








生物技术通报 ›› 2023, Vol. 39 ›› Issue (9): 176-182.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0275
王腾辉1( ), 葛雯冬1, 罗雅方1, 范震宇1, 王玉书1,2(
), 葛雯冬1, 罗雅方1, 范震宇1, 王玉书1,2( )
)
收稿日期:2023-03-24
出版日期:2023-09-26
发布日期:2023-10-24
通讯作者:
王玉书,女,博士,教授,研究方向:园艺植物分子生物学;E-mail: wangys1019@126.com作者简介:王腾辉,女,硕士研究生,研究方向:生物化学与分子生物学;E-mail: 372838418@qq.com
基金资助:
WANG Teng-hui1( ), GE Wen-dong1, LUO Ya-fang1, FAN Zhen-yu1, WANG Yu-shu1,2(
), GE Wen-dong1, LUO Ya-fang1, FAN Zhen-yu1, WANG Yu-shu1,2( )
)
Received:2023-03-24
Published:2023-09-26
Online:2023-10-24
摘要:
筛选羽衣甘蓝白色叶形成相关的关键基因和位点以及心叶颜色性状的遗传规律,以羽衣甘蓝红叶亲本WR、白叶亲本WB及其构建的F2分离群体为试验材料,从F2群体中挑选红叶和白叶植株各20株,分别构建2个DNA混合池,对子代混合池和亲本分别开展50×和20×覆盖深度的全基因组重测序,定位白叶性状关联区间,并根据基因注释信息预测候选基因。结果表明,重测序获得3 987 718个单核酸多态性(SNP)标记,获得位于第2、3和9染色体的8个显著关联区间,根据基因注释和功能分析,筛选到8个候选基因(Bol030253、Bol029431、Bol012077、Bol007709、Bol030318、Bol030235、Bol030286和Bol005195)。将8个基因确定为羽衣甘蓝白叶的候选基因,可能在羽衣甘蓝叶片颜色形成过程中起着重要作用。
王腾辉, 葛雯冬, 罗雅方, 范震宇, 王玉书. 基于极端混合池(BSA)全基因组重测序的羽衣甘蓝白色叶基因定位[J]. 生物技术通报, 2023, 39(9): 176-182.
WANG Teng-hui, GE Wen-dong, LUO Ya-fang, FAN Zhen-yu, WANG Yu-shu. Gene Mapping of Kale White Leaves Based on Whole Genome Re-sequencing of Extreme Mixed Pool(BSA)[J]. Biotechnology Bulletin, 2023, 39(9): 176-182.
| 群体 Group | 总株数 Total plants | 红叶株数 No. of red-leaf plants | 白叶株数 No. of white-leaf plants | 红叶∶白叶 Red leaf∶White leaf | 卡方值 χ2 | 检验值 χ20.05 |
|---|---|---|---|---|---|---|
| WR | 20 | 20 | 0 | - | - | - |
| WB | 20 | 0 | 20 | - | - | - |
| F1 | 20 | 20 | 0 | - | - | - |
| F2 | 1 300 | 982 | 318 | 3.09∶1 | 0.20 | 3.84 |
表1 ‘WR’和‘WB’杂交组合后代分离比例
Table 1 Segregation of the crosses between ‘WR’ and ‘WB’ in progeny populations
| 群体 Group | 总株数 Total plants | 红叶株数 No. of red-leaf plants | 白叶株数 No. of white-leaf plants | 红叶∶白叶 Red leaf∶White leaf | 卡方值 χ2 | 检验值 χ20.05 |
|---|---|---|---|---|---|---|
| WR | 20 | 20 | 0 | - | - | - |
| WB | 20 | 0 | 20 | - | - | - |
| F1 | 20 | 20 | 0 | - | - | - |
| F2 | 1 300 | 982 | 318 | 3.09∶1 | 0.20 | 3.84 |
| 样品 Sample | Clean reads数 Number of clean reads | 比对reads数 Mapped reads | 参考基因组比对 Map ratio/% | 平均覆盖深度 Average cover depth | 基因组覆盖度 Coverage/% | 碱基数质量值大于Q20/% |
|---|---|---|---|---|---|---|
| W12 | 122 570 282 | 117 542 515 | 95.90 | 29.03 | 88.05 | 98.35 |
| W06 | 156 059 918 | 148 403 662 | 95.09 | 35.68 | 88.38 | 98.17 |
| L-bulk | 370 780 056 | 352 633 985 | 95.11 | 81.94 | 91.05 | 98.19 |
| B-bulk | 324 884 700 | 307 268 196 | 94.58 | 73.98 | 91.12 | 97.81 |
表2 不同样品测序比对结果
Table 2 Sequencing comparison results of different samples
