zea mays,Zea mays subsp. mays | 3.30 Å | 2018 | 5ZJI | [ | |
| PSII-FCP supercomplex | Chaetoceros gracilis | 3.02 Å | 2019 | 6JLU | [ | |
| C2S2M2L2-type PSII-LHCII supercomplex | Chlamydomonas reinhardtii | 3.40 Å | 2019 | 6KAD | [ | |
| C2S2-type PSII-LHCII supercomplex | Chlamydomonas reinhardtii | 2.70 Å | 2019 | 6KAC | [ | |
| Photosystem I | Chlamydomonas reinhardtii | 3.30 Å | 2019 | 6IJO | [ | |
| Cytochrome b6f complex | Spinacia oleracea | 3.58 Å | 2019 | 6RQF | [ | |
| Phycobilisome | Porphyridium purpureum | 2.80 Å | 2019 | 6KGX | [ | |
| NDH | Thermosynechococcus vestitus BP-1 | 3.10 Å | 2019 | 6NBY | [ | |
| Photosystem I | Dunaliella salina | 3.20 Å | 2020 | 6RHZ | [ | |
| PSI-FCPI supercomplex | Chaetoceros gracilis | 2.40 Å | 2020 | 6L4U | [ | |
| Photosystem I complex | Synechocystis sp. PCC 6803 substr. Kazusa | 3.10 Å | 2020 | 6UZV | [ | |
| Fd-NDH-1L complex | Thermosynechococcus elongatus BP-1 | 3.20 Å | 2020 | 6L7O | [ | |
| NDH-1LdelV complex | Thermosynechococcus elongatus BP-1 | 3.60 Å | 2020 | 6L7P | [ | |
| PSI-NDH supercomplex | Hordeum vulgare subsp. spontaneum | 4.50 Å | 2021 | 7F9O | [ | |
| Chloroplast NDH complex | Hordeum vulgare subsp. spontaneum | 3.70 Å | 2021 | 7EU3 | [ | |
| PSI-LHCI-Lhca5 supercomplex | Hordeum vulgare subsp. spontaneum | 3.40 Å | 2021 | 7EW6 | [ | |
| PSI-LHCI-Lhca6 supercomplex | Hordeum vulgare subsp. spontaneum | 3.88 Å | 2021 | 7EWK | [ | |
| Immunity | NLR complex | Arabidopsis thaliana | 3.70 Å | 2019 | 6J5W | [ |
| NLR RPP1 LRR-ID domain in complex with ATR1 | Hyaloperonospora arabidopsidis Emoy2,Arabidopsis thaliana | 3.16 Å | 2020 | 7CRB | [ | |
| Activated Roq1 resistosome | Nicotiana benthamiana,Xanthomonas euvesicatoria | 3.80 Å | 2020 | 7JLU | [ | |
| Transportation | atOSCA3.1 channel | Arabidopsis thaliana | 4.80 Å | 2018 | 5Z1F | [ |
| atOSCA1.1 channel | Arabidopsis thaliana | 3.52 Å | 2018 | 6JPF | [ | |
| Activated ion channel OSCA1.2 | Arabidopsis thaliana | 3.10 Å | 2018 | 6MGV | [ | |
| Cation channel | Arabidopsis thaliana | 3.68 Å | 2018 | 6IJZ | [ | |
| Ion channel OSCA1.2 | Oryza sativa | 4.90 Å | 2019 | 6OCE | [ | |
| MSL1 | Arabidopsis thaliana | 3.06 Å | 2020 | 6VXM | [ |
| [1] |
Henderson R, Unwin PN. Three-dimensional model of purple membrane obtained by electron microscopy[J]. Nature, 1975, 257(5521):28-32.
doi: 10.1038/257028a0 URL |
| [2] |
Unwin PN, Henderson R. Molecular structure determination by electron microscopy of unstained crystalline specimens[J]. J Mol Biol, 1975, 94(3):425-440.
pmid: 1236957 |
| [3] |
Henderson R, Baldwin JM, Ceska TA, et al. Model for the structure of bacteriorhodopsin based on high-resolution electron cryo-microscopy[J]. J Mol Biol, 1990, 213(4):899-929.
pmid: 2359127 |
| [4] |
Henderson R. The potential and limitations of neutrons, electrons and X-rays for atomic resolution microscopy of unstained biological molecules[J]. Q Rev Biophys, 1995, 28(2):171-193.
pmid: 7568675 |
| [5] |
Adrian M, Dubochet J, Lepault J, et al. Cryo-electron microscopy of viruses[J]. Nature, 1984, 308(5954):32-36.
doi: 10.1038/308032a0 URL |
| [6] |
Downing KH. Observations of restricted beam-induced specimen motion with small-spot illumination[J]. Ultramicroscopy, 1988, 24(4):387-397.
