








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 60-78.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1358
收稿日期:2025-12-11
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
周科,男,博士,副研究员,研究方向 :作物风味与营养品质;E-mail: kezhou@hainanu.edu.cn作者简介:姜喆卉,女,博士,助理研究员,研究方向 :作物风味与营养品质;E-mail: 996991@hainanu.edu.cn基金资助:
JIANG Zhe-hui( ), WANG Xiao-long, WANG Shou-chuang(
), WANG Xiao-long, WANG Shou-chuang( ), ZHOU Ke(
), ZHOU Ke( )
)
Received:2025-12-11
Published:2026-03-26
Online:2026-04-23
摘要:
番茄作为全球消费量最大的果蔬之一,其风味品质直接影响消费体验与市场价值。随着我国人均收入水平提升及消费结构升级,市场对高品质番茄的需求日益迫切。然而,受限于风味性状遗传调控的复杂性及检测技术的局限性,加之传统育种长期优先考虑产量、抗病性与耐贮运性等农艺性状,导致番茄果实风味品质普遍下降,难以满足消费需求。因此,深度解析风味代谢物的生物合成机制及其遗传调控网络,已成为精准改良番茄风味的关键突破口。本文系统综述了近年来国内外在番茄风味代谢物的合成代谢途径解析及其潜在遗传调控机制研究方面取得的前沿进展,重点探讨了基于多组学整合分析策略的番茄风味遗传改良技术体系:通过全面采集番茄种质资源的表型组、代谢组、转录组及基因组等多维度组学数据,结合机器学习与生物信息学分析方法,精准挖掘控制番茄风味形成的关键功能基因与调控位点,并利用现代分子育种技术手段,实现优质风味番茄新品种的高效定向选育。此外,本文还深入剖析了当前番茄风味育种研究在风味成分量化评价标准体系构建、风味性状遗传调控机制解析以及多性状(风味‒产量‒抗性等)协同改良等方面存在的关键技术瓶颈,并针对性地提出了未来研究的发展方向与策略建议。综上所述,本文旨在推动番茄风味育种理念从传统的“生产者导向”向“消费者导向”转型,为番茄及其他作物风味性状的精准遗传改良提供理论依据与技术路径,进而促进我国农业与种业的高质量发展。
姜喆卉, 王小龙, 王守创, 周科. 番茄风味物质代谢途径解析与分子育种研究进展[J]. 生物技术通报, 2026, 42(3): 60-78.
JIANG Zhe-hui, WANG Xiao-long, WANG Shou-chuang, ZHOU Ke. Advances in the Elucidation of Metabolic Pathways and Molecular Breeding for Tomato Flavor[J]. Biotechnology Bulletin, 2026, 42(3): 60-78.
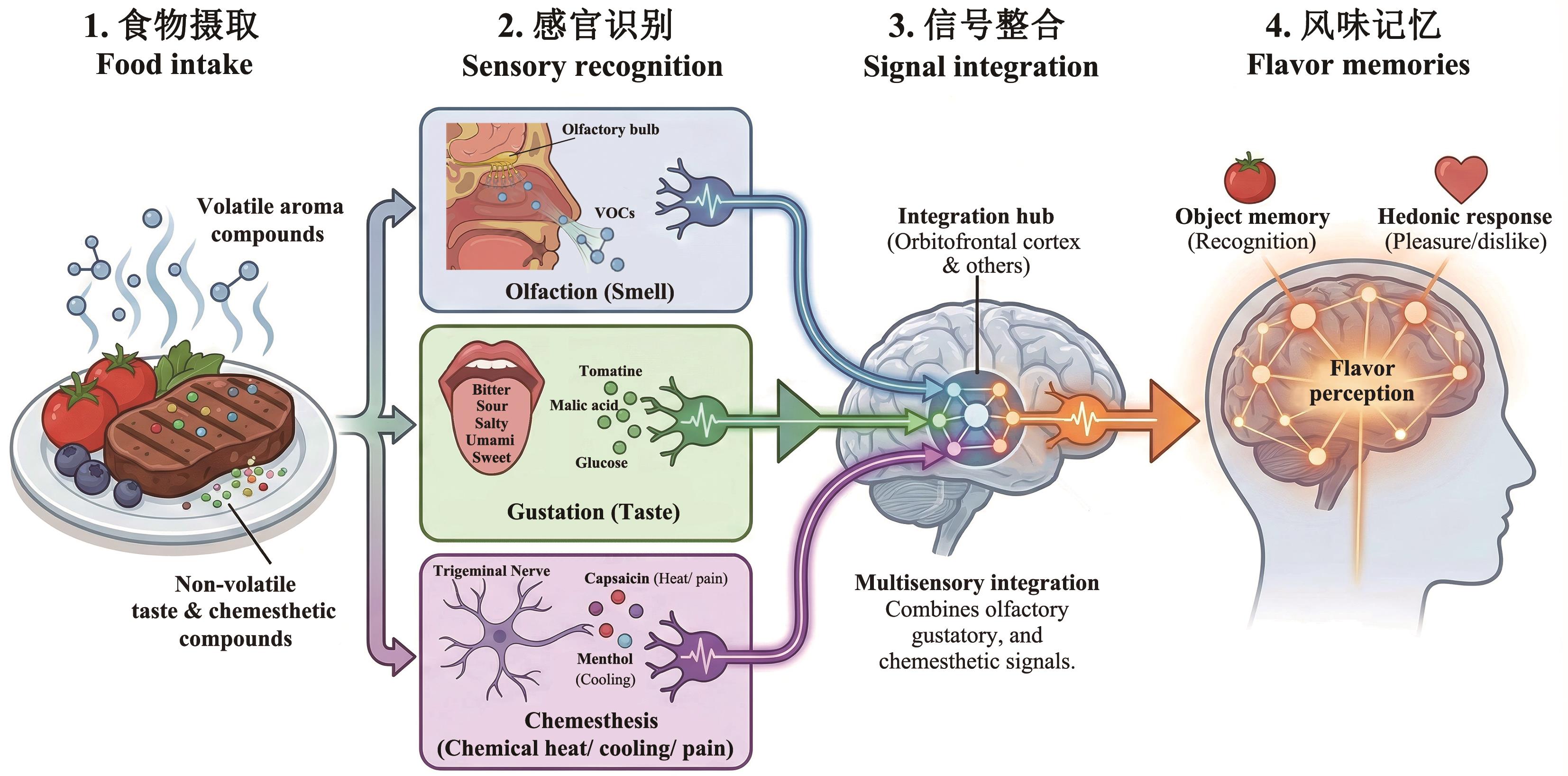
图1 人类风味感知机制的多模态交互示意图人体形成风味感知的生物学机制共分为4个阶段:(1)食物摄取:食物提供挥发性香气化合物及非挥发性味觉与化学物理觉识别物质;(2)感官识别:通过鼻腔嗅觉、口腔味觉(酸甜苦咸鲜)以及三叉神经介导的化学物理觉(如辣椒素的痛热感、薄荷的凉感)进行多维度感知;(3)信号整合:外周神经将信号传入大脑,在味觉皮层形成味觉,随后大脑对多感官信号进行跨模态整合;(4)风味记忆:最终转化为客观的物体识别(如辨认出是番茄)与主观的享乐反应(即喜爱或厌恶的情绪),形成完整的风味体验。VOCs:挥发性化合物。该示意图由Germini3软件辅助修改
Fig. 1 Schematic diagram of multimodal interactions in human flavor perception mechanismsThe biological mechanism of flavor perception in humans comprises four stages: 1) Food intake: Food provides volatile aromatic compounds and non-volatile substances for taste and chemoreceptor identification. 2) Sensory recognition: Multidimensional perception occurs through nasal olfaction, oral taste (sweet, sour, bitter, salty, and umami), and chemoreceptors mediated by the trigeminal nerve (e.g., pain/heat sensation from capsaicin, and cooling sensation from menthol). 3) Signal integration: Signals from peripheral neural circuits enter the brain via the solitary nucleus in the brainstem, travel through the parabrachial nucleus and thalamus to the primary gustatory cortex to form taste perception, then project to regions like the amygdala and orbitofrontal cortex for cross-modal integration of multisensory signals. 4) Flavor memory: This ultimately translates into objective object recognition (e.g., identifying a tomato) and subjective hedonic responses (i.e., emotional preferences like liking or disliking), forming a complete flavor experience. VOCs: Volatile organic compounds. This schematic diagram was modified with the assistance of Germini3 software
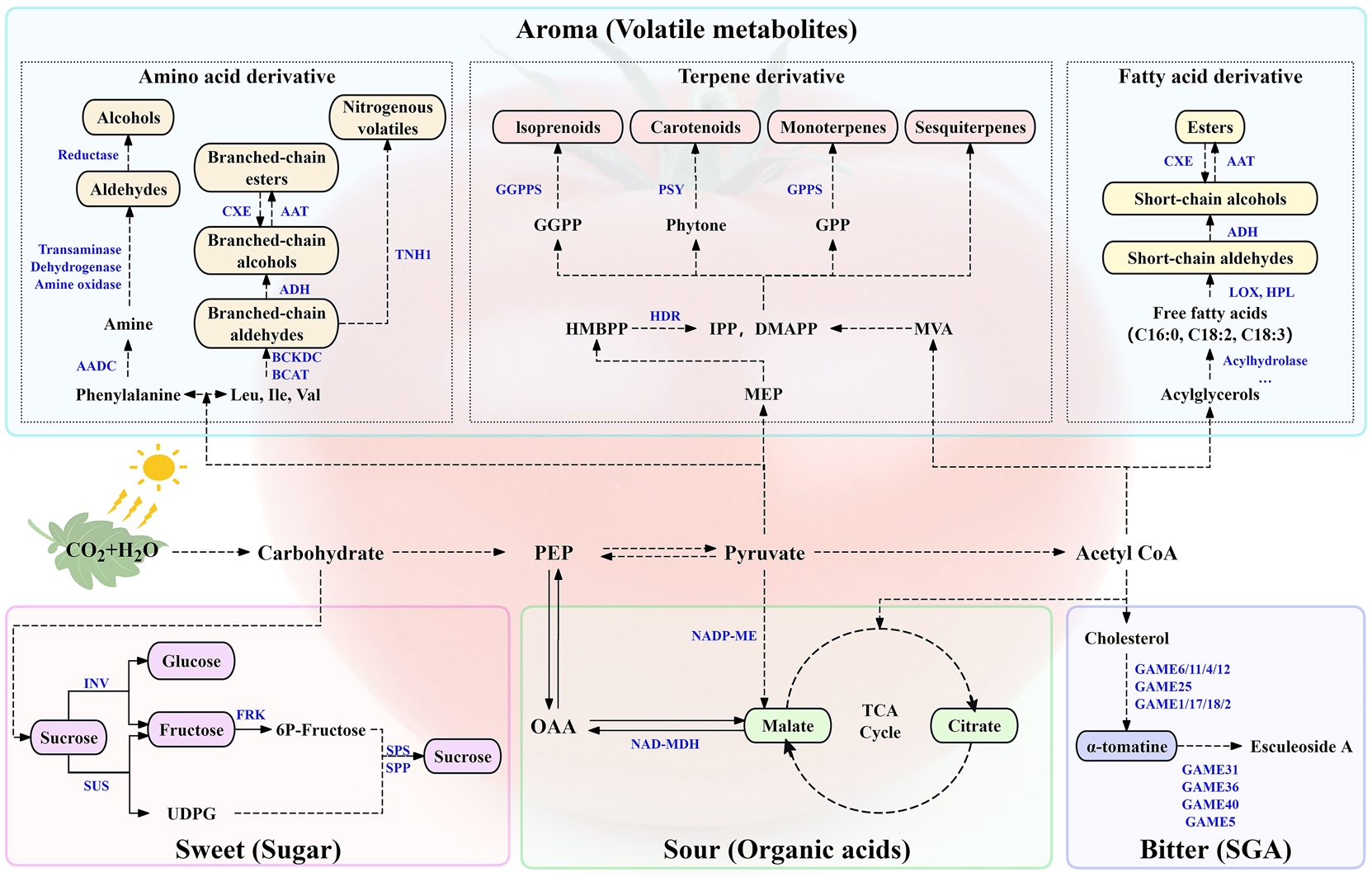
图2 番茄糖类、有机酸类、苦味、挥发性代谢物相关的初生及次生代谢途径本图基于KEGG注释及文献参考构建,虚线箭头涉及多步反应。AADC:芳香族氨基酸脱羧酶;AAT:醇酰基转移酶;ADH:醇脱氢酶;BCAT:支链氨基转移酶;BCKDC:支链α-酮酸脱羧酶;CXE:羧甲基酯酶;DMAPP:二甲基烯丙基二磷酸;FRK:果糖激酶;GAME1/2/4/5/6/11/12/17/18/25/31/36/40:甾体糖苷生物碱代谢酶1/2/4/5/6/11/12/17/18/25/31/36/40;GGPP:香叶基香叶基二磷酸;GGPPS:香叶基香叶基二磷酸合成酶;GPP:香叶基二磷酸;GPPS:GPP合成酶;HDR:1-羟基-2-甲基-2-(E)-丁烯基-4-二磷酸还原酶;HMBPP:1-羟基-2-甲基-2-(E)-丁烯基-4-二磷酸;HPL:氢过氧化物裂解酶;Ile:异亮氨酸;INV:转化酶;Leu:亮氨酸;LOX:脂氧合酶;IPP:异戊烯基二磷酸;MEP:甲基-D-赤藓醇磷酸盐;MVA:甲羟戊酸;NAD-MDH:NAD-苹果酸脱氢酶;NADP-ME:NADP-苹果酸酶;OAA:草酰乙酸;PEP:磷酸烯醇式丙酮酸;PSY:八氢番茄红素合酶;SGA:甾体糖苷生物碱;SPP:蔗糖磷酸酯酶;SPS:蔗糖磷酸合酶;SUS:蔗糖合酶;TCA cycle:三羧酸循环;TNH1:四氢噻唑烷-N-羟化酶1;UDPG:二磷酸尿苷葡萄糖;VAL:缬氨酸
Fig. 2 Primary and secondary metabolic pathways related to sugars, organic acids, bitterness, and volatile metabolites in tomato (Solanum lycopersicum)This figure was created based on KEGG annotations and references, and the dotted line arrow is involved multi-step reactions. AADC: Aromatic amino acid decarboxylase. AAT: Alcohol acyltransferase. ADH: Alcohol dehydrogenase. BCAT: Branched-chain aminotransferase. BCKDC: Branched-chain α-ketoacid decarboxylase. CXE: Carboxymethylesterase. DMAPP: Dimethylallyl diphosphate. FRK: Fructokinase. GAME1/2/4/5/6/11/12/17/18/25/31/36/40: Glycoalkaloid metabolism enzyme 1/2/4/5/6/11/12/17/18/25/31/36/40. GGPP: Geranylgeranyl diphosphate. GGPPS: Geranylgeranyl diphosphate synthase. GPP: Geranyl diphosphate. GPPS: GPP synthase. HDR: 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate reductase. HMBPP: 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate. HPL: Hydroperoxide lyase. Ile: Isoleucine. INV: Invertase. Leu: Leucine. LOX: Lipoxygenase. IPP: Isopentenyl diphosphate. MEP: 2-C-methyl-D-erythritol-4-phosphate. MVA: Mevalonate. NAD-MDH: NAD-malate dehydrogenase. NADP-ME: NADP-malic enzyme. OAA: Oxaloacetic acid. PEP: Phosphoenolpyruvate. PSY: Phytoene synthase. SGA: Steroidal glycoalkaloids. SPP: Sucrose phosphate phosphatase. SPS: Sucrose phosphate synthase. sus: sucrose synthase. TCA cycle: Tricarboxylic acid cycle. TNH1: Tetrahydrothiazolidine N-hydroxylase 1. UDPG: Uridine diphosphate glucose. Val: Valine
物质类别 Primary pathway | 代谢物 Metabolites | 相关基因 Related genes | 风味贡献Attributes | 参考文献 Reference |
|---|---|---|---|---|
糖类 Sugars | 果糖、葡萄糖 Fructose, glucose | Lin5, STP11,SWEET, SlNAP2, SlFgr, SlGLK2, SlVPE5 | Positive | [ |
有机酸 Organic acids | 苹果酸 Malic acid | SlME, SlTDT, SlALMT9, SlMIR164A | Positive | [ |
| 柠檬酸 Citric acid | SlCS, SlAco, SlPEPCK | Positive | [ | |
生物碱类 Alkaloids | 甾体糖苷生物碱 Steroidal alkaloids | GAME1, GAME2, GAME5-9, GAME11, GAME12 GAME17, GAME18, GAME25, GAME31 | Negative | [ |
