








生物技术通报 ›› 2024, Vol. 40 ›› Issue (4): 97-109.doi: 10.13560/j.cnki.biotech.bull.1985.2023-1025
李兴容1,2( ), 谭志兵3, 赵燕3, 李曜魁2, 赵炳然1,2(
), 谭志兵3, 赵燕3, 李曜魁2, 赵炳然1,2( ), 唐丽1,2(
), 唐丽1,2( )
)
收稿日期:2023-11-02
出版日期:2024-04-26
发布日期:2024-04-30
通讯作者:
赵炳然,男,博士,研究员,研究方向:植物遗传学与品种培育;E-mail: brzhao652@hhrrc.ac.cn;作者简介:李兴容,女,硕士研究生,研究方向:水稻重金属积累与耐受;E-mail: L3141429315@163.com
基金资助:
LI Xing-rong1,2( ), TAN Zhi-bing3, ZHAO Yan3, LI Yao-kui2, ZHAO Bing-ran1,2(
), TAN Zhi-bing3, ZHAO Yan3, LI Yao-kui2, ZHAO Bing-ran1,2( ), TANG Li1,2(
), TANG Li1,2( )
)
Received:2023-11-02
Published:2024-04-26
Online:2024-04-30
摘要:
【目的】部分稻米镉超标严重影响我国粮食质量安全,本研究旨在鉴定对稻米镉积累有调控作用的基因,为阻控稻米镉积累提供新的基因资源。【方法】通过逆转录PCR和RACE技术,克隆了水稻低亲和性阳离子转运蛋白基因家族的一个新成员OsLCT3。通过生物信息学方法对OsLCT3的自然变异进行分析,对OsLCT3蛋白的理化性质进行预测;利用实时荧光定量 PCR分析其全生育期表达模式及对镉、锰、铁胁迫的响应;采用融合报告基因定位法探究OsLCT3的亚细胞定位。通过苗期镉胁迫水培以及镉污染土壤的成熟期植株各部位的镉及二价矿质金属元素测定,分析敲除OsLCT3对水稻二价阳离子运输的影响。此外,通过酵母细胞的异源功能互补验证OsLCT3 对酵母镉耐受性的影响。【结果】OsLCT3仅存在于部分水稻品种中,编码区全长1 263 bp,根据编码区变异其氨基酸序列可划分为5个单倍型,编码的蛋白具有12个跨膜结构域,与小麦、节节麦的LCT亲缘关系较近,与水稻OsLCT2的同源性仅52%,与籼稻、粳稻两个亚种OsLCT1的同源性分别为49%、47%。OsLCT3在生长发育的各时期均在根部高表达,在扬花期的表达最高,且在根部的表达受镉胁迫及过量铁、锰的抑制,蛋白定位于质膜。与野生型相比,oslct3敲除系苗期株高降低,地上部的镉、铁、锌含量降低,根部铁含量升高,其他元素含量不变;根部镉、铁、锌向地上部的转运率降低。大田条件下,oslct3敲除系成熟期的茎叶和糙米镉含量较野生型均显著下降,稻草和糙米锰、铜、铁、锌含量较对照无显著差异。表达OsLCT3导致酵母对镉胁迫更敏感。【结论】OsLCT3参与根部的镉、铁、锌向地上部的运输,正调控稻米镉积累。
李兴容, 谭志兵, 赵燕, 李曜魁, 赵炳然, 唐丽. 水稻低亲和性阳离子转运蛋白基因OsLCT3的克隆与功能研究[J]. 生物技术通报, 2024, 40(4): 97-109.
LI Xing-rong, TAN Zhi-bing, ZHAO Yan, LI Yao-kui, ZHAO Bing-ran, TANG Li. Cloning and Functional Analysis of OsLCT3, a Low-affinity Cation Transporter Gene of Rice[J]. Biotechnology Bulletin, 2024, 40(4): 97-109.
