








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 187-202.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1227
刘淼1( ), 林涛2, 贾乐松1, 胡丰1, 李涛1, 李志万1, 刘美芳1, 郑方燕1(
), 林涛2, 贾乐松1, 胡丰1, 李涛1, 李志万1, 刘美芳1, 郑方燕1( ), 崔龙1(
), 崔龙1( )
)
收稿日期:2025-11-14
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
崔龙,男,博士,副研究员,研究方向 :蔬菜学;E-mail: jingling5161@126.com作者简介:刘淼,男,硕士,研究方向:生物信息学;E-mail: 2033251151@qq.com基金资助:
LIU Miao1( ), LIN Tao2, JIA Le-song1, HU Feng1, LI Tao1, LI Zhi-wan1, LIU Mei-fang1, ZHENG Fang-yan1(
), LIN Tao2, JIA Le-song1, HU Feng1, LI Tao1, LI Zhi-wan1, LIU Mei-fang1, ZHENG Fang-yan1( ), CUI Long1(
), CUI Long1( )
)
Received:2025-11-14
Published:2026-03-26
Online:2026-04-23
摘要:
番茄(Solanum lycopersicum)果实在野生种到栽培种的演化与驯化过程中逐渐呈现出色泽多样性。野生番茄果实的颜色特征(如绿色或黄色)是其长期适应性演化的选择结果,赋予植株关键生态适应能力;而栽培番茄中丰富的颜色变异则主要受到人工驯化过程中人类选择偏好的影响。本文系统梳理了番茄果实色泽形成的遗传调控机制,重点聚焦于类胡萝卜素、叶绿素以及花青素三大核心代谢途径中关键基因的调控网络,深入阐释了自然选择与人工选择在驱动果实颜色多样性形成过程中的交互作用。目前,对番茄果实色泽形成的机理研究已形成较为完整且系统的认知体系。调控番茄红色、黄色、粉色、绿色及紫色等主要果实色泽表型的主效基因已基本明确,相关研究正从单一结构基因的功能解析,逐步拓展至涵盖RIN、NOR等成熟调控因子,以及MYB、bHLH等家族转录因子在内的复杂调控网络的系统解析。多组学技术的整合应用,进一步阐明了果实色泽形成过程中的多层次动态调控机制。现代育种技术的快速发展,特别是基因编辑技术的应用,进一步拓宽了果实颜色的变异范围。此外,色泽形成与糖、酸、香气等重要品质性状之间的协同调控日益受到关注,这表明果实颜色不仅是单纯的视觉特征,更是衡量果实综合品质的重要指示性状。未来研究可从分子调控、多性状耦合、育种转化等多维度系统推进,构建从基础理论到育种应用的全链条研究体系。
刘淼, 林涛, 贾乐松, 胡丰, 李涛, 李志万, 刘美芳, 郑方燕, 崔龙. 从野生到栽培:番茄果实色泽的演化与调控机制[J]. 生物技术通报, 2026, 42(3): 187-202.
LIU Miao, LIN Tao, JIA Le-song, HU Feng, LI Tao, LI Zhi-wan, LIU Mei-fang, ZHENG Fang-yan, CUI Long. From Wild to Cultivated: Evolution and Regulatory Mechanisms of Tomato Fruit Color[J]. Biotechnology Bulletin, 2026, 42(3): 187-202.
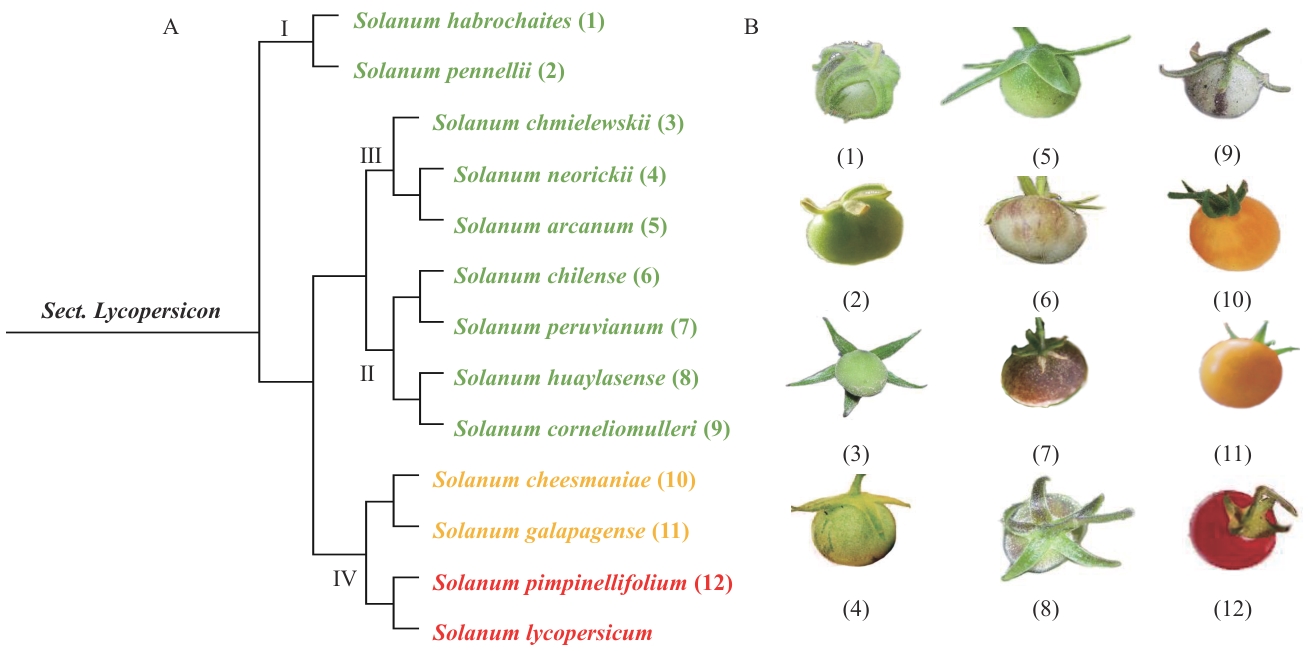
图1 野生番茄的进化分类与果实颜色A:番茄(Sect. Lycopersicon)系统发育树,该发育树呈现了不同番茄物种的演化分支关系,共分为Ⅰ、Ⅱ、Ⅲ、Ⅳ 4个主要分支类群;B:番茄属不同物种果实的表型图,按照序号顺序对应了发育树中的果实表型,展示了不同演化分支物种的果实表型特征。番茄果实图片来自网站https://tomato.international/w/index.php/メインページ
Fig. 1 Evolutionary classification and fruit color of wild tomatoes (Solanum lycopersicum)A: Phylogenetic tree of tomato (Sect. Lycopersicon). This phylogenetic tree illustrates the evolutionary branching relationships among different tomato species, divided into four major clades: I, II, Ⅲ, and Ⅳ. B: Phenotypic images of fruits from different tomato species, arranged in sequential order, correspond to the fruit phenotypes in the phylogenetic tree, illustrating the phenotypic characteristics of fruits from different evolutionary clades. The tomato fruit pictures are sourced from the website:https://tomato.international/w/index.php/メインページ
物种名称 Species name | 果实直径 Fruit diameter (cm) | 心室数 Number of locules | 成熟时果皮颜色 Skin color when ripe | 条纹(纵贯) Stripes (vertical) |
|---|---|---|---|---|
| S. habrochaites | 1‒1.5 | 2 | 均匀淡紫色 | 深绿色 |
| S. pennellii | 1‒1.3 | 2 | 均匀绿色 | 无 |
| S. chmielewskii | 1‒1.3 | 2 | 绿色 | 深绿色 |
| S. neorickii | 1‒1.3 | 2 | 绿色 | 深绿色 |
| S. arcanum | 1‒1.4 | 2 | 绿色 | 深绿色变紫色 |
| S. chilense | 1‒1.5 | 2‒5 | 绿白色 | 紫色 |
| S. peruvianum | 1‒1.5 | 2 | 白紫色/紫色 | 绿色或紫色 |
| S. huaylasense | 1‒1.4 | 2 | 绿色 | 深绿色或紫色 |
| S. corneliomulleri | 0.9‒1.3 | 2 | 绿色/绿白色 | 深绿色或紫色 |
| S. cheesmaniae | 0.6‒1.4 | 2 | 黄色/橙色 | 无 |
| S. galapagense | 0.6‒1.1 | 2 | 橙色 | 无 |
| S. pimpinellifolium | ≈1 | 2 | 鲜红色 | 无 |
| S. lycopersicum | 1.5‒10+ | 通常≥2 | 通常红色 | 通常无 |
表1 番茄属野生种与栽培种果实形态特征
Table 1 Fruit morphological characteristics of wild and cultivated species in the genus Solanum
物种名称 Species name | 果实直径 Fruit diameter (cm) | 心室数 Number of locules | 成熟时果皮颜色 Skin color when ripe | 条纹(纵贯) Stripes (vertical) |
|---|---|---|---|---|
| S. habrochaites | 1‒1.5 | 2 | 均匀淡紫色 | 深绿色 |
| S. pennellii | 1‒1.3 | 2 | 均匀绿色 | 无 |
| S. chmielewskii | 1‒1.3 | 2 | 绿色 | 深绿色 |
| S. neorickii | 1‒1.3 | 2 | 绿色 | 深绿色 |
| S. arcanum | 1‒1.4 | 2 | 绿色 | 深绿色变紫色 |
| S. chilense | 1‒1.5 | 2‒5 | 绿白色 | 紫色 |
| S. peruvianum | 1‒1.5 | 2 | 白紫色/紫色 | 绿色或紫色 |
| S. huaylasense | 1‒1.4 | 2 | 绿色 | 深绿色或紫色 |
| S. corneliomulleri | 0.9‒1.3 | 2 | 绿色/绿白色 | 深绿色或紫色 |
| S. cheesmaniae | 0.6‒1.4 | 2 | 黄色/橙色 | 无 |
| S. galapagense | 0.6‒1.1 | 2 | 橙色 | 无 |
| S. pimpinellifolium | ≈1 | 2 | 鲜红色 | 无 |
| S. lycopersicum | 1.5‒10+ | 通常≥2 | 通常红色 | 通常无 |
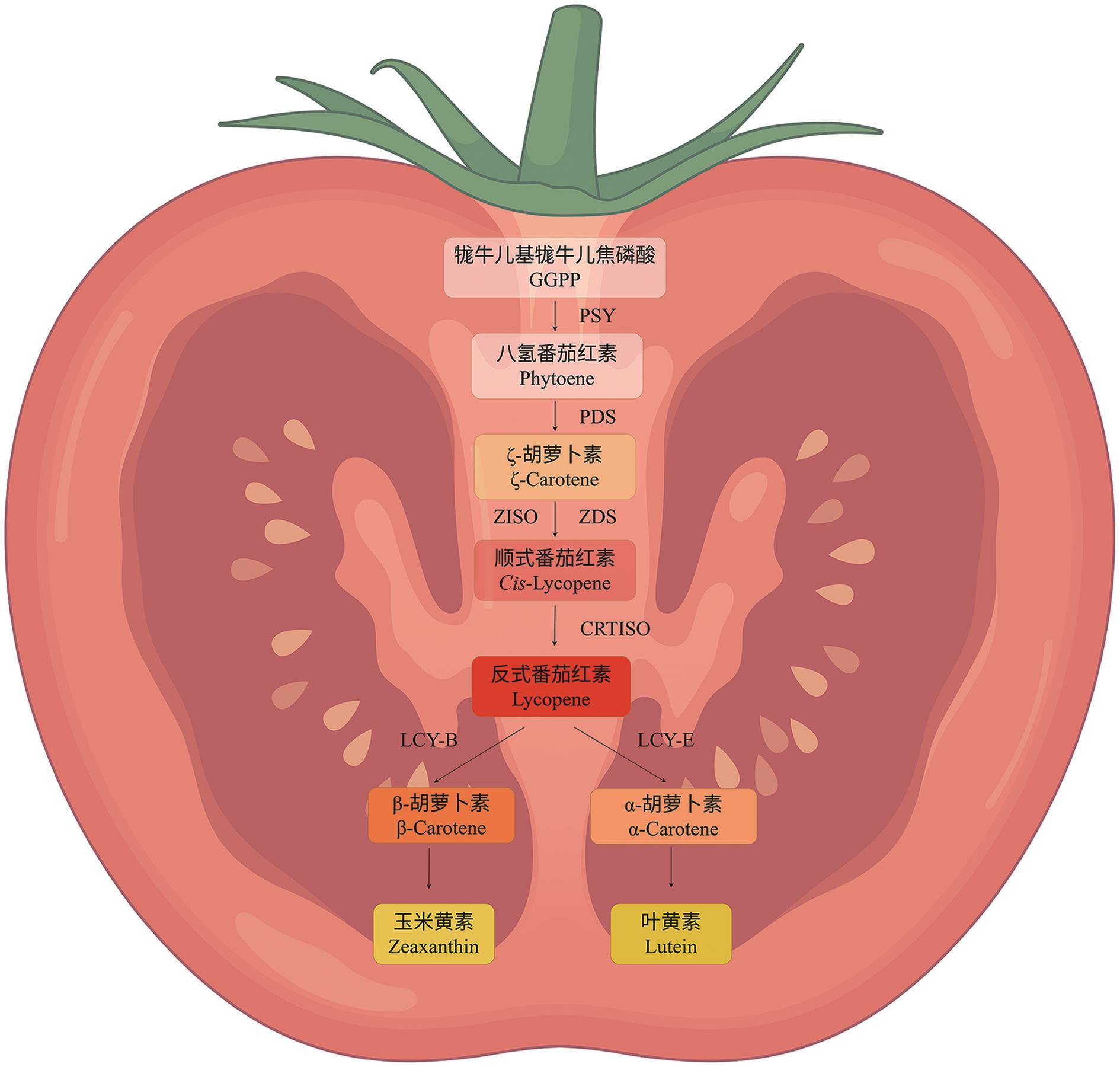
图2 番茄类胡萝卜素合成途径类胡萝卜素生物合成途径示意图,从GGPP开始,经PSY、PDS、ZISO、ZDS、CRTISO等一系列酶促反应,最终分支生成β-胡萝卜素和α-胡萝卜素的过程。本文图均由Figdraw绘制
Fig. 2 Biosynthetic pathway of tomato carotenoidSchematic of the carotenoid biosynthetic pathway. Starting from GGPP, a series of enzyme-catalyzed reactions occur via PSY, PDS, ZISO, ZDS, and CRTISO. The pathway eventually branches to produce β-carotene and α-carotene. Figures in this article are all created by Figdraw
基因 Gene | 作用机制与功能 Mechanisms of action and functions | 导致的表型 Resulting phenotype | 参考文献 Reference |