pmid: 3363743 |
| [7] |
Hayward SB, Glaeser RM. Radiation damage of purple membrane at low temperature[J]. Ultramicroscopy, 1979, 04(2):201-210.
pmid: 473421 |
| [8] |
Frank J. Classification of macromolecular assemblies studied as ‘single particles’[J]. Q Rev Biophys, 1990, 23(3):281-329.
pmid: 2204955 |
| [9] |
Russo CJ, Passmore LA. Controlling protein adsorption on graphene for cryo-EM using low-energy hydrogen plasmas[J]. Nat Methods, 2014, 11(6):649-652.
doi: 10.1038/NMETH.2931 |
| [10] |
Pantelic RS, Meyer JC, Kaiser U, et al. Graphene oxide:a substrate for optimizing preparations of frozen-hydrated samples[J]. J Struct Biol, 2010, 170(1):152-156.
doi: 10.1016/j.jsb.2009.12.020 pmid: 20035878 |
| [11] |
Russo CJ, Passmore LA. Electron microscopy:Ultrastable gold substrates for electron cryomicroscopy[J]. Science, 2014, 346(6215):1377-1380.
doi: 10.1126/science.1259530 URL |
| [12] |
Huang XJ, Zhang L, Wen ZL, et al. Amorphous nickel titanium alloy film:a new choice for cryo electron microscopy sample preparation[J]. Prog Biophys Mol Biol, 2020, 156:3-13.
doi: 10.1016/j.pbiomolbio.2020.07.009 URL |
| [13] |
Fan HC, Wang B, Zhang Y, et al. A cryo-electron microscopy support film formed by 2D crystals of hydrophobin HFBI[J]. Nat Commun, 2021, 12(1):7257.
doi: 10.1038/s41467-021-27596-8 URL |
| [14] |
Ravelli RBG, Nijpels FJT, Henderikx RJM, et al. Cryo-EM structures from sub-nl volumes using pin-printing and jet vitrification[J]. Nat Commun, 2020, 11(1):2563.
doi: 10.1038/s41467-020-16392-5 URL |
| [15] |
Noble AJ, Wei H, Dandey VP, et al. Reducing effects of particle adsorption to the air-water interface in cryo-EM[J]. Nat Methods, 2018, 15(10):793-795.
doi: 10.1038/s41592-018-0139-3 pmid: 30250056 |
| [16] |
Bharat TAM, Scheres SHW. Resolving macromolecular structures from electron cryo-tomography data using subtomogram averaging in RELION[J]. Nat Protoc, 2016, 11(11):2054-2065.
doi: 10.1038/nprot.2016.124 URL |
| [17] |
Punjani A, Rubinstein JL, Fleet DJ, et al. cryoSPARC:algorithms for rapid unsupervised cryo-EM structure determination[J]. Nat Methods, 2017, 14(3):290-296.
doi: 10.1038/nmeth.4169 pmid: 28165473 |
| [18] |
Tegunov D, Cramer P. Real-time cryo-electron microscopy data preprocessing with warp[J]. Nat Methods, 2019, 16(11):1146-1152.
doi: 10.1038/s41592-019-0580-y URL |
| [19] |
Wu CL, Huang XJ, Cheng J, et al. High-quality, high-throughput cryo-electron microscopy data collection via beam tilt and astigmatism-free beam-image shift[J]. J Struct Biol, 2019, 208(3):107396.
doi: 10.1016/j.jsb.2019.09.013 URL |
| [20] |
Liao MF, Cao EH, Julius D, et al. Structure of the TRPV1 ion channel determined by electron cryo-microscopy[J]. Nature, 2013, 504(7478):107-112.
doi: 10.1038/nature12822 URL |
| [21] |
Yip KM, Fischer N, Paknia E, et al. Atomic-resolution protein structure determination by cryo-EM[J]. Nature, 2020, 587(7832):157-161.
doi: 10.1038/s41586-020-2833-4 URL |
| [22] |
Nakane T, Kotecha A, Sente A, et al. Single-particle cryo-EM at atomic resolution[J]. Nature, 2020, 587(7832):152-156.
doi: 10.1038/s41586-020-2829-0 URL |
| [23] | Wan RX, Bai R, Yan CY, et al. Structures of the catalytically activated yeast spliceosome reveal the mechanism of branching[J]. Cell, 2019, 177(2):339-351. e13. |
| [24] |
Wang N, Zhao DM, Wang JL, et al. Architecture of African swine fever virus and implications for viral assembly[J]. Science, 2019, 366(6465):640-644.
doi: 10.1126/science.aaz1439 pmid: 31624094 |
| [25] |
Andrés G, Charro D, Matamoros T, et al. The cryo-EM structure of African swine fever virus unravels a unique architecture comprising two icosahedral protein capsids and two lipoprotein membranes[J]. J Biol Chem, 2020, 295(1):1-12.