| 样品 Sample | Clean reads数 Number of clean reads | 比对reads数 Mapped reads | 参考基因组比对 Map ratio/% | 平均覆盖深度 Average cover depth | 基因组覆盖度 Coverage/% | 碱基数质量值大于Q20/% |
|---|---|---|---|---|---|---|
| W12 | 122 570 282 | 117 542 515 | 95.90 | 29.03 | 88.05 | 98.35 |
| W06 | 156 059 918 | 148 403 662 | 95.09 | 35.68 | 88.38 | 98.17 |
| L-bulk | 370 780 056 | 352 633 985 | 95.11 | 81.94 | 91.05 | 98.19 |
| B-bulk | 324 884 700 | 307 268 196 | 94.58 | 73.98 | 91.12 | 97.81 |
| 变异位点信息 Variation sites information | WR | WB | R-bulk | B-bulk |
|---|---|---|---|---|
| 基因区间 Intergenic region | 525 758 | 529 072 | 555 294 | 555 413 |
| 非同义突变 Non-synonymous | 266 962 | 270 929 | 283 962 | 283 910 |
| 同义突变 Synonymous | 150 618 | 152 757 | 159 396 | 159 535 |
| 起始密码子丢失 Start lost | 611 | 628 | 672 | 668 |
| 终止密码子丢失 Stop_lost | 6 645 | 6 703 | 7 028 | 7 075 |
| 终止密码子获得 Stop_Gained | 13 829 | 13 994 | 14 600 | 14 596 |
| 5'非翻译区 UTR_5_Prime | 489 | 484 | 511 | 512 |
| SNP总数 SNP total | 964 912 | 979 644 | 1 021 463 | 1 021 699 |
表3 变异位点注释统计表
Table 3 Annotation of variation sites
| 变异位点信息 Variation sites information | WR | WB | R-bulk | B-bulk |
|---|---|---|---|---|
| 基因区间 Intergenic region | 525 758 | 529 072 | 555 294 | 555 413 |
| 非同义突变 Non-synonymous | 266 962 | 270 929 | 283 962 | 283 910 |
| 同义突变 Synonymous | 150 618 | 152 757 | 159 396 | 159 535 |
| 起始密码子丢失 Start lost | 611 | 628 | 672 | 668 |
| 终止密码子丢失 Stop_lost | 6 645 | 6 703 | 7 028 | 7 075 |
| 终止密码子获得 Stop_Gained | 13 829 | 13 994 | 14 600 | 14 596 |
| 5'非翻译区 UTR_5_Prime | 489 | 484 | 511 | 512 |
| SNP总数 SNP total | 964 912 | 979 644 | 1 021 463 | 1 021 699 |
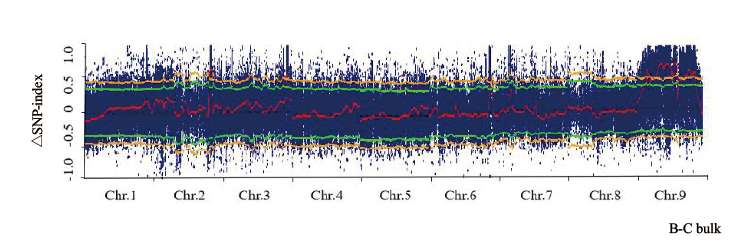
图1 所有染色体的△SNP-index图 红色曲线表示滑窗后的△SNP-index值,黄色线表示99%置信区间值
Fig. 1 △SNP-index map of all chromosomes The red curve represents the Delta SNP-index value after the sliding window, the yellow curve represents the 99% confidence interva
| 染色体 Chromosome | 起始位置 Start/bp | 终止位置 End/bp | 关联区域大小 Size/Mb | 基因数量 No. of genes |
|---|---|---|---|---|
| Chr.2 | 35 100 000 | 37 050 000 | 3.95 | 20 |
| Chr.3 | 51 600 000 | 52 800 000 | 1.20 | 38 |
| Chr.9 | 5 850 000 | 7 850 000 | 2.00 | 20 |
| Chr.9 | 9 150 000 | 10 600 000 | 1.45 | 14 |
| Chr.9 | 11 900 000 | 21 650 000 | 9.75 | 199 |
| Chr.9 | 22 000 000 | 24 250 000 | 2.25 | 16 |
| Chr.9 | 28 100 000 | 33 050 000 | 4.95 | 151 |
| Chr.9 | 33 250 000 | 35 350 000 | 2.10 | 53 |
| 总计Total | — | — | 27.65 | 510 |
表4 SNP位点关联分析获得的关联区域
Table 4 Correlation regions obtained by SNP correlation analysis methods
| 染色体 Chromosome | 起始位置 Start/bp | 终止位置 End/bp | 关联区域大小 Size/Mb | 基因数量 No. of genes |
|---|---|---|---|---|
| Chr.2 | 35 100 000 | 37 050 000 | 3.95 | 20 |
