








• 技术与方法 •
曾亚丹1,2,3, 毕佳俊1,2,3, 李兴龙1,2,4, 常悦1,2,4, 角德灵1,2,3, 魏红江1,2,3, 赵恒1,2,3( )
)
收稿日期:2025-12-24
出版日期:2026-05-08
通讯作者:
赵恒hengzhao2014@126.com基金资助:
ZENG Ya-dan1,2,3, BI Jia-jun1,2,3, LI Xing-long1,2,4, CHANG Yue1,2,4, JIAO De-ling1,2,3, WEI Hong-jiang1,2,3, ZHAO Heng1,2,3( )
)
Received:2025-12-24
Published:2026-05-08
摘要:
目的 系统评估CRISPR-Cas12a(Cpf1)系统在猪细胞中介导基因定点插入(knock-in)的潜能,并为其在猪基因组精准编辑中的应用提供实验依据与理论支撑。 方法 构建基于Acidaminococcus sp. Cas12a(AsCas12a)与Lachnospiraceae bacterium Cas12a(LbCas12a)的表达载体,并针对猪基因组安全位点Rosa26与AAVS1设计特异性crRNA,旨在比较两种Cas12a变体的切割效能。 结果 在AAVS1位点,AsCas12a的两条crRNA编辑效率分别为2.6%与1.2%,而LbCas12a对应的效率分别为6.5%与2.5%;在Rosa26位点,AsCas12a与LbCas12a的编辑效率分别为5.0%与12.7%,LbCas12a在猪源细胞中展现出相对较高的编辑活性。在此基础上,本研究构建了基于同源重组(HDR)、同源介导末端连接(HMEJ)、微同源介导末端连接(MMEJ)及非同源末端连接(NHEJ)4种策略的敲入载体,并评估了其整合效率。结果表明,上述4种修复途径均可介导目标基因在Rosa26和AAVS1位点的定点整合,其中HDR效率为35.0%、HMEJ(25.0%)、NHEJ与MMEJ均为15.0%。 结论 基于Cas12a基因编辑系统,在猪细胞中成功构建了4种定点插入策略,并对其效率进行了系统评估,该研究成果为拓展Cas12a系统在猪基因工程领域的应用提供了重要的实验依据和参考资料。
曾亚丹, 毕佳俊, 李兴龙, 常悦, 角德灵, 魏红江, 赵恒. Cas12a介导的猪细胞定点插入体系构建与效率评估[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1408.
ZENG Ya-dan, BI Jia-jun, LI Xing-long, CHANG Yue, JIAO De-ling, WEI Hong-jiang, ZHAO Heng. Construction and Efficiency Evaluation of the Cas12a-mediated Site-specific Knock-in System in Pig Cells[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1408.
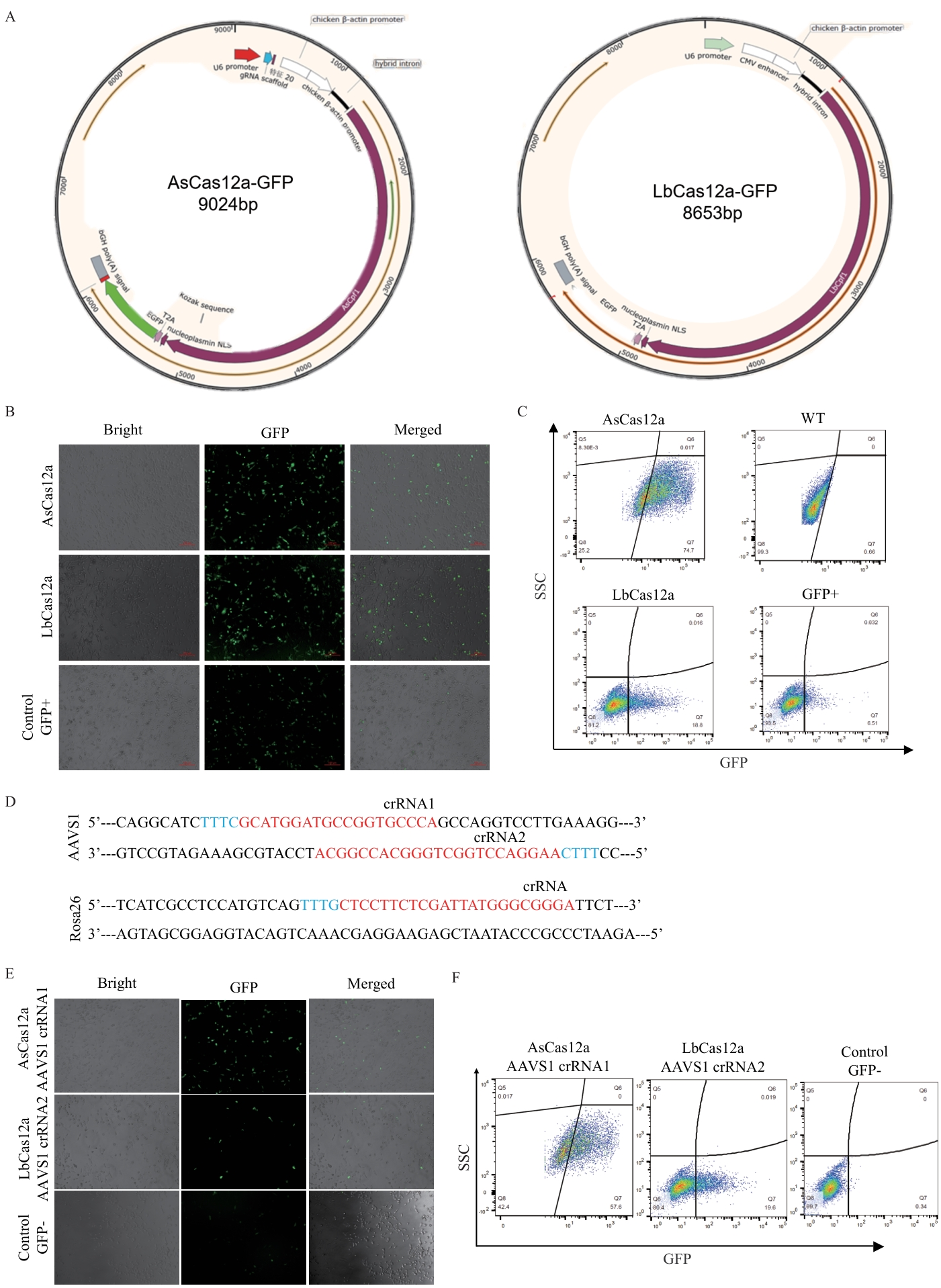
图1 AsCas12a和LbCas12a载体的构建和功能验证A:构建的AsCas12a、LbCas12a载体图谱;B:AsCas12a、LbCas12a载体转染PIEC后48 h在荧光显微镜下观察到细胞呈现绿色荧光;C:转染细胞的流式分析结果。AsCas12a和LbCas12a质粒转染细胞后,分别有74.7%和18.8%的细胞呈现绿色荧光;D:AAVS1和Rosa26基因靶位点设计示意图;E:AsCas12a和LbCas12a靶向AAVS1位点的细胞荧光图;F:细胞流式分析