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') | 引物用途 Primer usage |
|---|---|---|
| T3-CDS-F | TATCTTATGGACTGGTCTGTAGC | OsLCT3 CDS扩增、测序 |
| T3-CDS-R | TCTGTTGTCTGATGATGATTTTG | OsLCT3 CDS amplification and sequencing |
| T3-C-F | ATGGTGGCCCTTTCTATCCA | OsLCT3 RACE核心序列扩增 |
| T3-C-R | TCACTGTATCTCCAAAACCA | OsLCT3 RACE core sequence amplification |
| T3-3-F2 | TTGCCTCTGCAATAGTTGCC | OsLCT3 3' RACE 扩增 |
| T3-3-F3 | GTACTGTAATGGTTACACTTAG | OsLCT3 3' RACE amplification |
| T3-5-R2 | GAGGAAATACCAAGCCACCATC | OsLCT3 5' RACE 扩增 |
| T3-5-R3 | CGAAGTGATACATCCCTCAACCTG | OsLCT3 5' RACE amplification |
| OsLCT3-F2 | ATGGTGGCTTGGTATTTCCTC | OsLCT3 DNA片段扩增 |
| OsLCT3-R2 | CACCTCATTGGCATTACTGTCC | Amplification of OsLCT3 DNA fragments |
| OsLCT3-F3 | GAGATCCGTGCGAAAGTGCT | OsLCT3 DNA片段扩增 Amplification of OsLCT3 DNA fragments |
| OsLCT3-R3 | AGAGTGAGTGAAAAGCCCGA | |
| OsActin1-F | TGCTATGTACGTCGCCATCCAG | OsActin1 DNA片段扩增 |
| OsActin1-R | AATGAGTAACCACGCTCCGTCA | Amplification of OsActin1 DNA fragments |
| T3-qP-F | GGTGGTTCTGGGAAGACTAAA | OsLCT3 RT-qPCR分析 |
| T3-qP-R | ATGGAGAGCAGACCTGATATTG | OsLCT3 RT-qPCR analysis |
| OsActin1-qP-F | CAACACCCCTGCTATGTACG | OsActin1 RT-qPCR分析 |
| OsActin1-qP-R | CATCACCAGAGTCCAACACAA | OsActin1 RT-qPCR analysis |
| T3P1-F | GGCAGTACATTGGAAGCTTGTCTG | OsLCT3 敲除载体靶位点1 接头引物 |
| T3P1-R | AAACCAGACAAGCTTCCAATGTAC | OsLCT3 knockout vector target site 1 adaptor primer |
| T3P2-F | GCCGTCAATTACAACGCCACCGG | OsLCT3 敲除载体靶位点2 接头引物 |
| T3P2-R | AAACCCGGTGGCGTTGTAATTGA | OsLCT3 knockout vector target site 2 adapter primers |
| T3-p1132-F | cgcggtggcggccgctctagaATGGTGGCCCTTTCTATCCAT | OsLCT3亚细胞定位 |
| T3-p1132-R | gataagcttgatatcgaattcCTGTATCTCCAAAACCACCGCG | OsLCT3 subcellular localization |
| pYES2-T3-F | gggaatattaagcttggtaccATGGTGGCCCTTTCTATCCAT | OsLCT3酵母载体构建 |
| YES2-T3-R | gatggatatctgcagaattcTCACTGTATCTCCAAAACCACCG | OsLCT3 yeast vector construction |
