








生物技术通报 ›› 2025, Vol. 41 ›› Issue (8): 1-10.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0300
• 综述与专论 • 下一篇
收稿日期:2025-03-20
出版日期:2025-08-26
发布日期:2025-08-14
通讯作者:
李佳楠,女,博士,教授,研究方向 :食品生物技术;E-mail: lydian_l@163.com作者简介:蔡如凤,女,硕士研究生,研究方向 :蛋白质工程、生物食品技术;E-mail: 2363763510@qq.com
基金资助:
CAI Ru-feng( ), YANG Yu-xuan, YU Ji-zheng, LI Jia-nan(
), YANG Yu-xuan, YU Ji-zheng, LI Jia-nan( )
)
Received:2025-03-20
Published:2025-08-26
Online:2025-08-14
摘要:
蛋白质功能与其三维结构间存在着密不可分的关联,这一认知长期引领着生命科学领域的探索方向。科学家们为解析蛋白质结构投入了大量精力,而蛋白质测序技术的迅猛发展,使得序列数据呈指数级增长,与结构研究进展之间的差距日益显著。过去十几年间,人工智能技术的蓬勃发展为这一困境带来了转机,其以深度学习、神经网络等核心算法为支撑,推动蛋白质工程迎来了全新变革。借助人工智能技术,新一代蛋白质结构预测和设计方法取得重大突破。这些基于先进算法的工具,极大地提高了蛋白质结构建模的准确性和速度。它们不仅助力结构生物学、药物研发等领域的发展,还为蛋白质合成提供了关键依据。除此之外,人工智能正推动蛋白质研究从“结构解析”向“逆向设计”转型。通过构建序列-结构-功能的多维度关联模型,研究人员能够基于特定功能需求,反向设计具有预期结构的蛋白质序列。从而更精准地设计蛋白质序列,为生物合成开辟新路径。本综述聚焦于人工智能在蛋白质工程中的核心作用,阐述了蛋白质工程目前所面临的挑战和传统蛋白结构解析方法所面临的瓶颈,并以此引入介绍了基于人工智能的结构预测工具的发展,分析其在蛋白质合成中的应用;探讨人工智能驱动下,从结构解析到合成蛋白的算法革命及未来潜在方向,以期为该领域的研究提供参考。
蔡如凤, 杨宇轩, 于基正, 李佳楠. 人工智能重塑蛋白质工程:从结构解析到合成生物学的算法革命[J]. 生物技术通报, 2025, 41(8): 1-10.
CAI Ru-feng, YANG Yu-xuan, YU Ji-zheng, LI Jia-nan. Artificial Intelligence Transforms Protein Engineering: From Structural Analysis to Synthetic Biology through Algorithmic Advancements[J]. Biotechnology Bulletin, 2025, 41(8): 1-10.
模型 Model | 发布时间 Release time | CASP参赛版本 CASP entry version | GDT_TS中位数 GDT_TS med-number | 覆盖UniProt比例 Coverage ratio of UniProt | 关键技术革新 Key technological innovation |
|---|---|---|---|---|---|
| AlphaFold1 | 2018 | CASP13 | 68.5 | 35% | 残基距离图预测 |
| AlphaFold2 | 2020 | CASP14 | 92.4 | 98% | Evoformer架构 |
| AlphaFold3 | 2024 | - | - | 全结构域 | 多聚体建模 |
表 1 AlphaFold系列模型性能对比
Table 1 Comparison of performance across AlphaFold model series
模型 Model | 发布时间 Release time | CASP参赛版本 CASP entry version | GDT_TS中位数 GDT_TS med-number | 覆盖UniProt比例 Coverage ratio of UniProt | 关键技术革新 Key technological innovation |
|---|---|---|---|---|---|
| AlphaFold1 | 2018 | CASP13 | 68.5 | 35% | 残基距离图预测 |
| AlphaFold2 | 2020 | CASP14 | 92.4 | 98% | Evoformer架构 |
| AlphaFold3 | 2024 | - | - | 全结构域 | 多聚体建模 |
| 特征 Characteristic | AlphaFold2 | RoseTTAFold |
|---|---|---|
| 计算资源 | 128 TPU v3 (4 d/蛋白) | 4 GPU (8 h/蛋白) |
| 核心架构 | Evoformer+结构模块 | 三轨Transformer |
| 动态结构预测 | 单构象输出 | 支持构象系综生成 |
| 膜蛋白预测精度 | TM-score 0.72 | TM-score 0.81 |
| 开源程度 | 部分开源 | 全代码公开 |
表2 RoseTTAFold与AlphaFold2的技术对比
Table 2 Technical comparison between RoseTTAFold and AlphaFold2