doi: 10.1074/jbc.AC119.011196 URL |
| [26] |
Liang YL, Khoshouei M, Radjainia M, et al. Phase-plate cryo-EM structure of a class B GPCR-G-protein complex[J]. Nature, 2017, 546(7656):118-123.
doi: 10.1038/nature22327 URL |
| [27] | de Waal P, Zhou XE, He YZ, et al. Structural identification of phosphorylation codes for arrestin recruitment by G protein-coupled receptors[J]. Protein Sci, 2017, 26:133-133. |
| [28] |
Du J, Wang DJ, Fan HC, et al. Structures of human mGlu2 and mGlu7 homo- and heterodimers[J]. Nature, 2021, 594(7864):589-593.
doi: 10.1038/s41586-021-03641-w URL |
| [29] |
Kang YY, Kuybeda O, de Waal PW, et al. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein[J]. Nature, 2018, 558(7711):553-558.
doi: 10.1038/s41586-018-0215-y URL |
| [30] |
Liang YL, Khoshouei M, Glukhova A, et al. Phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor-Gs complex[J]. Nature, 2018, 555(7694):121-125.
doi: 10.1038/nature25773 URL |
| [31] |
Qiao AN, Han S, Li XM, et al. Structural basis of G s and G i recognition by the human glucagon receptor[J]. Science, 2020, 367(6484):1346-1352.
doi: 10.1126/science.aaz5346 URL |
| [32] |
Zhang Y, Sun BF, Feng D, et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein[J]. Nature, 2017, 546(7657):248-253.
doi: 10.1038/nature22394 URL |
| [33] | Zhou XE, He YZ, de Waal PW, et al. Identification of phosphorylation codes for arrestin recruitment by G protein-coupled receptors[J]. Cell, 2017, 170(3):457-469. e13. |
| [34] | Gong HR, Li J, Xu A, et al. An electron transfer path connects subunits of a mycobacterial respiratory super complex[J]. Science, 2018, 362(6418):eaat8923. |
| [35] |
Zong S, Wu M, Gu JK, et al. Structure of the intact 14-subunit human cytochrome c oxidase[J]. Cell Res, 2018, 28(10):1026-1034.
doi: 10.1038/s41422-018-0071-1 URL |
| [36] |
Zhu GL, Zeng H, Zhang SB, et al. A 3. 3 Å-resolution structure of hyperthermophilic respiratory complex III reveals the mechanism of its thermal stability[J]. Angew Chem Int Ed Engl, 2020, 59(1):343-351.
doi: 10.1002/anie.v59.1 URL |
| [37] |
Wrapp D, Wang NS, Corbett KS, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation[J]. Science, 2020, 367(6483):1260-1263.
doi: 10.1126/science.abb2507 URL |
| [38] |
Turo\u0148ová B, Sikora M, Schürmann C, et al. In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges[J]. Science, 2020, 370(6513):203-208.
doi: 10.1126/science.abd5223 URL |
| [39] | Long SW, Olsen RJ, Christensen PA, et al. Molecular architecture of early dissemination and massive second wave of the SARS-CoV-2 virus in a major metropolitan area[J]. medRxiv, 2020:2020 Sep 29;2020. 09. 22. 20199125. |
| [40] | Yao HP, Song YT, Chen Y, et al. Molecular architecture of the SARS-CoV-2 virus[J]. Cell, 2020, 183(3):730-738. e13. |
| [41] | Walls AC, Park YJ, Tortorici MA, et al. Structure, function, antigenicity of the SARS-CoV-2 spike glycoprotein[J]. Cell, 2020, 181(2):281-292. e6. |
| [42] |
Ke ZL, Oton J, Qu K, et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions[J]. Nature, 2020, 588(7838):498-502.
doi: 10.1038/s41586-020-2665-2 URL |
| [43] | Wang Q, Wu JQ, Wang HF, et al. Structural basis for RNA replication by the SARS-CoV-2 polymerase[J]. Cell, 2020, 182(2):417-428. e13. |
| [44] |
Lucić V, Förster F, Baumeister W. Structural studies by electron tomography:from cells to molecules[J]. Annu Rev Biochem, 2005, 74:833-865.
doi: 10.1146/biochem.2005.74.issue-1 URL |
| [45] |
Danev R, Baumeister W. Expanding the boundaries of cryo-EM with phase plates[J]. Curr Opin Struct Biol, 2017, 46:87-94.
doi: 10.1016/j.sbi.2017.06.006 URL |
| [46] |
Wang HW, Fan X. Challenges and opportunities in cryo-EM with phase plate[J]. Curr Opin Struct Biol, 2019, 58:175-182.
