








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 85-94.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0535
收稿日期:2024-06-03
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
沈川作者简介:沈川,男,博士,副教授,研究方向:植物与病原菌互作;E-mail: chuan_shen@aku.edu.cn
基金资助:
SHEN Chuan1( ), LI Xia1, QIN Jian-feng2, DUAN Long-fei2, LIU Jia3
), LI Xia1, QIN Jian-feng2, DUAN Long-fei2, LIU Jia3
Received:2024-06-03
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】通过构建魔芋cDNA文库,筛选转录因子WRKY72的互作蛋白,为解析WRKY72调控魔芋抗病的分子机制提供帮助。【方法】以胡萝卜软腐果胶杆菌侵染不同阶段的魔芋为实验材料,提取总RNA等量混匀后构建魔芋cDNA文库,使用同源重组的方法构建转录因子WRKY72的诱饵载体,采用酵母双杂交技术筛选候选靶标蛋白,并进行一对一验证。【结果】酵母文库滴度为1.6×107 CFU/mL,插入片段的平均长度在1 000 bp左右,诱饵载体在酵母中经验证无自激活。诱饵载体pGBKT7-WRKY72筛库获得了41个阳性互作蛋白,包括转运蛋白、通道蛋白、热休克蛋白、叶绿素结合蛋白和过氧化物酶等。对这些互作蛋白进行GO和KEGG功能富集分析,最后选取12个互作蛋白进行一对一酵母双杂交验证。【结论】成功构建了高质量的魔芋文库并筛选到了转录因子WRKY72的互作蛋白,为解析魔芋抗软腐病机制提供了理论基础。
沈川, 李夏, 覃剑锋, 段龙飞, 刘佳. 基于软腐病菌诱导的魔芋酵母双杂交文库筛选WRKY72互作蛋白[J]. 生物技术通报, 2025, 41(1): 85-94.
SHEN Chuan, LI Xia, QIN Jian-feng, DUAN Long-fei, LIU Jia. Screening for WRKY72-interacting Proteins Using a Yeast Two-hybrid Library Derived from Soft Rot Pathogen-induced Amorphophallus konjac[J]. Biotechnology Bulletin, 2025, 41(1): 85-94.
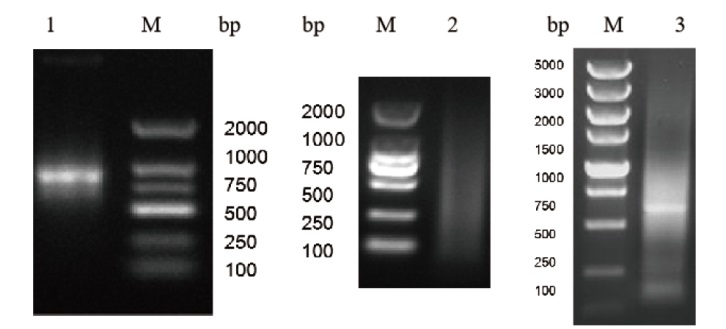
图1 魔芋总RNA、mRNA和双链cDNA的电泳图 1:总RNA电泳图;2:mRNA纯化电泳图;3:双链cDNA电泳图。M:DNA marker
Fig. 1 Electrophoresis of total RNA, mRNA and ds cDNA isolated from konjac(A. konjac) 1: Electrophoresis of total RNA; 2: electrophoresis of purified mRNA; 3: electrophoresis of ds cDNA; M: DNA marker
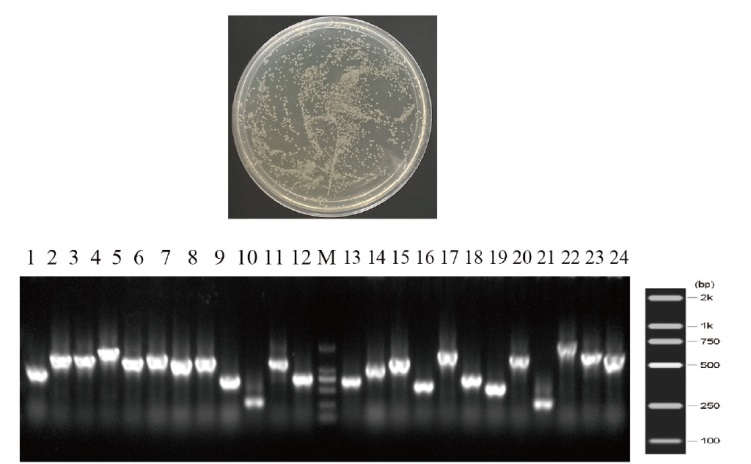
图2 文库滴度检测及PCR检测文库插入cDNA片段 1-24:PCR鉴定24个随机的次级cDNA文库克隆。M:DNA marker
Fig. 2 Library titer detection and PCR detection of inserted fragments in the library 1-24: PCR detection of 24 clones randomly selected from the secondary cDNA library. M: DL2000 DNA marker
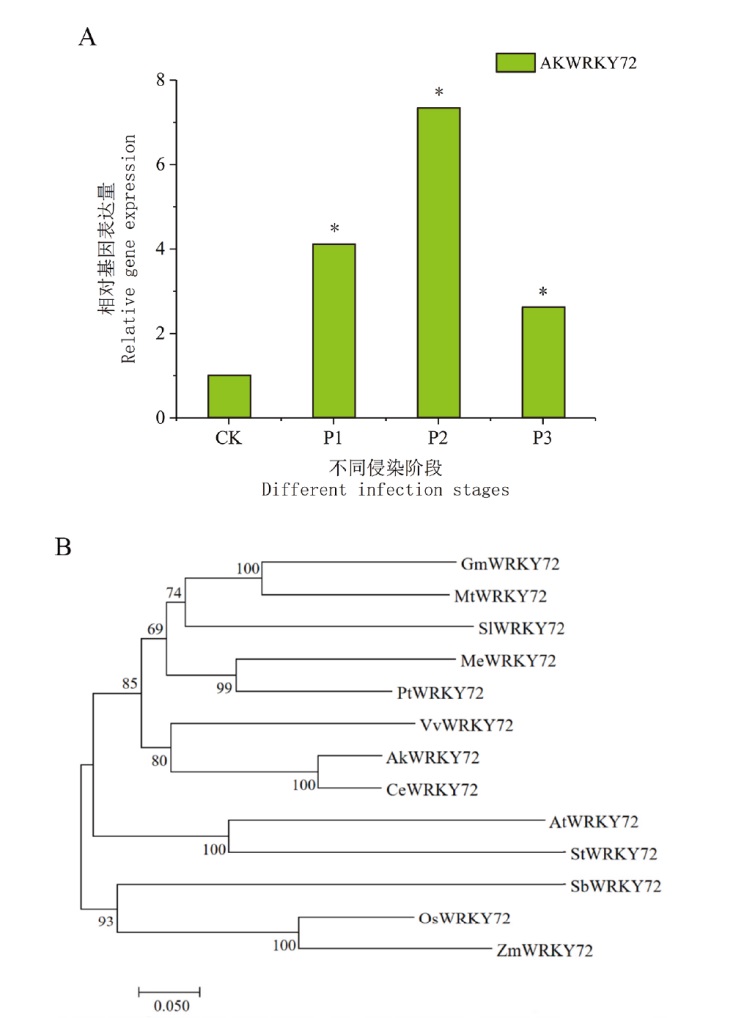
图3 AkWRKY72的生物信息学分析 A:软腐菌侵染魔芋不同阶段AkWRKY72的表达量,P1表示换头期前,P2表示换头期后,P3表示成熟期;B:不同物种中WRKY72蛋白的进化树分析。“*”代表P<0.05
Fig. 3 Bioinformatics analysis of AkWRKY72 A: Expressions of AkWRKY72 in konjac infected with soft rot at different stages, P1 refers to the period before heading, P2 refers to the period after heading, and P3 refers to the mature period. B: Evolutionary tree analysis of WRKY72 proteins in different species. “*” indicates P<0.05