| 特征 Characteristic | AlphaFold2 | RoseTTAFold |
|---|---|---|
| 计算资源 | 128 TPU v3 (4 d/蛋白) | 4 GPU (8 h/蛋白) |
| 核心架构 | Evoformer+结构模块 | 三轨Transformer |
| 动态结构预测 | 单构象输出 | 支持构象系综生成 |
| 膜蛋白预测精度 | TM-score 0.72 | TM-score 0.81 |
| 开源程度 | 部分开源 | 全代码公开 |

图1 不同方法得到的β2-AR三维结构A:X-ray法,PDB ID为4G8R;B:NMR法,PDB ID为6KR8;C:cryo-EM法,PDB ID为8GGI;D:AlphaFold预测结果,编号为AF-P07550
Fig. 1 Three-dimensional structures of β2-AR obtained by different methodsA: X-ray method, with PDB ID 4G8R; B: NMR method, with PDB ID 6KR8; C: cryo-EM method, with PDB ID 8GGI; D: AlphaFold prediction result, with the number AF-P07550
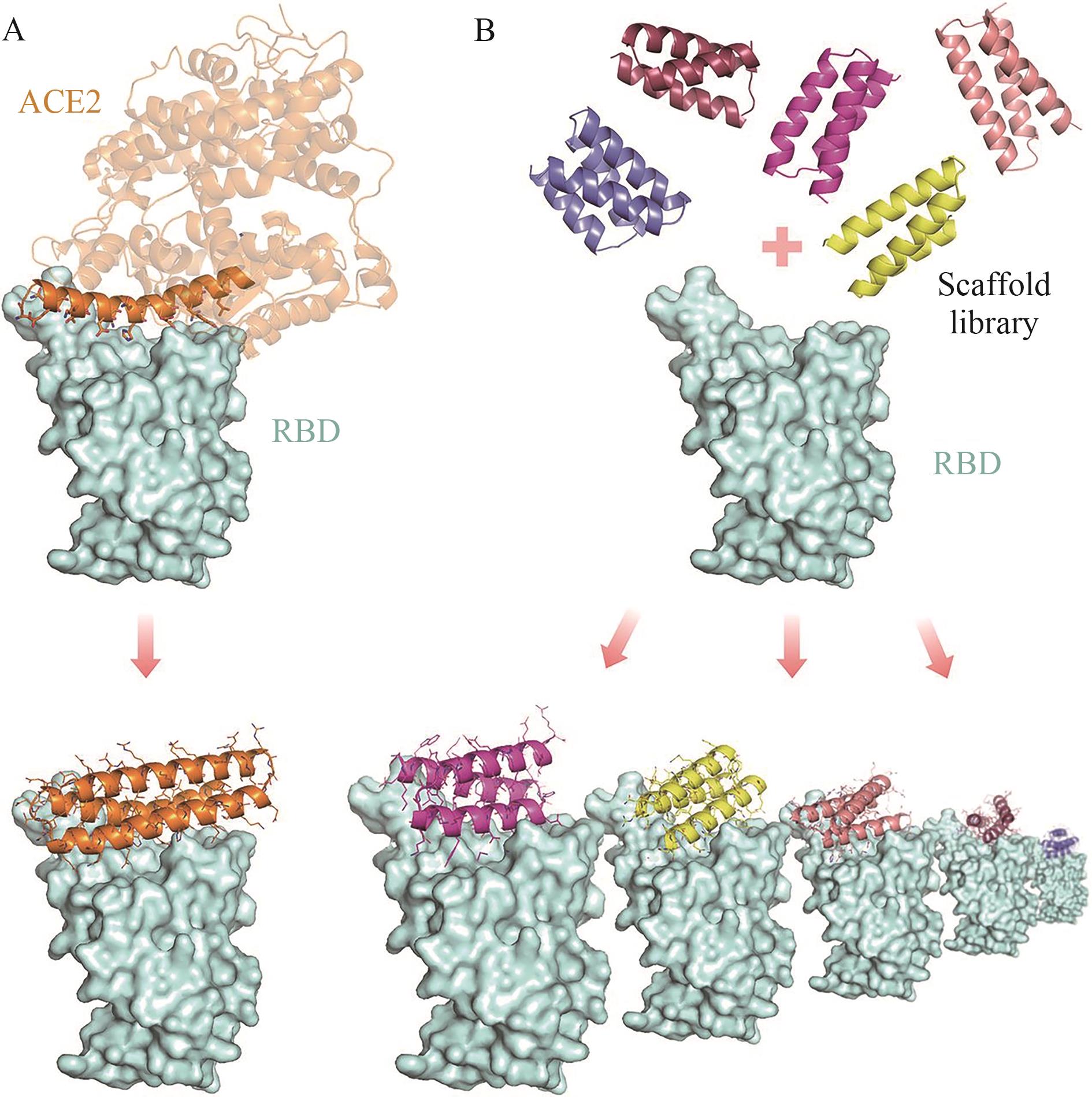
图2 计算设计方法概述A:包含ACE2螺旋的螺旋蛋白的设计;B:大规模从头设计小螺旋支架,然后进行RIF对接以识别形状和化学互补结合模式
Fig. 2 Overview diagram of computational design methodA: Design of helical proteins containing ACE2 helix. B:Large-scale head-to-tail design of small helical scaffolds followed by RIF docking to identify shape and chemical complementary binding patterns
| [1] | Woolfson DN. A brief history of de novo protein design: minimal, rational, and computational [J]. J Mol Biol, 2021, 433(20): 167160. |
| [2] | Simon I, Magyar C. Assortment of frontiers in protein science [J]. Int J Mol Sci, 2022, 23(7): 3685. |
| [3] | Zhuravlev PI, Papoian GA. Protein functional landscapes, dynamics, allostery: a tortuous path towards a universal theoretical framework [J]. Q Rev Biophys, 2010, 43(3): 295-332. |
| [4] | AlQuraishi M. Machine learning in protein structure prediction [J]. Curr Opin Chem Biol, 2021, 65: 1-8. |
| [5] | Zhou BX, Tan Y, Hu YT, et al. Protein engineering in the deep learning era [J]. mLife, 2024, 3(4): 477-491. |
| [6] | 潘杰, 中山皓博. 人工智能及其在生命科学中的应用与展望 [J]. 山东师范大学学报: 自然科学版, 2024, 39(2): 117-142. |
| Pan J, Akihiro Nakayama. Artificial intelligence and its application and prospect in life sciences [J]. J Shandong Norm Univ Nat Sci Ed, 2024, 39(2): 117-142. | |
| [7] | Gupta V, Liao WK, Choudhary A, et al. Evolution of artificial intelligence for application in contemporary materials science [J]. MRS Commun, 2023, 13(5): 754-763. |
| [8] | Pederson T. Protein structure: has levinthal’s paradox “folded”? [J]. FASEB J, 2021, 35(3): e21416. |
| [9] | Wang NN, Dong J, Ouyang DF. AI-directed formulation strategy design initiates rational drug development [J]. J Control Release, 2025, 378: 619-636. |
| [10] | Senior AW, Evans R, Jumper J, et al. Protein structure prediction using multiple deep neural networks in the 13th Critical Assessment of Protein Structure Prediction (CASP13) [J]. Proteins, 2019, 87(12): 1141-1148. |