氨基酸类 Amino Acids | 脯氨酸 Proline | SlP5CS1, SlP5CR | Positive | [ |
天冬酰胺、γ-氨基丁酸 Asparagine, GABA | GDSL esterase/lipase | Unknown | [ | |
谷氨酰胺、苏氨酸 Glutamine, threonine | Unknown | |||
醛类 Aldehydes | 反式-2-己烯醛 Trans-2-Hexenal | LeHPL | Positive | [ |
| 苯乙醛 Phenylacetaldehyde | LeAADCIA, LeAADCIB, LeAADC2 | Positive | [ | |
| 苯甲醛 Benzaldehyde | Positive | |||
反式-2-庚烯醛 Trans-2-Heptenal | Unknown | |||
| 己醛 Hexanal | TomloxC | Unknown | [ | |
| 庚醛 Heptaldehyde | Unknown | |||
| 壬醛 Nonylaldehyde | Negative | |||
反式-2-戊烯醛 Trans-2-Pentenal | SlscADH1 | Unknown | [ | |
| 顺式柠檬醛 Neral | Unknown | |||
| 香叶醛 Geranial | SlADH2 | Unknown | [ | |
| 水杨醛 Salicylaldehyde | Negative | |||
| β-环柠檬醛 β-Cyclocitral | LeCCD4 | Unknown | [ | |
含氮硫类挥发性代谢物 Nitrogen- and sulfur-containing volatile metabolites | 2-异丁基噻唑 2-Isobutylthiazole | SlTNH1 | Positive | [ |
3-甲基丁醛肟 3-methylbutyraldehyde oxime | Negative | [ | ||
| 苯乙腈 Benzylcyanide | Positive | |||
| 3-甲基丁腈 3-methylbutanenitrile | Negative | [ | ||
醇类 Alcohols | 2-苯乙醇 2-Phenylethanol | LeAADCIA, LeAADC1B, LeAADC2, PPEAT, FLORAL4 | Positive | [ |
| 1-戊烯-3-醇 1-Penten-3-ol | Unknown | |||
| 3-甲基丁醇 3-Methylbutanol | Unknown | |||
| 1-辛烯-3-醇 1-Octen-3-ol | SlFAD7 | Negative | [ | |
反式-3-己烯-1-醇 Trans-3-hexen-1-ol | SlADH1 | Unknown | [ | |
| 1-戊醇 1-Pentanol | SlFAD7, Sl-LIP8 | Unknown | [ | |
反式-2-己烯-1-醇 E-2-hexen-1-ol | Positive | |||
| 己醇 Hexylalcohol | Sl-LIP8 | Unknown | [ | |
顺式-3-己烯-1-醇 Z-3-hexen-1-ol | ADH2 | Unknown | [ | |
| 芳樟醇 Linalool | TPS3, TPS5, TPS20, TPS37, TPS39, SlMYB75 | Unknown | [ | |
| 橙花叔醇 Nerolidol | Unknown | [ | ||
2-甲基-1-丁醇 2-Methyl-1-butanol | BCKDC, SlBCAT1, SlBCAT2, AAT1 | Unknown | [ | |
3-甲基-1-丁醇 3-Methyl-1-butanol | Positive | |||
酚类 Phenols | 丁香酚 Eugenol | Negative | ||
| 愈创木酚 Guaiacol | E8 | Unknown | [ | |
酯类 Esters | 水杨酸甲酯 Methyl salicylate | SlSAMT | Negative | [ |
顺式-3-己烯基乙酸酯 cis-3-hexenylacetate | Unknown | |||
| 乙酸己酯 Hexylacetate | Unknown | |||
酮类 Ketones | 1-戊烯-3-酮 1-Penten-3-one | Unknown | ||
香叶基丙酮、β-紫罗兰酮 Geranylacetone, β-Ionone | LeCCD1A, LeCCD1B | Positive | [ | |
6-甲基-5-庚烯-2-酮 6-Methyl-5-hepten-2-one | Positive | |||
| 烷类Alkanes | 1-硝基-2-苯乙烷 1-nitro-2-Phenylethane | SlTNH1 | Positive | [ |
1-硝基-3-甲基丁烷 1-nitro-3-methylbutane | SlTNH1 | Negative | [ |
表1 番茄风味相关代谢物及基因列表
Table 1 List of tomato flavor-related metabolites and gene
物质类别 Primary pathway | 代谢物 Metabolites | 相关基因 Related genes | 风味贡献Attributes | 参考文献 Reference |
|---|---|---|---|---|
糖类 Sugars | 果糖、葡萄糖 Fructose, glucose | Lin5, STP11,SWEET, SlNAP2, SlFgr, SlGLK2, SlVPE5 | Positive | [ |
有机酸 Organic acids | 苹果酸 Malic acid | SlME, SlTDT, SlALMT9, SlMIR164A | Positive | [ |
| 柠檬酸 Citric acid | SlCS, SlAco, SlPEPCK | Positive | [ | |
生物碱类 Alkaloids | 甾体糖苷生物碱 Steroidal alkaloids | GAME1, GAME2, GAME5-9, GAME11, GAME12 GAME17, GAME18, GAME25, GAME31 | Negative | [ |
氨基酸类 Amino Acids | 脯氨酸 Proline | SlP5CS1, SlP5CR | Positive | [ |
天冬酰胺、γ-氨基丁酸 Asparagine, GABA | GDSL esterase/lipase | Unknown | [ | |
谷氨酰胺、苏氨酸 Glutamine, threonine | Unknown | |||
醛类 Aldehydes | 反式-2-己烯醛 Trans-2-Hexenal | LeHPL | Positive | [ |
| 苯乙醛 Phenylacetaldehyde | LeAADCIA, LeAADCIB, LeAADC2 | Positive | [ | |
| 苯甲醛 Benzaldehyde | Positive | |||
反式-2-庚烯醛 Trans-2-Heptenal | Unknown | |||
| 己醛 Hexanal | TomloxC | Unknown | [ | |
| 庚醛 Heptaldehyde | Unknown | |||
| 壬醛 Nonylaldehyde | Negative | |||
反式-2-戊烯醛 Trans-2-Pentenal | SlscADH1 | Unknown | [ | |
| 顺式柠檬醛 Neral | Unknown | |||
| 香叶醛 Geranial | SlADH2 | Unknown | [ | |
| 水杨醛 Salicylaldehyde | Negative | |||
| β-环柠檬醛 β-Cyclocitral | LeCCD4 | Unknown | [ | |
含氮硫类挥发性代谢物 Nitrogen- and sulfur-containing volatile metabolites | 2-异丁基噻唑 2-Isobutylthiazole | SlTNH1 | Positive | [ |
3-甲基丁醛肟 3-methylbutyraldehyde oxime | Negative | [ | ||
| 苯乙腈 Benzylcyanide | Positive | |||
| 3-甲基丁腈 3-methylbutanenitrile | Negative | [ | ||
醇类 Alcohols | 2-苯乙醇 2-Phenylethanol | LeAADCIA, LeAADC1B, LeAADC2, PPEAT, FLORAL4 | Positive | [ |
| 1-戊烯-3-醇 1-Penten-3-ol | Unknown | |||
| 3-甲基丁醇 3-Methylbutanol | Unknown | |||
| 1-辛烯-3-醇 1-Octen-3-ol | SlFAD7 | Negative | [ | |
反式-3-己烯-1-醇 Trans-3-hexen-1-ol | SlADH1 | Unknown | [ | |
| 1-戊醇 1-Pentanol | SlFAD7, Sl-LIP8 | Unknown | [ | |
反式-2-己烯-1-醇 E-2-hexen-1-ol | Positive | |||
| 己醇 Hexylalcohol | Sl-LIP8 | Unknown | [ | |
顺式-3-己烯-1-醇 Z-3-hexen-1-ol | ADH2 | Unknown | [ | |
| 芳樟醇 Linalool | TPS3, TPS5, TPS20, TPS37, TPS39, SlMYB75 | Unknown | [ | |
| 橙花叔醇 Nerolidol | Unknown | [ | ||
2-甲基-1-丁醇 2-Methyl-1-butanol | BCKDC, SlBCAT1, SlBCAT2, AAT1 | Unknown | [ | |
3-甲基-1-丁醇 3-Methyl-1-butanol | Positive | |||
酚类 Phenols | 丁香酚 Eugenol | Negative | ||
| 愈创木酚 Guaiacol | E8 | Unknown | [ | |
酯类 Esters | 水杨酸甲酯 Methyl salicylate | SlSAMT | Negative | [ |
顺式-3-己烯基乙酸酯 cis-3-hexenylacetate | Unknown | |||
| 乙酸己酯 Hexylacetate | Unknown | |||
酮类 Ketones | 1-戊烯-3-酮 1-Penten-3-one | Unknown | ||
香叶基丙酮、β-紫罗兰酮 Geranylacetone, β-Ionone | LeCCD1A, LeCCD1B | Positive | [ | |
6-甲基-5-庚烯-2-酮 6-Methyl-5-hepten-2-one | Positive | |||
| 烷类Alkanes | 1-硝基-2-苯乙烷 1-nitro-2-Phenylethane | SlTNH1 | Positive | [ |
1-硝基-3-甲基丁烷 1-nitro-3-methylbutane | SlTNH1 | Negative | [ |
研究策略 Research strategy | 研究逻辑 Approach | 技术手段 Technical means | 研究场景 Research context | 优势 Advantages | 局限性 Limitations | 代表性基因与功能 Representative genes and functions | 参考文献Reference |
|---|---|---|---|---|---|---|---|
正向遗传学 Forward genetics | 从表型到基因 | 图位克隆/QTL定位 | 两个极端表型亲本杂交衍生的群体(如F2、RIL、NIL等) | 1. 检测效能高; 2. 遗传背景简单; 3. 可直接用于育种转育 | 1. 构建群体耗时; 2. 多样性低; 3. 定位区间较大 | Lin5,细胞壁转化酶,直接调控果实可溶性固形物(TSS)与甜度 | [ |