Fig. 1 Construction and functional validation of AsCas12a and LbCas12a vectorsA: Profile of the constructed AsCas12a and LbCas12a vectors. B: Green fluorescence was observed in the cells at 48 h after transfecting PIEC cells with AsCas12a and LbCas12a vectors under a fluorescence microscope. C: Flow cytometry analysis result of transfected cells. The 74.7% and 18.8% of the cells, respectively, showed green fluorescence, after transfection with AsCas12a and LbCas12a plasmids. D: Schematic diagram of the design of gene targeting at AAVS1 and Rosa26 loci. E: Fluorescence images of cells targeting the AAVS1 locus with AsCas12a and LbCas12a. F: Flow cytometry analysis of the cells
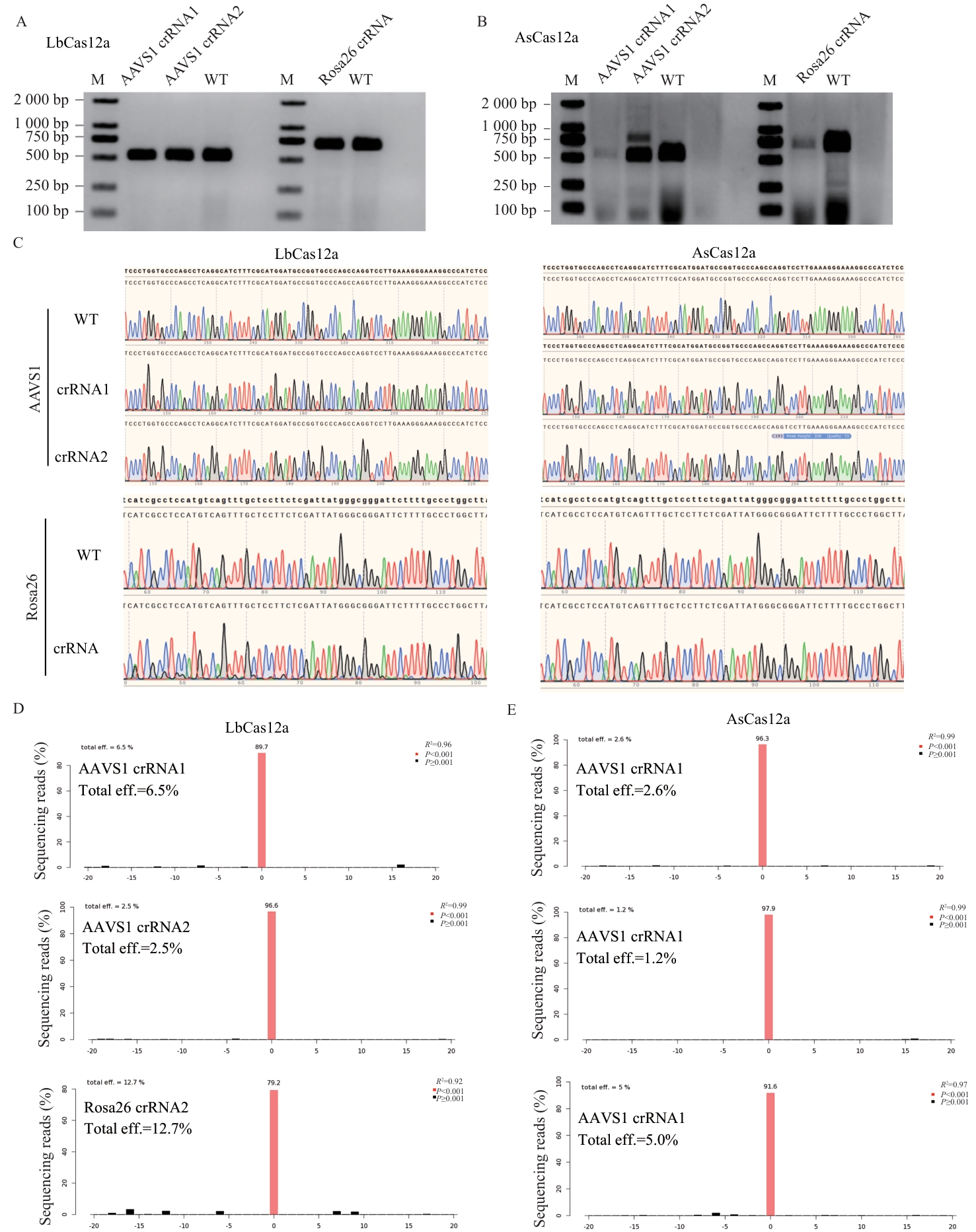
图2 AsCas12a和LbCas12a在AAVS1和Rosa26基因靶位点编辑效率分析A:LbCas12a AAVS1 crRNA1、LbCas12a AAVS1 crRNA2和LbCas12a Rosa26 crRNA转染细胞基因组的PCR结果图(M:Marker 2 000,WT: wild type)。B:AsCas12a AAVS1 crRNA1、AsCas12a AAVS1 crRNA2和AsCas12a Rosa26crRNA转染细胞基因组的PCR结果图(M:Marker 2 000,WT: wild type)。C:PCR测序结果的峰值图。D:TIDE分析LbCas12a AAVS1 crRNA1、LbCas12a AAVS1 crRNA2 PCR测序结果。E:AsCas12a AAVS1 crRNA1、AsCas12a AAVS1 crRNA2和AsCas12a Rosa26 crRNA PCR扩增产物的TIDE分析
Fig. 2 Analysis of editing efficiency of AsCas12a and LbCas12a at AAVS1 and Rosa26 lociA: PCR results of the genomic DNA from the cells transfected with LbCas12a AAVS1 crRNA1, LbCas12a AAVS1 crRNA2, and LbCas12a Rosa26 crRNA (M: Marker 2 000. WT: wild