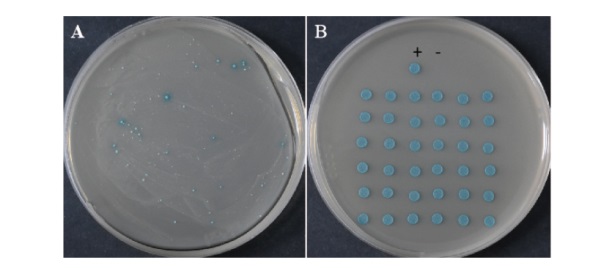
图5 魔芋AkWRKY72互作蛋白的筛选 A:转化的酵母细胞涂布TDO/X/3'AT 5 mmol/L平板;B:转化的酵母细胞点QDO/X平板。+:PGBKT7-53和PGADT7-T共转化阳性对照,-:PGBKT7-lam和PGADT7-T共转化阴性对照
Fig. 5 Screening of candidate interaction protein for AkWRKY72 in konjac A: Transformed yeast cells coated with TDO/X/3'AT 5 mM plates. B: Transformed yeast cells spotted with QDO/X plates. +: PGBKT7-53 and PGADT7-T co-transformed positive control, -: PGBKT7-lam and PGADT7-T co-transformed negative control
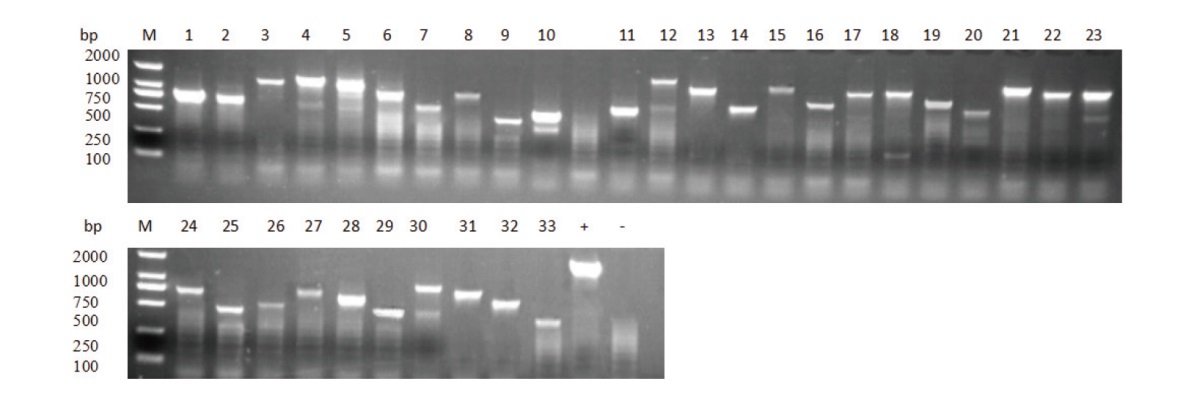
图6 筛选阳性克隆菌落PCR产物琼脂糖凝胶电泳图 1-33:菌落PCR产物。+:阳性对照,-:阴性对照
Fig. 6 Agarose gel electrophoresis of PCR products of screened positive clones 1-33: PCR products of colonies. +: Positive control, -: negative control
| 编号 No. | 功能注释 Function annotation | 序列号 Accession | |
|---|---|---|---|
| 1 | 平衡核苷酸转运蛋白1样蛋白 | Equilibrative nucleotide transporter 1-like | AH13G08570.1 |
| 2 | 蛋白酶体亚基 β-6型 | Proteasome subunit beta type-6 | XM_021751824.2 |
| 3 | 水通道蛋白 PIP2-6 | Aquaporin PIP2-6 | XM_010921795.3 |
| 4 | 叶绿素 a-b 结合蛋白 | Chlorophyll a-b binding protein | XM_009391104.2 |
| 5 | 类热休克蛋白质 | Class I heat shock protein-like | XM_037569548.1 |
| 6 | 小泛素相关修饰物1 | Small ubiquitin-related modifier 1 | XM_009414468.2 |
| 7 | 磷酸核酮糖3-异构酶 | Ribulose-phosphate 3-epimerase | XM_020248600.1 |
| 8 | TVP38/TMEm64家族膜蛋白sr0305 | TVP38/TMEM64 family membrane protein slr0305-like | XM_039134159.1 |
| 9 | 小分子热休克蛋白 | Small heat shock protein | XM_008813654.3 |
| 10 | 蛋白质转运蛋白Sec61亚基 | Protein transport protein Sec61 subunit gamma | XM_037612338.1 |
| 11 | 液泡蛋白分选相关蛋白13C | Vacuolar protein sorting-associated protein 13C | XM_039279221.1 |
| 12 | 光系统 I 反应中心亚基 | Photosystem I reaction center subunit II | XM_009402563.2 |
| 13 | 延伸因子1-a | Elongation factor 1-alpha | XM_031625179.2 |
| 14 | 类 γ-谷氨酰水解酶2 | Gamma-glutamyl hydrolase 2-like | XM_059806763.1 |
| 15 | 31kD 核糖核蛋白 | 31 kD ribonucleoprotein | XM_008791202.4 |
| 16 | 类似苹果酸脱氢酶 | Malate dehydrogenase, glyoxysomal-like | XM_008780993.4 |
| 17 | 铁氧化还原蛋白 | Ferredoxin | XM_051344602.1 |
| 18 | 叶绿素 a-b 结合蛋白4 | Chlorophyll a-b binding protein 4 | XM_020255110.1 |