全基因组关联分析 GWAS | 自然种质群体 | 1. 分辨率高; 2. 遗传背景丰富; 3. 多性状同步分析; 4. 无需构建杂交群体 | 1. 易受群体结构干扰; 2. 假阳性率高; 3. 计算与检测成本高 | STP1,糖转运蛋白,STP1Insertion 中SSC水平约是STP1Deletion 的1.2倍 | [ | ||
反向遗传学 Reverse genetics | 从基因到表型 | RNA干扰 RNAi | 基因敲降 | 避免基因敲除导致胚胎致死问题 | 1. 抑制不彻底; 2. 易脱靶; 3. 多代遗传稳定性问题; 4. 目前面临监管政策限制 | HT1,己糖转运蛋白,沉默HT1基因的果实中己糖积累降低55% | [ |
基因编辑 CRISPR/Cas9 | 基因敲除 | 1. 定点精准编辑; 2. 实现单基因; 3. 多代稳定遗传 | 1. 基因敲除存在致死风险; 2. 功能冗余干扰; 3. 面临监管政策限制 | SlSWEET12c,糖转运蛋白,其敲除突变体植株中果糖和葡萄糖含量降低约40% | [ | ||
过表达 Overexpression | 获得功能 | 1. 直接验证基因的正向调控功能; 2. 克服功能冗余问题 | 1. 异位表达干扰植株正常生理; 2. 非自然遗传状态; 3. 共抑制现象; 4. 多代遗传稳定性问题; 5. 面临监管政策限制 | SlTST3a,液泡糖转运蛋白,过表达植株中果糖和葡萄糖含量升高约1.3倍 | [ |
表2 正向与反向遗传学在番茄糖含量调控关键基因解析方面的比较分析
Table 2 Comparative analysis of forward genetics and reverse genetics in the identification of key genes regulating sugar content in tomatoes
研究策略 Research strategy | 研究逻辑 Approach | 技术手段 Technical means | 研究场景 Research context | 优势 Advantages | 局限性 Limitations | 代表性基因与功能 Representative genes and functions | 参考文献Reference |
|---|---|---|---|---|---|---|---|
正向遗传学 Forward genetics | 从表型到基因 | 图位克隆/QTL定位 | 两个极端表型亲本杂交衍生的群体(如F2、RIL、NIL等) | 1. 检测效能高; 2. 遗传背景简单; 3. 可直接用于育种转育 | 1. 构建群体耗时; 2. 多样性低; 3. 定位区间较大 | Lin5,细胞壁转化酶,直接调控果实可溶性固形物(TSS)与甜度 | [ |
全基因组关联分析 GWAS | 自然种质群体 | 1. 分辨率高; 2. 遗传背景丰富; 3. 多性状同步分析; 4. 无需构建杂交群体 | 1. 易受群体结构干扰; 2. 假阳性率高; 3. 计算与检测成本高 | STP1,糖转运蛋白,STP1Insertion 中SSC水平约是STP1Deletion 的1.2倍 | [ | ||
反向遗传学 Reverse genetics | 从基因到表型 | RNA干扰 RNAi | 基因敲降 | 避免基因敲除导致胚胎致死问题 | 1. 抑制不彻底; 2. 易脱靶; 3. 多代遗传稳定性问题; 4. 目前面临监管政策限制 | HT1,己糖转运蛋白,沉默HT1基因的果实中己糖积累降低55% | [ |
基因编辑 CRISPR/Cas9 | 基因敲除 | 1. 定点精准编辑; 2. 实现单基因; 3. 多代稳定遗传 | 1. 基因敲除存在致死风险; 2. 功能冗余干扰; 3. 面临监管政策限制 | SlSWEET12c,糖转运蛋白,其敲除突变体植株中果糖和葡萄糖含量降低约40% | [ | ||
过表达 Overexpression | 获得功能 | 1. 直接验证基因的正向调控功能; 2. 克服功能冗余问题 | 1. 异位表达干扰植株正常生理; 2. 非自然遗传状态; 3. 共抑制现象; 4. 多代遗传稳定性问题; 5. 面临监管政策限制 | SlTST3a,液泡糖转运蛋白,过表达植株中果糖和葡萄糖含量升高约1.3倍 | [ |
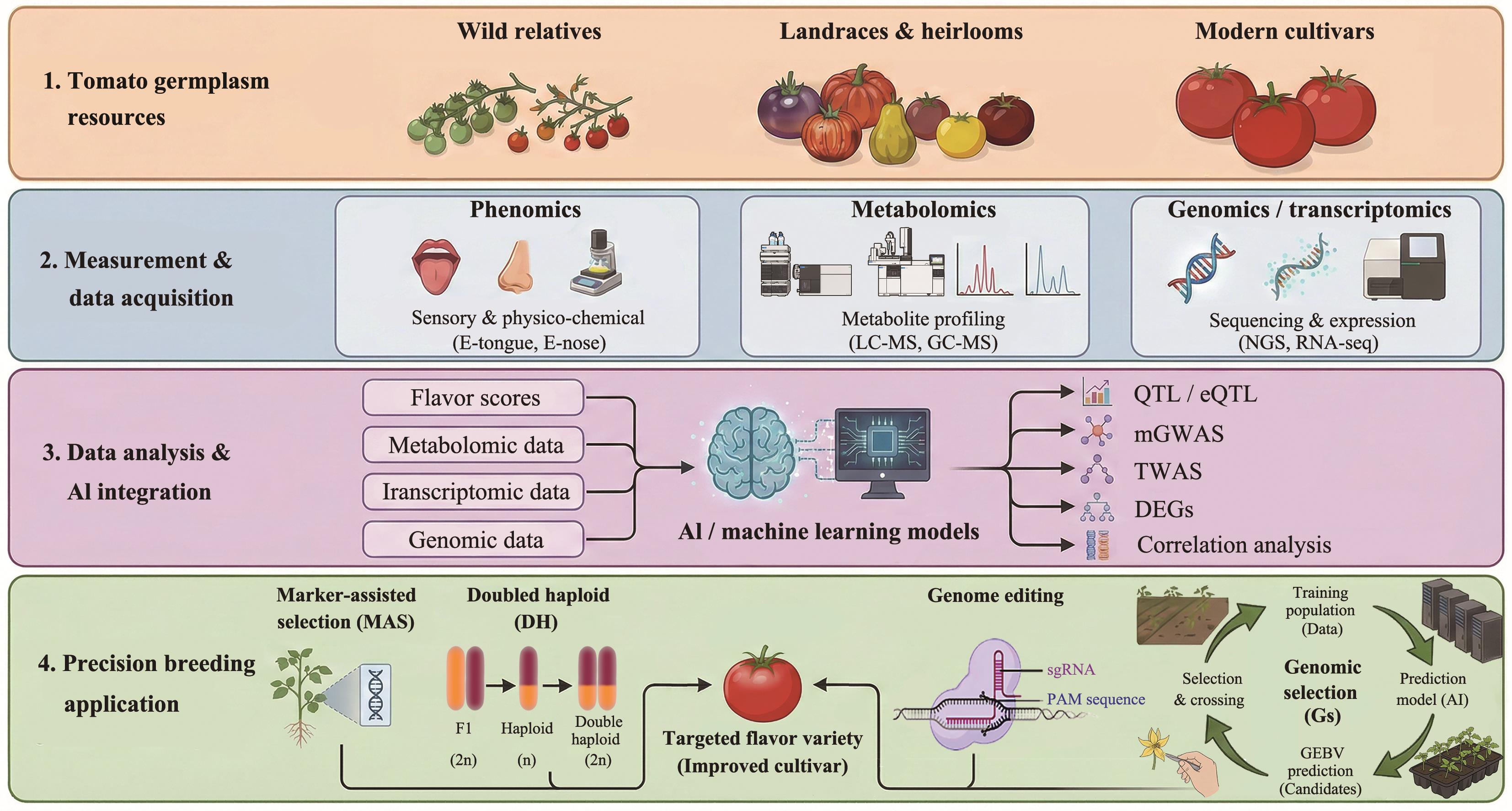
图3 多组学大数据驱动下的番茄风味性状精准改良技术路线人工智能驱动的番茄精准育种流程可分为4个阶段:(1)资源整合:从野生近缘种、地方品种到现代栽培种,建立丰富的番茄种质资源库;(2)多组学数据采集:利用电子舌、电子鼻(表型组)、质谱分析(代谢组)及高通量测序(基因组/转录组)获取海量生物信息;(3)AI集成分析:将多维度数据输入机器学习模型,进行QTL定位、mGWAS关联分析及差异表达基因(DEGs)鉴定,挖掘调控风味的关键位点;(4)育种应用:结合分子标记辅助选择(MAS)、双单倍体技术、基因编辑技术及全基因组选择(GS)技术,高效培育风味优异的新品种。LC-MS:液相色谱‒质谱联用技术;GC-MS:气相色谱‒质谱联用技术;NGS:高通量测序技术;QTL:数量性状位点;eQTL:表达量性状位点;mGWAS:代谢组全基因组关联研究;TWAS:转录组全基因组关联分析;DEGs:差异表达基因。该示意图由Germini3软件辅助修改
Fig. 3 Precision improvement strategy for tomato flavor traits driven by multi-omics big dataThe AI-driven precision breeding process for tomatoes comprises four stages: 1) Integration germplasm resources: Establish a comprehensive tomato germplasm repository spanning wild relatives, landraces, and modern cultivars. 2) Multi-omics data collection: Acquire vast biological information through electric tongue/nose (phenomics), mass spectrometry (metabolomics), and high-throughput sequencing (genomics/transcriptomics). 3) AI-integrated analysis: Input multidimensional data into machine learning models to perform QTL mapping, mGWAS association analysis, and differential gene expression (DEG) identification, uncovering key loci regulating flavor. 4) Breeding applications: Combine molecular marker-assisted selection (MAS), double haploid technology, gene editing techniques, and genome-wide selection (GS) to efficiently develop new varieties with superior flavor. LC-MS: Liquid chromatography-mass spectrometry. GC-MS: Gas chromatography-mass spectrometry. NGS: Next-generation sequencing. QTL: Quantitative trait locus. eQTL: Expression quantitative trait locus. mGWAS: Metabolome-wide association study. TWAS: Transcriptome-wide association study. DEGs: Differentially expressed genes. This schematic diagram was modified with the assistance of Germini3 software
| [1] | Du MM, Sun CL, Deng L, et al. Molecular breeding of tomato: Advances and challenges [J]. J Integr Plant Biol, 2025, 67(3): 669-721. |
| [2] | Razifard H, Ramos A, Della Valle AL, et al. Genomic evidence for complex domestication history of the cultivated tomato in Latin America [J]. Mol Biol Evol, 2020, 37(4): 1118-1132. |
| [3] | 李君明, 项朝阳, 王孝宣, 等. “十三五”我国番茄产业现状及展望 [J]. 中国蔬菜, 2021(2): 13-20. |
| Li JM, Xiang CY, Wang XX, et al. Current situation of tomato industry in China during ‘the thirteenth Five-Year Plan’ period and future prospect [J]. China Veg, 2021(2): 13-20. | |
| [4] | Klee HJ, Tieman DM. The genetics of fruit flavour preferences [J]. Nat Rev Genet, 2018, 19(6): 347-356. |
| [5] | Wang SC, Qiang Q, Xiang LJ, et al. Targeted approaches to improve tomato fruit taste [J]. Hortic Res, 2023, 10(1): uhac229. |
| [6] | Kaur G, Abugu M, Tieman D. The dissection of tomato flavor: biochemistry, genetics, and omics [J]. Front Plant Sci, 2023, 14: 1144113. |
| [7] | Wang Y, Sun CL, Ye ZB, et al. The genomic route to tomato breeding: Past, present, and future [J]. Plant Physiol, 2024, 195(4): 2500-2514. |
| [8] | Shen SQ, Zhan CS, Yang CK, et al. Metabolomics-centered mining of plant metabolic diversity and function: Past decade and future perspectives [J]. Mol Plant, 2023, 16(1): 43-63. |
| [9] | Li Y, Chen Y, Zhou L, et al. MicroTom metabolic network: rewiring tomato metabolic regulatory network throughout the growth cycle [J]. Mol Plant, 2020, 13(8): 1203-1218. |
| [10] | Lee H, MacPherson LJ, Parada CA, et al. Rewiring the taste system [J]. Nature, 2017, 548(7667): 330-333. |
| [11] | Green BG, Dalton P, Byrnes N, et al. Chemesthesis: chemical touch in food and eating [M]. Hoboken: Wiley-Blackwell, 2016: 293. |
| [12] | Mombaerts P. The human repertoire of odorant receptor genes and pseudogenes [J]. Annu Rev Genom Hum Genet, 2001, 2: 493-510. |