| Chr.3 | 51 600 000 | 52 800 000 | 1.20 | 38 |
| Chr.9 | 5 850 000 | 7 850 000 | 2.00 | 20 |
| Chr.9 | 9 150 000 | 10 600 000 | 1.45 | 14 |
| Chr.9 | 11 900 000 | 21 650 000 | 9.75 | 199 |
| Chr.9 | 22 000 000 | 24 250 000 | 2.25 | 16 |
| Chr.9 | 28 100 000 | 33 050 000 | 4.95 | 151 |
| Chr.9 | 33 250 000 | 35 350 000 | 2.10 | 53 |
| 总计Total | — | — | 27.65 | 510 |
| 基因ID Gene ID | 染色体位置 Chromosomal location/bp | 基因功能注释 Gene function annotation |
|---|---|---|
| Bol030253 | Chr.9:19341984-19344641 | 五肽重复(PPR)超家族蛋白Pentatricopeptide repeat(PPR)superfamily protein |
| Bol029431 | Chr.9:18884039-18885364 | 跨膜蛋白161AB蛋白Transmembrane protein 161AB protein |
| Bol012077 | Chr.9:12469942-12472086 | 跨膜蛋白,推测(DUF3537)Transmembrane protein, putative(DUF3537) |
| Bol007709 | Chr.9:14272763-14273749 | 跨膜蛋白Transmembrane protein |
| Bol030318 | Chr.9:20636604-20637310 | 跨膜蛋白Transmembrane protein |
| Bol030235 | Chr.9:19061948-19069105 | 胡萝卜素通过隐黄质转化为玉米黄质Carotene to zeaxanthin via cryptoxanthin |
| Bol030286 | Chr.9:19934118-19938365 | 编码一种定位于叶绿体和线粒体的FtsH蛋白酶Encodes an FtsH protease that is localized to the chloroplast and the mitochondrion |
| Bol005195 | Chr.9:14477658-14478362 | 叶绿体atp合酶δ亚基ATP synthase subunit delta, chloroplast |
表5 候选基因功能注释
Table 5 Functional annotation of candidate genes
| 基因ID Gene ID | 染色体位置 Chromosomal location/bp | 基因功能注释 Gene function annotation |
|---|---|---|
| Bol030253 | Chr.9:19341984-19344641 | 五肽重复(PPR)超家族蛋白Pentatricopeptide repeat(PPR)superfamily protein |
| Bol029431 | Chr.9:18884039-18885364 | 跨膜蛋白161AB蛋白Transmembrane protein 161AB protein |
| Bol012077 | Chr.9:12469942-12472086 | 跨膜蛋白,推测(DUF3537)Transmembrane protein, putative(DUF3537) |
| Bol007709 | Chr.9:14272763-14273749 | 跨膜蛋白Transmembrane protein |
| Bol030318 | Chr.9:20636604-20637310 | 跨膜蛋白Transmembrane protein |
| Bol030235 | Chr.9:19061948-19069105 | 胡萝卜素通过隐黄质转化为玉米黄质Carotene to zeaxanthin via cryptoxanthin |
| Bol030286 | Chr.9:19934118-19938365 | 编码一种定位于叶绿体和线粒体的FtsH蛋白酶Encodes an FtsH protease that is localized to the chloroplast and the mitochondrion |
| Bol005195 | Chr.9:14477658-14478362 | 叶绿体atp合酶δ亚基ATP synthase subunit delta, chloroplast |
| [1] | 孙日飞, 张淑江, 章时蕃, 等. 紫红色大白菜种质的创新研究[J]. 园艺学报, 2006, 33(5): 1032. |
| Sun RF, Zhang SJ, Zhang SF, et al. Research on creation of purple Chinese cabbage germplasm[J]. Acta Hortic Sin, 2006, 33(5): 1032. | |
| [2] |
Li HB, Zhu LX, Yuan GG, et al. Fine mapping and candidate gene analysis of an anthocyanin-rich gene, BnaA.PL1, conferring purple leaves in Brassica napus L.[J]. Mol Genet Genomics, 2016, 291(4): 1523-1534.