| 19 | 脂磷酸磷酸酶 δ | Lipid phosphate phosphatase delta | XM_010936202.3 |
| 20 | 硫氧还蛋白 H 型 | Thioredoxin H-type | XM_062288332.1 |
| 21 | LHCII III 型叶绿素 a-b 结合蛋白 | Chlorophyll a-b binding protein of LHCII type III | XM_039974480.1 |
| 22 | 核酮糖-1,5-二磷酸羧化酶/加氧酶小亚基 | Ribulose-1,5-bisphosphate carboxylase/oxygenase small subunit | AF034479.1 |
| 23 | 大核糖体亚基蛋白 eL39 | Large ribosomal subunit protein eL39 | XM_062364610.1 |
| 24 | LHCII 类1型叶绿素 a-b 结合蛋白 | Chlorophyll a-b binding protein of LHCII type 1-like | XM_009391104.2 |
| 25 | 叶绿素 a-b 结合蛋白8 | Chlorophyll a-b binding protein 8 | XM_057956264.1 |
| 26 | 镁-原卟啉 IX 单甲酯氧化环化酶 | Magnesium-protoporphyrin IX monomethyl ester Oxidative cyclase | XM_018819649.1 |
| 27 | 类赖氨酸组氨酸转运蛋白 | Lysine histidine transporter 1-like | XM_020737698.1 |
| 28 | 乙酰鸟氨酸脱乙酰基酶 | Acetylornithine deacetylase | XM_010935186.3 |
| 29 | 过氧化物酶21 | Peroxidase 21 | XM_009393918.1 |
| 30 | 核转录因子 Y 亚基 A-7 | Nuclear transcription factor Y subunit A-7 | XM_021736266.2 |
| 31 | 谷氨酸 tRNA 连接酶 | Glutamate-tRNA ligase | XM_058220292.1 |
| 32 | Rubisco激活酶(rca1) | Rubisco activase(rca1) | AF338237.1 |
| 33 | 光系统 I 叶绿素 a/b 结合蛋白2 | Photosystem I chlorophyll a/b-binding protein 2 | XM_042535174.1 |
| 34 | 叶绿素a-b结合蛋白 CP29.1 | Chlorophyll a-b binding protein CP29.1 | XM_062333380.1 |
| 35 | 热休克蛋白 | Heat shock protein | XM_010930652.3 |
| 36 | 丝氨酸/苏氨酸蛋白磷酸酶2A | Serine/threonine protein phosphatase 2A | XM_017844298.3 |
| 37 | 鸟氨酸氨基转移酶 | Ornithine transcarbamylase | XM_043855639.1 |
| 38 | 60S 核糖体蛋白质 L34-like | 60S ribosomal protein L34-like | XM_039037995.1 |
| 39 | 60S 核糖体蛋白质 L18-2-like | 60S ribosomal protein L18-2-like | XM_039120108.1 |
| 40 | 谷氨酰胺合成酶 | Glutamine synthetase | XM_010940494.3 |
| 41 | 琥珀酸脱氢酶 | Succinate dehydrogenase | XM_020232587.1 |
表1 酵母双杂筛选获得的AkWRKY72互作蛋白的功能注释
Table 1 Functional annotation of AkWRKY72-interacting proteins screened by yeast two-hybrid system
| 编号 No. | 功能注释 Function annotation | 序列号 Accession | |
|---|---|---|---|
| 1 | 平衡核苷酸转运蛋白1样蛋白 | Equilibrative nucleotide transporter 1-like | AH13G08570.1 |
| 2 | 蛋白酶体亚基 β-6型 | Proteasome subunit beta type-6 | XM_021751824.2 |
| 3 | 水通道蛋白 PIP2-6 | Aquaporin PIP2-6 | XM_010921795.3 |
| 4 | 叶绿素 a-b 结合蛋白 | Chlorophyll a-b binding protein | XM_009391104.2 |