type). B: PCR results of the genomic DNA from the cells transfected with AsCas12a AAVS1 crRNA1, AsCas12a AAVS1 crRNA2, and AsCas12a Rosa26 crRNA (M: Marker 2 000, WT: wild type). C: Peak plot of the PCR sequencing results. D: TIDE analysis of the sequencing results from LbCas12a AAVS1 crRNA1 and LbCas12a AAVS1 crRNA2 PCR. E: TIDE analysis of the PCR amplification products of AsCas12a AAVS1 crRNA1, AsCas12a AAVS1 crRNA2, and AsCas12a Rosa26 crRNA
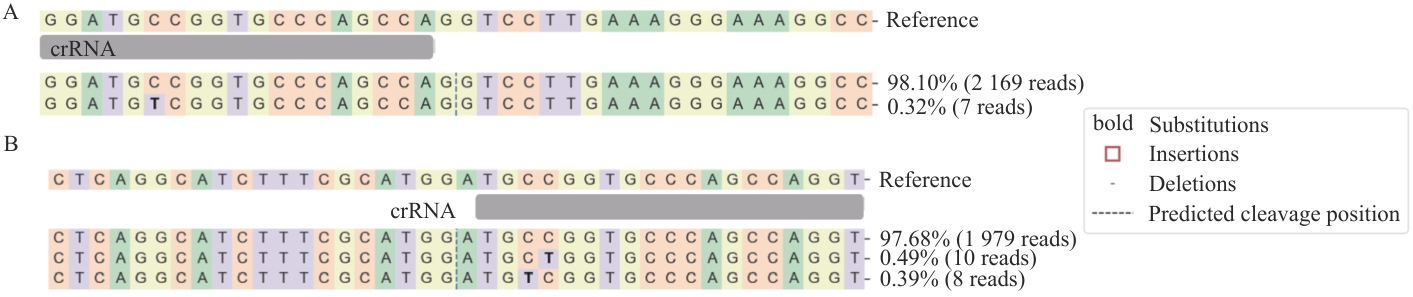
图3 通过FastNGS分析LbCas12a系统在猪PIEC细胞中的编辑准确性和突变性A:LbCas12a介导的AAVS1基因crRNA1靶位点PCR产物的高通量测序分析,共获得2 176条测序reads。B:LbCas12a介导的Rosa26基因crRNA2靶位点PCR产物的高通量测序分析,1 997条reads。加粗字体表示碱基突变;红色方框表示碱基插入;短横线(-)表示碱基缺失;虚线指示预测的编辑位点
Fig. 3 Editing Accuracy and Mutagenesis of the LbCas12a System in Pig PIEC Cells Analyzed by FastNGSA: High-throughput sequencing analysis of the PCR products from the AAVS1 gene crRNA1 target site mediated by LbCas12a, with a total of 2 176 sequencing reads obtained. B: High-throughput sequencing analysis of the PCR products from the Rosa26 gene crRNA2 target site mediated by LbCas12a, with 1 997 sequencing reads obtained. Bold characters indicate base mutations; red boxes indicate nucleotide insertions; hyphens (-) indicate deletions; and dotted lines denote predicted cleavage/editing sites
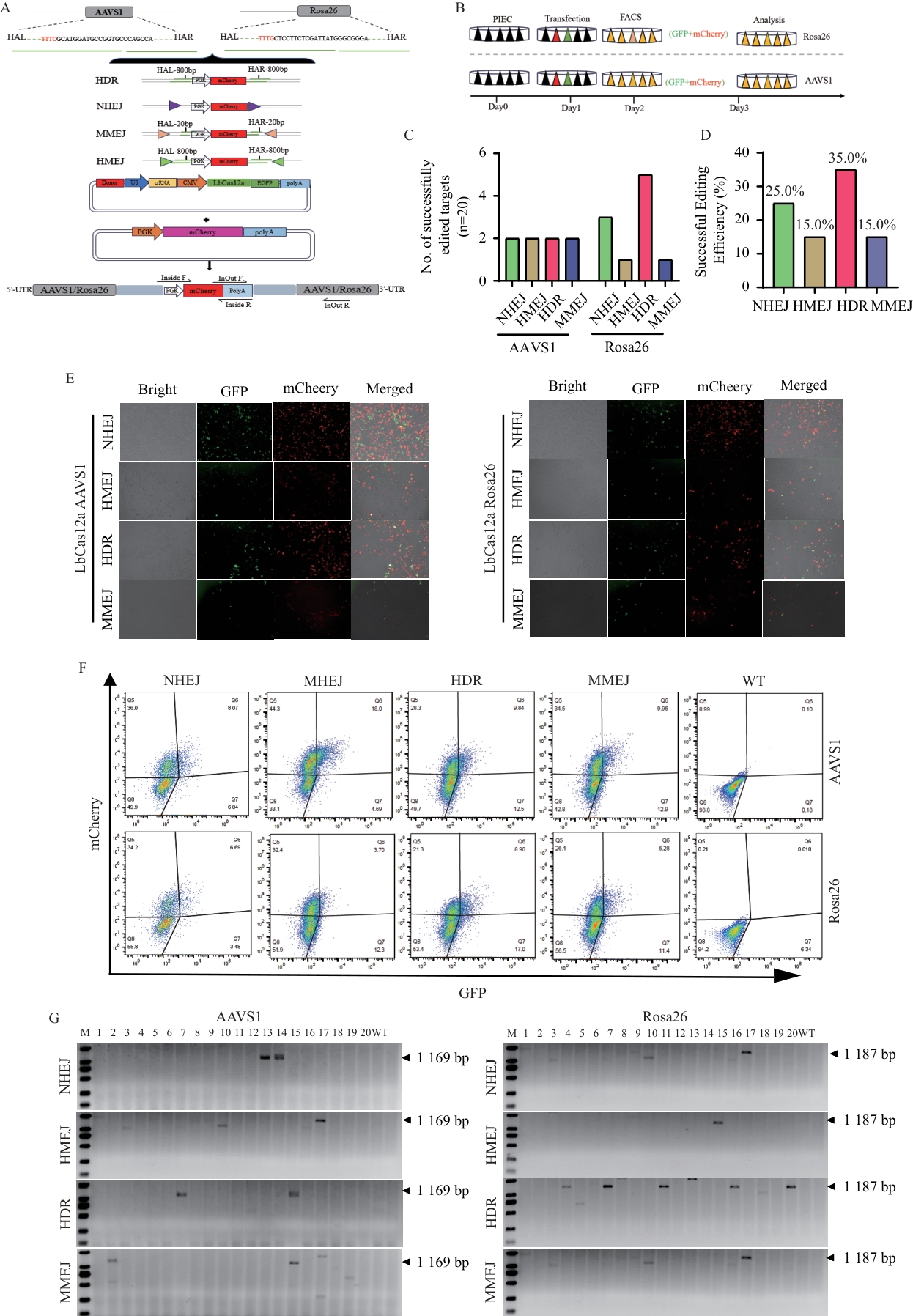