|---|---|---|---|
| NOR | NAC家族转录因子,促进类胡萝卜素积累,调控乙烯合成 | 突变后果实不成熟,呈橙色 | [ |
| CNR | SBP-box转录因子,调控成熟启动,影响类胡萝卜素积累 | 突变后果实不成熟,颜色为黄色或绿色 | [ |
| RIN | MADS-box转录因子,正向调控成熟过程,促进乙烯合成和类胡萝卜素积累 | 突变后果实不成熟,颜色为黄色或绿色 | [ |
| SlAP2a | 果实成熟的负调控因子,影响类胡萝卜素合成 | 果实呈橙色,果肉颜色变浅 | [ |
| TAGL1 | 表达下调,促进叶绿素合成和叶绿体发育,同时抑制类胡萝卜素积累 | 未成熟果实出现绿色条纹,成熟后为黄色条纹 | [ |
| SlMYB72 | R2R3-MYB转录因子,负调控叶绿素合成,正调控类胡萝卜素和类黄酮代谢 | 下调表达导致果实出现深绿色斑点(绿果期)和黄斑(红果期) | [ |
| HY5 | 光信号通路转录因子,促进类胡萝卜素和花青素积累 | 突变后果实颜色变浅,呈浅红色 | [ |
| SlIDI1/YFT3 | 在质体MEP途径中催化IPP和DMAPP的可逆异构化,为类胡萝卜素合成提供必需的前体底物。类异戊二烯代谢通路中的关键酶 | 功能丧失突变导致酶失活,DMAPP前体严重匮乏,番茄红素合成几乎完全阻断,果实呈黄色。过表达则提供更充足前体,果实红色加深 | [ |
| SlMCT | MEP途径中的关键酶,萜类化合物生物合成的第2步。该基因突变导致酶功能异常,影响下游类胡萝卜素和激素合成 | 类胡萝卜素和叶绿素含量显著降低,果实变黄 | [ |
| PSY1 | 类胡萝卜素合成的限速酶,催化番茄红素合成第一步 | 敲除果实呈黄绿色 | [ |
| PDS | 类胡萝卜素生物合成中的一种关键酶,可将植物烯转化为ζ-胡萝卜素 | PDS沉默的果实出现黄色表型 | [ |
| ZDS | 在类胡萝卜素合成途径中,负责将ζ-胡萝卜素转化为链孢红素,是生成番茄红素的关键步骤之一 | ZDS功能缺失导致果实呈黄色/橙色 | [ |
| CRTISO | 催化胡萝卜素去饱和过程中的顺反异构化,合成全反式番茄红素和下游环化类胡萝卜素(如β-胡萝卜素、叶黄素)的关键酶 | 突变果实呈橙色 | [ |
| CYC-B | 编码番茄红素β-环化酶,将番茄红素转化为β-胡萝卜素 | 功能获得突变使β-胡萝卜素大增,果肉呈橙色 | [ |
| og/ogc | beta基因的无效等位基因。发生移码突变,无法产生有功能的番茄红素β-环化酶 | 果实呈深红色 | [ |
| SlCCD4b | 催化类胡萝卜素在C9-C10位点裂解,生成挥发性芳香物质;负调控番茄果实中β-胡萝卜素和番茄红素的积累 | 过表达导致果实呈深橙色 | [ |
| ClpR1 | 敲低ClpR1会削弱Clp蛋白酶活性,导致蛋白质降解受阻,引发蛋白质折叠应激;进而上调ClpB3和OR/OR-like等分子伴侣基因表达,促进DXS和PSY酶稳定性,增加β-胡萝卜素积累,同时抑制番茄红素积累 | 沉默导致成熟果实呈橙色 | [ |
| Delta | 催化番茄红素生成δ-胡萝卜素 | 突变体成熟果实为橙色 | [ |
| WRKY32 | RNAi生成品系的番茄果实中,乙烯信号传导减少,导致乙烯释放受到抑制,染色体发育延迟,类胡萝卜素积累减少 | 果实表型呈黄色 | [ |
| YFT1 | 调控乙烯合成、信号转导、质体发育和类胡萝卜素积累 | 突变体果实呈黄色 | [ |
| SNAC9 | 正向调控番茄果实成熟过程中的类胡萝卜素代谢 | 敲除后果实成熟延迟,果实颜色变浅 | [ |
| SlNAP7 | 影响番茄叶绿体发育和番茄红素积累 | 沉默使得果皮和果肉呈现黄色 | [ |
| DXS1 | 催化MEP途径第一步,合成质体类异戊二烯的关键限速酶 | 绿熟期果实呈白色,红熟期颜色变浅 | [ |
| CRY1a | 促进果实中类胡萝卜素的积累 | 过表达果实颜色加深,红熟期呈深红色;敲除果实颜色变浅 | [ |
| GH | 质体末端氧化酶,参与类胡萝卜素合成 | 突变体果实为橙黄色 | [ |
| hp-3 | 催化玉米黄质向紫黄质的转化,影响类胡萝卜素(尤其是番茄红素)的合成与储存能力 | 果实颜色加深(红色增强) | [ |
| SlBEL11 | 作为转录抑制因子,直接结合下游基因番茄红素β-环化酶2的启动子,抑制其转录 | 沉默时果实成熟时呈黄色 | [ |
| SlCHRC | 色粒体相关类胡萝卜素结合蛋白(CHRC)调节色粒体发育和类胡萝卜素积累 | 敲除导致果实成熟延迟,类胡萝卜素含量降低,果实表型更绿 | [ |
| SlDML2 | DNA去甲基化酶,影响类胡萝卜素(如β-胡萝卜素和番茄红素)积累 | 突变体果实保持绿色 | [ |
SlBBX20 (Solyc01g110180) | 促进类胡萝卜素合成 | 过表达使未成熟果实呈深绿色,成熟果实颜色加深 | [ |
| SlCMB1 | MADS-box转录因子,正调控果实成熟;促进乙烯合成和类胡萝卜素积累;与RIN、TAGL1等蛋白互作 | 抑制后果实成熟延迟,颜色变浅,呈浅橙或橙色 | [ |
表2 类胡萝卜素相关调控基因
Table 2 Carotenoid-related regulatory genes
基因 Gene | 作用机制与功能 Mechanisms of action and functions | 导致的表型 Resulting phenotype | 参考文献 Reference |
|---|---|---|---|
| NOR | NAC家族转录因子,促进类胡萝卜素积累,调控乙烯合成 | 突变后果实不成熟,呈橙色 | [ |
| CNR | SBP-box转录因子,调控成熟启动,影响类胡萝卜素积累 | 突变后果实不成熟,颜色为黄色或绿色 | [ |
| RIN | MADS-box转录因子,正向调控成熟过程,促进乙烯合成和类胡萝卜素积累 | 突变后果实不成熟,颜色为黄色或绿色 | [ |
| SlAP2a | 果实成熟的负调控因子,影响类胡萝卜素合成 | 果实呈橙色,果肉颜色变浅 | [ |
| TAGL1 | 表达下调,促进叶绿素合成和叶绿体发育,同时抑制类胡萝卜素积累 | 未成熟果实出现绿色条纹,成熟后为黄色条纹 | [ |
| SlMYB72 | R2R3-MYB转录因子,负调控叶绿素合成,正调控类胡萝卜素和类黄酮代谢 | 下调表达导致果实出现深绿色斑点(绿果期)和黄斑(红果期) | [ |
| HY5 | 光信号通路转录因子,促进类胡萝卜素和花青素积累 | 突变后果实颜色变浅,呈浅红色 | [ |
| SlIDI1/YFT3 | 在质体MEP途径中催化IPP和DMAPP的可逆异构化,为类胡萝卜素合成提供必需的前体底物。类异戊二烯代谢通路中的关键酶 | 功能丧失突变导致酶失活,DMAPP前体严重匮乏,番茄红素合成几乎完全阻断,果实呈黄色。过表达则提供更充足前体,果实红色加深 | [ |
| SlMCT | MEP途径中的关键酶,萜类化合物生物合成的第2步。该基因突变导致酶功能异常,影响下游类胡萝卜素和激素合成 | 类胡萝卜素和叶绿素含量显著降低,果实变黄 | [ |
| PSY1 | 类胡萝卜素合成的限速酶,催化番茄红素合成第一步 | 敲除果实呈黄绿色 | [ |
| PDS | 类胡萝卜素生物合成中的一种关键酶,可将植物烯转化为ζ-胡萝卜素 | PDS沉默的果实出现黄色表型 | [ |
| ZDS | 在类胡萝卜素合成途径中,负责将ζ-胡萝卜素转化为链孢红素,是生成番茄红素的关键步骤之一 | ZDS功能缺失导致果实呈黄色/橙色 | [ |
| CRTISO | 催化胡萝卜素去饱和过程中的顺反异构化,合成全反式番茄红素和下游环化类胡萝卜素(如β-胡萝卜素、叶黄素)的关键酶 | 突变果实呈橙色 | [ |
| CYC-B | 编码番茄红素β-环化酶,将番茄红素转化为β-胡萝卜素 | 功能获得突变使β-胡萝卜素大增,果肉呈橙色 | [ |
| og/ogc | beta基因的无效等位基因。发生移码突变,无法产生有功能的番茄红素β-环化酶 | 果实呈深红色 | [ |
| SlCCD4b | 催化类胡萝卜素在C9-C10位点裂解,生成挥发性芳香物质;负调控番茄果实中β-胡萝卜素和番茄红素的积累 | 过表达导致果实呈深橙色 | [ |
| ClpR1 | 敲低ClpR1会削弱Clp蛋白酶活性,导致蛋白质降解受阻,引发蛋白质折叠应激;进而上调ClpB3和OR/OR-like等分子伴侣基因表达,促进DXS和PSY酶稳定性,增加β-胡萝卜素积累,同时抑制番茄红素积累 | 沉默导致成熟果实呈橙色 | [ |
| Delta | 催化番茄红素生成δ-胡萝卜素 | 突变体成熟果实为橙色 | [ |
| WRKY32 | RNAi生成品系的番茄果实中,乙烯信号传导减少,导致乙烯释放受到抑制,染色体发育延迟,类胡萝卜素积累减少 | 果实表型呈黄色 | [ |
| YFT1 | 调控乙烯合成、信号转导、质体发育和类胡萝卜素积累 | 突变体果实呈黄色 | [ |
| SNAC9 | 正向调控番茄果实成熟过程中的类胡萝卜素代谢 | 敲除后果实成熟延迟,果实颜色变浅 | [ |
| SlNAP7 | 影响番茄叶绿体发育和番茄红素积累 | 沉默使得果皮和果肉呈现黄色 | [ |
| DXS1 | 催化MEP途径第一步,合成质体类异戊二烯的关键限速酶 | 绿熟期果实呈白色,红熟期颜色变浅 | [ |
| CRY1a | 促进果实中类胡萝卜素的积累 | 过表达果实颜色加深,红熟期呈深红色;敲除果实颜色变浅 | [ |
| GH | 质体末端氧化酶,参与类胡萝卜素合成 | 突变体果实为橙黄色 | [ |
| hp-3 | 催化玉米黄质向紫黄质的转化,影响类胡萝卜素(尤其是番茄红素)的合成与储存能力 | 果实颜色加深(红色增强) | [ |
| SlBEL11 | 作为转录抑制因子,直接结合下游基因番茄红素β-环化酶2的启动子,抑制其转录 | 沉默时果实成熟时呈黄色 | [ |
| SlCHRC | 色粒体相关类胡萝卜素结合蛋白(CHRC)调节色粒体发育和类胡萝卜素积累 | 敲除导致果实成熟延迟,类胡萝卜素含量降低,果实表型更绿 | [ |
| SlDML2 | DNA去甲基化酶,影响类胡萝卜素(如β-胡萝卜素和番茄红素)积累 | 突变体果实保持绿色 | [ |
SlBBX20 (Solyc01g110180) | 促进类胡萝卜素合成 | 过表达使未成熟果实呈深绿色,成熟果实颜色加深 | [ |
| SlCMB1 | MADS-box转录因子,正调控果实成熟;促进乙烯合成和类胡萝卜素积累;与RIN、TAGL1等蛋白互作 | 抑制后果实成熟延迟,颜色变浅,呈浅橙或橙色 | [ |