| [11] | 阎隆飞, 孙之荣. 蛋白质分子结构 [M]. 北京: 清华大学出版社, 1999: 211-213. |
| Yan LF, Sun ZR. Protein molecular structure [M]. Beijing: Tsinghua University Press, 1999: 211-213. | |
| [12] | Berman HM, Battistuz T, Bhat TN, et al. The protein data bank [J]. Nucleic Acids Research, 2000, 28(1): 235-242. |
| [13] | 宁正元, 林世强. 蛋白质结构的预测及其应用 [J]. 福建农业大学学报, 2006, 35(3): 308-313. |
| Ning ZY, Lin SQ. Protein structure prediction and its application [J]. J Fujian Agric For Univ Nat Sci Ed, 2006, 35(3): 308-313. | |
| [14] | 杜宗阳. 蛋白质与RNA三级结构预测算法研究 [D]. 天津: 南开大学, 2022. |
| Du ZY. Research on algorithms for protein and RNA tertiary structure prediction [D]. Tianjin: Nankai University, 2022. | |
| [15] | 张晓凯, 张丛丛, 刘忠民, 等. 冷冻电镜技术的应用与发展 [J]. 科学技术与工程, 2019, 19(24): 9-17. |
| Zhang XK, Zhang CC, Liu ZM, et al. Application and development of cryo-electron microscopy technology [J]. Sci Technol Eng, 2019, 19(24): 9-17. | |
| [16] | 郭贝一, 郭晓强. AlphaFold和蛋白质结构预测 [J]. 科学, 2024, 76(5): 39-44. |
| Guo BY, Guo XQ. AlphaFold and protein structure prediction [J]. Science, 2024, 76(5): 39-44. | |
| [17] | Shaw DE, Maragakis P, Lindorff-Larsen K, et al. Atomic-level characterization of the structural dynamics of proteins [J]. Science, 2010, 330(6002): 341-346. |
| [18] | Lindorff-Larsen K, Piana S, Dror RO, et al. How fast-folding proteins fold [J]. Science, 2011, 334(6055): 517-520. |
| [19] | Karplus M, McCammon JA. Molecular dynamics simulations of biomolecules [J]. Nat Struct Biol, 2002, 9(9): 646-652. |
| [20] | Jumper J, Evans R, Pritzel A, et al. Highly accurate protein structure prediction with AlphaFold [J]. Nature, 2021, 596(7873): 583-589. |
| [21] | Varadi M, Anyango S, Deshpande M, et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models [J]. Nucleic Acids Res, 2022, 50(d1): D439-D444. |
| [22] | Li Y, Zhang CX, Feng CJ, et al. Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction [J]. Nat Commun, 2023, 14(1): 5745. |
| [23] | Dauparas J, Anishchenko I, Bennett N, et al. Robust deep learning-based protein sequence design using ProteinMPNN [J]. Science, 2022, 378(6615): 49-56. |
| [24] | Baek M, DiMaio F, Anishchenko I, et al. Accurate prediction of protein structures and interactions using a three-track neural network [J]. Science, 2021, 373(6557): 871-876. |
| [25] | Lisanza SL, Gershon JM, Tipps SWK, et al. Multistate and functional protein design using RoseTTAFold sequence space diffusion [J]. Nat Biotechnol, 2024. |