图4 LbCas12a介导AAVS1和Rosa26基因靶位点在4种不同敲入路径的效率评估A:Cas12a介导HR、HMEJ、MMEJ和NHEJ四种基因敲入路径的机制示意图。B:Cas12a介导4种基因敲入路径的细胞筛选流程图。C:Cas12a介导HR、NHEJ、MMEJ、HMEJ四种敲入途径在AAVS1和Rosa26位点定点整合的单细胞阳性个数统计。D:Cas12a介导HR、NHEJ、MMEJ、HMEJ四种敲入途径在AAVS1和Rosa26位点定点整合的单细胞编辑效率结果。E:LbCas12a介导AAVS1基因crRNA1靶位点和Rosa26基因crRNA靶位点质粒转染PIEC细胞48小时后的荧光结果。F:LbCas12a介导AAVS1基因crRNA1靶位点和Rosa26基因crRNA靶位点质粒转染PIEC细胞48小时后的流式分选检测结果。G:LbCas12a介导AAVS1和Rosa26基因crRNA1靶位点的NHEJ、HMEJ、HDR和MMEJ修复途径PCR鉴定结果。图中的1-10和11-20为两次转染后每个单细胞经独立PCR扩增的凝胶图,WT:野生型DNA组
Fig. 4 Efficiency evaluation of the four different gene knock-in pathways at AAVS1 and Rosa26 gene loci mediated by LbCas12aA: Schematic diagram of the mechanisms of Cas12a-mediated HR, NHEJ, MMEJ, and HMEJ gene knock-in pathways. B: Cell screening flowchart for the four Cas12a-mediated gene knock-in pathways. C: Statistics of the number of positive single cells for targeted integration via Cas12a-mediated HR, NHEJ, MMEJ, and HMEJ pathways at the AAVS1 and Rosa26 loci. D: Targeted integration efficiency results for Cas12a-mediated HR, NHEJ, MMEJ, and HMEJ repair pathways at the AAVS1 and Rosa26 loci in single cells. E: Fluorescence results at 48 h after plasmid transfection targeting the AAVS1 crRNA1 locus and Rosa26 crRNA locus in PIEC cells mediated by LbCas12a. F: Flow cytometry results at 48 h after plasmid transfection targeting the AAVS1 crRNA1 locus and Rosa26 crRNA locus in PIEC cells mediated by LbCas12a. G: PCR-identified results of NHEJ, HMEJ, HDR, and MMEJ repair pathways mediated by LbCas12a at the crRNA1 target sites of AAVS1 and Rosa26. The gel images (lane 1-10 and 11-20) indicate independent PCR amplifications from single cells across two separate transfections. WT: Wild-type DNA control