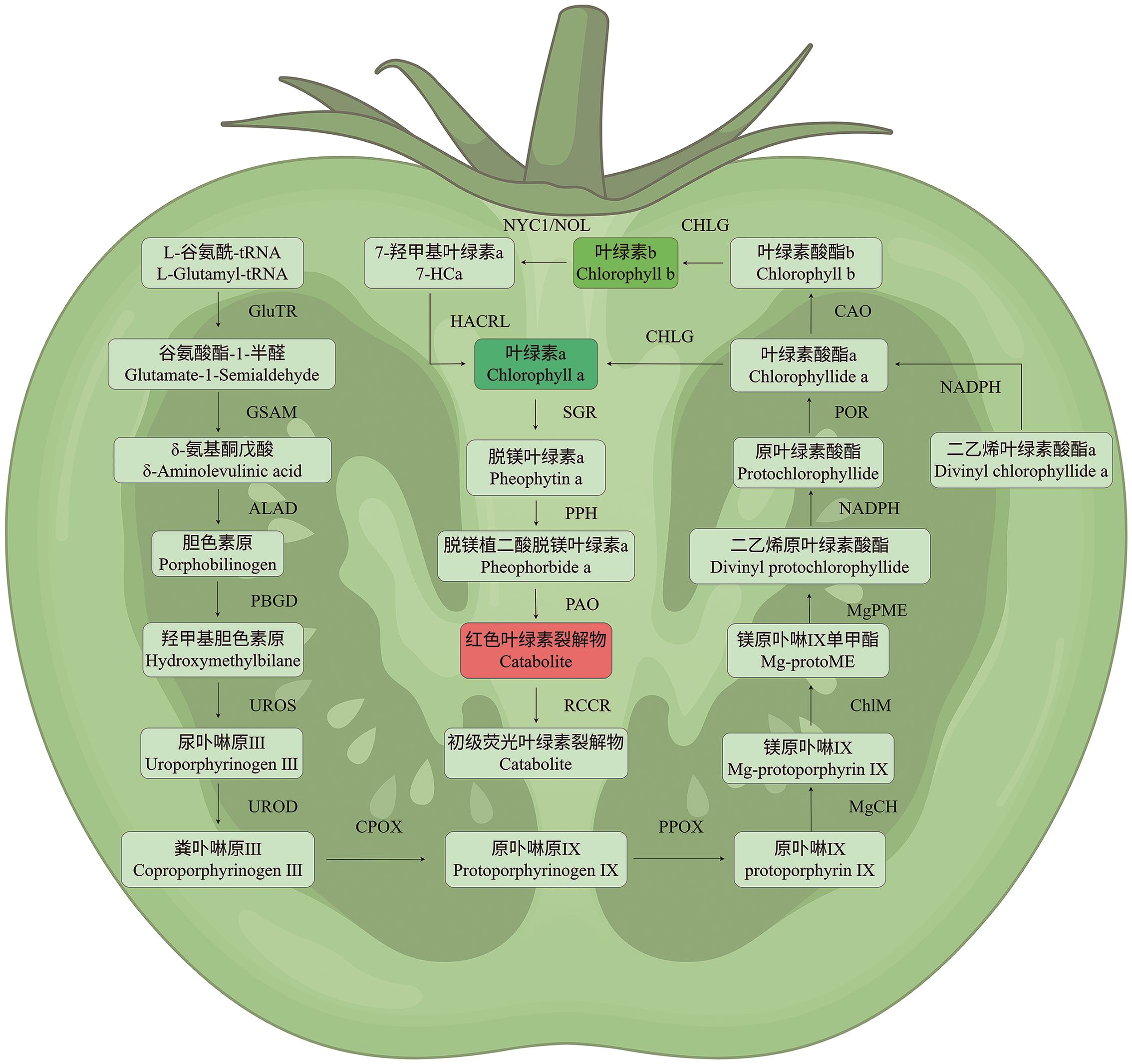
图3 叶绿素代谢途径植物光合色素的核心代谢网络,叶绿素从起始物质L-谷氨酰-tRNA开始的完整生物合成过程,涉及多步酶促反应及关键中间产物,最终生成叶绿素a和叶绿素b。随后,叶绿素b依次经过NYCI/NOL、HACR、SGR、PPH、PAO、RCCR等酶的催化作用,最终形成初级荧光叶绿素裂解物
Fig. 3 Chlorophyll metabolic pathwayThe core metabolic network of plant photosynthetic pigments: The complete biosynthetic process of chlorophyll starting from the precursor L-glutamyl-tRNA involves multiple enzymatic steps and key intermediate products, ultimately yielding chlorophyll a and chlorophyll b. Subsequently, chlorophyll b undergoes catalysis by enzymes such as NYCI/NOL, HACR, SGR, PPH, PAO, and RCCR, ultimately forming the primary fluorescent chlorophyll catabolite
基因 Gene | 作用机制与功能 Mechanisms of action and functions | 导致的表型 Resulting phenotype | 参考文献 Reference |
|---|---|---|---|
| SlSGR1 | 促进果实成熟过程中叶绿素的降解 | 突变后果实保持绿色,成熟后呈褐色 | [ |
| SlGLK2 | 调控叶绿体发育,影响叶绿素分布 | 过表达果实肩部持续深绿色,形成“内绿果”;功能缺失导致果实肩部绿色变浅 | [ |
| SlBEL2 | 负调控叶绿体发育和叶绿素合成,影响果实绿色肩部形成 | 过表达果肩绿色消失,果实整体呈浅绿色;敲除SlBEL2果肩绿色加深 | [ |
| SlZHD17 | 影响番茄果实叶绿体的发育,促进了叶绿素的积累 | 下调导致未成熟和绿熟阶段果实更绿,成熟时期明显色素沉着不均匀 | [ |
| SlARF10 | 正向调控一些叶绿素合成相关基因表达,促进叶绿体发育和光合作用 | 过表达后果实在绿熟期呈深绿色;RNAi敲低后果实在绿熟期呈浅绿色 | [ |
| SlBL4 | 负调控叶绿素合成与叶绿体发育;抑制细胞壁代谢相关基因 | 下调后果实颜色略深绿,成熟后果实着色不均,不完全变红 | [ |
| SlBEL11 | 负调控叶绿素合成与叶绿体发育 | 沉默后未熟果实叶绿素显著积累,果色深绿;成熟延迟,出现斑驳绿粉相间表型 | [ |
| SlAPRR2-LIKE | 调控质体数目与面积,影响叶绿素与类胡萝卜素积累 | 过表达增加质体数与色素含量,果色深绿 | [ |
| TKN2 | Class I KNOX转录因子;正调控叶绿体发育 | 功能获得性突变体果实叶绿素含量增加,果色深绿且均匀 | [ |
| TKN4 | Class I KNOX转录因子;正调控叶绿体发育梯度 | 功能缺失突变体(ug)果实肩部绿色缺失,成熟后果实颜色更均匀 | [ |
| HP1 | 负调控光形态建成;突变导致叶绿体和质体区室增大 | 突变体叶绿素和类胡萝卜素含量增加,果色更深 | [ |
| DR12/SlARF4 | 属于ARF转录因子家族,参与生长素响应;抑制其表达导致叶绿素含量增加、叶绿体数量增多、果实成熟不均匀 | 未熟果实呈深绿色;成熟果实呈斑驳状 | [ |
| SlGRAS9 | 负调控叶绿素积累和叶绿体发育 | 敲除或敲低后果实呈深绿色 | [ |
| SlELP2L | 功能缺失会导致番茄果实叶绿素降解受阻 | 基因沉默(RNAi)后,番茄果实在未成熟阶段呈现深绿色 | [ |
| TDET1/HP-2 | 负调控光形态建成和光信号传导;影响绿体和花青素积累 | 突变导致未成熟果实深绿色 | [ |
| SlDXR | MEP途径限速酶,调控类胡萝卜素、叶绿素、类黄酮合成;影响叶绿体发育 | 突变体果实呈橙黄色,果肉为黄/白色 | [ |
| SlMYB72 | R2R3-MYB转录因子,负调控叶绿素合成,正调控类胡萝卜素和类黄酮代谢 | 下调表达导致果实出现深绿色斑点(绿果期)和黄斑(红果期) | [ |
| su | 调控质体分裂;突变导致叶片和绿色果实中质体数量减少、体积增大(巨型叶绿体) | su-1/su-3突变体的未成熟果实呈浅绿色 | [ |
| SlKN5 | 与BLH1-clade蛋白形成复合物,抑制叶绿体生物发生、叶绿素合成和赤霉素降解相关基因的表达;在果实发育早期抑制叶绿素积累 | 敲除导致未成熟果实颜色更深绿 | [ |
表3 叶绿素相关调控基因
Table 3 Chlorophyll-related regulatory genes
基因 Gene | 作用机制与功能 Mechanisms of action and functions | 导致的表型 Resulting phenotype | 参考文献 Reference |
|---|---|---|---|
| SlSGR1 | 促进果实成熟过程中叶绿素的降解 | 突变后果实保持绿色,成熟后呈褐色 | [ |
| SlGLK2 | 调控叶绿体发育,影响叶绿素分布 | 过表达果实肩部持续深绿色,形成“内绿果”;功能缺失导致果实肩部绿色变浅 | [ |
| SlBEL2 | 负调控叶绿体发育和叶绿素合成,影响果实绿色肩部形成 | 过表达果肩绿色消失,果实整体呈浅绿色;敲除SlBEL2果肩绿色加深 | [ |
| SlZHD17 | 影响番茄果实叶绿体的发育,促进了叶绿素的积累 | 下调导致未成熟和绿熟阶段果实更绿,成熟时期明显色素沉着不均匀 | [ |
| SlARF10 | 正向调控一些叶绿素合成相关基因表达,促进叶绿体发育和光合作用 | 过表达后果实在绿熟期呈深绿色;RNAi敲低后果实在绿熟期呈浅绿色 | [ |
| SlBL4 | 负调控叶绿素合成与叶绿体发育;抑制细胞壁代谢相关基因 | 下调后果实颜色略深绿,成熟后果实着色不均,不完全变红 | [ |
| SlBEL11 | 负调控叶绿素合成与叶绿体发育 | 沉默后未熟果实叶绿素显著积累,果色深绿;成熟延迟,出现斑驳绿粉相间表型 | [ |
| SlAPRR2-LIKE | 调控质体数目与面积,影响叶绿素与类胡萝卜素积累 | 过表达增加质体数与色素含量,果色深绿 | [ |
| TKN2 | Class I KNOX转录因子;正调控叶绿体发育 | 功能获得性突变体果实叶绿素含量增加,果色深绿且均匀 | [ |
| TKN4 | Class I KNOX转录因子;正调控叶绿体发育梯度 | 功能缺失突变体(ug)果实肩部绿色缺失,成熟后果实颜色更均匀 | [ |
| HP1 | 负调控光形态建成;突变导致叶绿体和质体区室增大 | 突变体叶绿素和类胡萝卜素含量增加,果色更深 | [ |
| DR12/SlARF4 | 属于ARF转录因子家族,参与生长素响应;抑制其表达导致叶绿素含量增加、叶绿体数量增多、果实成熟不均匀 | 未熟果实呈深绿色;成熟果实呈斑驳状 | [ |
| SlGRAS9 | 负调控叶绿素积累和叶绿体发育 | 敲除或敲低后果实呈深绿色 | [ |
| SlELP2L | 功能缺失会导致番茄果实叶绿素降解受阻 | 基因沉默(RNAi)后,番茄果实在未成熟阶段呈现深绿色 | [ |
| TDET1/HP-2 | 负调控光形态建成和光信号传导;影响绿体和花青素积累 | 突变导致未成熟果实深绿色 | [ |
| SlDXR | MEP途径限速酶,调控类胡萝卜素、叶绿素、类黄酮合成;影响叶绿体发育 | 突变体果实呈橙黄色,果肉为黄/白色 | [ |
| SlMYB72 | R2R3-MYB转录因子,负调控叶绿素合成,正调控类胡萝卜素和类黄酮代谢 | 下调表达导致果实出现深绿色斑点(绿果期)和黄斑(红果期) | [ |
| su | 调控质体分裂;突变导致叶片和绿色果实中质体数量减少、体积增大(巨型叶绿体) | su-1/su-3突变体的未成熟果实呈浅绿色 | [ |
| SlKN5 | 与BLH1-clade蛋白形成复合物,抑制叶绿体生物发生、叶绿素合成和赤霉素降解相关基因的表达;在果实发育早期抑制叶绿素积累 | 敲除导致未成熟果实颜色更深绿 | [ |
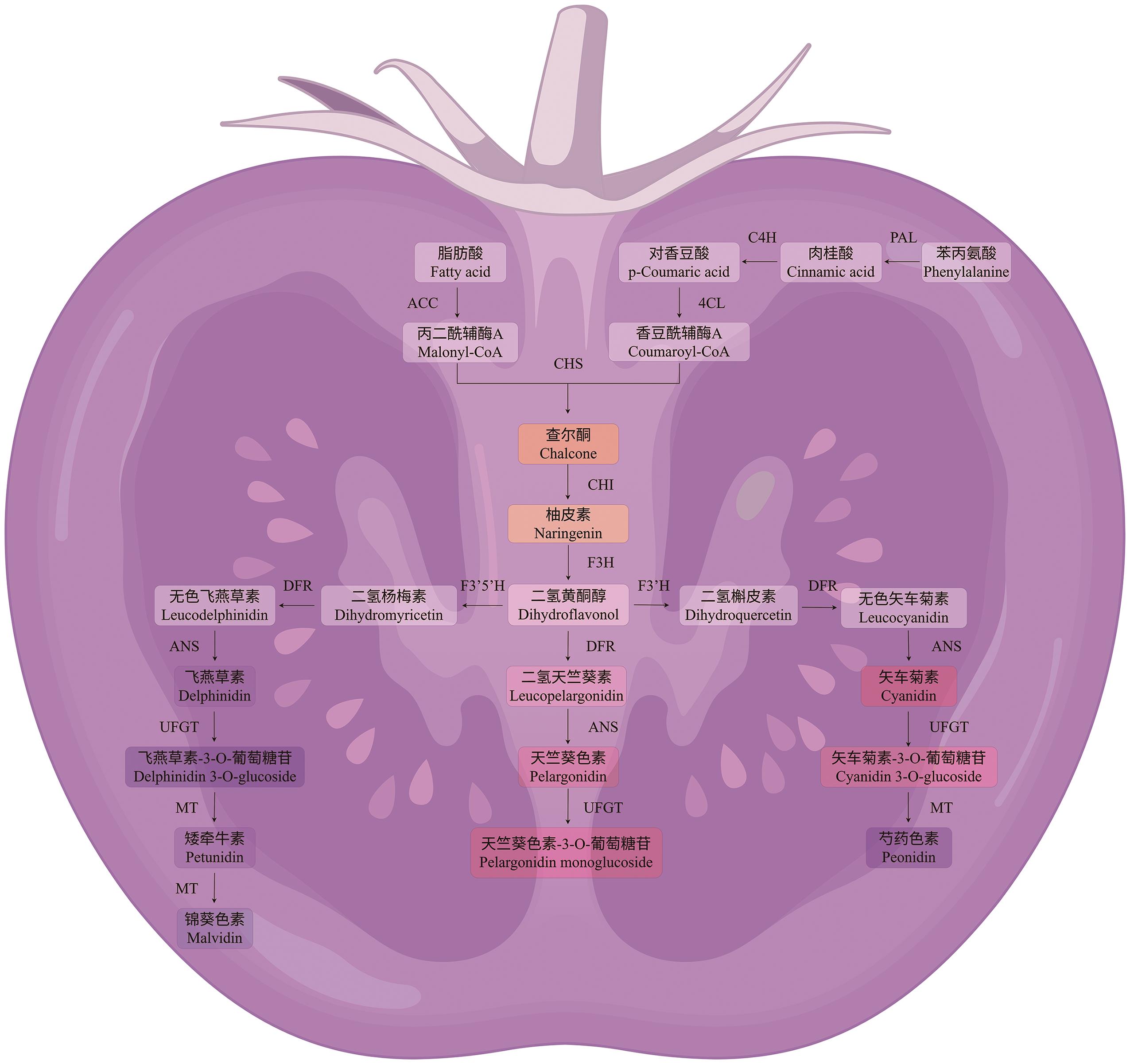
图4 番茄花青素合成途径黄酮类色素(飞燕草素、天竺葵素、矢车菊素及其衍生物)的生物合成过程,关键酶包括PAL、C4H、4CL、CHS、CHI、F3H、F3'5'H、F3'H、DFR、ANS、UFGT及甲基转移酶(MT),催化底物逐步转化为各类黄酮色素
Fig. 4 Anthocyanin biosynthesis pathway in tomatoThe biosynthesis of flavonoid pigments (delphinidin, pelargonidin, cyanidin, and their derivatives) involves the progressive conversion of catalytic substrates into various flavonoid pigments, with key enzymes including PAL, C4H, 4CL, CHS, CHI, F3H, F3'5'H, F3'H, DFR, ANS, UFGT, and methyltransferases (MT)
基因 Gene | 主要作用机制与功能 Mechanisms of action and functions | 导致的表型 Resulting phenotype | 参考文献 Reference |
|---|---|---|---|
| SlAN2-like | R2R3-MYB转录因子是花青素合成的关键主调控因子 | InR型:果皮紫色;过表达:果皮与果肉均紫色;敲除:紫色减少 | [ |
| SlJAF13 | 直接结合并激活SlAN1启动子(G-box基序),促进花青素合成;形成负反馈调控回路,精细调控花青素积累;是MBW复合体的关键成分 | 突变体果实无花青素积累,呈绿色;互补株系(SlJAF13-Com):恢复花青素积累,果皮呈紫色斑点 | [ |