| HPT-F | GCTCCATACAAGCCAACCACG | Hpt片段扩增 |
| HPT-R | CCTGCCTGAAACCGAACTGC | Hpt fragment amplification |
| CAS9-F | CGAGACGAACGGTGAGACTGGTG | Cas9片段扩增 |
| CAS9-R | GGTGCTTGTTGTAGGCGGAGAGG | Cas9 fragment amplification |
| OsLCT3-CAS-F | CTGCGGTATATCATCCCAATGCT | OsLCT3的靶点PCR扩增 |
| OsLCT3-CAS-R | CAACTATGGTGAAAACTTCGGCG | PCR amplification of OsLCT3 targets |
| pYES2-T3-eGFP-F | gggaatattaagcttggtaccATGGTGGCCCTTTCTATCCAT | OsLCT3-eGFP酵母载体构建 |
| pYES2-T3-eGFP-R | tgatggatatctgcagaattcTTACTTGTACAGCTCGTCCATGCC | OsLCT3-eGFP yeast vector construction |
表1 本研究所用的引物及序列
Table 1 Primers and sequences used in this study
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') | 引物用途 Primer usage |
|---|---|---|
| T3-CDS-F | TATCTTATGGACTGGTCTGTAGC | OsLCT3 CDS扩增、测序 |
| T3-CDS-R | TCTGTTGTCTGATGATGATTTTG | OsLCT3 CDS amplification and sequencing |
| T3-C-F | ATGGTGGCCCTTTCTATCCA | OsLCT3 RACE核心序列扩增 |
| T3-C-R | TCACTGTATCTCCAAAACCA | OsLCT3 RACE core sequence amplification |
| T3-3-F2 | TTGCCTCTGCAATAGTTGCC | OsLCT3 3' RACE 扩增 |
| T3-3-F3 | GTACTGTAATGGTTACACTTAG | OsLCT3 3' RACE amplification |
| T3-5-R2 | GAGGAAATACCAAGCCACCATC | OsLCT3 5' RACE 扩增 |
| T3-5-R3 | CGAAGTGATACATCCCTCAACCTG | OsLCT3 5' RACE amplification |
| OsLCT3-F2 | ATGGTGGCTTGGTATTTCCTC | OsLCT3 DNA片段扩增 |
| OsLCT3-R2 | CACCTCATTGGCATTACTGTCC | Amplification of OsLCT3 DNA fragments |
| OsLCT3-F3 | GAGATCCGTGCGAAAGTGCT | OsLCT3 DNA片段扩增 Amplification of OsLCT3 DNA fragments |
| OsLCT3-R3 | AGAGTGAGTGAAAAGCCCGA | |
| OsActin1-F | TGCTATGTACGTCGCCATCCAG | OsActin1 DNA片段扩增 |
| OsActin1-R | AATGAGTAACCACGCTCCGTCA | Amplification of OsActin1 DNA fragments |
| T3-qP-F | GGTGGTTCTGGGAAGACTAAA | OsLCT3 RT-qPCR分析 |
| T3-qP-R | ATGGAGAGCAGACCTGATATTG | OsLCT3 RT-qPCR analysis |
| OsActin1-qP-F | CAACACCCCTGCTATGTACG | OsActin1 RT-qPCR分析 |
| OsActin1-qP-R | CATCACCAGAGTCCAACACAA | OsActin1 RT-qPCR analysis |
| T3P1-F | GGCAGTACATTGGAAGCTTGTCTG | OsLCT3 敲除载体靶位点1 接头引物 |