| 5 | 类热休克蛋白质 | Class I heat shock protein-like | XM_037569548.1 |
| 6 | 小泛素相关修饰物1 | Small ubiquitin-related modifier 1 | XM_009414468.2 |
| 7 | 磷酸核酮糖3-异构酶 | Ribulose-phosphate 3-epimerase | XM_020248600.1 |
| 8 | TVP38/TMEm64家族膜蛋白sr0305 | TVP38/TMEM64 family membrane protein slr0305-like | XM_039134159.1 |
| 9 | 小分子热休克蛋白 | Small heat shock protein | XM_008813654.3 |
| 10 | 蛋白质转运蛋白Sec61亚基 | Protein transport protein Sec61 subunit gamma | XM_037612338.1 |
| 11 | 液泡蛋白分选相关蛋白13C | Vacuolar protein sorting-associated protein 13C | XM_039279221.1 |
| 12 | 光系统 I 反应中心亚基 | Photosystem I reaction center subunit II | XM_009402563.2 |
| 13 | 延伸因子1-a | Elongation factor 1-alpha | XM_031625179.2 |
| 14 | 类 γ-谷氨酰水解酶2 | Gamma-glutamyl hydrolase 2-like | XM_059806763.1 |
| 15 | 31kD 核糖核蛋白 | 31 kD ribonucleoprotein | XM_008791202.4 |
| 16 | 类似苹果酸脱氢酶 | Malate dehydrogenase, glyoxysomal-like | XM_008780993.4 |
| 17 | 铁氧化还原蛋白 | Ferredoxin | XM_051344602.1 |
| 18 | 叶绿素 a-b 结合蛋白4 | Chlorophyll a-b binding protein 4 | XM_020255110.1 |
| 19 | 脂磷酸磷酸酶 δ | Lipid phosphate phosphatase delta | XM_010936202.3 |
| 20 | 硫氧还蛋白 H 型 | Thioredoxin H-type | XM_062288332.1 |
| 21 | LHCII III 型叶绿素 a-b 结合蛋白 | Chlorophyll a-b binding protein of LHCII type III | XM_039974480.1 |
| 22 | 核酮糖-1,5-二磷酸羧化酶/加氧酶小亚基 | Ribulose-1,5-bisphosphate carboxylase/oxygenase small subunit | AF034479.1 |
| 23 | 大核糖体亚基蛋白 eL39 | Large ribosomal subunit protein eL39 | XM_062364610.1 |
| 24 | LHCII 类1型叶绿素 a-b 结合蛋白 | Chlorophyll a-b binding protein of LHCII type 1-like | XM_009391104.2 |
| 25 | 叶绿素 a-b 结合蛋白8 | Chlorophyll a-b binding protein 8 | XM_057956264.1 |
| 26 | 镁-原卟啉 IX 单甲酯氧化环化酶 | Magnesium-protoporphyrin IX monomethyl ester Oxidative cyclase | XM_018819649.1 |
| 27 | 类赖氨酸组氨酸转运蛋白 | Lysine histidine transporter 1-like | XM_020737698.1 |
| 28 | 乙酰鸟氨酸脱乙酰基酶 | Acetylornithine deacetylase | XM_010935186.3 |
| 29 | 过氧化物酶21 | Peroxidase 21 | XM_009393918.1 |
| 30 | 核转录因子 Y 亚基 A-7 | Nuclear transcription factor Y subunit A-7 | XM_021736266.2 |
| 31 | 谷氨酸 tRNA 连接酶 | Glutamate-tRNA ligase | XM_058220292.1 |
| 32 | Rubisco激活酶(rca1) | Rubisco activase(rca1) | AF338237.1 |