| aft | 在果皮中激活花青素合成,受光照和低温诱导 | 果皮出现紫色斑点 | [ |
| Abg | SlAN2like的功能性等位基因,通过正确剪接激活花青素合成 | 可以从紫色斑点的果实到强烈均匀的乌黑色 | [ |
| atv | 编码SlMYB-ATV,花青素抑制因子,其功能丧失突变导致花青素合成增强。与Aft或Abg协同作用增强果皮花青素积累 | 单独存在时颜色变化不明显,与Aft或abg结合后果皮呈深紫色 | [ |
| SlANT1 | MYB转录因子,调控花青素合成、修饰与转运相关基因的表达,促进花青素积累 | 果实表皮出现紫色斑点 | [ |
| SlAN2(SlMYB75) | R2R3-MYB转录因子是番茄响应环境诱导花青素合成的主要调控因子 | 过表达使果实和果肉由红变橙黄;过表达导致番茄营养组织和果实中花青素显著积累,呈现紫色 | [ |
| SlMYB12 | 调控类黄酮合成途径,特别是柚皮素查尔酮的积累;表达下调导致果皮中黄色类黄酮缺失 | 敲除导致果皮透明,果实呈粉红色 | [ |
| CHS | 类黄酮合成路径第一步关键酶;催化柚皮素查尔酮形成 | 沉默导致果实色泽暗沉、个头较小且颜色更偏深红 | [ |
SlBBX20 (Solyc12g089240) | 正向调节水果类黄酮积累 | 突变体果实呈橙黄色 | [ |
| SlWRKY14 | 调节番茄果实色素沉着,调控类胡萝卜素和类黄酮代谢途径 | 突变体果实更红,过表达果实呈橙黄色 | [ |
| NF-YB | 参与类黄酮合成调控 | 沉默导致果皮无色,果实呈粉红色 | [ |
| SlMYB72 | R2R3-MYB转录因子,负调控叶绿素合成,正调控类胡萝卜素和类黄酮代谢 | 下调表达导致果实出现深绿色斑点(绿果期)和黄斑(红果期) | [ |
| HY5 | 光信号通路转录因子,促进类胡萝卜素和花青素积累 | 突变后果实颜色变浅,呈浅红色 | [ |
表4 类黄酮相关调控基因
Table 4 Flavonoid-related regulatory genes
基因 Gene | 主要作用机制与功能 Mechanisms of action and functions | 导致的表型 Resulting phenotype | 参考文献 Reference |
|---|---|---|---|
| SlAN2-like | R2R3-MYB转录因子是花青素合成的关键主调控因子 | InR型:果皮紫色;过表达:果皮与果肉均紫色;敲除:紫色减少 | [ |
| SlJAF13 | 直接结合并激活SlAN1启动子(G-box基序),促进花青素合成;形成负反馈调控回路,精细调控花青素积累;是MBW复合体的关键成分 | 突变体果实无花青素积累,呈绿色;互补株系(SlJAF13-Com):恢复花青素积累,果皮呈紫色斑点 | [ |
| aft | 在果皮中激活花青素合成,受光照和低温诱导 | 果皮出现紫色斑点 | [ |
| Abg | SlAN2like的功能性等位基因,通过正确剪接激活花青素合成 | 可以从紫色斑点的果实到强烈均匀的乌黑色 | [ |
| atv | 编码SlMYB-ATV,花青素抑制因子,其功能丧失突变导致花青素合成增强。与Aft或Abg协同作用增强果皮花青素积累 | 单独存在时颜色变化不明显,与Aft或abg结合后果皮呈深紫色 | [ |
| SlANT1 | MYB转录因子,调控花青素合成、修饰与转运相关基因的表达,促进花青素积累 | 果实表皮出现紫色斑点 | [ |
| SlAN2(SlMYB75) | R2R3-MYB转录因子是番茄响应环境诱导花青素合成的主要调控因子 | 过表达使果实和果肉由红变橙黄;过表达导致番茄营养组织和果实中花青素显著积累,呈现紫色 | [ |
| SlMYB12 | 调控类黄酮合成途径,特别是柚皮素查尔酮的积累;表达下调导致果皮中黄色类黄酮缺失 | 敲除导致果皮透明,果实呈粉红色 | [ |
| CHS | 类黄酮合成路径第一步关键酶;催化柚皮素查尔酮形成 | 沉默导致果实色泽暗沉、个头较小且颜色更偏深红 | [ |
SlBBX20 (Solyc12g089240) | 正向调节水果类黄酮积累 | 突变体果实呈橙黄色 | [ |
| SlWRKY14 | 调节番茄果实色素沉着,调控类胡萝卜素和类黄酮代谢途径 | 突变体果实更红,过表达果实呈橙黄色 | [ |
| NF-YB | 参与类黄酮合成调控 | 沉默导致果皮无色,果实呈粉红色 | [ |
| SlMYB72 | R2R3-MYB转录因子,负调控叶绿素合成,正调控类胡萝卜素和类黄酮代谢 | 下调表达导致果实出现深绿色斑点(绿果期)和黄斑(红果期) | [ |
| HY5 | 光信号通路转录因子,促进类胡萝卜素和花青素积累 | 突变后果实颜色变浅,呈浅红色 | [ |
| [1] | Jenkins JA. The origin of the cultivated tomato [J]. Econ Bot, 1948, 2(4): 379-392. |
| [2] | Blanca J, Cañizares J, Cordero L, et al. Variation revealed by SNP genotyping and morphology provides insight into the origin of the tomato [J]. PLoS One, 2012, 7(10): e48198. |
| [3] | Yoshiyama Y, Wakabayashi Y, Mercer KL, et al. Natural genetic variation in dynamic photosynthesis is correlated with stomatal anatomical traits in diverse tomato species across geographical habitats [J]. J Exp Bot, 2024, 75(21): 6762-6777. |
| [4] | van Andel T, Vos RA, Michels E, et al. Sixteenth-century tomatoes in Europe: who saw them, what they looked like, and where they came from [J]. PeerJ, 2022, 10: e12790. |
| [5] | 赵凌侠, 李景富. 番茄起源、传播及分类的回顾 [J]. 作物品种资源, 1999(3): 29-31. |
| Zhao LX, Li JF. Review on the origin, spread and classification of tomato [J]. China Seed Ind, 1999(3): 29-31. | |
| [6] | Mary F Willson CJW. The evolution of fruit color in fleshy-fruited plants [J]. Am Nat, 1990, 136(6): 790-809. |
| [7] | Kapoor L, Simkin AJ, George Priya Doss C, et al. Fruit ripening: dynamics and integrated analysis of carotenoids and anthocyanins [J]. BMC Plant Biol, 2022, 22(1): 27. |
| [8] | Zuriaga E, Blanca J, Nuez F. Classification and phylogenetic relationships in Solanum section Lycopersicon based on AFLP and two nuclear gene sequences [J]. Genet Resour Crop Evol, 2009, 56(5): 663-678. |
| [9] | Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution [J]. Nature, 2012, 485(7400): 635-641. |
| [10] | Lin T, Zhu GT, Zhang JH, et al. Genomic analyses provide insights into the history of tomato breeding [J]. Nat Genet, 2014, 46(11): 1220-1226. |
| [11] | Zhu GT, Wang SC, Huang ZJ, et al. Rewiring of the fruit metabolome in tomato breeding [J]. Cell, 2018, 172(1/2): 249-261.e12. |
| [12] | Gao L, Gonda I, Sun HH, et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor [J]. Nat Genet, 2019, 51(6): 1044-1051. |
| [13] | Alonge M, Wang XG, Benoit M, et al. Major impacts of widespread structural variation on gene expression and crop improvement in tomato [J]. Cell, 2020, 182(1): 145-161.e23. |
| [14] | Zhou Y, Zhang ZY, Bao ZG, et al. Graph pangenome captures missing heritability and empowers tomato breeding [J]. Nature, 2022, 606(7914): 527-534. |
| [15] | Yu XF, Qu MH, Shi YN, et al. Chromosome-scale genome assemblies of wild tomato relatives Solanum habrochaites and Solanum galapagense reveal structural variants associated with stress tolerance and terpene biosynthesis [J]. Hortic Res, 2022, 9: uhac139. |