| [13] | Bushdid C, Magnasco MO, Vosshall LB, et al. Humans can discriminate more than 1 trillion olfactory stimuli [J]. Science, 2014, 343(6177): 1370-1372. |
| [14] | Bauchet G, Grenier S, Samson N, et al. Identification of major loci and genomic regions controlling acid and volatile content in tomato fruit: implications for flavor improvement [J]. New Phytol, 2017, 215(2): 624-641. |
| [15] | Colantonio V, Ferrão LFV, Tieman DM, et al. Metabolomic selection for enhanced fruit flavor [J]. Proc Natl Acad Sci U S A, 2022, 119(7): e2115865119. |
| [16] | Tieman D, Bliss P, McIntyre LM, et al. The chemical interactions underlying tomato flavor preferences [J]. Curr Biol, 2012, 22(11): 1035-1039. |
| [17] | Tieman D, Zhu GT, Resende MFR Jr, et al. A chemical genetic roadmap to improved tomato flavor [J]. Science, 2017, 355(6323): 391-394. |
| [18] | Zhang ZH, Ye WZ, Li C, et al. Volatilomics-based discovery of key volatiles affecting flavor quality in tomato [J]. Foods, 2024, 13(6): 879. |
| [19] | Klee HJ, Giovannoni JJ. Genetics and control of tomato fruit ripening and quality attributes [J]. Annu Rev Genet, 2011, 45: 41-59. |
| [20] | Tao JJ, Wang YX, Fernie AR, et al. Transcriptomic and epigenetic signatures of tomato fruit after postharvest UV-C irradiation are associated with the maintenance of fruit quality [J]. Plant Biotechnol J, 2025, 23(10): 4500-4502. |
| [21] | D’Aoust MA, Yelle S, Nguyen-Quoc B. Antisense inhibition of tomato fruit sucrose synthase decreases fruit setting and the sucrose unloading capacity of young fruit [J]. Plant Cell, 1999, 11(12): 2407-2418. |
| [22] | Klann EM, Hall B, Bennett AB. Antisense acid invertase (TIV1) gene alters soluble sugar composition and size in transgenic tomato fruit [J]. Plant Physiol, 1996, 112(3): 1321-1330. |
| [23] | Cai HX, Liang MY, Qin X, et al. Tonoplast sugar transporters coordinately regulate tomato fruit development and quality [J]. Plant Commun, 2025, 6(5): 101314. |
| [24] | Zhu GT, Wang SC, Huang ZJ, et al. Rewiring of the fruit metabolome in tomato breeding [J]. Cell, 2018, 172(1/2): 249-261.e12. |
| [25] | Zhang JZ, Lyu HJ, Chen J, et al. Releasing a sugar brake generates sweeter tomato without yield penalty [J]. Nature, 2024, 635(8039): 647-656. |
| [26] | Nguyen CV, Vrebalov JT, Gapper NE, et al. Tomato GOLDEN2 - LIKE transcription factors Reveal molecular gradients that function during fruit development and ripening [J]. Plant Cell, 2014, 26(2): 585-601. |
| [27] | Powell ALT, Nguyen CV, Hill T, et al. Uniform ripening encodes a Golden 2 - like transcription factor regulating tomato fruit chloroplast development [J]. Science, 2012, 336(6089): 1711-1715. |
| [28] | Wang Y, Shi CM, Ge PF, et al. A 21-bp InDel in the promoter of STP1 selected during tomato improvement accounts for soluble solid content in fruits [J]. Hortic Res, 2023, 10(3): uhad009. |
| [29] | Qin GZ, Zhu Z, Wang WH, et al. A tomato vacuolar invertase inhibitor mediates sucrose metabolism and influences fruit ripening [J]. Plant Physiol, 2016, 172(3): 1596-1611. |
| [30] | Sagar M, Chervin C, Mila I, et al. SlARF4, an auxin response factor involved in the control of sugar metabolism during tomato fruit development [J]. Plant Physiol, 2013, 161(3): 1362-1374. |
| [31] | Anthon GE, LeStrange M, Barrett DM. Changes in pH, acids, sugars and other quality parameters during extended vine holding of ripe processing tomatoes [J]. J Sci Food Agric, 2011, 91(7): 1175-1181. |
| [32] | Paolo D, Bianchi G, Morelli CF, et al. Impact of drying techniques, seasonal variation and organic growing on flavor compounds profiles in two Italian tomato varieties [J]. Food Chem, 2019, 298: 125062. |
| [33] | Scarano A, Gerardi C, Sommella E, et al. Engineering the polyphenolic biosynthetic pathway stimulates metabolic and molecular changes during fruit ripening in “Bronze” tomato [J]. Hortic Res, 2022, 9: uhac097. |
| [34] | Agius C, von Tucher S, Poppenberger B, et al. Quantification of sugars and organic acids in tomato fruits [J]. MethodsX, 2018, 5: 537-550. |
| [35] | Centeno DC, Osorio S, Nunes-Nesi A, et al. Malate plays a crucial role in starch metabolism, ripening, and soluble solid content of tomato fruit and affects postharvest softening [J]. Plant Cell, 2011, 23(1): 162-184. |
| [36] | Lobit P, Genard M, Soing P, et al. Modelling malic acid accumulation in fruits: relationships with organic acids, potassium, and temperature [J]. J Exp Bot, 2006, 57(6): 1471-1483. |
| [37] | Ma BQ, Chen J, Zheng HY, et al. Comparative assessment of sugar and malic acid composition in cultivated and wild apples [J]. Food Chem, 2015, 172: 86-91. |
| [38] | Cebolla-Cornejo J, Roselló S, Valcárcel M, et al. Evaluation of genotype and environment effects on taste and aroma flavor components of Spanish fresh tomato varieties [J]. J Agric Food Chem, 2011, 59(6): 2440-2450. |
| [39] | Ye J, Wang X, Hu TX, et al. An InDel in the promoter of Al-ACTIVATED MALATE TRANSPORTER9 selected during tomato domestication determines fruit malate contents and aluminum tolerance [J]. Plant Cell, 2017, 29(9): 2249-2268. |
| [40] | Lu Y, Zhu HL. The regulation of nutrient and flavor metabolism in tomato fruit [J]. Veg Res, 2022, 2(1): 1-14. |
| [41] | Martínez-Esteso MJ, Sellés-Marchart S, Lijavetzky D, et al. A DIGE-based quantitative proteomic analysis of grape berry flesh development and ripening reveals key events in sugar and organic acid metabolism [J]. J Exp Bot, 2011, 62(8): 2521-2569. |