| T3P1-R | AAACCAGACAAGCTTCCAATGTAC | OsLCT3 knockout vector target site 1 adaptor primer |
| T3P2-F | GCCGTCAATTACAACGCCACCGG | OsLCT3 敲除载体靶位点2 接头引物 |
| T3P2-R | AAACCCGGTGGCGTTGTAATTGA | OsLCT3 knockout vector target site 2 adapter primers |
| T3-p1132-F | cgcggtggcggccgctctagaATGGTGGCCCTTTCTATCCAT | OsLCT3亚细胞定位 |
| T3-p1132-R | gataagcttgatatcgaattcCTGTATCTCCAAAACCACCGCG | OsLCT3 subcellular localization |
| pYES2-T3-F | gggaatattaagcttggtaccATGGTGGCCCTTTCTATCCAT | OsLCT3酵母载体构建 |
| YES2-T3-R | gatggatatctgcagaattcTCACTGTATCTCCAAAACCACCG | OsLCT3 yeast vector construction |
| HPT-F | GCTCCATACAAGCCAACCACG | Hpt片段扩增 |
| HPT-R | CCTGCCTGAAACCGAACTGC | Hpt fragment amplification |
| CAS9-F | CGAGACGAACGGTGAGACTGGTG | Cas9片段扩增 |
| CAS9-R | GGTGCTTGTTGTAGGCGGAGAGG | Cas9 fragment amplification |
| OsLCT3-CAS-F | CTGCGGTATATCATCCCAATGCT | OsLCT3的靶点PCR扩增 |
| OsLCT3-CAS-R | CAACTATGGTGAAAACTTCGGCG | PCR amplification of OsLCT3 targets |
| pYES2-T3-eGFP-F | gggaatattaagcttggtaccATGGTGGCCCTTTCTATCCAT | OsLCT3-eGFP酵母载体构建 |
| pYES2-T3-eGFP-R | tgatggatatctgcagaattcTTACTTGTACAGCTCGTCCATGCC | OsLCT3-eGFP yeast vector construction |
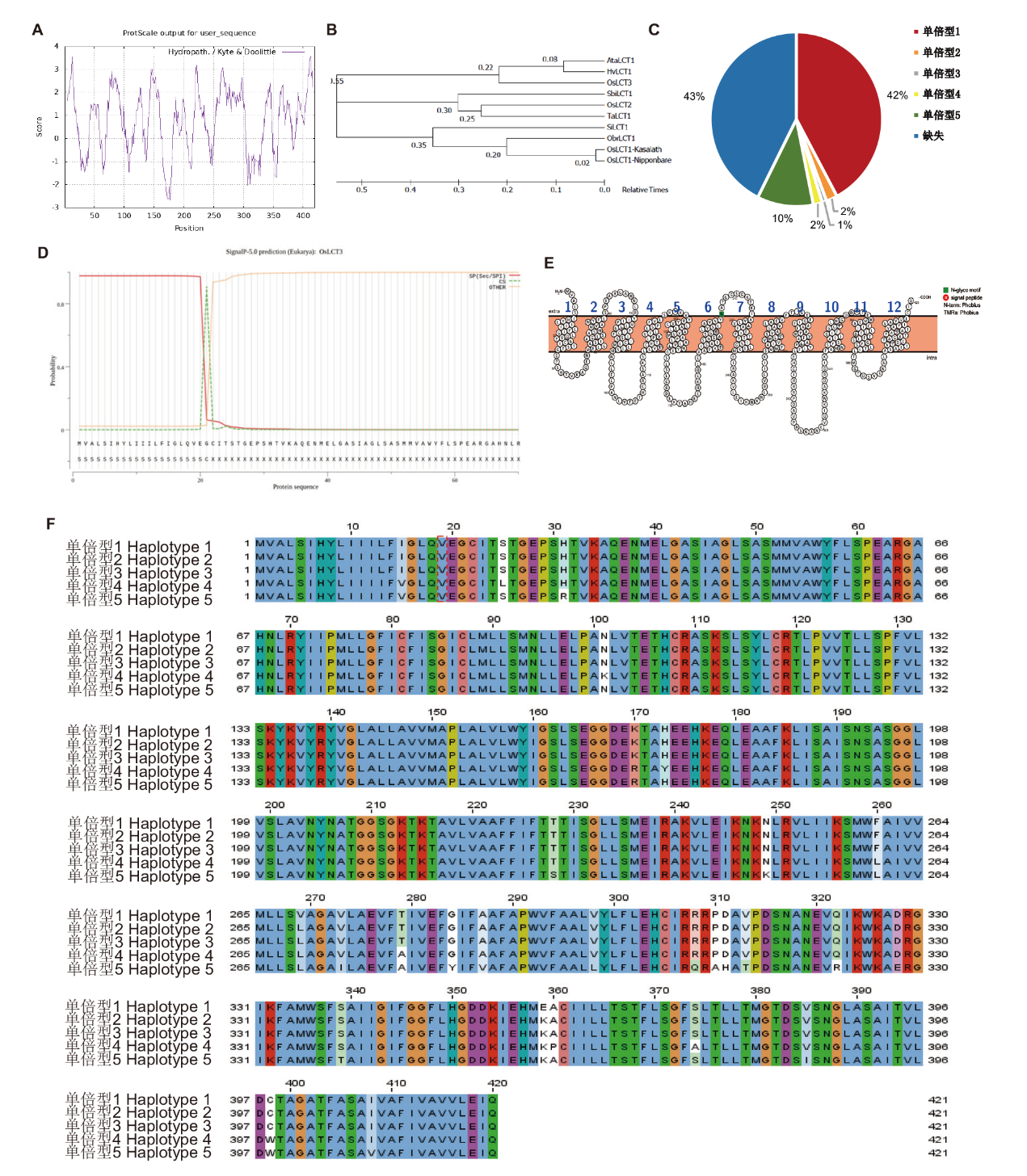
图2 OsLCT3 的生物信息学分析 A:OsLCT3蛋白的疏水性分析;B:LCT家族成员的系统进化分析;C:OsLCT3单倍型的分布频率;D:OsLCT3蛋白的信号肽预测;E:OsLCT3跨膜结构域分析;F:OsLCT3单倍型分析;Ta: 小麦;Sbi: 高粱;Os: 水稻;Si: 小米;Ata: 节节麦;Hv: 大麦;Obr: 短花野生稻
Fig. 2 Bioinformatics analysis for OsLCT3 A: Hydrophobicity of OsLCT3 protein. B: Phylogenetic analysis of LCT family members. C: Distribution frequency of the OsLCT3 haplotype. D: Signaling peptide prediction of OsLCT3 protein. E: OsLCT3 transmembrane domain analysis. F: OsLCT3 haplotype analysis. Ta: Triticum aestivum; Sbi: Sorghum bicolor; Os: Oryza sativa; Si: Setaria italica; Ata: Aegilops tauschii; Hv: Hordeum vulgare; Obr: Oryza brachyantha
| 蛋白质 Protein | 氨基酸数 Amino acid No./aa | 分子式 Formula | 分子量 Molecular weight/kD | 等电点 pI | 不稳定指数 Instability index | 平均疏水性 Hydropathy index | 正 / 负电残基数 Acid-base amino acid | 脂溶指数 Aliphatic index |