| [1] | Zetsche B, Gootenberg JS, Abudayyeh OO, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system [J]. Cell, 2015, 163(3): 759-771. |
| [2] | 王炀坤, 纪世新, 苏莹莹, 等. 基因编辑工具CRISPR-Cas9与CRISPR-Cas12a的比较与应用 [J]. 长春师范大学学报, 2023, 42(12): 107-112. |
| Wang YK, Ji SX, Su YY, et al. The diversity and applications of CRISPR-Cas9 and CRISPR-Cas12a [J]. J Changchun Norm Univ, 2023, 42(12): 107-112. | |
| [3] | Fonfara I, Richter H, Bratovič M, et al. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA [J]. Nature, 2016, 532(7600): 517-521. |
| [4] | Hsu PD, Scott DA, Weinstein JA, et al. DNA targeting specificity of RNA-guided Cas9 nucleases [J]. Nat Biotechnol, 2013, 31(9): 827-832. |
| [5] | Doudna JA, Charpentier E. The new frontier of genome engineering with CRISPR-Cas9 [J]. Science, 2014, 346(6213): 1258096. |
| [6] | Li JF, Norville JE, Aach J, et al. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9 [J]. Nat Biotechnol, 2013, 31(8): 688-691. |
| [7] | Ran FA, Hsu PD, Wright J, et al. Genome engineering using the CRISPR-Cas9 system [J]. Nat Protoc, 2013, 8(11): 2281-2308. |
| [8] | Mali P, Yang LH, Esvelt KM, et al. RNA-guided human genome engineering via Cas9 [J]. Science, 2013, 339(6121): 823-826. |
| [9] | Lin S, Staahl BT, Alla RK, et al. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery [J]. eLife, 2014, 3: e04766. |
| [10] | Schubert MS, Thommandru B, Woodley J, et al. Optimized design parameters for CRISPR Cas9 and Cas12a homology-directed repair [J]. Sci Rep, 2021, 11: 19482. |