doi: 10.1007/s00438-016-1199-7 URL |
| [3] |
Guo N, Wu J, Zheng SN, et al. Anthocyanin profile characterization and quantitative trait locus mapping in zicaitai(Brassica rapa L. ssp. chinensis var. purpurea)[J]. Mol Breeding, 2015, 35(5): 113.
doi: 10.1007/s11032-015-0237-1 URL |
| [4] |
Zhu PF, Cheng MM, Feng X, et al. Mapping of Pi, a gene conferring pink leaf in ornamental kale(Brassica oleracea L. var. acephala DC)[J]. Euphytica, 2016, 207(2): 377-385.
doi: 10.1007/s10681-015-1555-4 URL |
| [5] |
He Q, Lu QQ, He YT, et al. Dynamic changes of the anthocyanin biosynthesis mechanism during the development of heading Chinese cabbage(Brassica rapa L.) and Arabidopsis under the control of BrMYB2[J]. Front Plant Sci, 2020, 11: 593766.
doi: 10.3389/fpls.2020.593766 URL |
| [6] | 叶沈华, 马晓伟, 杨杰, 等. 甘蓝型油菜叶片白化分子机理研究[J]. 植物遗传资源学报, 2023, 1: 1-18 |
| Ye SH, Ma XW, Yang J, et al. Molecular mechanisms of albino leaves in Brassica napus[J]. J Plant Gen Res, 2023, 1: 1-18. | |
| [7] |
Jiang XF, Zhao H, Guo F, et al. Transcriptomic analysis reveals mechanism of light-sensitive albinism in tea plant Camellia sinensis ‘Huangjinju ’[J]. BMC Plant Biol, 2020, 20(1): 216.
doi: 10.1186/s12870-020-02425-0 |
| [8] | 王容, 李博, 刘刚, 等. 大麦阶段性低温诱导白化突变体的转录组分析[J]. 麦类作物学报, 2023, 43(1): 64-74. |
| Wang R, Li B, Liu G, et al. Transcription analysis of stage specific low temperature induced albinism in barley[J]. J Triticeae Crops, 2023, 43(1): 64-74. | |
| [9] |
Lu HF, Lin T, Klein J, et al. QTL-seq identifies an early flowering QTL located near flowering locus T in cucumber[J]. Theor Appl Genet, 2014, 127(7): 1491-1499.
doi: 10.1007/s00122-014-2313-z URL |
| [10] |
Illa-Berenguer E, Van Houten J, Huang ZJ, et al. Rapid and reliable identification of tomato fruit weight and locule number loci by QTL-seq[J]. Theor Appl Genet, 2015, 128(7): 1329-1342.
doi: 10.1007/s00122-015-2509-x pmid: 25893466 |
| [11] |
张尧锋, 张冬青, 余华胜, 等. 基于极端混合池(BSA)全基因组重测序的甘蓝型油菜有限花序基因定位[J]. 中国农业科学, 2018, 51(16): 3029-3039.
doi: 10.3864/j.issn.0578-1752.2018.16.001 |
| Zhang YF, Zhang DQ, Yu HS, et al. Location and mapping of the determinate growth habit of Brassica napus by bulked segregant analysis(BSA)using whole genome re-sequencing[J]. Sci Agric Sin, 2018, 51(16): 3029-3039. | |
| [12] |
乔军, 刘婧, 李素文, 等. 基于极端混合池全基因组重测序的茄子萼下果色基因预测[J]. 园艺学报, 2022, 49(3): 613-621.
doi: 10.16420/j.issn.0513-353x.2021-0018 |
| Qiao J, Liu J, Li SW, et al. Prediction of fruit color genes under the Calyx of eggplant based on genome-wide resequencing in an extreme mixing pool[J]. Acta Hortic Sin, 2022, 49(3): 613-621. | |
| [13] |
严昕, 项超, 刘荣, 等. 基于BSA-seq技术对豌豆花色基因的精细定位[J]. 作物学报, 2023, 49(4): 1006-1015.