|---|---|---|---|---|---|---|---|---|
| OsLCT3 | 420 | C2088H3315N511O567S22 | 45.355 | 6.51 | 25.72 | 0.777 | 28/30 | 121.95 |
表2 OsLCT3 蛋白的理化性质
Table 2 Physicochemical properties of OsLCT3 protein
| 蛋白质 Protein | 氨基酸数 Amino acid No./aa | 分子式 Formula | 分子量 Molecular weight/kD | 等电点 pI | 不稳定指数 Instability index | 平均疏水性 Hydropathy index | 正 / 负电残基数 Acid-base amino acid | 脂溶指数 Aliphatic index |
|---|---|---|---|---|---|---|---|---|
| OsLCT3 | 420 | C2088H3315N511O567S22 | 45.355 | 6.51 | 25.72 | 0.777 | 28/30 | 121.95 |
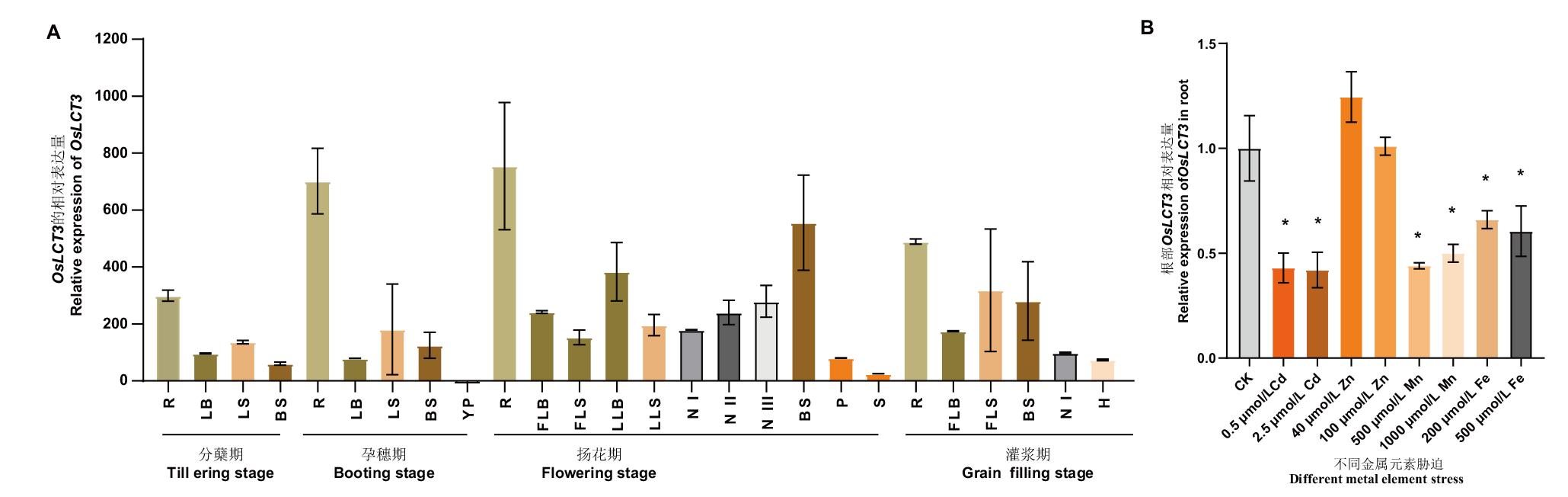
图3 OsLCT3的表达模式 A:OsLCT3在不同生育期的表达模式分析;孕穗期幼穗的 OsLCT3表达水平被设置为 1,采用 2-△△Ct 法估算各个时期各个组织部位中 OsLCT3的相对表达水平(R:根;LB:叶片;LS:叶鞘;BS:茎基部;YP:幼穗;FLB:旗叶叶片;FLS:旗叶叶鞘;LLB:低位叶片;LLS:低位叶鞘;N I:倒一茎节;N II:倒二茎节;N III:倒三茎节;P:穗梗;S:小穗;H:颖壳);B: OsLCT3对不同金属元素胁迫7 d的响应。标准差基于3个生物学重复。数据分析采用单因素方差分析,t- test 检验,*表示相同条件下与对照相比,在0.05水平上差异显著
Fig. 3 Expression patterns of OsLCT3 A: Expression analysis of OsLCT3 at different growth stages. The expression level of OsLCT3 in young panicle of booting stage was set to 1, and the 2-△△ Ct method was used to estimate the relative expression of OsLCT3 in each tissue at each stage(R: Root. LB: Leaf blade. LS: Leaf sheath. BS: Basal stem. YP: Young panicle. FLB: Flag leaf blade. FLS: Flag leaf sheath. LLB: Lower leaf blade. LLS: Lower leaf sheath. N I: Node I. N II: Node II. N III: Node III. P: Peduncle. S: Spikelet. H: Husk). B: Response of OsLCT3 to stress of different metal elements for 7 d. Standard deviation is based on three biological replicates. One-way ANOVA was used for data analysis,* indicating a significant difference at the level of 0.05 compared with the control by student’s t- test

图4 OsLCT3的亚细胞定位 eGFP:增强绿色荧光蛋白;mCherry:红色荧光蛋白;Bright field:明场;Merge:荧光整合;比例尺=10 μm