| [16] | Li N, He Q, Wang J, et al. Super-pangenome analyses highlight genomic diversity and structural variation across wild and cultivated tomato species [J]. Nat Genet, 2023, 55(5): 852-860. |
| [17] | Delgado-Vargas F, Jiménez AR, Paredes-López O. Natural pigments: carotenoids, anthocyanins, and betalains—characteristics, biosynthesis, processing, and stability [J]. Crit Rev Food Sci Nutr, 2000, 40(3): 173-289. |
| [18] | Rosati C, Diretto G, Giuliano G. Biosynthesis and engineering of carotenoids and apocarotenoids in plants: state of the art and future prospects [J]. Biotechnol Genet Eng Rev, 2009, 26(1): 139-162. |
| [19] | 金凤媚, 薛俊, 郏艳红, 等. 番茄果实颜色相关基因的研究进展 [J]. 天津农业科学, 2006, 12(4): 3-6. |
| Jin FM, Xue J, Jia YH, et al. Research advance of tomato fruit color related gene [J]. Tianjin Agric Sci, 2006, 12(4): 3-6. | |
| [20] | Egea I, Barsan C, Bian WP, et al. Chromoplast differentiation: current status and perspectives [J]. Plant Cell Physiol, 2010, 51(10): 1601-1611. |
| [21] | 孟凡娟, 许向阳, 李景富. 番茄果实色素含量和表面颜色相关性研究 [J]. 东北农业大学学报, 2006, 37(4): 459-462. |
| Meng FJ, Xu XY, Li JF. Study on correlation of the tomato fruit’s pigment content and surface colour [J]. J Northeast Agric Univ, 2006, 37(4): 459-462. | |
| [22] | 阮美颖, 叶青静, 周国治, 等. 不同颜色樱桃番茄果实成熟过程中色素的变化 [J]. 浙江农业科学, 2013, 54(5): 526-528. |
| Ruan MY, Ye QJ, Zhou GZ, et al. Changes of pigment during fruit ripening of cherry tomato with different colors [J]. J Zhejiang Agric Sci, 2013, 54(5): 526-528. | |
| [23] | 王仁杰, 夏海波, 蔡红明, 等. 不同颜色樱桃番茄品质研究 [J]. 食品工业, 2024, 45(1): 104-108. |
| Wang RJ, Xia HB, Cai HM, et al. Study on quality of different color cherry tomatoes [J]. Food Ind, 2024, 45(1): 104-108. | |
| [24] | Kang A, George KW, Wang G, et al. Isopentenyl diphosphate (IPP)-bypass mevalonate pathways for isopentenol production [J]. Metab Eng, 2016, 34: 25-35. |
| [25] | Fray RG, Grierson D. Identification and genetic analysis of normal and mutant phytoene synthase genes of tomato by sequencing, complementation and co-suppression [J]. Plant Mol Biol, 1993, 22(4): 589-602. |
| [26] | Wang TL, He QY, Wang CY, et al. The expression profile of genes related to carotenoid biosynthesis in pepper under abiotic stress reveals a positive correlation with plant tolerance [J]. Life, 2024, 14(12): 1659. |
| [27] | Rosati C, Aquilani R, Dharmapuri S, et al. Metabolic engineering of beta-carotene and lycopene content in tomato fruit [J]. Plant J, 2000, 24(3): 413-420. |
| [28] | Gao Y, Zhu N, Zhu XF, et al. Diversity and redundancy of the ripening regulatory networks revealed by the fruit ENCODE and the new CRISPR/Cas9 CNR and NOR mutants [J]. Hortic Res, 2019, 6: 39. |
| [29] | Zhu HH, Yang JL, Chen WW. Epigenetic insights into an epimutant colorless non-ripening: from fruit ripening to stress responses [J]. Front Plant Sci, 2024, 15: 1440120. |
| [30] | Wang RF, Lammers M, Tikunov Y, et al. The rin, nor and Cnr spontaneous mutations inhibit tomato fruit ripening in additive and epistatic manners [J]. Plant Sci, 2020, 294: 110436. |
| [31] | Chung MY, Vrebalov J, Alba R, et al. A tomato (Solanum lycopersicum) APETALA2/ERF gene, SlAP2a, is a negative regulator of fruit ripening [J]. Plant J, 2010, 64(6): 936-947. |
| [32] | Liu GZ, Li CX, Yu HY, et al. GREEN STRIPE, encoding methylated TOMATO AGAMOUS-LIKE 1, regulates chloroplast development and Chl synthesis in fruit [J]. New Phytol, 2020, 228(1): 302-317. |
| [33] | Wu MB, Xu X, Hu XW, et al. SlMYB72 regulates the metabolism of chlorophylls, carotenoids, and flavonoids in tomato fruit [J]. Plant Physiol, 2020, 183(3): 854-868. |
| [34] | Jia HM, Xu YP, Deng YW, et al. Key transcription factors regulate fruit ripening and metabolite accumulation in tomato [J]. Plant Physiol, 2024, 195(3): 2256-2273. |
| [35] | Li WZ, Chen LL, Zhao WH, et al. Mutation of YFT3, an isomerase in the isoprenoid biosynthetic pathway, impairs its catalytic activity and carotenoid accumulation in tomato fruit [J]. Hortic Res, 2024, 11(9): uhae202. |
| [36] | Yu MY, Xie YG, Qian ZL, et al. A single nucleotide substitution in the SlMCT gene contributes to great morphological alternations in tomato [J]. Mol Hortic, 2025, 5(1): 49. |
| [37] | Yang TX, Ali M, Lin LH, et al. Recoloring tomato fruit by CRISPR/Cas9-mediated multiplex gene editing [J]. Hortic Res, 2023, 10(1): uhac214. |
| [38] | Romero I, Tikunov Y, Bovy A. Virus-induced gene silencing in detached tomatoes and biochemical effects of phytoene desaturase gene silencing [J]. J Plant Physiol, 2011, 168(10): 1129-1135. |