doi: 10.1016/j.sbi.2019.06.013 URL |
| [47] | Bieber A, Capitanio C, Wilfling F, et al. Sample preparation by 3D-correlative focused ion beam milling for high-resolution cryo-electron tomography[J]. J Vis Exp, 2021(176):e62886. |
| [48] | Moravcová J, Pinkas M, Holbová R, et al. Preparation and cryo-FIB micromachining of Saccharomyces cerevisiae for cryo-electron tomography[J]. J Vis Exp, 2021(177):e62351. |
| [49] |
Zhang JG, Zhang DY, Sun L, et al. VHUT-cryo-FIB, a method to fabricate frozen hydrated lamellae from tissue specimens for in situ cryo-electron tomography[J]. J Struct Biol, 2021, 213(3):107763.
doi: 10.1016/j.jsb.2021.107763 URL |
| [50] | Hu H, Santiveri M, Wadhwa N, et al. Structural basis of torque generation in the bi-directional bacterial flagellar motor[J]. Trends Biochem Sci, 2021, S0968-0004(21)00139-0. |
| [51] |
Namba K. A proposed gear mechanism for torque generation in the flagellar motor[J]. Nat Struct Mol Biol, 2020, 27(11):1004-1006.
doi: 10.1038/s41594-020-00514-0 URL |
| [52] |
Zhao XW, Norris SJ, Liu J. Molecular architecture of the bacterial flagellar motor in cells[J]. Biochemistry, 2014, 53(27):4323-4333.
doi: 10.1021/bi500059y URL |
| [53] | Zhu SW, Qin Z, Wang JY, et al. In situ structural analysis of the spirochetal flagellar motor by cryo-electron tomography[J]. Methods Mol Biol, 2017, 1593:229-242. |
| [54] |
Dimchev G, Amiri B, Fäβler F, et al. Computational toolbox for ultrastructural quantitative analysis of filament networks in cryo-ET data[J]. J Struct Biol, 2021, 213(4):107808.
doi: 10.1016/j.jsb.2021.107808 pmid: 34742832 |
| [55] |
Sazzed S, Song JH, Kovacs JA, et al. Tracing actin filament bundles in three-dimensional electron tomography density maps of hair cell stereocilia[J]. Molecules, 2018, 23(4):882.
doi: 10.3390/molecules23040882 URL |
| [56] | Kittisopikul M, Shimi T, Tatli M, et al. Computational analyses reveal spatial relationships between nuclear pore complexes and specific lamins[J]. J Cell Biol, 2021, 220(4):e202007082. |
| [57] |
Mahamid J, Pfeffer S, Schaffer M, et al. Visualizing the molecular sociology at the HeLa cell nuclear periphery[J]. Science, 2016, 351(6276):969-972.
doi: 10.1126/science.aad8857 pmid: 26917770 |
| [58] |
Zhang YQ, Li S, Zeng C, et al. Molecular architecture of the luminal ring of the Xenopus laevis nuclear pore complex[J]. Cell Res, 2020, 30(6):532-540.
doi: 10.1038/s41422-020-0320-y URL |
| [59] | 孙飞. 生物三维电子显微成像技术展望[M]. 中国科学院创新发展研究中心. 中国生命健康2035-技术预见. 北京: 科学出版社, 2019:241-256. |
| Sun Fei. Prospect of biological three-dimensional electron microscopy imaging technology[M]. Center for Innovation and Development Chinese Academy of Sciences. Technology Foresight Towards 2035 in China: Life and Health. Science Press. 2019:241-256. | |
| [60] |
Anderson S, Anderson HL, Bashall A, et al. Assembly and crystal structure of a photoactive array of five porphyrins[J]. Angew Chem Int Ed Engl, 1995, 34(10):1096-1099.
doi: 10.1002/(ISSN)1521-3773 URL |
| [61] |
Matsumura H, Xie Y, Shirakata S, et al. Crystal structures of C4 form maize and quaternary complex of E. coli phosphoenolpyruvate carboxylases[J]. Structure, 2002, 10(12):1721-1730.
doi: 10.1016/S0969-2126(02)00913-9 URL |
| [62] |
Toporik H, Khmelnitskiy A, Dobson Z, et al. The structure of a red-shifted photosystem I reveals a red site in the core antenna[J]. Nat Commun, 2020, 11:5279.
doi: 10.1038/s41467-020-18884-w URL |
| [63] |
Pi X, Tian LR, Dai HE, et al. Unique organization of photosystem I-light-harvesting super complex revealed by cryo-EM from a red alga[J]. Proc Natl Acad Sci USA, 2018, 115(17):4423-4428.
doi: 10.1073/pnas.1722482115 URL |
| [64] |
Kubota-Kawai H, Burton-Smith RN, Tokutsu R, et al. Ten antenna proteins are associated with the core in the supramolecular organization of the photosystem I super complex in Chlamydomonas reinhardtii[J]. J Biol Chem, 2019, 294(12):4304-4314.
doi: 10.1074/jbc.RA118.006536 pmid: 30670590 |
| [65] |
Su XD, Ma J, Pan XW, et al. Antenna arrangement and energy transfer pathways of a green algal photosystem-I-LHCI super complex[J]. Nat Plants, 2019, 5(3):273-281.