| [42] | Sweetman C, Deluc LG, Cramer GR, et al. Regulation of malate metabolism in grape berry and other developing fruits [J]. Phytochemistry, 2009, 70(11/12): 1329-1344. |
| [43] | Etienne A, Génard M, Lobit P, et al. What controls fleshy fruit acidity? A review of malate and citrate accumulation in fruit cells [J]. J Exp Bot, 2013, 64(6): 1451-1469. |
| [44] | Morgan MJ, Osorio S, Gehl B, et al. Metabolic engineering of tomato fruit organic acid content guided by biochemical analysis of an introgression line [J]. Plant Physiol, 2012, 161(1): 397-407. |
| [45] | Gai WX, Yuan LD, Yang F, et al. Genome-wide variants and optimal allelic combinations for citric acid in tomato [J]. Hortic Res, 2024, 11(5): uhae070. |
| [46] | Gai WX, Yang F, Yuan LD, et al. Multiple-model GWAS identifies optimal allelic combinations of quantitative trait loci for malic acid in tomato [J]. Hortic Res, 2023, 10(4): uhad021. |
| [47] | Vranová E, Coman D, Gruissem W. Network analysis of the MVA and MEP pathways for isoprenoid synthesis [J]. Annu Rev Plant Biol, 2013, 64: 665-700. |
| [48] | Sawai S, Ohyama K, Yasumoto S, et al. Sterol side chain reductase 2 is a key enzyme in the biosynthesis of cholesterol, the common precursor of toxic steroidal glycoalkaloids in potato [J]. Plant Cell, 2014, 26(9): 3763-3774. |
| [49] | Liu YM, Liu XW, Li YG, et al. Potato steroidal glycoalkaloids: properties, biosynthesis, regulation and genetic manipulation [J]. Mol Hortic, 2024, 4: 43. |
| [50] | Nakayasu M, Umemoto N, Ohyama K, et al. A dioxygenase catalyzes steroid 16 α-hydroxylation in steroidal glycoalkaloid biosynthesis [J]. Plant Physiol, 2017, 175(1): 120-133. |
| [51] | Umemoto N, Nakayasu M, Ohyama K, et al. Two cytochrome P450 monooxygenases catalyze early hydroxylation steps in the potato steroid glycoalkaloid biosynthetic pathway [J]. Plant Physiol, 2016, 171(4): 2458-2467. |
| [52] | Boccia M, Kessler D, Seibt W, et al. A scaffold protein manages the biosynthesis of steroidal defense metabolites in plants [J]. Science, 2024, 386(6728): 1366-1372. |
| [53] | Jozwiak A, Panda S, Akiyama R, et al. A cellulose synthase-like protein governs the biosynthesis of Solanum alkaloids [J]. Science, 2024, 386(6728): eadq5721. |
| [54] | Sonawane PD, Heinig U, Panda S, et al. Short-chain dehydrogenase/reductase governs steroidal specialized metabolites structural diversity and toxicity in the genus Solanum [J]. Proc Natl Acad Sci USA, 2018, 115(23): E5419-E5428. |
| [55] | Sonawane PD, Gharat SA, Jozwiak A, et al. A BAHD-type acyltransferase concludes the biosynthetic pathway of non-bitter glycoalkaloids in ripe tomato fruit [J]. Nat Commun, 2013, 14: 4540. |
| [56] | Sonawane PD, Jozwiak A, Barbole R, et al. 2-oxoglutarate-dependent dioxygenases drive expansion of steroidal alkaloid structural diversity in the genus Solanum [J]. New Phytol, 2022, 234(4): 1394-1410. |
| [57] | Cárdenas PD, Sonawane PD, Heinig U, et al. Pathways to defense metabolites and evading fruit bitterness in genus Solanum evolved through 2-oxoglutarate-dependent dioxygenases [J]. Nat Commun, 2019, 10: 5169. |
| [58] | Cárdenas PD, Sonawane PD, Pollier J, et al. GAME9 regulates the biosynthesis of steroidal alkaloids and upstream isoprenoids in the plant mevalonate pathway [J]. Nat Commun, 2016, 7: 10654. |
| [59] | Bai F, Wu MB, Huang W, et al. Removal of toxic steroidal glycoalkaloids and bitterness in tomato is controlled by a complex epigenetic and genetic network [J]. Sci Adv, 2025, 11(8): eads9601. |
| [60] | Tieman D, Zeigler M, Schmelz E, et al. Functional analysis of a tomato salicylic acid methyl transferase and its role in synthesis of the flavor volatile methyl salicylate[J]. Plant J, 2010, 62(1): 113-123. |
| [61] | Rambla JL, Tikunov YM, Monforte AJ, et al. The expanded tomato fruit volatile landscape [J]. J Exp Bot, 2014, 65(16): 4613-4623. |
| [62] | Garbowicz K, Liu ZY, Alseekh S, et al. Quantitative trait loci analysis identifies a prominent gene involved in the production of fatty acid-derived flavor volatiles in tomato [J]. Mol Plant, 2018, 11(9): 1147-1165. |
| [63] | Distefano M, Cincotta F, Giuffrida F, et al. Preharvest applications of monopotassium phosphate to improve fruit quality and volatilome composition in cold-stored cherry tomato [J]. Hortic Plant J, 2025, 11(3): 1231-1247. |
| [64] | Zhao JT, Sauvage C, Zhao JH, et al. Meta-analysis of genome-wide association studies provides insights into genetic control of tomato flavor [J]. Nat Commun, 2019, 10: 1534. |
| [65] | Wang YT, Liu RL, Huang SW, et al. Effects of potassium application on flavor compounds of cherry tomato fruits [J]. J Plant Nutr, 2009, 32(9): 1451-1468. |
| [66] | Kochevenko A, Araújo WL, Maloney GS, et al. Catabolism of branched chain amino acids supports respiration but not volatile synthesis in tomato fruits [J]. Mol Plant, 2012, 5(2): 366-375. |
| [67] | Li X, Tieman D, Alseekh S, et al. Natural variations in the Sl-AKR9 aldo/keto reductase gene impact fruit flavor volatile and sugar contents [J]. Plant J, 2023, 115(4): 1134-1150. |
| [68] | Tikunov YM, Roohanitaziani R, Meijer-Dekens F, et al. The genetic and functional analysis of flavor in commercial tomato: the FLORAL4 gene underlies a QTL for floral aroma volatiles in tomato fruit [J]. Plant J, 2020, 103(3): 1189-1204. |
| [69] | Tikunov YM, Molthoff J, de Vos RCH, et al. NON-SMOKY GLYCOSYLTRANSFERASE1 prevents the release of smoky aroma from tomato fruit [J]. Plant Cell, 2013, 25(8): 3067-3078. |