| [26] | Mirdita M, Schütze K, Moriwaki Y, et al. ColabFold: making protein folding accessible to all [J]. Nat Methods, 2022, 19(6): 679-682. |
| [27] | Velloso JPL, Kovacs AS, Pires DEV, et al. AI-driven GPCR analysis, engineering, and targeting [J]. Curr Opin Pharmacol, 2024, 74: 102427. |
| [28] | Singh I, Seth A, Billesbølle CB, et al. Structure-based discovery of conformationally selective inhibitors of the serotonin transporter [J]. Cell, 2023, 186(10): 2160-2175.e17. |
| [29] | Pándy-Szekeres G, Caroli J, Mamyrbekov A, et al. GPCRdb in 2023: state-specific structure models using AlphaFold2 and new ligand resources [J]. Nucleic Acids Res, 2023, 51(d1): D395-D402. |
| [30] | 王子佳, 郭卫娜, 郭巧珍, 等. β2-肾上腺素受体激动剂在神经退行性病变相关认知障碍中的作用 [J]. 中国医学科学院学报, 2022, 44(6): 1112-1116. |
| Wang ZJ, Guo WN, Guo QZ, et al. Role of β2-adrenergic receptor agonist in the cognitive impairment associated with neurodegenerative diseases [J]. Acta Academiae Medicinae Sinicae, 2022, 44(6): 1112-1116. | |
| [31] | Ramos BP, Colgan LA, Nou E, et al. β2 adrenergic agonist, clenbuterol, enhances working memory performance in aging animals [J]. Neurobiol Aging, 2008, 29(7): 1060-1069. |
| [32] | Humphreys IR, Pei J, Baek M, et al. Computed structures of core eukaryotic protein complexes [J]. Science, 2021, 374(6573): eabm4805. |
| [33] | Goverde CA, Wolf B, Khakzad H, et al. De novo protein design by inversion of the AlphaFold structure prediction network [J]. Protein Sci, 2023, 32(6): e4653. |
| [34] | Love AC, Prescher JA. Seeing (and using) the light: recent developments in bioluminescence technology [J]. Cell Chem Biol, 2020, 27(8): 904-920. |
| [35] | Jiang TY, Du LP, Li MY. Lighting up bioluminescence with coelenterazine: strategies and applications [J]. Photochem Photobiol Sci, 2016, 15(4): 466-480. |
| [36] | Yeh AH, Norn C, Kipnis Y, et al. De novo design of luciferases using deep learning [J]. Nature, 2023, 614(7949): 774-780. |
| [37] | Rohl CA, Strauss CEM, Misura KMS, et al. Protein structure prediction using Rosetta [J]. Meth Enzymol, 2004, 383: 66-93. |
| [38] | Dou JY, Vorobieva AA, Sheffler W, et al. De novo design of a fluorescence-activating β-barrel [J]. Nature, 2018, 561(7724): 485-491. |
| [39] | Cao L, Goreshnik I, Coventry B, et al. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors [J]. Science, 2020, 370(6515): 426-431. |