doi: 10.1038/s41477-019-0380-5 URL |
| [66] |
Perez-Boerema A, Klaiman D, Caspy I, et al. Structure of a minimal photosystem I from the green alga Dunaliella salina[J]. Nat Plants, 2020, 6(3):321-327.
doi: 10.1038/s41477-020-0611-9 pmid: 32123351 |
| [67] |
Nagao R, Kato K, Ifuku K, et al. Structural basis for assembly and function of a diatom photosystem I-light-harvesting super complex[J]. Nat Commun, 2020, 11(1):2481.
doi: 10.1038/s41467-020-16324-3 pmid: 32424145 |
| [68] |
Pan XW, Ma J, Su XD, et al. Structure of the maize photosystem I super complex with light-harvesting complexes I and II[J]. Science, 2018, 360(6393):1109-1113.
doi: 10.1126/science.aat1156 URL |
| [69] | Shen L, Tang K, Wang W, et al. Architecture of the chloroplast PSI-NDH super complex in Hordeum vulgare[J]. Nature, 2021: 2021 Dec 8. |
| [70] |
Zhang CL, Shuai J, Ran ZX, et al. Structural insights into NDH-1 mediated cyclic electron transfer[J]. Nat Commun, 2020, 11(1):888.
doi: 10.1038/s41467-020-14732-z URL |
| [71] |
Wei XP, Su XD, Cao P, et al. Structure of spinach photosystem II-LHCII super complex at 3. 2 Å resolution[J]. Nature, 2016, 534(7605):69-74.
doi: 10.1038/nature18020 URL |
| [72] |
Su XD, Ma J, Wei XP, et al. Structure and assembly mechanism of plant C2S2M2-type PSII-LHCII super complex[J]. Science, 2017, 357(6353):815-820.
doi: 10.1126/science.aan0327 URL |
| [73] |
van Bezouwen LS, Caffarri S, Kale RS, et al. Subunit and chlorophyll organization of the plant photosystem II super complex[J]. Nat Plants, 2017, 3:17080.
doi: 10.1038/nplants.2017.80 pmid: 28604725 |
| [74] | Pi X, Zhao SH, Wang WD, et al. The pigment-protein network of a diatom photosystem II-light-harvesting antenna super complex[J]. Science, 2019, 365(6452):eaax4406. |
| [75] |
Sheng X, Watanabe A, Li AJ, et al. Structural insight into light harvesting for photosystem II in green algae[J]. Nat Plants, 2019, 5(12):1320-1330.
doi: 10.1038/s41477-019-0543-4 pmid: 31768031 |
| [76] |
Malone LA, Qian P, Mayneord GE, et al. Cryo-EM structure of the spinach cytochrome b 6 f complex at 3. 6 Å resolution[J]. Nature, 2019, 575(7783):535-539.
doi: 10.1038/s41586-019-1746-6 URL |
| [77] |
Laughlin TG, Bayne AN, Trempe JF, et al. Structure of the complex I-like molecule NDH of oxygenic photosynjournal[J]. Nature, 2019, 566(7744):411-414.
doi: 10.1038/s41586-019-0921-0 URL |
| [78] |
Zhang J, Ma JF, Liu DS, et al. Structure of phycobilisome from the red alga Griffithsia Pacifica[J]. Nature, 2017, 551(7678):57-63.
doi: 10.1038/nature24278 URL |
| [79] |
Ma JF, You X, Sun S, et al. Structural basis of energy transfer in Porphyridium purpureum phycobilisome[J]. Nature, 2020, 579(7797):146-151.
doi: 10.1038/s41586-020-2020-7 URL |
| [80] |
Li MJ, Ma JF, Li XM, et al. In situ cryo-ET structure of phycobilisome-photosystem II super complex from red alga[J]. eLife, 2021, 10:e69635.
doi: 10.7554/eLife.69635 URL |
| [81] |
Zhu JK. Abiotic stress signaling and responses in plants[J]. Cell, 2016, 167(2):313-324.
doi: 10.1016/j.cell.2016.08.029 URL |
| [82] |
Song W, Forderer A, Yu DL, et al. Structural biology of plant defence[J]. New Phytol, 2021, 229(2):692-711.
doi: 10.1111/nph.v229.2 URL |
| [83] |
Wang W, Feng BM, Zhou JM, et al. Plant immune signaling:advancing on two frontiers[J]. J Integr Plant Biol, 2020, 62(1):2-24.
doi: 10.1111/jipb.v62.1 URL |
| [84] |
Wang JZ, Chai JJ. Structural insights into the plant immune receptors PRRs and NLRs[J]. Plant Physiol, 2020, 182(4):1566-1581.