| [39] | McQuinn RP, Gapper NE, Gray AG, et al. Manipulation of ZDS in tomato exposes carotenoid- and ABA-specific effects on fruit development and ripening [J]. Plant Biotechnol J, 2020, 18(11): 2210-2224. |
| [40] | Tal Isaacson GR. Cloning of tangerine from tomato reveals a carotenoid isomerase essential for the production of β-carotene and xanthophylls in plants [J]. Plant Cell, 2002, 14(2): 333-342. |
| [41] | Gil Ronen LC. An alternative pathway to β-carotene formation in plant chromoplasts discovered by map-based cloning of beta and old-gold color mutations in tomato [J]. Proc Natl Acad Sci U S A, 2000, 97(20): 11102-11107. |
| [42] | Yoo HJ, Chung MY, Lee HA, et al. Natural overexpression of carotenoid cleavage dioxygenase 4 in tomato alters carotenoid flux [J]. Plant Physiol, 2023, 192(2): 1289-1306. |
| [43] | D’Andrea L, Simon-Moya M, Llorente B, et al. Interference with Clp protease impairs carotenoid accumulation during tomato fruit ripening [J]. J Exp Bot, 2018, 69(7): 1557-1568. |
| [44] | Ronen G, Cohen M, Zamir D, et al. Regulation of carotenoid biosynthesis during tomato fruit development: expression of the gene for lycopene Epsilon-cyclase is down-regulated during ripening and is elevated in the mutant Delta [J]. Plant J, 1999, 17(4): 341-351. |
| [45] | Zhao WH, Li YH, Fan SZ, et al. The transcription factor WRKY32 affects tomato fruit colour by regulating YELLOW FRUITED-TOMATO 1, a core component of ethylene signal transduction [J]. J Exp Bot, 2021, 72(12): 4269-4282. |
| [46] | Zhao WH, Gao L, Li YH, et al. Yellow-fruited phenotype is caused by 573 bp insertion at 5' UTR of YFT1 allele in yft1 mutant tomato [J]. Plant Sci, 2020, 300: 110637. |
| [47] | Feng Y, Kou XH, Yuan S, et al. CRISPR/Cas9-mediated SNAC9 mutants reveal the positive regulation of tomato ripening by SNAC9 and the mechanism of carotenoid metabolism regulation [J]. Hortic Res, 2023, 10(4): uhad019. |
| [48] | Fu DQ, Meng LH, Zhu BZ, et al. Silencing of the SlNAP7 gene influences plastid development and lycopene accumulation in tomato [J]. Sci Rep, 2016, 6: 38664. |
| [49] | García-Alcázar M, Giménez E, Pineda B, et al. Albino T-DNA tomato mutant reveals a key function of 1-deoxy-D-xylulose-5-phosphate synthase (DXS1) in plant development and survival [J]. Sci Rep, 2017, 7: 45333. |
| [50] | Liu CC, Ahammed GJ, Wang GT, et al. Tomato CRY1a plays a critical role in the regulation of phytohormone homeostasis, plant development, and carotenoid metabolism in fruits [J]. Plant Cell Environ, 2018, 41(2): 354-366. |
| [51] | Barr J, White WS, Chen L, et al. The GHOST terminal oxidase regulates developmental programming in tomato fruit [J]. Plant Cell Environ, 2004, 27(7): 840-852. |
| [52] | Galpaz N, Wang Q, Menda N, et al. Abscisic acid deficiency in the tomato mutant high-pigment 3 leading to increased plastid number and higher fruit lycopene content [J]. Plant J, 2008, 53(5): 717-730. |
| [53] | He Y, Wang Y, Zhang MZ, et al. SlBEL11 affects tomato carotenoid accumulation by regulating SlLCY-b2 [J]. Front Nutr, 2022, 9: 1062006. |
| [54] | Wang Y, Tian C, Na QT, et al. The role of SlCHRC in carotenoid biosynthesis and plastid development in tomato fruit [J]. Int J Biol Macromol, 2024, 281(Pt 3): 136354. |
| [55] | Niu QF, Xu YP, Huang H, et al. Two transcription factors play critical roles in mediating epigenetic regulation of fruit ripening in tomato [J]. Proc Natl Acad Sci USA, 2025, 122(15): e2422798122. |
| [56] | Xiong C, Luo D, Lin AH, et al. A tomato B-box protein SlBBX20 modulates carotenoid biosynthesis by directly activating PHYTOENE SYNTHASE 1, and is targeted for 26S proteasome-mediated degradation [J]. New Phytol, 2019, 221(1): 279-294. |
| [57] | Zhang JL, Hu ZL, Yao QY, et al. A tomato MADS-box protein, SlCMB1, regulates ethylene biosynthesis and carotenoid accumulation during fruit ripening [J]. Sci Rep, 2018, 8(1): 3413. |
| [58] | Nguyen TNP, Sung J. Light spectral-ranged specific metabolisms of plant pigments [J]. Metabolites, 2024, 15(1): 1. |
| [59] | Wu Y, Jin X, Liao WB, et al. 5-aminolevulinic acid (ALA) alleviated salinity stress in cucumber seedlings by enhancing chlorophyll synthesis pathway [J]. Front Plant Sci, 2018, 9: 635. |
| [60] | Akram NA, Ashraf M. Regulation in plant stress tolerance by a potential plant growth regulator, 5-aminolevulinic acid [J]. J Plant Growth Regul, 2013, 32(3): 663-679. |
| [61] | Li DD, Yu FS, Zhang YZ, et al. Integrative analysis of different low-light-tolerant cucumber lines in response to low-light stress [J]. Front Plant Sci, 2022, 13: 1093859. |
| [62] | Tanaka R, Tanaka A. Tetrapyrrole biosynthesis in higher plants [J]. Annu Rev Plant Biol, 2007, 58: 321-346. |