| 33 | 光系统 I 叶绿素 a/b 结合蛋白2 | Photosystem I chlorophyll a/b-binding protein 2 | XM_042535174.1 |
| 34 | 叶绿素a-b结合蛋白 CP29.1 | Chlorophyll a-b binding protein CP29.1 | XM_062333380.1 |
| 35 | 热休克蛋白 | Heat shock protein | XM_010930652.3 |
| 36 | 丝氨酸/苏氨酸蛋白磷酸酶2A | Serine/threonine protein phosphatase 2A | XM_017844298.3 |
| 37 | 鸟氨酸氨基转移酶 | Ornithine transcarbamylase | XM_043855639.1 |
| 38 | 60S 核糖体蛋白质 L34-like | 60S ribosomal protein L34-like | XM_039037995.1 |
| 39 | 60S 核糖体蛋白质 L18-2-like | 60S ribosomal protein L18-2-like | XM_039120108.1 |
| 40 | 谷氨酰胺合成酶 | Glutamine synthetase | XM_010940494.3 |
| 41 | 琥珀酸脱氢酶 | Succinate dehydrogenase | XM_020232587.1 |
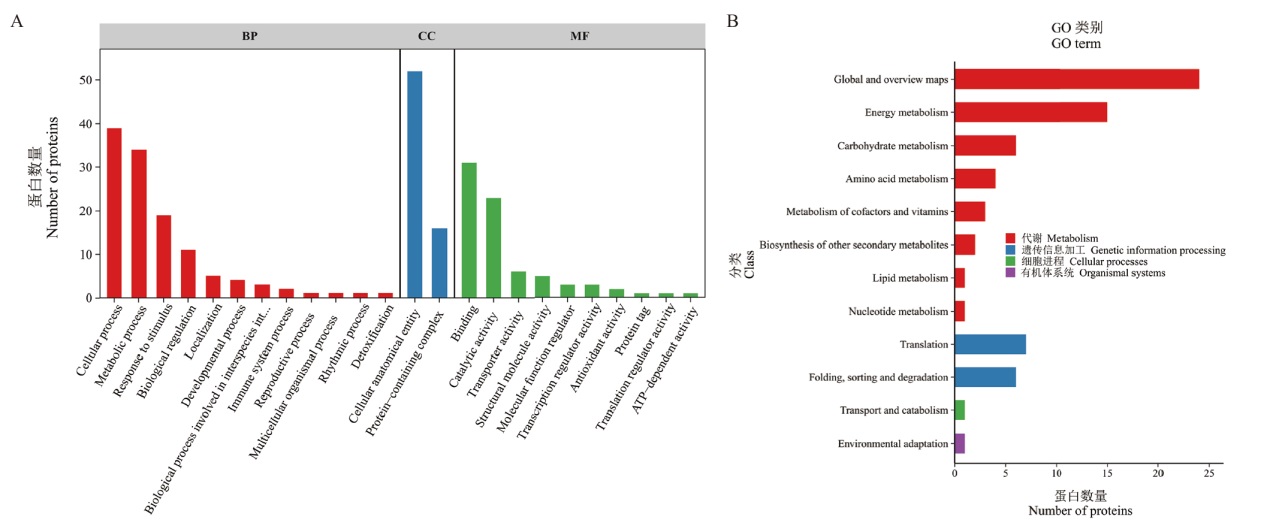
图7 AkWRKY72互作蛋白的功能富集分析 A:GO功能富集分析;B:KEGG功能富集分析
Fig. 7 Functional enrichment analysis of AkWRKY72-interacting proteins A: GO functional enrichment analysis. B: KEGG functional enrichment analysis
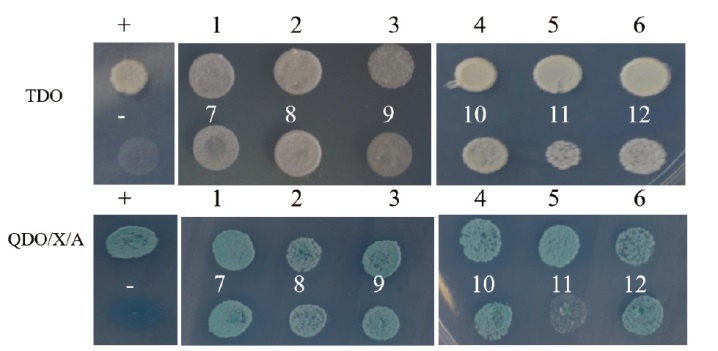
图8 AkWRKY72互作蛋白的一对一验证 +:PGBKT7-53和PGADT7-T共转化阳性对照;-:PGBKT7-lam和PGADT7-T共转化阴性对照。1:过氧化物酶21;2:核转录因子Y亚基 A-7;3:热休克蛋白;4:延伸因子1-a;5:铁氧化还原蛋白;6:泡蛋白分选相关蛋白13C;7:水通道蛋白 PIP2-6;8:叶绿素 a-b 结合蛋白4;9:丝氨酸/苏氨酸蛋白磷酸酶2A;10:叶绿素 a-b 结合蛋白8;11:蛋白质转运蛋白 Sec61亚基;12:类赖氨酸组氨酸转运蛋白
Fig. 8 One-to-one verification of interaction protein for AkWRKY72 +: PGBKT7-53 and PGADT7-T co-transformed positive control; -: PGBKT7-lam and PGADT7-T co-transformed negative control. 1: Peroxidase 21; 2: nuclear transcription factor Y subunit A-7; 3: 18.1 kD class I heat shock protein; 4: elongation factor 1-alpha; 5: ferredoxin; 6: vacuolar protein sorting-associated protein 13C; 7: aquaporin PIP2-6; 8: chlorophyll a-b binding protein 4; 9: serine/threonine protein phosphatase 2A; 10: chlorophyll a-b binding protein 8; 11: protein transport protein Sec61 subunit gamma; 12: lysine histidine transporter 1-like
| [1] | 王昊, 李昆志. 魔芋属植物分子生物学研究现状与展望[J]. 分子植物育种, 2022, 20(5): 1530-1536. |
| Wang H, Li KZ. Research status and prospect in molecular biology of Amorphophallus[J]. Molecular Plant Breeding, 2022, 20(5): 1530-1536. | |
| [2] | 王健, 乐超银, 郭政宏, 等. 魔芋软腐病致病机理及其生物防治[J]. 安徽农业科学, 2009, 37(30): 14746-14748. |
| Wang J, Le CY, Guo ZH, et al. Pathogenic mechanism and biological control of konjac soft rot disease[J]. J Anhui Agric Sci, 2009, 37(30): 14746-14748. | |