| [70] | Zhou F, Pichersky E. The complete functional characterisation of the terpene synthase family in tomato [J]. New Phytol, 2020, 226(5): 1341-1360. |
| [71] | Pérez-Pérez J, Minguillón S, Kabbas-Piñango E, et al. Metabolic crosstalk between hydroxylated monoterpenes and salicylic acid in tomato defense response against bacteria [J]. Plant Physiol, 2024, 195(3): 2323-2338. |
| [72] | Paetzold H, Garms S, Bartram S, et al. The isogene 1-deoxy-D-xylulose 5-phosphate synthase 2 controls isoprenoid profiles, precursor pathway allocation, and density of tomato trichomes [J]. Mol Plant, 2010, 3(5): 904-916. |
| [73] | Liao P, Chen XJ, Wang MF, et al. Improved fruit α-tocopherol, carotenoid, squalene and phytosterol contents through manipulation of Brassica juncea 3-HYDROXY-3-METHYLGLUTARYL-COA SYNTHASE1 in transgenic tomato [J]. Plant Biotechnol J, 2018, 16(3): 784-796. |
| [74] | Yang C, Gao X, Jiang Y, et al. Synergy between methylerythritol phosphate pathway and mevalonate pathway for isoprene production in Escherichia coli [J]. Metab Eng, 2016, 37: 79-91. |
| [75] | Reddy UK, Karnatam KS, Talavera-Caro A, et al. Uncovering the genetic architecture of pungency, carotenoids, and flavor in Capsicum chinense via TWAS-mGWAS integration and spatial transcriptomics [J]. Hortic Res, 2025, 12(12): uhaf243. |
| [76] | Liu Y, Qu WJ, Liu YX, et al. Impact of different peeling treatments on the isomerization and micellization of carotenoids and the flavor in tomato pulp [J]. Food Chem, 2025, 476: 143452. |
| [77] | Cheng GT, Li YS, Qi SM, et al. SlCCD1A enhances the aroma quality of tomato fruits by promoting the synthesis of carotenoid-derived volatiles [J]. Foods, 2021, 10(11): 2678. |
| [78] | Gao Y, Bian HX, Wang HQ, et al. The m6A reader SlYTH1 regulates flavor-related volatiles biosynthesis via affecting mRNA stability and translation in tomato fruit [J]. Plant Physiol, 2025, 198(3): kiaf245. |
| [79] | Bian HX, Song PZ, Gao Y, et al. The m6A reader SlYTH2 negatively regulates tomato fruit aroma by impeding the translation process [J]. Proc Natl Acad Sci USA, 2024, 121(28): e2405100121. |
| [80] | Zhang XS, Feng CY, Wang MN, et al. Plasma membrane-localized SlSWEET7a and SlSWEET14 regulate sugar transport and storage in tomato fruits [J]. Hortic Res, 2021, 8: 186. |
| [81] | Li FM, Lin JS, Ahiakpa JK, et al. ZF protein C2H2-71 regulates the soluble solids content in tomato by inhibiting LIN5 [J]. J Integr Agric, 2025, 24(6): 2190-2202. |
| [82] | Wang BK, Li N, Huang SY, et al. Enhanced soluble sugar content in tomato fruit using CRISPR/Cas9-mediated SlINVINH1 and SlVPE5 gene editing [J]. PeerJ, 2021, 9: e12478. |
| [83] | Ma XM, Zhang YJ, Turečková V, et al. The NAC transcription factor SlNAP2 regulates leaf senescence and fruit yield in tomato [J]. Plant Physiol, 2018, 177(3): 1286-1302. |
| [84] | Lin LK, Yuan KL, Huang YD, et al. A WRKY transcription factor PbWRKY40 from Pyrus betulaefolia functions positively in salt tolerance and modulating organic acid accumulation by regulating PbVHA-B1 expression [J]. Environ Exp Bot, 2022, 196: 104782. |
| [85] | Liu RL, Li BQ, Qin GZ, et al. Identification and functional characterization of a tonoplast dicarboxylate transporter in tomato (Solanum lycopersicum) [J]. Front Plant Sci, 2017, 8: 186. |
| [86] | Osorio S, Vallarino JG, Szecowka M, et al. Alteration of the interconversion of pyruvate and malate in the plastid or cytosol of ripening tomato fruit invokes diverse consequences on sugar but similar effects on cellular organic acid, metabolism, and transitory starch accumulation [J]. Plant Physiol, 2013, 161(2): 628-643. |
| [87] | Liu YY, Hu HR, Yang RJ, et al. Current advances in the biosynthesis, metabolism, and transcriptional regulation of α-tomatine in tomato [J]. Plants, 2023, 12(18): 3289. |
| [88] | Gharat SA, Jozwiak A, Rogachev I, et al. Complex engineering of Solanum alkaloids structural diversity in Nicotiana benthamiana [J]. Plant Biotechnol J, 2026, 24(1): 328-345. |
| [89] | Liu ZQ, Wu JX, Wang LC, et al. Integration of transcriptome and metabolome reveals regulatory mechanisms of volatile flavor formation during tomato fruit ripening [J]. Hortic Plant J, 2025, 11(2): 680-692. |
| [90] | Moummou H, Tonfack LB, Chervin C, et al. Functional characterization of SlscADH1, a fruit-ripening-associated short-chain alcohol dehydrogenase of tomato [J]. J Plant Physiol, 2012, 169(15): 1435-1444. |
| [91] | Zhang C, Duan WY, Chen KS, et al. Transcriptome and methylome analysis reveals effects of ripening on and off the vine on flavor quality of tomato fruit [J]. Postharvest Biol Technol, 2020, 162: 111096. |
| [92] | Gao Y, Lin YJ, Xu M, et al. The role and interaction between transcription factor NAC-NOR and DNA demethylase SlDML2 in the biosynthesis of tomato fruit flavor volatiles [J]. New Phytol, 2022, 235(5): 1913-1926. |
| [93] | Ye W, Zhang WM, Liu TM, et al. Improvement of ethanol production in Saccharomyces cerevisiae by high-efficient disruption of the ADH2 gene using a novel recombinant TALEN vector [J]. Front Microbiol, 2016, 7: 1067. |
| [94] | Amaya I, Roldán-Guerra FJ, Ordóñez-Díaz JL, et al. Differential expression of CCD4(4B) drives natural variation in fruit carotenoid content in strawberry (Fragaria spp.) [J]. Plant Biotechnol J, 2025, 23(3): 679-691. |
| [95] | Cao X, Du R, Xu YC, et al. Phytoene synthases 1 modulates tomato fruit quality through influencing the metabolic flux between carotenoid and flavonoid pathways [J]. Hortic Plant J, 2024, 10(6): 1383-1397. |