doi: 10.3724/SP.J.1006.2023.24055 |
| Yan X, Xiang C, Liu R, et al. Fine mapping of pea flower color genes based on BSA-seq technology[J]. Acta Agron Sin, 2023, 49(4): 1006-1015. | |
| [14] |
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data[J]. Bioinformatics, 2014, 30(15): 2114-2120.
doi: 10.1093/bioinformatics/btu170 pmid: 24695404 |
| [15] |
Li H, Handsaker B, Wysoker A, et al. The sequence alignment/map format and SAMtools[J]. Bioinformatics, 2009, 25(16): 2078-2079.
doi: 10.1093/bioinformatics/btp352 pmid: 19505943 |
| [16] |
Mckenna A, Hanna M, Banks E, et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data[J]. Genome Res, 2010, 20(9): 1297-1303.
doi: 10.1101/gr.107524.110 pmid: 20644199 |
| [17] |
Wang K, Li MY, Hakonarson H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data[J]. Nucleic Acids Research, 2010, 38(16): e164.
doi: 10.1093/nar/gkq603 URL |
| [18] |
Takagi H, Abe A, Yoshida K, et al. QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations[J]. Plant J, 2013, 74(1): 174-183.
doi: 10.1111/tpj.2013.74.issue-1 URL |
| [19] |
Song B, Stöcklin J, Armbruster WS, et al. Reversible colour change in leaves enhances pollinator attraction and reproductive success in Saururus chinensis(Saururaceae)[J]. Ann bot, 2018, 121(4): 641-650.
doi: 10.1093/aob/mcx195 URL |
| [20] |
Sun JF, Gong YB, Renner SS, et al. Multifunctional bracts in the dove tree Davidia involucrata(Nyssaceae: Cornales): rain protection and pollinator attraction[J]. Am Nat, 2008, 171(1): 119-124.
doi: 10.1086/523953 URL |
| [21] |
Maiwald D, Dietzmann A, Jahns P, et al. Knock-out of the genes coding for the Rieske protein and the ATP-synthase delta-subunit of Arabidopsis. Effects on photosynthesis, thylakoid protein composition, and nuclear chloroplast gene expression[J]. Plant Physiol, 2003, 133(1): 191-202.
doi: 10.1104/pp.103.024190 URL |
| [22] |
Colcombet J, Lopez-Obando M, Heurtevin L, et al. Systematic study of subcellular localization of Arabidopsis PPR proteins confirms a massive targeting to organelles[J]. RNA Biol, 2013, 10(9): 1557-1575.
doi: 10.4161/rna.26128 pmid: 24037373 |
| [23] |
Liu XY, Jiang RC, Wang Y, et al. ZmPPR26, a DYW-type pentatricopeptide repeat protein, is required for C-to-U RNA editing at atpA-1148 in maize chloroplasts[J]. J Exp Bot, 2021, 72(13): 4809-4821.
doi: 10.1093/jxb/erab185 URL |
| [24] |
刘胜坤, 孙世磊, 董鲁朋, 等. 玉米黄叶基因ZmNPPR5的克隆[J]. 植物遗传资源学报, 2023, 24(2): 388-395.
doi: 10.13430/j.cnki.jpgr.20220918002 |
| Liu SK, Sun SL, Dong LP, et al. Cloning of a new yellow leaf gene ZmNPPR5 in maize[J]. J Plant Genet Resour, 2023, 24(2): 388-395. | |
| [25] |
Wagner R, von Sydow L, Aigner H, et al. Deletion of FtsH11 protease has impact on chloroplast structure and function in Arabidopsis thaliana when grown under continuous light[J]. Plant Cell Environ, 2016, 39(11): 2530-2544.
doi: 10.1111/pce.v39.11 URL |
| [26] |
Xia H, Zhou Y, Lin Z, et al. Characterization and functional validation of β-carotene hydroxylase AcBCH genes in Actinidia chinensis[J]. Hortic Res, 2022, 9: uhac063.
doi: 10.1093/hr/uhac063 URL |
| [27] |
Diretto G, Wwlsch R, Tavazza R et al. Silencing of beta-carotene hydroxylase increases total carotenoid and beta-carotene levels in potato tubers[J]. BMC Plant Biol, 2007, 7(1): 11.