| [3] | Wei HY, Yang M, Ke YG, et al. Comparative physiological and transcriptomic profiles reveal regulatory mechanisms of soft rot disease resistance in Amorphophallus spp[J]. Physiol Mol Plant Pathol, 2022, 118: 101807. |
| [4] |
Eulgem T, Rushton PJ, Robatzek S, et al. The WRKY superfamily of plant transcription factors[J]. Trends Plant Sci, 2000, 5(5): 199-206.
doi: 10.1016/s1360-1385(00)01600-9 pmid: 10785665 |
| [5] |
Eulgem T, Rushton PJ, Schmelzer E, et al. Early nuclear events in plant defence signalling: rapid gene activation by WRKY transcription factors[J]. EMBO J, 1999, 18(17): 4689-4699.
doi: 10.1093/emboj/18.17.4689 pmid: 10469648 |
| [6] |
Jiang JJ, Ma SH, Ye NH, et al. WRKY transcription factors in plant responses to stresses[J]. J Integr Plant Biol, 2017, 59(2): 86-101.
doi: 10.1111/jipb.12513 |
| [7] | Liu F, Li XX, Wang MR, et al. Interactions of WRKY15 and WRKY33 transcription factors and their roles in the resistance of oilseed rape to Sclerotinia infection[J]. Plant Biotechnol J, 2018, 16(4): 911-925. |
| [8] | Javed T, Gao SJ. WRKY transcription factors in plant defense[J]. Trends Genet, 2023, 39(10): 787-801. |
| [9] | Zhou R, Dong YH, Liu X, et al. JrWRKY21 interacts with JrPTI5L to activate the expression of JrPR5L for resistance to Colletotrichum gloeosporioides in walnut[J]. Plant J, 2022, 111(4): 1152-1166. |
| [10] | Wang X, Qiao QH, Zhao KK, et al. PbWRKY18 promotes resistance against black spot disease by activation of the chalcone synthase gene PbCHS3 in pear[J]. Plant Sci, 2024, 341: 112015. |
| [11] | Dang FF, Lin JH, Li YJ, et al. SlWRKY30 and SlWRKY81 synergistically modulate tomato immunity to Ralstonia solanacearum by directly regulating SlPR-STH2[J]. Hortic Res, 2023, 10(5): uhad050. |
| [12] | Xiao SH, Ming YQ, Hu Q, et al. GhWRKY41 forms a positive feedback regulation loop and increases cotton defence response against Verticillium dahliae by regulating phenylpropanoid metabolism[J]. Plant Biotechnol J, 2023, 21(5): 961-978. |
| [13] |
Bludau I, Aebersold R. Proteomic and interactomic insights into the molecular basis of cell functional diversity[J]. Nature Reviews Molecular Cell Biology, 2020, 21(6): 327-340.