| [96] | Howe GA, Lee GI, Itoh A, et al. Cytochrome P450-dependent metabolism of oxylipins in tomato. cloning and expression of allene oxide synthase and fatty acid hydroperoxide lyase [J]. Plant Physiol, 2000, 123(2): 711-724. |
| [97] | Maloney GS, Kochevenko A, Tieman DM, et al. Characterization of the branched-chain amino acid aminotransferase enzyme family in tomato [J]. Plant Physiol, 2010, 153(3): 925-936. |
| [98] | Wu Q, Tao XY, Ai XZ, et al. Contribution of abscisic acid to aromatic volatiles in cherry tomato (Solanum lycopersicum L.) fruit during postharvest ripening [J]. Plant Physiol Biochem, 2018, 130: 205-214. |
| [99] | Singh RK, Srivastava S, Chidley HG, et al. Overexpression of mango alcohol dehydrogenase (MiADH1) mimics hypoxia in transgenic tomato and alters fruit flavor components [J]. Agri Gene, 2018, 7: 23-33. |
| [100] | Gao L, Gonda I, Sun HH, et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor [J]. Nat Genet, 2019, 51(6): 1044-1051. |
| [101] | Li X, Tieman D, Liu ZM, et al. Identification of a lipase gene with a role in tomato fruit short-chain fatty acid-derived flavor volatiles by genome-wide association [J]. Plant J, 2020, 104(3): 631-644. |
| [102] | Simkin AJ, Schwartz SH, Auldridge M, et al. The tomato carotenoid cleavage dioxygenase 1 genes contribute to the formation of the flavor volatiles β-ionone, pseudoionone, and geranylacetone [J]. Plant J, 2004, 40(6): 882-892. |
| [103] | Jian W, Cao HH, Yuan S, et al. SlMYB75, an MYB-type transcription factor, promotes anthocyanin accumulation and enhances volatile aroma production in tomato fruits [J]. Hortic Res, 2019, 6: 22. |
| [104] | Sun YG, He YQ, Wang HX, et al. Genome-wide identification and expression analysis of GDSL esterase/lipase genes in tomato [J]. J Integr Agric, 2022, 21(2): 389-406. |
| [105] | Zhang HX, Mo FL, Li D, et al. Genome-wide identification and expression analysis of the GT8 gene family in tomato (Solanum lycopersicum) and the functional of SlGolS1 under cold stress [J]. Sci Hortic, 2024, 338: 113686. |
| [106] | Akiyama R, Nakayasu M, Umemoto N, et al. Tomato E8 encodes a C-27 hydroxylase in metabolic detoxification of α-tomatine during fruit ripening [J]. Plant Cell Physiol, 2021, 62(5): 775-783. |
| [107] | Chen M, Tian FY, Xie Y, et al. A comprehensive study of carboxylesterases involved in the degradation of volatile esters in horticultural crops [J]. Ind Crops Prod, 2025, 236: 122039. |
| [108] | Fridman E, Carrari F, Liu YS, et al. Zooming in on a quantitative trait for tomato yield using interspecific introgressions [J]. Science, 2004, 305(5691): 1786-1789. |
| [109] | McCurdy DW, Dibley S, Cahyanegara R, et al. Functional characterization and RNAi-mediated suppression reveals roles for hexose transporters in sugar accumulation by tomato fruit [J]. Mol Plant, 2010, 3(6): 1049-1063. |
| [110] | Liu H, Zhang JQ, Zhang RR, et al. SlMYB1R1-SlSWEET12c module synergistically promotes sugar accumulation in tomato fruits [J]. Plant J, 2025, 121(4): e70062. |
| [111] | Carviel JL, Al-Daoud F, Neumann M, et al. Forward and reverse genetics to identify genes involved in the age-related resistance response in Arabidopsis thaliana [J]. Mol Plant Pathol, 2009, 10(5): 621-634. |
| [112] | Lippman ZB, Semel Y, Zamir D. An integrated view of quantitative trait variation using tomato interspecific introgression lines [J]. Curr Opin Genet Dev, 2007, 17(6): 545-552. |
| [113] | Eshed Y, Zamir D. An introgression line population of Lycopersicon pennellii in the cultivated tomato enables the identification and fine mapping of yield-associated QTL [J]. Genetics, 1995, 141(3): 1147-1162. |
| [114] | Jaganathan D, Bohra A, Thudi M, et al. Fine mapping and gene cloning in the post-NGS era: advances and prospects [J]. Theor Appl Genet, 2020, 133(5): 1791-1810. |
| [115] | Fang CY, Luo J. Metabolic GWAS-based dissection of genetic bases underlying the diversity of plant metabolism [J]. Plant J, 2019, 97(1): 91-100. |
| [116] | Sauvage C, Segura V, Bauchet G, et al. Genome-wide association in tomato reveals 44 candidate loci for fruit metabolic traits [J]. Plant Physiol, 2014, 165(3): 1120-1132. |
| [117] | Li N, He Q, Wang J, et al. Super-pangenome analyses highlight genomic diversity and structural variation across wild and cultivated tomato species [J]. Nat Genet, 2023, 55(5): 852-860. |
| [118] | Fernandez-Pozo N, Zheng Y, Snyder SI, et al. The tomato expression atlas [J]. Bioinformatics, 2017, 33(15): 2397-2398. |
| [119] | Zouine M, Maza E, Djari A, et al. TomExpress, a unified tomato RNA-Seq platform for visualization of expression data, clustering and correlation networks [J]. Plant J, 2017, 92(4): 727-735. |
| [120] | Osorio S, Alba R, Nikoloski Z, et al. Integrative comparative analyses of transcript and metabolite profiles from pepper and tomato ripening and development stages uncovers species-specific patterns of network regulatory behavior [J]. Plant Physiol, 2012, 159(4): 1713-1729. |
| [121] | Alseekh S, Tohge T, Wendenberg R, et al. Identification and mode of inheritance of quantitative trait loci for secondary metabolite abundance in tomato [J]. Plant Cell, 2015, 27(3): 485-512. |
| [122] | 谢剑平, 毛健, 张启东, 等. 风味科学的研究内涵与前沿挑战 [J]. 食品科学技术学报, 2025, 43(1): 1-16. |
| Xie JP, Mao J, Zhang QD, et al. Research connotation, frontiers and challenges of flavor science [J]. J Food Sci Technol, 2025, 43(1): 1-16. | |
| [123] | Lou HC, Li SJ, Shi ZH, et al. Engineering source-sink relations by prime editing confers heat-stress resilience in tomato and rice [J]. Cell, 2025, 188(2): 530-549.e20. |