| [11] | Maruyama T, Dougan SK, Truttmann MC, et al. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining [J]. Nat Biotechnol, 2015, 33(5): 538-542. |
| [12] | Tan JT, Zhao YC, Wang B, et al. Efficient CRISPR/Cas9-based plant genomic fragment deletions by microhomology-mediated end joining [J]. Plant Biotechnol J, 2020, 18(11): 2161-2163. |
| [13] | Yao X, Wang X, Hu XD, et al. Homology-mediated end joining-based targeted integration using CRISPR/Cas9 [J]. Cell Res, 2017, 27(6): 801-814. |
| [14] | Huang J, Rowe D, Subedi P, et al. CRISPR-Cas12a induced DNA double-strand breaks are repaired by multiple pathways with different mutation profiles in Magnaporthe oryzae [J]. Nat Commun, 2022, 13: 7168. |
| [15] | Cao L, Dong Y, Sun KW, et al. Experimental animal models for moyamoya disease: a species-oriented scoping review [J]. Front Surg, 2022, 9: 929871. |
| [16] | Gawel K, Langlois M, Martins T, et al. Seizing the moment: Zebrafish epilepsy models [J]. Neurosci Biobehav Rev, 2020, 116: 1-20. |
| [17] | Ely ZA, Mathey-Andrews N, Naranjo S, et al. A prime editor mouse to model a broad spectrum of somatic mutations in vivo [J]. Nat Biotechnol, 2024, 42(3): 424-436. |
| [18] | Lunney JK, Van Goor A, Walker KE, et al. Importance of the pig as a human biomedical model [J]. Sci Transl Med, 2021, 13(621): eabd5758. |
| [19] | Hou NP, Du XG, Wu S. Advances in pig models of human diseases [J]. Anim Models Exp Med, 2022, 5(2): 141-152. |