doi: 10.1104/pp.19.01252 URL |
| [85] | 赵燕波, 郭婉兰, 张丹丹, 等. 植物免疫受体病原体识别机制的结构生物学研究进展[J]. 福建师范大学学报:自然科学版, 2020, 36(2):12-24. |
| Zhao YB, Guo WL, Zhang DD, et al. Progress in structural basis for pathogen recognition mechanism of plant immune receptors[J]. J Fujian Norm Univ Nat Sci Ed, 2020, 36(2):12-24. | |
| [86] | 邓一文, 刘裕强, 王静, 等. 农作物抗病虫研究的战略思考[J]. 中国科学:生命科学, 2021, 51:1435-1446. |
| Deng YW, Liu YQ, Wang J, et al. Strategic thinking and research on crop disease and pest resistance in China[J]. Sci Sin Vitae, 2021, 51:1435-1446. | |
| [87] |
Jones JDG, Dangl JL. The plant immune system[J]. Nature, 2006, 444(7117):323-329.
doi: 10.1038/nature05286 URL |
| [88] | Jones JDG, Vance RE, Dangl JL. Intracellular innate immune surveillance devices in plants and animals[J]. Science, 2016, 354(6316):aaf6395. |
| [89] | Ngou BPM, Ahn HK, Ding PT, et al. Mutual potentiation of plant immunity by cell-surface and intracellular receptors[J]. Nature, 2021, 592(7852):110-115. |
| [90] | Yuan MH, Jiang ZY, Bi GZ, et al. Pattern-recognition receptors are required for NLR-mediated plant immunity[J]. Nature, 2021, 592(7852):105-109. |
| [91] |
Pruitt RN, Gust AA, Nürnberger T. Plant immunity unified[J]. Nat Plants, 2021, 7(4):382-383.
doi: 10.1038/s41477-021-00903-3 URL |
| [92] | Chang M, Chen H, Liu F, et al. PTI and ETI:convergent pathways with diverse elicitors[J]. Trends Plant Sci, 2021: 2021 Dec 1;S1360-2021 Dec 1;S1385(21)00319-8. |
| [93] |
Boutrot F, Zipfel C. Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance[J]. Annu Rev Phytopathol, 2017, 55:257-286.
doi: 10.1146/annurev-phyto-080614-120106 pmid: 28617654 |
| [94] |
Liu ZX, Wu Y, Yang F, et al. BIK1 interacts with PEPRs to mediate ethylene-induced immunity[J]. Proc Natl Acad Sci USA, 2013, 110(15):6205-6210.
doi: 10.1073/pnas.1215543110 URL |
| [95] |
Iizasa E, Mitsutomi M, Nagano Y. Direct binding of a plant LysM receptor-like kinase, LysM RLK1/CERK1, to chitin in vitro[J]. J Biol Chem, 2010, 285(5):2996-3004.
doi: 10.1074/jbc.M109.027540 URL |
| [96] |
Hayafune M, Berisio R, Marchetti R, et al. Chitin-induced activation of immune signaling by the rice receptor CEBiP relies on a unique sandwich-type dimerization[J]. Proc Natl Acad Sci USA, 2014, 111(3):E404-E413.
doi: 10.1073/pnas.1312099111 URL |
| [97] |
Wang JZ, Chai JJ. Molecular actions of NLR immune receptors in plants and animals[J]. Sci China Life Sci, 2020, 63(9):1303-1316.
doi: 10.1007/s11427-019-1687-6 URL |
| [98] | Bentham A, Burdett H, Anderson PA, et al. Animal NLRs provide structural insights into plant NLR function[J]. Ann Bot, 2017, 119(5):827-702. |
| [99] | Wang JZ, Wang J, Hu MJ, et al. Ligand-triggered allosteric ADP release primes a plant NLR complex[J]. Science, 2019, 364(6435):eaav5868. |
| [100] | Wang JZ, Hu MJ, Wang J, et al. Reconstitution and structure of a plant NLR resistosome conferring immunity[J]. Science, 2019, 364(6435):eaav5870. |
| [101] | Bi GZ, Su M, Li N, et al. The ZAR1 resistosome is a calcium-permeable channel triggering plant immune signaling[J]. Cell, 2021, 184(13):3528-3541. e12. |
| [102] | Ma SC, Lapin D, Liu L, et al. Direct pathogen-induced assembly of an NLR immune receptor complex to form a holoenzyme[J]. Science, 2020, 370(3521):eabe3069. |
| [103] | Martin R, Qi TC, Zhang HB, et al. Structure of the activated ROQ1 resistosome directly recognizing the pathogen effector XopQ[J]. Science, 2020, 370(3521):eabd9993. |
| [104] |
Brutus A, Sicilia F, Macone A, et al. A domain swap approach reveals a role of the plant wall-associated kinase 1(WAK1)as a receptor of oligogalacturonides[J]. Proc Natl Acad Sci USA, 2010, 107(20):9452-9457.