| [63] | Hörtensteiner S. Chlorophyll degradation during senescence [J]. Annu Rev Plant Biol, 2006, 57: 55-77. |
| [64] | Tanaka A, Ito H. Chlorophyll degradation and its physiological function [J]. Plant Cell Physiol, 2025, 66(2): 139-152. |
| [65] | Nguyen CV, Vrebalov JT, Gapper NE, et al. Tomato GOLDEN2-LIKE transcription factors reveal molecular gradients that function during fruit development and ripening [J]. Plant Cell, 2014, 26(2): 585-601. |
| [66] | Niu XL, Li HL, Li R, et al. Transcription factor SlBEL2 interferes with GOLDEN2-LIKE and influences green shoulder formation in tomato fruits [J]. Plant J, 2022, 112(4): 982-997. |
| [67] | Shi Y, Pang XQ, Liu WJ, et al. SlZHD17 is involved in the control of chlorophyll and carotenoid metabolism in tomato fruit [J]. Hortic Res, 2021, 8(1): 259. |
| [68] | Yuan YJ, Mei LH, Wu MB, et al. SlARF10, an auxin response factor, is involved in chlorophyll and sugar accumulation during tomato fruit development [J]. J Exp Bot, 2018, 69(22): 5507-5518. |
| [69] | Yan F, Gao YS, Pang XQ, et al. BEL1-LIKE HOMEODOMAIN4 regulates chlorophyll accumulation, chloroplast development, and cell wall metabolism in tomato fruit [J]. J Exp Bot, 2020, 71(18): 5549-5561. |
| [70] | Meng LH, Fan ZQ, Zhang Q, et al. BEL1-LIKE HOMEODOMAIN 11 regulates chloroplast development and chlorophyll synthesis in tomato fruit [J]. Plant J, 2018, 94(6): 1126-1140. |
| [71] | Nadakuduti SS, Holdsworth WL, Klein CL, et al. KNOX genes influence a gradient of fruit chloroplast development through regulation of GOLDEN2-LIKE expression in tomato [J]. Plant J, 2014, 78(6): 1022-1033. |
| [72] | Cookson PJ, Kiano JW, Shipton CA, et al. Increases in cell elongation, plastid compartment size and phytoene synthase activity underlie the phenotype of the high pigment-1 mutant of tomato [J]. Planta, 2003, 217(6): 896-903. |
| [73] | Jones B, Frasse P, Olmos E, et al. Down-regulation of DR12, an auxin-response-factor homolog, in the tomato results in a pleiotropic phenotype including dark green and blotchy ripening fruit [J]. Plant J, 2002, 32(4): 603-613. |
| [74] | Shi Y, Hu GJ, Wang Y, et al. The SlGRAS9-SlZHD17 transcriptional cascade regulates chlorophyll and carbohydrate metabolism contributing to fruit quality traits in tomato [J]. New Phytol, 2024, 241(6): 2540-2557. |
| [75] | Zhu MK, Li YL, Chen GP, et al. Silencing SlELP2L, a tomato Elongator complex protein 2-like gene, inhibits leaf growth, accelerates leaf, sepal senescence and produces dark-green fruit [J]. Sci Rep, 2015, 5: 7693. |
| [76] | Mustilli AC, Fenzi F, Ciliento R, et al. Phenotype of the tomato high pigment-2 mutant is caused by a mutation in the tomato homolog of DEETIOLATED1 [J]. Plant Cell, 1999, 11(2): 145-157. |
| [77] | 马星云, 范冰丽, 唐光彩, 等. DXR调控番茄叶绿体发育、花色与果实着色机制初探 [J]. 园艺学报, 2024, 51(6): 1241-1255. |
| Ma XY, Fan BL, Tang GC, et al. Preliminary study on the mechanism of DXR regulating chloroplast development flower color and fruit coloring in tomato [J]. Acta Hortic Sin, 2024, 51(6): 1241-1255. | |
| [78] | Daniel Forth KAP. The suffulta mutation in tomato reveals a novel method of plastid replication during fruit ripening [J]. J Exp Bot, 2006, 57(9): 1971-1979. |
| [79] | Ezura K, Lu Y, Suzuki Y, et al. Class II knotted-like homeodomain protein SlKN5 with BEL1-like homeodomain proteins suppresses fruit greening in tomato fruit [J]. Plant J, 2024, 118(6): 2037-2054. |
| [80] | Tanaka Y, Sasaki N, Ohmiya A. Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids [J]. Plant J, 2008, 54(4): 733-749. |
| [81] | Li L, Ye J, Li HH, et al. Characterization of metabolites and transcripts involved in flower pigmentation in Primula vulgaris [J]. Front Plant Sci, 2020, 11: 572517. |
| [82] | Zhang Y, Butelli E, Martin C. Engineering anthocyanin biosynthesis in plants [J]. Curr Opin Plant Biol, 2014, 19: 81-90. |
| [83] | Zhang BP, Hülskamp M. Evolutionary analysis of MBW function by phenotypic rescue in Arabidopsis thaliana [J]. Front Plant Sci, 2019, 10: 375. |
| [84] | Sun CL, Deng L, Du MM, et al. A transcriptional network promotes anthocyanin biosynthesis in tomato flesh [J]. Mol Plant, 2020, 13(1): 42-58. |
| [85] | Chen YZ, Kim P, Kong LZ, et al. A dual-function transcription factor, SlJAF13, promotes anthocyanin biosynthesis in tomato [J]. J Exp Bot, 2022, 73(16): 5559-5580. |
| [86] | Menconi J, Perata P, Gonzali S. In pursuit of purple: anthocyanin biosynthesis in fruits of the tomato clade [J]. Trends Plant Sci, 2024, 29(5): 589-604. |
| [87] | Mes PJ, Boches P, Myers JR, et al. Characterization of tomatoes expressing anthocyanin in the fruit [J]. J Amer Soc Hort Sci, 2008, 133(2): 262-269. |