| [20] | Huang WZ, Xiong TL, Zhao YT, et al. Computational prediction and experimental validation identify functionally conserved lncRNAs from zebrafish to human [J]. Nat Genet, 2024, 56(1): 124-135. |
| [21] | 董战旗, 秦琪, 张新铃, 等. 家蚕CRISPR/Cpf1基因编辑系统建立 [J]. 生物工程学报, 2021, 37(12): 4342-4350. |
| Dong ZQ, Qin Q, Zhang XL, et al. Development of a CRISPR/Cpf1 gene editing system in silkworm Bombyx mori [J]. Chin J Biotechnol, 2021, 37(12): 4342-4350. | |
| [22] | Tang KY, Zhou LQ, Tian XL, et al. Cas12a-knock-in mice for multiplexed genome editing, disease modelling and immune-cell engineering [J]. Nat Biomed Eng, 2025, 9(8): 1290-1308. |
| [23] | Sun RR, Zhao YQ, Wang WJ, et al. Nonspecific interactions between Cas12a and dsDNA located downstream of the PAM mediate target search and assist AsCas12a for DNA cleavage [J]. Chem Sci, 2023, 14(14): 3839-3851. |
| [24] | Bernabé-Orts JM, Casas-Rodrigo I, Minguet EG, et al. Assessment of Cas12a-mediated gene editing efficiency in plants [J]. Plant Biotechnol J, 2019, 17(10): 1971-1984. |
| [25] | Zhang LY, Li G, Zhang YX, et al. Boosting genome editing efficiency in human cells and plants with novel LbCas12a variants [J]. Genome Biol, 2023, 24: 102. |
| [26] | Anders C, Niewoehner O, Duerst A, et al. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease [J]. Nature, 2014, 513(7519): 569-573. |
| [27] | Li P, Zhang LJ, Li ZF, et al. Cas12a mediates efficient and precise endogenous gene tagging via MITI microhomology-dependent targeted integrations [J]. Cell Mol Life Sci, 2020, 77(19): 3875-3884. |