Fig. 4 Subcellular localization of OsLCT3 eGFP: Enhanced green fluorescence. mCherry: Red fluorescent protein. Bright field: Bright field. Merge: Fluorescence integration. Scale bar=10 μm
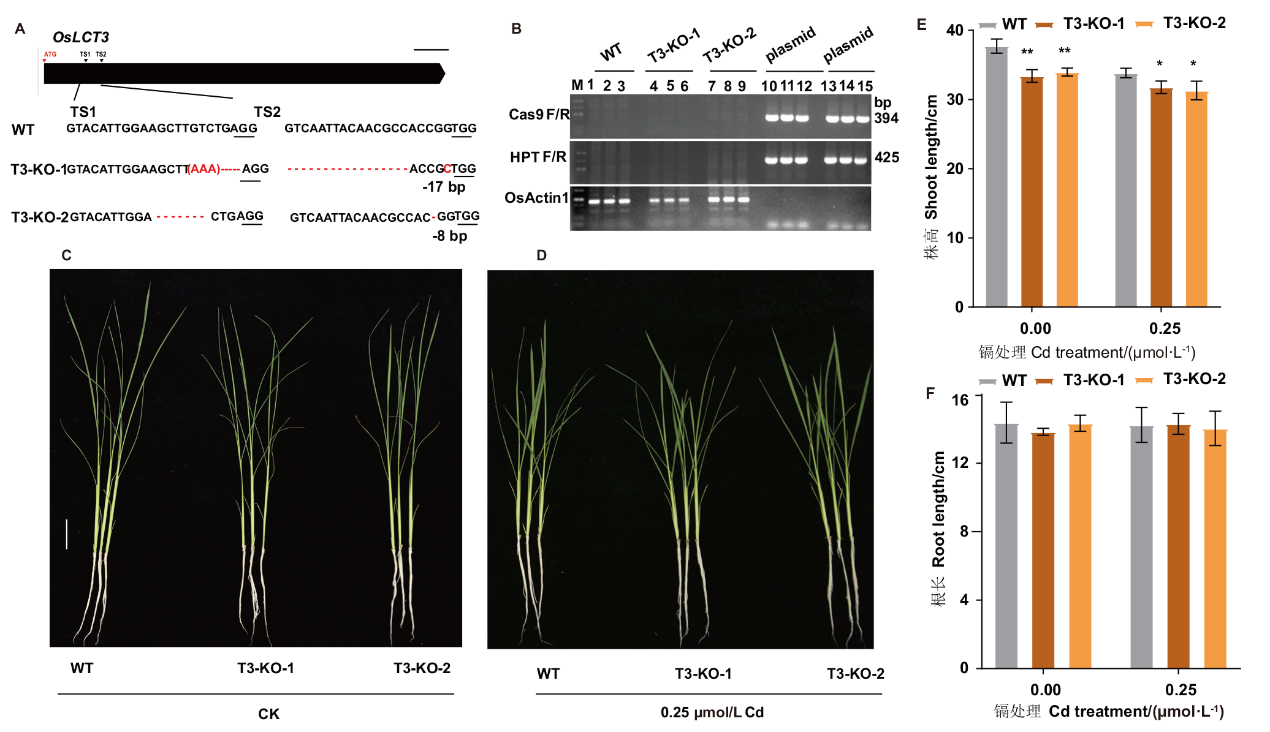
图5 oslct3和WT水培条件下的生长情况 A-B:oslct3纯合突变体的鉴定;C-D:WT和敲除系的表型;E:株高;F:根长;*表示相同条件下与野生型相比,0.05水平上差异显著,**表示相同条件与下野生型相比,0.01水平上差异显著(t-test 检验),下同。比例尺=5 cm
Fig. 5 Growth of oslct3 and WT in hydroponic growth A-B: Identification of oslct3 homozygous mutants. C-D: Phenotypes of WT and knockout lines. E: Plant height. F: Root length. * or ** indicate statistically significant difference in comparison to WT at P< 0.05 or P< 0.01 by student’s t test, the same applies hereinafter. Scale bar=5 cm
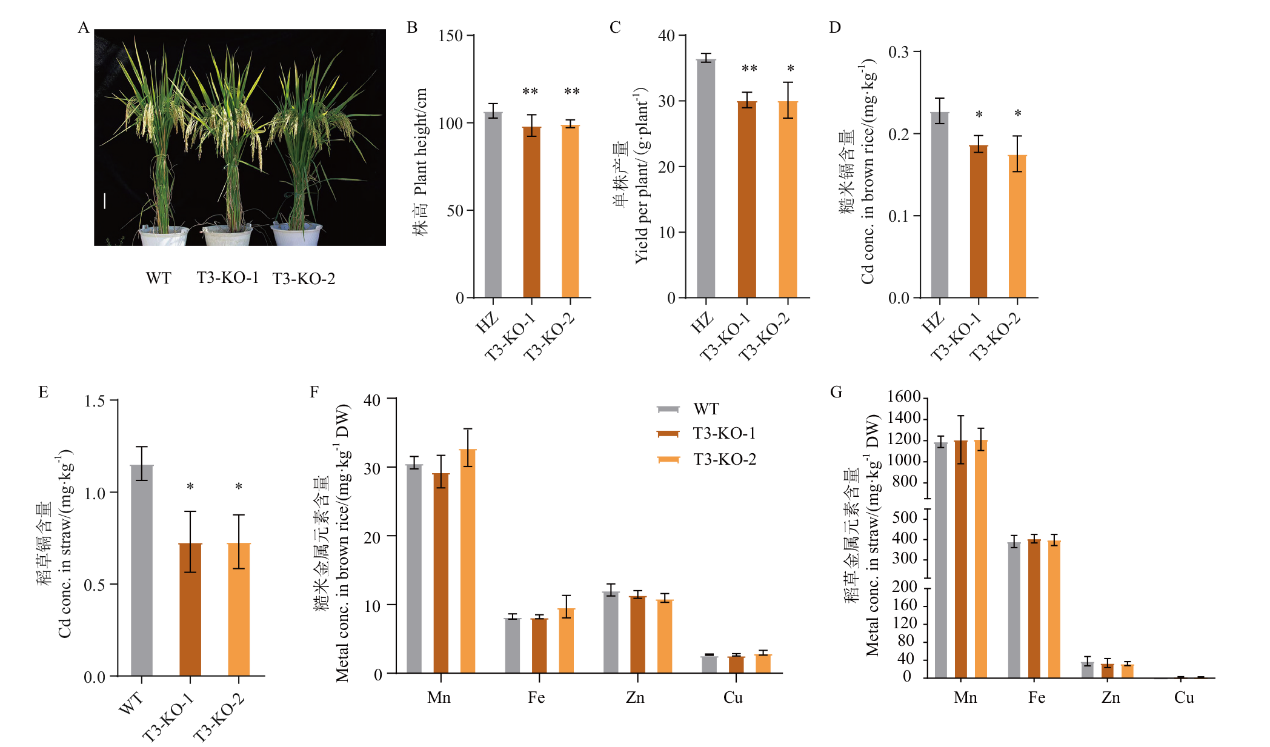
图7 Cd污染大田条件下oslct3 产量、稻草和糙米的金属含量分析 A:成熟期植株表型;B:株高;C: 单株产量;D-E:糙米、稻草Cd含量;F-G:糙米、稻草金属元素含量。比例尺=10 cm
Fig. 7 Grain yield of oslct3 and metal contents in straw and brown rice under Cd-contaminated fields A: Plant phenotype at maturity. B: Plant height. C: Yield per plant. D-E: Cd content in brown rice and straw. F-G: Brown rice and straw metal element content. Scale bar=10 cm

图8 OsLCT3 对酵母 Cd 耐受性的影响鉴定 A:OsLCT3 在酵母中Cd耐受性表型鉴定;B:OsLCT3在酵母中的耐Cd性生长曲线分析
Fig. 8 Identification of effects of OsLCT3 on the tolerance of yeast to Cd stress A: OsLCT3 Cd tolerance phenotypic identification in yeast. B: Cd tolerance growth curve analysis of OsLCT3 in yeast
| [1] |
Jamers A, Blust R, De Coen W, et al. An omics based assessment of cadmium toxicity in the green alga Chlamydomonas reinhardtii[J]. Aquat Toxicol, 2013, 126: 355-364.
doi: 10.1016/j.aquatox.2012.09.007 pmid: 23063003 |
| [2] | 汪鹏, 王静, 陈宏坪, 等. 我国稻田系统镉污染风险与阻控[J]. 农业环境科学学报, 2018, 37(7): 1409-1417. |
| Wang P, Wang J, Chen HP, et al. Cadmium risk and mitigation in paddy systems in China[J]. J Agro Environ Sci, 2018, 37(7): 1409-1417. | |
| [3] |
Clemens S, Ma JF. Toxic heavy metal and metalloid accumulation in crop plants and foods[J]. Annu Rev Plant Biol, 2016, 67: 489-512.