| [88] | Cao X, Qiu Z, Wang X, et al. A putative R3 MYB repressor is the candidate gene underlying atroviolacium, a locus for anthocyanin pigmentation in tomato fruit [J]. J Exp Bot, 2017, 68(21/22): 5745-5758. |
| [89] | Zhu MZ, Zhao R, Wu HY, et al. Integrated transcriptome and metabolome analysis provides insights into anthocyanin biosynthesis in Cichorium intybus L [J]. BMC Plant Biol, 2025, 25(1): 409. |
| [90] | Kiferle C, Fantini E, Bassolino L, et al. Tomato R2R3-MYB proteins SlANT1 and SlAN2: same protein activity, different roles [J]. PLoS One, 2015, 10(8): e0136365. |
| [91] | 张振, 徐志璇, 王丽娜, 等. 过表达SlMYB75对番茄幼苗、果实及种子的影响 [J]. 山东农业大学学报: 自然科学版, 2019, 50(6): 937-943. |
| Zhang Z, Xu ZX, Wang LN, et al. Effects of overexpression of SlMYB75 on tomato seedlings, fruits and seeds [J]. J Shandong Agric Univ Nat Sci Ed, 2019, 50(6): 937-943. | |
| [92] | Ballester AR, Molthoff J, de Vos R, et al. Biochemical and molecular analysis of pink tomatoes: deregulated expression of the gene encoding transcription factors SlMYB12 leads to pink tomato fruit color [J]. Plant Physiol, 2010, 152(1): 71-84. |
| [93] | Schijlen EG, de Vos CH, Martens S, et al. RNA interference silencing of Chalcone synthase, the first step in the flavonoid biosynthesis pathway, leads to parthenocarpic tomato fruits [J]. Plant Physiol, 2007, 144(3): 1520-1530. |
| [94] | Shiose L, Dos Reis Moreira J, Lira BS, et al. A tomato B-box protein regulates plant development and fruit quality through the interaction with PIF4, HY5, and RIN transcription factors [J]. J Exp Bot, 2024, 75(11): 3368-3387. |
| [95] | Li YY, Meng FL, You HW, et al. SlWRKY14 integrates carotenoid and flavonoid biosynthetic pathways to regulate coloration and quality of tomato fruits [J]. Cell Rep, 2025, 44(10): 116369. |
| [96] | Wang JF, Li GB, Li CX, et al. NF-Y plays essential roles in flavonoid biosynthesis by modulating histone modifications in tomato [J]. New Phytol, 2021, 229(6): 3237-3252. |
| [97] | Ilahy R, Tlili I, Siddiqui MW, et al. Inside and beyond color: comparative overview of functional quality of tomato and watermelon fruits [J]. Front Plant Sci, 2019, 10: 769. |
| [98] | Beyer P, Al-Babili S, Ye XD, et al. Golden Rice: introducing the beta-carotene biosynthesis pathway into rice endosperm by genetic engineering to defeat vitamin A deficiency [J]. J Nutr, 2002, 132(3): 506S-510S. |
| [99] | Elitzur T, Yakir E, Quansah L, et al. Banana MaMADS transcription factors are necessary for fruit ripening and molecular tools to promote shelf-life and food security [J]. Plant Physiol, 2016, 171(1): 380-391. |
| [100] | Cui L, Zheng FY, Li CX, et al. Defective mutations in STAY-GREEN 1, PHYTOENE SYNTHASE 1, and MYB12 genes lead to formation of green ripe fruit in tomato [J]. J Exp Bot, 2024, 75(11): 3322-3336. |
| [101] | Polturak G, Aharoni A. Advances and future directions in betalain metabolic engineering [J]. New Phytol, 2019, 224(4): 1472-1478. |
| [102] | Orzaez D, Medina A, Torre S, et al. A visual reporter system for virus-induced gene silencing in tomato fruit based on anthocyanin accumulation [J]. Plant Physiol, 2009, 150(3): 1122-1134. |
| [1] | 尹跃, 秦小雅, 米佳, 安巍, 何军, 张锋锋. 枸杞FBN基因家族鉴定及与类胡萝卜素代谢的相关性分析[J]. 生物技术通报, 2026, 42(3): 338-348. |
| [2] | 李亚妮, 韩鸿宇, 耿梦爽, 米若兰, 王韦琪, 于文静, 孟宪文, 李传友. ChiC基因调控番茄灰霉病抗性的机制研究[J]. 生物技术通报, 2026, 42(3): 255-262. |
| [3] | 程云霞, 张俊红, 叶杰. 番茄果实可溶性固形物积累的遗传调控研究进展[J]. 生物技术通报, 2026, 42(3): 145-155. |
| [4] | 颜晨琳, 李凡, 闫春婷, 程蛟文, 胡开林, 叶志彪, 宋建文. 番茄果实形态发育相关基因研究进展[J]. 生物技术通报, 2026, 42(3): 172-186. |
| [5] | 马莹莹, 游惠婉, 郑积荣, 汪俏梅, 刘丽红. 基于PSY多层级调控的园艺作物品质形成机制研究进展[J]. 生物技术通报, 2026, 42(3): 96-110. |
| [6] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [7] | 胡秋玲, 陈灵, 黄嘉怡, 赵梓乔, 潘璐怡, 刘慧丽, 刘太波. 多胺调控果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 203-212. |
| [8] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [9] | 杜丹, 郭翔, 胡鑫, 潘宇. 质体发育调控果实成熟与品质的研究进展[J]. 生物技术通报, 2026, 42(3): 48-59. |
| [10] | 姜喆卉, 王小龙, 王守创, 周科. 番茄风味物质代谢途径解析与分子育种研究进展[J]. 生物技术通报, 2026, 42(3): 60-78. |
| [11] | 王潇奕, 李金焱, 邢醒, 朱鸿亮. 基于乙烯响应筛选调控番茄成熟且影响呼吸的基因及其功能分析[J]. 生物技术通报, 2026, 42(3): 275-282. |
| [12] | 李迎辉, 王杨博涵, 周浩博, 卢心如, 张珂欣, 于洋, 李传友, 孙传龙. 番茄VPE基因家族鉴定和抗逆功能分析[J]. 生物技术通报, 2026, 42(3): 263-274. |
| [13] | 袁梦博, 赵光伟, 贺玉花, 黄祥, 徐永阳, 张健, 孔维虎, 田小琴, 户克云, 唐伶俐. CmCRC基因影响甜瓜性别分化的功能研究[J]. 生物技术通报, 2026, 42(3): 294-301. |
| [14] | 陈玲嫣, 李伟勋, 逄欣欣, 高翔, 焦绪瑶. 拟杆菌遗传操作工具箱:从传统方法到合成生物学前沿[J]. 生物技术通报, 2026, 42(2): 51-64. |
| [15] | 吕呈聪, 衡蒙, 陈思琪, 金雪花. 彩色马蹄莲花青素苷转运相关ZhGSTF的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 161-169. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||