doi: 10.1186/1471-2229-7-11 |
| [28] | 宋江华, 张立新. 植物跨膜蛋白研究进展[J]. 生物学杂志, 2009, 26(6): 62-64. |
| Song JH, Zhang LX. Progress on the transmembrane protein in plants[J]. J Biol, 2009, 26(6): 62-64. |
| [1] | 吴元明, 林佳怡, 柳雨汐, 李丹婷, 张宗琼, 郑晓明, 逄洪波. 基于BSA-seq和RNA-seq挖掘水稻株高相关QTL[J]. 生物技术通报, 2023, 39(8): 173-184. |
| [2] | 方澜, 黎妍妍, 江健伟, 成胜, 孙正祥, 周燚. 盘龙参内生真菌胞内细菌7-2H的分离鉴定和促生特性研究[J]. 生物技术通报, 2023, 39(8): 272-282. |
| [3] | 郭少华, 毛会丽, 刘征权, 付美媛, 赵平原, 马文博, 李旭东, 关建义. 一株鱼源致病性嗜水气单胞菌XDMG的全基因组测序及比较基因组分析[J]. 生物技术通报, 2023, 39(8): 291-306. |
| [4] | 张志霞, 李天培, 曾虹, 朱稀贤, 杨天雄, 马斯楠, 黄磊. 冰冷杆菌PG-2的基因组测序及生物信息学分析[J]. 生物技术通报, 2023, 39(3): 290-300. |
| [5] | 和梦颖, 刘文彬, 林震鸣, 黎尔彤, 汪洁, 金小宝. 一株抗革兰阳性菌的戈登氏菌WA4-43全基因组测序与分析[J]. 生物技术通报, 2023, 39(2): 232-242. |
| [6] | 张傲洁, 李青云, 宋文红, 颜少慧, 唐爱星, 刘幽燕. 基于苯酚降解的粪产碱杆菌Alcaligenes faecalis JF101的全基因组分析[J]. 生物技术通报, 2023, 39(10): 292-303. |
| [7] | 王帅, 吕鸿睿, 张昊, 吴占文, 肖翠红, 孙冬梅. 解磷菌PSB-R全基因组测序鉴定及其解磷特性分析[J]. 生物技术通报, 2023, 39(1): 274-283. |
| [8] | 周诗晨, 仪治本, 王馨翊, 杨晓颖, 孙丽娜, 栾维江, 梁闪闪. 高粱双粒突变体Dgs的遗传分析与基因定位[J]. 生物技术通报, 2022, 38(7): 171-177. |
| [9] | 罗雅方, 朱春花, 肖钰婷, 李方全, 张江, 王玉书. 羽衣甘蓝类黄酮糖基转移酶基因的筛选及分析[J]. 生物技术通报, 2022, 38(11): 194-201. |
| [10] | 张泽颖, 范清锋, 邓云峰, 韦廷舟, 周正富, 周建, 王劲, 江世杰. 一株高产脂肪酶菌株WCO-9全基因组测序及比较基因组分析[J]. 生物技术通报, 2022, 38(10): 216-225. |
| [11] | 薛清, 杜虹锐, 薛会英, 王译浩, 王暄, 李红梅. 苜蓿滑刃线虫线粒体基因组及其系统发育研究[J]. 生物技术通报, 2021, 37(7): 98-106. |
| [12] | 陈体强, 徐晓兰, 石林春, 钟礼义. 紫芝栽培品种‘武芝2号’(‘紫芝S2’)全基因组测序及分析[J]. 生物技术通报, 2021, 37(11): 42-56. |
| [13] | 郭鹤宝, 王星, 何山文, 张晓霞. 表型特征结合基因组分析鉴定不同菌落形态Bacillus velezensis ACCC 19742[J]. 生物技术通报, 2020, 36(2): 142-148. |
| [14] | 韩立杰, 才宏伟. 高粱粒重遗传研究进展[J]. 生物技术通报, 2019, 35(5): 15-27. |
| [15] | 李晓凯 ,王贵 ,乔贤 ,范一星 ,张磊 ,马宇浩 ,聂瑞雪 ,王瑞军 ,何利兵 ,苏蕊. 全基因组测序在重要家畜上的研究进展[J]. 生物技术通报, 2018, 34(6): 11-21. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||