doi: 10.1073/pnas.1000675107 URL |
| [105] |
Choi J, Tanaka K, Cao YR, et al. Identification of a plant receptor for extracellular ATP[J]. Science, 2014, 343(6168):290-294.
doi: 10.1126/science.343.6168.290 URL |
| [106] |
Wang LM, Wilkins KA, Davies JM. Arabidopsis DORN1 extracellular ATP receptor;activation of plasma membrane K +-and Ca2 +-permeable conductances[J]. New Phytol, 2018, 218(4):1301-1304.
doi: 10.1111/nph.2018.218.issue-4 URL |
| [107] |
Gong ZZ, Xiong LM, Shi HZ, et al. Plant abiotic stress response and nutrient use efficiency[J]. Sci China Life Sci, 2020, 63(5):635-674.
doi: 10.1007/s11427-020-1683-x URL |
| [108] | 朱健康, 倪建平. 植物非生物胁迫信号转导及应答[J]. 中国稻米, 2016, 22(6):52-60. |
| Zhu JK, Ni JP. Abiotic stress signaling and responses in plants[J]. China Rice, 2016, 22(6):52-60. | |
| [109] |
Yuan F, Yang HM, Xue Y, et al. OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in Arabidopsis[J]. Nature, 2014, 514(7522):367-371.
doi: 10.1038/nature13593 URL |
| [110] |
Basu D, Haswell ES. Plant mechanosensitive ion channels:an ocean of possibilities[J]. Curr Opin Plant Biol, 2017, 40:43-48.
doi: 10.1016/j.pbi.2017.07.002 URL |
| [111] |
Lee CP, Maksaev G, Jensen GS, et al. MSL1 is a mechanosensitive ion channel that dissipates mitochondrial membrane potential and maintains redox homeostasis in mitochondria during abiotic stress[J]. Plant J, 2016, 88(5):809-825.
doi: 10.1111/tpj.2016.88.issue-5 URL |
| [112] |
Zhang MF, Wang DL, Kang YL, et al. Structure of the mechanosensitive OSCA channels[J]. Nat Struct Mol Biol, 2018, 25(9):850-858.
doi: 10.1038/s41594-018-0117-6 URL |
| [113] |
Jojoa-Cruz S, Saotome K, Murthy SE, et al. Cryo-EM structure of the mechanically activated ion channel OSCA1. 2[J]. eLife, 2018, 7:e41845.
doi: 10.7554/eLife.41845 URL |
| [114] |
Murthy SE, Dubin AE, Whitwam T, et al. OSCA/TMEM63 are an evolutionarily conserved family of mechanically activated ion channels[J]. eLife, 2018, 7:e41844.
doi: 10.7554/eLife.41844 URL |
| [115] |
Liu X, Wang JW, Sun LF. Structure of the hyperosmolality-gated calcium-permeable channel OSCA1. 2[J]. Nat Commun, 2018, 9(1):5060.
doi: 10.1038/s41467-018-07564-5 URL |
| [116] |
Maity K, Heumann JM, McGrath AP, et al. Cryo-EM structure of OSCA1. 2 from Oryza sativa elucidates the mechanical basis of potential membrane hyperosmolality gating[J]. Proc Natl Acad Sci USA, 2019, 116(28):14309-14318.
doi: 10.1073/pnas.1900774116 URL |
| [117] | Li YW, Hu YF, Wang JW, et al. Structural insights into a plant mechanosensitive ion channel MSL1[J]. Cell Rep, 2020, 30(13):4518-4527. e3. |
| [118] |
Deng ZQ, Maksaev G, Schlegel AM, et al. Structural mechanism for gating of a eukaryotic mechanosensitive channel of small conductance[J]. Nat Commun, 2020, 11(1):3690.
doi: 10.1038/s41467-020-17538-1 URL |
| [119] |
Clark MD, Contreras GF, Shen R, et al. Electromechanical coupling in the hyperpolarization-activated K + channel KAT1[J]. Nature, 2020, 583(7814):145-149.
doi: 10.1038/s41586-020-2335-4 URL |
| [120] |
Li SY, Yang F, Sun DM, et al. Cryo-EM structure of the hyperpolarization-activated inwardly rectifying potassium channel KAT1 from Arabidopsis[J]. Cell Res, 2020, 30(11):1049-1052.
doi: 10.1038/s41422-020-00407-3 URL |
| [121] |
Gao F, Han XW, Wu JH, et al. A heat-activated calcium-permeable channel—Arabidopsis cyclic nucleotide-gated ion channel 6—is involved in heat shock responses[J]. Plant J, 2012, 70(6):1056-1069.
doi: 10.1111/tpj.2012.70.issue-6 URL |
| [122] |
Ma Y, Dai XY, Xu YY, et al. COLD1 confers chilling tolerance in rice[J]. Cell, 2015, 160(6):1209-1221.