doi: 10.1038/s41580-020-0231-2 pmid: 32235894 |
| [14] | 赵小明, 李增义, 崔鸣, 等. 安康魔芋软腐病防治技术初步研究[J]. 西北农业学报, 2021, 30(8): 1263-1270. |
| Zhao XM, Li ZY, Cui M, et al. Preliminary study on soft rot control technology of konjac in Ankang[J]. Acta Agric Boreali Occidentalis Sin, 2021, 30(8): 1263-1270. | |
| [15] |
黄幸, 丁峰, 彭宏祥, 等. 植物WRKY转录因子家族研究进展[J]. 生物技术通报, 2019, 35(12): 129-143.
doi: 10.13560/j.cnki.biotech.bull.1985.2019-0626 |
|
Huang X, Ding F, Peng HX, et al. Research progress on family of plant WRKY transcription factors[J]. Biotechnol Bull, 2019, 35(12): 129-143.
doi: 10.13560/j.cnki.biotech.bull.1985.2019-0626 |
|
| [16] |
王婷, 葛怀娜, 郭宏. 酵母双杂交技术应用进展[J]. 生物技术进展, 2015, 5(5): 392-396.
doi: 10.3969/j.issn.2095-2341.2015.05.12 |
|
Wang T, Ge HN, Guo H. Progress on application of yeast two-hybrid technique[J]. Curr Biotechnol, 2015, 5(5): 392-396.
doi: 10.3969/j.issn.2095-2341.2015.05.12 |
|
| [17] | 李琳琳, 王超群, 李春霞, 等. 木薯MeHsfB3a互作蛋白的筛选和鉴定[J]. 分子植物育种, 2023, 21(12): 3850-3856. |
| Li LL, Wang CQ, Li CX, et al. Screening and identification of interacting protein MeHsfB3a from cassava[J]. Mol Plant Breed, 2023, 21(12): 3850-3856. | |
| [18] | 王佳莉, 梁晓宇, 柯宇航, 等. 白粉菌侵染橡胶树叶片的酵母双杂交文库构建及应用[J/OL]. 植物病理学报, 2024: 1-12. |
| Wang JL, Liang XY, Ke YH, et al. Construction and application of a yeast two-hybrid library of rubber tree leaves infected with Oidium heveae[J/OL]. Acta Phytopathologica Sinica, 2024: 1-12. | |
| [19] | 戴凡炜, 王振江, 罗国庆, 等. 青枯菌诱导的桑树根部酵母双杂交文库构建和MaTGA1互作蛋白的筛选及鉴定[J]. 植物生理学报, 2023, 59(11): 2126-2134. |
| Dai FW, Wang ZJ, Luo GQ, et al. Construction of yeast two-hybrid cDNA library induced by Ralstonia solanacearum and interaction protein screening and identification for MaTGA1 in mulberry roots[J]. Plant Physiology Journal, 2023, 59(11): 2126-2134. | |
| [20] |
Zheng ZY, Qamar SA, Chen ZX, et al. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens[J]. Plant J, 2006, 48(4): 592-605.
doi: 10.1111/j.1365-313X.2006.02901.x pmid: 17059405 |
| [21] |
Chi YJ, Yang Y, Zhou Y, et al. Protein-protein interactions in the regulation of WRKY transcription factors[J]. Mol Plant, 2013, 6(2): 287-300.
doi: 10.1093/mp/sst026 pmid: 23455420 |
| [22] | Bhattarai KK, Atamian HS, Kaloshian I, et al. WRKY72-type transcription factors contribute to basal immunity in tomato and Arabidopsis as well as gene-for-gene resistance mediated by the tomato R gene Mi-1[J]. Plant J, 2010, 63(2): 229-240. |
| [23] | Zheng LY, Gao S, Bai YJ, et al. NF-YC15 transcription factor activates ethylene biosynthesis and improves cassava disease resistance[J]. Plant Biotechnol J, 2024. DOI: 10.1111/pbi.14355. |
| [24] |
Durian G, Rahikainen M, Alegre S, et al. Protein phosphatase 2A in the regulatory network underlying biotic stress resistance in plants[J]. Front Plant Sci, 2016, 7: 812.
doi: 10.3389/fpls.2016.00812 pmid: 27375664 |
| [25] |
Campa A, Ferreira JJ. Gene coding for an elongation factor is involved in resistance against powdery mildew in common bean[J]. Theor Appl Genet, 2017, 130(5): 849-860.
doi: 10.1007/s00122-017-2864-x pmid: 28233030 |
| [26] | Wei YY, Liang S, Zhang YR, et al. MoSec61β, the beta subunit of Sec61, is involved in fungal development and pathogenicity, plant immunity, and ER-phagy in Magnaporthe oryzae[J]. Virulence, 2020, 11(1): 1685-1700. |
| [27] |
Giner A, Pascual L, Bourgeois M, et al. A mutation in the melon Vacuolar Protein Sorting 41prevents systemic infection of Cucumber Mosaic Virus[J]. Sci Rep, 2017, 7(1): 10471.