| [40] | 谭生龙. 基于序列的蛋白质功能分类系统的研究与设计 [J]. 科技创新与应用, 2016, 6(27): 68. |
| Tan SL. Research and design of protein function classification system based on sequence [J]. Technol Innov Appl, 2016, 6(27): 68. | |
| [41] | 叶玉珍, 丁达夫. 蛋白质骨架库的构建及其在功能蛋白质设计中的应用 [J]. 生物物理学报, 1999, 15(4): 751-757. |
| Ye YZ, Ding DF. Construction of protein scaffold database and itsapplications to functional protein design [J]. Acta Biophys Sin, 1999, 15(4): 751-757. | |
| [42] | Li YL, Jiao WT, Liu RH, et al. Expanding the sequence spaces of synthetic binding protein using deep learning-based framework ProteinMPNN [J]. Front Comput Sci, 2024, 19(5): 195903. |
| [43] | Sumida KH, Núñez-Franco R, Kalvet I, et al. Improving protein expression, stability, and function with ProteinMPNN [J]. J Am Chem Soc, 2024, 146(3): 2054-2061. |
| [44] | Watson JL, Juergens D, Bennett NR, et al. De novo design of protein structure and function with RFdiffusion [J]. Nature, 2023, 620(7976): 1089-1100. |
| [45] | Robins K. BPS2025-Enhancing antibody design using RFdiffusion and ProteinMPNN for novel intrabody generation [J]. Biophysical Journal, 2025, 124(3):217a-218a. |
| [46] | Whitesides GM. The origins and the future of microfluidics [J]. Nature, 2006, 442(7101): 368-373. |
| [47] | Snoek J, Larochelle H, Adams PR. Practical bayesian optimization of machine learning algorithms [J]. CoRR, 2012. |
| [48] | Abramson J, Adler J, Dunger J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold3 [J]. Nature, 2024, 630(8016): 493-500. |
| [49] | Nguyen E, Poli M, Durrant MG, et al. Sequence modeling and design from molecular to genome scale with Evo [J]. Science, 2024, 386(6723): eado9336. |
| [50] | 张裕, 周化岚, 张建国, 等. 无细胞蛋白质表达系统的优化与应用 [J]. 生命的化学, 2022, 42(8): 1493-1501. |
| Zhang Y, Zhou HL, Zhang JG, et al. Optimization and application of cell-free system for protein expression [J]. Chem Life, 2022, 42(8): 1493-1501. | |
| [51] | Mazzotti G, Hartmann D, Booth MJ. Precise, orthogonal remote-control of cell-free systems using photocaged nucleic acids [J]. J Am Chem Soc, 2023, 145(17): 9481-9487. |
| [52] | Graham F. Daily briefing: AlphaFold developers share Nobel prize in chemistry [J]. Nature, 2024. |
| [53] | Evseev P, Shneider M, Miroshnikov K. Evolution of phage tail sheath protein [J]. Viruses, 2022, 14(6): 1148. |
| [54] | Podgorski JM, Freeman K, Gosselin S, et al. A structural dendrogram of the actinobacteriophage major capsid proteins provides important structural insights into the evolution of capsid stability [J]. Structure, 2023, 31(3): 282-294.e5. |