| [124] | Dunkel A, Steinhaus M, Kotthoff M, et al. Nature’s chemical signatures in human olfaction: a foodborne perspective for future biotechnology [J]. Angew Chem Int Ed, 2014, 53(28): 7124-7143. |
| [125] | Liu SH, Shi TY, Yu JW, et al. Research on bitter peptides in the field of bioinformatics: a comprehensive review [J]. Int J Mol Sci, 2024, 25(18): 9844. |
| [126] | Ge XY, Zhou YJ, Li Q, et al. Machine learning for food flavor prediction and regulation: models, data integration, and future perspectives [J]. J Adv Res, 2025. DOI: 10.1016/j.jare.2025.10.018 . |
| [127] | Ferrão LFV, Sater H, Lyrene P, et al. Terpene volatiles mediates the chemical basis of blueberry aroma and consumer acceptability [J]. Food Res Int, 2022, 158: 111468. |
| [128] | Villena J, Moreno C, Roselló S, et al. Breeding tomato flavor: Modeling consumer preferences of tomato landraces [J]. Sci Hortic, 2023, 308: 111597. |
| [129] | Wei Z, Li SH, Jiang YJ, et al. Recent advances in implicit salt reduction innovative technologies: Innovative strategies and future challenges [J]. Food Res Int, 2025, 221: 117346. |
| [130] | Liang WT, Zhong RX, Ma ZG, et al. Intelligent sensory technologies, NIR spectroscopy and chemometrics combined with machine learning based on multi-source data fusion for comprehensive evaluation of Sinapis Semen in different processing degrees [J]. J Pharm Biomed Anal, 2026, 268: 117212. |
| [131] | Fu JJ, Hao YF, Li HH, et al. Integration of genomic selection with doubled-haploid evaluation in hybrid breeding: From GS 1.0 to GS 4.0 and beyond [J]. Mol Plant, 2022, 15(4): 577-580. |
| [132] | Duangjit J, Causse M, Sauvage C. Efficiency of genomic selection for tomato fruit quality [J]. Mol Breeding, 2016, 36(3): 29. |
| [133] | Cappetta E, Andolfo G, Di Matteo A, et al. Accelerating tomato breeding by exploiting genomic selection approaches [J]. Plants, 2020, 9(9): 1236. |
| [134] | Cappetta E, Andolfo G, Guadagno A, et al. Tomato genomic prediction for good performance under high-temperature and identification of loci involved in thermotolerance response [J]. Hortic Res, 2021, 8: 212. |
| [135] | Yamamoto E, Matsunaga H, Onogi A, et al. A simulation-based breeding design that uses whole-genome prediction in tomato [J]. Sci Rep, 2016, 6: 19454. |
| [136] | Xia XH, Cheng XH, Li R, et al. Advances in application of genome editing in tomato and recent development of genome editing technology [J]. Theor Appl Genet, 2021, 134(9): 2727-2747. |
| [137] | Li BS, Sun C, Li JY, et al. Targeted genome-modification tools and their advanced applications in crop breeding [J]. Nat Rev Genet, 2024, 25(9): 603-622. |
| [138] | Wang DD, Samsulrizal N, Yan C, et al. Characterisation of CRISPR mutants targeting genes modulating pectin degradation in ripening tomato [J]. Plant Physiol, 2018: pp.01187.2018. |
| [139] | Nonaka S, Arai C, Takayama M, et al. Efficient increase of ɣ-aminobutyric acid (GABA) content in tomato fruits by targeted mutagenesis [J]. Sci Rep, 2017, 7: 7057. |
| [140] | Cao XM, Wei CY, Duan WY, et al. Transcriptional and epigenetic analysis reveals that NAC transcription factors regulate fruit flavor ester biosynthesis [J]. Plant J, 2021, 106(3): 785-800. |
| [141] | Li Q, Sapkota M, van der Knaap E. Perspectives of CRISPR/Cas-mediated cis-engineering in horticulture: unlocking the neglected potential for crop improvement [J]. Hortic Res, 2020, 7: 36. |
| [142] | Nguyen NH, Bui TP, Le NT, et al. Disrupting Sc-uORFs of a transcription factor bZIP1 using CRISPR/Cas9 enhances sugar and amino acid contents in tomato (Solanum lycopersicum) [J]. Planta, 2023, 257(3): 57. |
| [143] | Rodríguez-Leal D, Lemmon ZH, Man J, et al. Engineering quantitative trait variation for crop improvement by genome editing [J]. Cell, 2017, 171(2): 470-480.e8. |
| [144] | Li XB, Xie JY, Dong C, et al. Efficient and heritable A-to-K base editing in rice and tomato [J]. Hortic Res, 2024, 11(1): uhad250. |
| [145] | Niu QF, Wu SQ, Xie HT, et al. Efficient A·T to G·C base conversions in dicots using adenine base editors expressed under the tomato EF1α promoter [J]. Plant Biotechnol J, 2023, 21(1): 5-7. |
| [1] | 刘淼, 林涛, 贾乐松, 胡丰, 李涛, 李志万, 刘美芳, 郑方燕, 崔龙. 从野生到栽培:番茄果实色泽的演化与调控机制[J]. 生物技术通报, 2026, 42(3): 187-202. |
| [2] | 李亚妮, 韩鸿宇, 耿梦爽, 米若兰, 王韦琪, 于文静, 孟宪文, 李传友. ChiC基因调控番茄灰霉病抗性的机制研究[J]. 生物技术通报, 2026, 42(3): 255-262. |
| [3] | 程云霞, 张俊红, 叶杰. 番茄果实可溶性固形物积累的遗传调控研究进展[J]. 生物技术通报, 2026, 42(3): 145-155. |
| [4] | 颜晨琳, 李凡, 闫春婷, 程蛟文, 胡开林, 叶志彪, 宋建文. 番茄果实形态发育相关基因研究进展[J]. 生物技术通报, 2026, 42(3): 172-186. |
| [5] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [6] | 杜丹, 郭翔, 胡鑫, 潘宇. 质体发育调控果实成熟与品质的研究进展[J]. 生物技术通报, 2026, 42(3): 48-59. |
| [7] | 李成泉, 史庆华, 杨晓玉. 园艺作物果实发育的miRNA调控网络:从分子机制到种质创新[J]. 生物技术通报, 2026, 42(3): 19-36. |
| [8] | 王潇奕, 李金焱, 邢醒, 朱鸿亮. 基于乙烯响应筛选调控番茄成熟且影响呼吸的基因及其功能分析[J]. 生物技术通报, 2026, 42(3): 275-282. |
| [9] | 李迎辉, 王杨博涵, 周浩博, 卢心如, 张珂欣, 于洋, 李传友, 孙传龙. 番茄VPE基因家族鉴定和抗逆功能分析[J]. 生物技术通报, 2026, 42(3): 263-274. |
| [10] | 刘欢, 郭发旭, 赵晓燕, 黄龙雨, 王健, 周国民, 张建华. 人工智能在DNA设计中的研究进展[J]. 生物技术通报, 2026, 42(1): 51-66. |
| [11] | 苏秀敏, 韩文清, 王佼, 李鹏, 王秋兰, 李万星, 曹晋军. 哈茨木霉M408的分离鉴定、生物学特性及对番茄早疫病的生防效果[J]. 生物技术通报, 2025, 41(9): 277-288. |
| [12] | 蔡如凤, 杨宇轩, 于基正, 李佳楠. 人工智能重塑蛋白质工程:从结构解析到合成生物学的算法革命[J]. 生物技术通报, 2025, 41(8): 1-10. |
| [13] | 王辉, 范灵熙, 孙纪录, 王苑, 伍宁丰, 田健, 关菲菲. 基于蛋白智能模型提升溶菌酶RPL187的热稳定性[J]. 生物技术通报, 2025, 41(7): 336-346. |
| [14] | 刘源, 赵冉, 卢振芳, 李瑞丽. 植物类胡萝卜素生物代谢途径及其功能研究进展[J]. 生物技术通报, 2025, 41(5): 23-31. |
| [15] | 侯亚涛, 李迎辉, 邓磊, 李常保, 李传友, 孙传龙. 番茄果重基因功能型分子标记的开发及群体基因型分析[J]. 生物技术通报, 2025, 41(4): 98-105. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||