doi: 10.1038/s41598-017-10783-3 pmid: 28874719 |
| [28] | Wu JT, Gao T, Hu JN, et al. Research advances in function and regulation mechanisms of plant small heat shock proteins(sHSPs)under environmental stresses[J]. Sci Total Environ, 2022, 825: 154054. |
| [29] | Li Q, Qin XJ, Qi JJ, et al. CsPrx25, a class III peroxidase in Citrus sinensis, confers resistance to Citrus bacterial canker through the maintenance of ROS homeostasis and cell wall lignification[J]. Hortic Res, 2020, 7(1): 192. |
| [30] |
Jiang ZH, Jin XJ, Yang M, et al. Barley stripe mosaic virus γb protein targets thioredoxin h-type 1 to dampen salicylic acid-mediated defenses[J]. Plant Physiol, 2022, 189(3): 1715-1727.
doi: 10.1093/plphys/kiac137 pmid: 35325212 |
| [31] | 牟彦洁. 拟南芥水通道蛋白AtPIP1;4和AtPIP2;4转运H2O2调控抗病性的分子机制[D]. 泰安: 山东农业大学, 2022. |
| Mu YJ. Arabidopsis thaliana aquaporin AtPIP1; 4 and AtPIP2; Molecular mechanism of H2O2 transporting 4 to regulate disease resistance[D]. Tai'an: Shandong Agricultural University, 2022. | |
| [32] | Yang H, Luo PG. Changes in photosynthesis could provide important insight into the interaction between wheat and fungal pathogens[J]. Int J Mol Sci, 2021, 22(16): 8865. |
| [1] | 邢丽南, 张艳芳, 葛明然, 赵令敏, 陈妍, 霍秀文. 山药DoWRKY40基因表达特征分析及互作蛋白筛选[J]. 生物技术通报, 2024, 40(8): 118-128. |
| [2] | 王秋月, 段鹏亮, 李海笑, 刘宁, 曹志艳, 董金皋. 玉米大斑病菌cDNA文库的构建及转录因子StMR1互作蛋白的筛选[J]. 生物技术通报, 2024, 40(6): 281-289. |
| [3] | 辛奇, 李压凡, 尹铮, 张晓丹, 陈霆, 刘晓华. 甘蔗CBL-CIPK基因家族的鉴定和表达分析[J]. 生物技术通报, 2024, 40(2): 197-211. |
| [4] | 任延靖, 张鲁刚, 赵孟良, 李江, 邵登魁. 白菜种子cDNA酵母文库的构建及BrTTG1互作蛋白的筛选及分析[J]. 生物技术通报, 2024, 40(2): 223-232. |
| [5] | 王子颖, 龙晨洁, 范兆宇, 张蕾. 利用酵母双杂交系统筛选水稻中与OsCRK5互作蛋白[J]. 生物技术通报, 2023, 39(9): 117-125. |
| [6] | 温晓蕾, 李建嫄, 李娜, 张娜, 杨文香. 小麦叶锈菌与小麦互作的酵母双杂交cDNA文库构建与应用[J]. 生物技术通报, 2023, 39(9): 136-146. |
| [7] | 韩浩章, 张丽华, 李素华, 赵荣, 王芳, 王晓立. 盐碱胁迫诱导的猴樟酵母cDNA文库构建及CbP5CS上游调控因子筛选[J]. 生物技术通报, 2023, 39(9): 236-245. |
| [8] | 王艺清, 王涛, 韦朝领, 戴浩民, 曹士先, 孙威江, 曾雯. 茶树SMAS基因家族的鉴定及互作分析[J]. 生物技术通报, 2023, 39(4): 246-258. |
| [9] | 侯筱媛, 车郑郑, 李姮静, 杜崇玉, 胥倩, 王群青. 大豆膜系统cDNA文库的构建及大豆疫霉效应子PsAvr3a互作蛋白的筛选[J]. 生物技术通报, 2023, 39(4): 268-276. |
| [10] | 王涛, 漆思雨, 韦朝领, 王艺清, 戴浩民, 周喆, 曹士先, 曾雯, 孙威江. CsPPR和CsCPN60-like在茶树白化叶片中的表达分析及互作蛋白验证[J]. 生物技术通报, 2023, 39(3): 218-231. |
| [11] | 蔡佳, 梁振宇, 黄瑜, 鲁义善, 施钢, 简纪常. 利用酵母双杂交系统筛选与鉴定石斑鱼EcBAG3互作因子[J]. 生物技术通报, 2022, 38(8): 77-83. |
| [12] | 刘娟, 朱春晓, 肖雪琼, 莫陈汨, 王高峰, 肖炎农. 淡紫紫孢菌亲环蛋白PlCYP6 互作蛋白的筛选[J]. 生物技术通报, 2021, 37(7): 137-145. |
| [13] | 杨华杰, 周玉萍, 范甜, 吕天晓, 谢楚萍, 田长恩. 拟南芥IQM4互作蛋白的筛选和鉴定[J]. 生物技术通报, 2021, 37(11): 190-196. |
| [14] | 张桐, 李智强, 伍国强. WRKY转录因子在植物逆境响应中的作用[J]. 生物技术通报, 2021, 37(10): 203-215. |
| [15] | 郝小花, 戴佳利, 暨文劲, 黄丹, 李东屏, 田连福. 水稻籽粒低镉蛋白LCD互作蛋白的筛选与鉴定[J]. 生物技术通报, 2020, 36(11): 21-29. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||