doi: 10.1146/annurev-arplant-043015-112301 pmid: 27128467 |
| [4] | Tang L, Li Y, Peng Y, et al. Breeding for rice cultivars with low cadmium accumulation[M]//Heavy Metal Toxicity and Tolerance in Plants: A Biological, Omics, and Genetic Engineering Approach. Chichester: John Wiley & Sons Ltd, 2023: 335-347. |
| [5] |
冀中英, 李曜魁, 孟前程, 等. 水稻积累及耐受镉和砷的分子机制与育种实践[J]. 植物遗传资源学报, 2023, 24(1): 75-85.
doi: 10.13430/j.cnki.jpgr.20220504002 |
| Ji ZY, Li YK, Meng QC, et al. Molecular mechanisms and breeding practices of accumulation and tolerance to cadmium and arsenic in rice[J]. J Plant Genet Resour, 2023, 24(1): 75-85. | |
| [6] |
Uraguchi S, Kamiya T, Clemens S, et al. Characterization of OsLCT1, a cadmium transporter from indica rice(Oryza sativa)[J]. Physiol Plant, 2014, 151(3): 339-347.
doi: 10.1111/ppl.12189 pmid: 24627964 |
| [7] |
Uraguchi S, Kamiya T, Sakamoto T, et al. Low-affinity cation transporter(OsLCT1)regulates cadmium transport into rice grains[J]. Proc Natl Acad Sci USA, 2011, 108(52): 20959-20964.
doi: 10.1073/pnas.1116531109 pmid: 22160725 |
| [8] | Kumar A. 水稻籽粒锌含量的全基因组关联分析及候选基因验证[D]. 北京: 中国农业科学院, 2021. |
| Kumar A. Genome-wide association analysis and validation of candidate genes[D]. Beijing: Chinese Academy of Agricultural Sciences, 2021. | |
| [9] | 李曜魁, 唐丽, 毛毕刚, 等. 籼稻低亲和阳离子转运蛋白基因OsLCT2的克隆与生物信息学分析[J]. 分子植物育种, 2016, 14(5): 1067-1074. |
| Li YK, Tang L, Mao BG, et al. Cloning and bioinformatics analysis of low-affinity cation transporter gene OsLCT2 in indica rice(Oryza sativa)[J]. Mol Plant Breed, 2016, 14(5): 1067-1074. | |
| [10] |
Tang L, Dong JY, Tan LT, et al. Overexpression of OsLCT2, a low-affinity cation transporter gene, reduces cadmium accumulation in shoots and grains of rice[J]. Rice, 2021, 14: 89.
doi: 10.1186/s12284-021-00530-8 pmid: 34693475 |
| [11] |
Schachtman DP, Kumar R, Schroeder JI, et al. Molecular and functional characterization of a novel low-affinity cation transporter(LCT1)in higher plants[J]. Proc Natl Acad Sci USA, 1997, 94(20): 11079-11084.
pmid: 9380762 |
| [12] |
Clemens S, Antosiewicz DM, Ward JM, et al. The plant cDNA LCT1 mediates the uptake of calcium and cadmium in yeast[J]. Proc Natl Acad Sci USA, 1998, 95(20): 12043-12048.
doi: 10.1073/pnas.95.20.12043 pmid: 9751787 |
| [13] |
Amtmann A, Fischer M, Marsh EL, et al. The wheat cDNA LCT1 generates hypersensitivity to sodium in a salt-sensitive yeast strain[J]. Plant Physiol, 2001, 126(3): 1061-1071.
pmid: 11457957 |
| [14] |
Antosiewicz DM, Hennig J. Overexpression of LCT1 in tobacco enhances the protective action of calcium against cadmium toxicity[J]. Environ Pollut, 2004, 129(2): 237-245.
doi: 10.1016/j.envpol.2003.10.025 pmid: 14987809 |
| [15] |
Wojas S, Ruszczyńska A, Bulska E, et al. Ca2+-dependent plant response to Pb2+ is regulated by LCT1[J]. Environ Pollut, 2007, 147(3): 584-592.
doi: 10.1016/j.envpol.2006.10.012 pmid: 17140712 |
| [16] |
Xia RZ, Zhou J, Cui HB, et al. Nodes play a major role in cadmium(Cd)storage and redistribution in low-Cd-accumulating rice(Oryza sativa L.) cultivars[J]. Sci Total Environ, 2023, 859: 160436.
doi: 10.1016/j.scitotenv.2022.160436 URL |
| [17] | 邱冠凯, 倪大虎, 韩忠民, 等. 水稻铁吸收与转运机理[J]. 土壤与作物, 2022, 11(2): 170-178. |
| Qiu GK, Ni DH, Han ZM, et al. Mechanisms of iron absorption and transport in rice: a review[J]. Soils Crops, 2022, 11(2): 170-178. | |
| [18] |
Tan LT, Qu MM, Zhu YX, et al. ZINC TRANSPORTER5 and ZINC TRANSPORTER9 function synergistically in zinc/cadmium uptake[J]. Plant Physiol, 2020, 183(3): 1235-1249.
doi: 10.1104/pp.19.01569 pmid: 32341004 |
| [19] |
薛欣月, 于雪然, 刘晓刚, 等. 水稻锌吸收、转运、累积机理研究进展[J]. 生物技术通报, 2022, 38(4): 29-43.