doi: 10.1016/j.cell.2015.01.046 URL |
| [123] |
Shi YT, Yang SH. COLD1:a cold sensor in rice[J]. Sci China Life Sci, 2015, 58(4):409-410.
doi: 10.1007/s11427-015-4831-6 URL |
| [124] |
Yang TB, Chaudhuri S, Yang LH, et al. A calcium/calmodulin-regulated member of the receptor-like kinase family confers cold tolerance in plants[J]. J Biol Chem, 2010, 285(10):7119-7126.
doi: 10.1074/jbc.M109.035659 URL |
| [125] |
Yang TB, Shad Ali G, Yang LH, et al. Calcium/calmodulin-regulated receptor-like kinase CRLK1 interacts with MEKK1 in plants[J]. Plant Signal Behav, 2010, 5(8):991-994.
doi: 10.4161/psb.5.8.12225 URL |
| [126] | Liu ZY, Jia YX, Ding YL, et al. Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during cold response[J]. Mol Cell, 2017, 66(1): 117-128. e5. |
| [127] | Feng W, Kita D, Peaucelle A, et al. The FERONIA receptor kinase maintains cell-wall integrity during salt stress through Ca2+ signaling[J]. Curr Biol, 2018, 28(5):666-675. e5. |
| [128] |
Tunyasuvunakool K, Adler J, Wu Z, et al. Highly accurate protein structure prediction for the human proteome[J]. Nature, 2021, 596(7873):590-596.
doi: 10.1038/s41586-021-03828-1 URL |
| [1] | 杨志晓, 侯骞, 刘国权, 卢志刚, 曹毅, 芶剑渝, 王轶, 林英超. 不同抗性烟草品系Rubisco及其活化酶对赤星病胁迫的响应[J]. 生物技术通报, 2023, 39(9): 202-212. |
| [2] | 刘奎, 李兴芬, 杨沛欣, 仲昭晨, 曹一博, 张凌云. 青杄转录共激活因子PwMBF1c的功能研究与验证[J]. 生物技术通报, 2023, 39(5): 205-216. |
| [3] | 魏明, 王欣玉, 伍国强, 赵萌. NAD依赖型去乙酰化酶SRT在植物表观遗传调控中的作用[J]. 生物技术通报, 2023, 39(4): 59-70. |
| [4] | 王琪, 胡哲, 富薇, 李光哲, 郝林. 伯克霍尔德氏菌GD17对黄瓜幼苗耐干旱的调节[J]. 生物技术通报, 2023, 39(3): 163-175. |
| [5] | 张红红, 方晓峰. 相分离调控植物胁迫感知和应答的研究进展[J]. 生物技术通报, 2023, 39(11): 44-53. |
| [6] | 阮航, 多浩源, 范文艳, 吕清晗, 姜述君, 朱生伟. AtERF49在拟南芥应答盐碱胁迫中的作用[J]. 生物技术通报, 2023, 39(1): 150-156. |
| [7] | 王楠楠, 王文佳, 朱强. 植物胁迫相关microRNA研究进展[J]. 生物技术通报, 2022, 38(8): 1-11. |
| [8] | 汤茜茜, 林楚宇, 陶增. 植物组蛋白去甲基化酶研究进展[J]. 生物技术通报, 2022, 38(7): 13-22. |
| [9] | 祖国蔷, 胡哲, 王琪, 李光哲, 郝林. Burkholderia sp. GD17对水稻幼苗镉耐受的调节[J]. 生物技术通报, 2022, 38(4): 153-162. |
| [10] | 李兵娟, 郑璐, 沈仁芳, 兰平. 拟南芥RPP1A参与幼苗生长的蛋白质组学分析[J]. 生物技术通报, 2022, 38(2): 10-20. |
| [11] | 黎猛, 陈跃, 胡凤荣. miR159-GAMYB途径调控植物生长发育的研究进展[J]. 生物技术通报, 2021, 37(9): 234-247. |
| [12] | 马旭辉, 陈茹梅, 柳小庆, 赵军, 张霞. 褪黑素对玉米幼苗根系发育和抗旱性的影响[J]. 生物技术通报, 2021, 37(2): 1-14. |
| [13] | 姚琼, 全林发, 徐淑, 董易之, 李文景, 池艳艳, 陈炳旭. 叉角厉蝽2个章鱼胺受体的基因克隆及化学农药对其表达的影响[J]. 生物技术通报, 2021, 37(10): 152-152. |
| [14] | 武欢, 卢珍红, 郝向阳, 王斌, 焦元辰, 杨春梅, 程春振. 非洲菊GjMnSOD基因的克隆及表达分析[J]. 生物技术通报, 2021, 37(10): 17-25. |
| [15] | 郑璐, 沈仁芳, 兰平. 植物非组蛋白赖氨酸乙酰化修饰的蛋白质组学研究进展[J]. 生物技术通报, 2021, 37(1): 77-89. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||