| [55] | Bisio H, Legendre M, Giry C, et al. Evolution of giant pandoravirus revealed by CRISPR/Cas9 [J]. Nat Commun, 2023, 14: 428. |
| [56] | Bian JH, Tan P, Nie T, et al. Optimizing enzyme thermostability by combining multiple mutations using protein language model [J]. mLife, 2024, 3(4): 492-504. |
| [57] | Liao YJ, Ma H, Wang ZY, et al. Rapid restoration of potent neutralization activity against the latest Omicron variant JN.1 via AI rational design and antibody engineering [J]. Proc Natl Acad Sci USA, 2025, 122(6): e2406659122. |
| [58] | Wu T, Chen XH, Fei YT, et al. Artificial metalloenzyme assembly in cellular compartments for enhanced catalysis [J]. Nat Chem Biol, 2025, 21(5): 779-789. |
| [59] | Jiang F, Li MC, Dong JJ, et al. A general temperature-guided language model to design proteins of enhanced stability and activity [J]. Sci Adv, 2024, 10(48): eadr2641. |
| [1] | 高婧, 陈益存, 高暝, 赵耘霄, 汪阳东. 植物单宁合成调控及其对环境的响应机制[J]. 生物技术通报, 2025, 41(7): 49-59. |
| [2] | 王辉, 范灵熙, 孙纪录, 王苑, 伍宁丰, 田健, 关菲菲. 基于蛋白智能模型提升溶菌酶RPL187的热稳定性[J]. 生物技术通报, 2025, 41(7): 336-346. |
| [3] | 吴娅, 姚润, 杨含婷, 刘微, 杨帅, 宋驰, 陈士林. 凤梨薄荷SDR基因家族全基因组鉴定及表达分析[J]. 生物技术通报, 2025, 41(5): 175-185. |
| [4] | 鲁天怡, 李爱朋, 费强. 生物合成聚乳酸研究进展[J]. 生物技术通报, 2025, 41(4): 47-60. |
| [5] | 李晓明, 尚秀华, 王有霜, 吴志华. 植物中苯并噁嗪类化合物的研究进展[J]. 生物技术通报, 2025, 41(4): 9-20. |
| [6] | 聂祝欣, 郭瑾, 乔子洋, 李微薇, 张学燕, 刘春阳, 王静. 黑果枸杞不同发育时期果实花色苷合成的转录组分析[J]. 生物技术通报, 2024, 40(8): 106-117. |
| [7] | 马小翔, 马泽源, 刘亚月, 周龙建, 和羿帆, 张翼. 仿突变生物合成调控对土曲霉C23-3次生代谢产物的影响[J]. 生物技术通报, 2024, 40(8): 275-287. |
| [8] | 沈真辉, 曹瑶, 杨林雷, 罗祥英, 子灵山, 陆青青, 李荣春. 金耳和毛韧革菌麦角硫因生物合成基因的克隆及生物信息学分析[J]. 生物技术通报, 2024, 40(7): 259-272. |
| [9] | 何玙冰, 付振浩, 李仁瀚, 刘秀霞, 刘春立, 杨艳坤, 李业, 白仲虎. 利用代谢工程在酿酒酵母中高效合成2-萘乙醇[J]. 生物技术通报, 2024, 40(7): 99-107. |
| [10] | 胡锦锦, 李素贞, 马旭辉, 柳小庆, 谢珊珊, 江海洋, 陈茹梅. 玉米花青素生物合成代谢调控[J]. 生物技术通报, 2024, 40(6): 34-44. |
| [11] | 张美玉, 赵玉斌, 王灵云, 宋元达, 赵新河, 任晓洁. 微藻破囊壶菌产功能性脂肪酸DHA研究进展[J]. 生物技术通报, 2024, 40(6): 81-94. |
| [12] | 李梦然, 叶伟, 李赛妮, 张维阳, 李建军, 章卫民. Lithocarols类化合物生物合成基因litI的表达及其启动子功能分析[J]. 生物技术通报, 2024, 40(6): 310-318. |
| [13] | 刘玉萍, 张维阳, 章卫民, 叶伟, 李冬利. Phomopsis tersa FS441聚酮杂萜类化合物生物合成基因启动子的鉴定[J]. 生物技术通报, 2024, 40(12): 248-255. |
| [14] | 纪宏超, 李正艳. 基于质谱的未知次生代谢物结构解析研究进展与展望[J]. 生物技术通报, 2024, 40(10): 76-85. |
| [15] | 王俊芳, 黄秋斌, 张飘丹, 张彭湃. Surfactin的结构、生物合成及其在生物防治中的作用[J]. 生物技术通报, 2024, 40(1): 100-112. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||