doi: 10.13560/j.cnki.biotech.bull.1985.2020-1484 |
| Xue XY, Yu XR, Liu XG, et al. Research progress in absorption, transportation and accumulation mechanism of zinc in rice[J]. Biotechnol Bull, 2022, 38(4): 29-43. | |
| [20] |
王璐瑶, 陈謇, 赵守清, 等. 水稻镉积累特性的生理和分子机制研究概述[J]. 植物学报, 2022, 57(2): 236-249.
doi: 10.11983/CBB21222 |
| Wang LY, Chen J, Zhao SQ, et al. Research progress of the physiological and molecular mechanisms of cadmium accumulation in rice[J]. Chin Bull Bot, 2022, 57(2): 236-249. | |
| [21] |
Tan LT, Zhu YX, Fan T, et al. OsZIP7 functions in xylem loading in roots and inter-vascular transfer in nodes to deliver Zn/Cd to grain in rice[J]. Biochem Biophys Res Commun, 2019, 512(1): 112-118.
doi: 10.1016/j.bbrc.2019.03.024 URL |
| [22] |
Takahashi R, Ishimaru Y, Shimo H, et al. The OsHMA2 transporter is involved in root-to-shoot translocation of Zn and Cd in rice[J]. Plant Cell Environ, 2012, 35(11): 1948-1957.
doi: 10.1111/pce.2012.35.issue-11 URL |
| [23] |
Hao XH, Zeng M, Wang J, et al. A node-expressed transporter OsCCX2 is involved in grain cadmium accumulation of rice[J]. Front Plant Sci, 2018, 9: 476.
doi: 10.3389/fpls.2018.00476 pmid: 29696032 |
| [24] | Luo JS, Huang J, Zeng DL, et al. A defensin-like protein drives cadmium efflux and allocation in rice[J]. Nat Commun, 2018, 9(1): 645. |
| [1] | 杨淇, 魏子迪, 宋娟, 童堃, 杨柳, 王佳涵, 刘海燕, 栾维江, 马轩. 水稻组蛋白H1三突变体的创建和转录组学分析[J]. 生物技术通报, 2024, 40(4): 85-96. |
| [2] | 李雪, 李容欧, 孔美懿, 黄磊. 解淀粉芽孢杆菌SQ-2对水稻的促生作用[J]. 生物技术通报, 2024, 40(2): 109-119. |
| [3] | 赵曜, 文朗, 骆少丹, 栗子杏, 刘超超. 番茄HMA基因家族的鉴定及SlHMA1镉转运功能研究[J]. 生物技术通报, 2024, 40(2): 212-222. |
| [4] | 邹修为, 岳佳妮, 李志宇, 戴良英, 李魏. 水稻热激转录因子HsfA2b调控非生物胁迫抗性的功能分析[J]. 生物技术通报, 2024, 40(2): 90-98. |
| [5] | 张超, 王子瑞, 孙亚丽, 毛馨晨, 唐家琪, 于恒秀. 水稻维生素B1合成相关基因OsTHIC的功能研究[J]. 生物技术通报, 2024, 40(2): 99-108. |
| [6] | 林鑫焱, 张传忠, 戴兵, 王馨珩, 刘剑锋, 温丽, 徐兴健, 方军. 水稻穗发芽遗传与分子机制的研究进展[J]. 生物技术通报, 2024, 40(1): 24-31. |
| [7] | 王子颖, 龙晨洁, 范兆宇, 张蕾. 利用酵母双杂交系统筛选水稻中与OsCRK5互作蛋白[J]. 生物技术通报, 2023, 39(9): 117-125. |
| [8] | 黄小龙, 孙贵连, 马丹丹, 闫慧清. 水稻幼苗酵母单杂文库构建及LAZY1上游调控因子筛选[J]. 生物技术通报, 2023, 39(9): 126-135. |
| [9] | 李雪琪, 张素杰, 于曼, 黄金光, 周焕斌. 基于CRISPR/CasX介导的水稻基因组编辑技术的建立[J]. 生物技术通报, 2023, 39(9): 40-48. |
| [10] | 吴元明, 林佳怡, 柳雨汐, 李丹婷, 张宗琼, 郑晓明, 逄洪波. 基于BSA-seq和RNA-seq挖掘水稻株高相关QTL[J]. 生物技术通报, 2023, 39(8): 173-184. |
| [11] | 姚莎莎, 王晶晶, 王俊杰, 梁卫红. 植物激素信号通路调控水稻粒型的分子机制[J]. 生物技术通报, 2023, 39(8): 80-90. |
| [12] | 李宇, 李素贞, 陈茹梅, 卢海强. 植物bHLH转录因子调控铁稳态的研究进展[J]. 生物技术通报, 2023, 39(7): 26-36. |
| [13] | 任沛东, 彭健玲, 刘圣航, 姚姿婷, 朱桂宁, 陆光涛, 李瑞芳. 沙福芽孢杆菌GX-H6的分离鉴定及对水稻细菌性条斑病的防病效果[J]. 生物技术通报, 2023, 39(5): 243-253. |
| [14] | 李怡君, 吴晨晨, 李睿, 王喆, 何山文, 韦善君, 张晓霞. 水稻内生细菌新资源分离培养方案探究[J]. 生物技术通报, 2023, 39(4): 201-211. |
| [15] | 卢振万, 李雪琪, 黄金光, 周焕斌. 利用胞嘧啶碱基编辑技术创制耐草甘膦水稻[J]. 生物技术通报, 2023, 39(2): 63-69. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||