| [28] | Allen D, Knop O, Itkowitz B, et al. CRISPR-Cas9 engineering of the RAG2 locus via complete coding sequence replacement for therapeutic applications [J]. Nat Commun, 2023, 14: 6771. |
| [29] | Van Vu T, Thi Hai Doan D, Kim J, et al. CRISPR/Cas-based precision genome editing via microhomology-mediated end joining [J]. Plant Biotechnol J, 2021, 19(2): 230-239. |
| [30] | Naqvi MM, Lee L, Montaguth OET, et al. CRISPR-Cas12a-mediated DNA clamping triggers target-strand cleavage [J]. Nat Chem Biol, 2022, 18(9): 1014-1022. |
| [1] | 李雅琦, 孙萌, 李秀丽, 魏静娜, 赵琳琳, 赵云平, 刘征辉, 苏蘩. 多种Cas12a蛋白普适性的高性能低成本荧光检测缓冲体系优化[J]. 生物技术通报, 2026, 42(4): 83-91. |
| [2] | 区琦, 冯瑶, 韦柳婷, 庄站伟, 赵云翔, 陈富美. 全基因组关联分析(GWAS)在母猪繁殖性状研究中的应用进展[J]. 生物技术通报, 2026, 42(4): 65-71. |
| [3] | 刘淼, 林涛, 贾乐松, 胡丰, 李涛, 李志万, 刘美芳, 郑方燕, 崔龙. 从野生到栽培:番茄果实色泽的演化与调控机制[J]. 生物技术通报, 2026, 42(3): 187-202. |
| [4] | 胡秋玲, 陈灵, 黄嘉怡, 赵梓乔, 潘璐怡, 刘慧丽, 刘太波. 多胺调控果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 203-212. |
| [5] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [6] | 袁梦博, 赵光伟, 贺玉花, 黄祥, 徐永阳, 张健, 孔维虎, 田小琴, 户克云, 唐伶俐. CmCRC基因影响甜瓜性别分化的功能研究[J]. 生物技术通报, 2026, 42(3): 294-301. |
| [7] | 陈玲嫣, 李伟勋, 逄欣欣, 高翔, 焦绪瑶. 拟杆菌遗传操作工具箱:从传统方法到合成生物学前沿[J]. 生物技术通报, 2026, 42(2): 51-64. |
| [8] | 刁辰洋, 崔有志, 李炳志. 靶向诱变介导的微生物进化技术研究进展[J]. 生物技术通报, 2025, 41(8): 11-21. |
| [9] | 余永霞, 杜再慧, 朱龙佼, 许文涛. 基因编辑技术在牛种中的应用及研究进展[J]. 生物技术通报, 2025, 41(8): 34-41. |
| [10] | 邓美壁, 严浪, 詹志田, 朱敏, 和玉兵. RUBY辅助的水稻高效CRISPR基因编辑[J]. 生物技术通报, 2025, 41(8): 65-73. |
| [11] | 霍贯中, 张欣濡, 田士军, 李君. CRISPR/Cas12a基因编辑技术在植物中的研究进展[J]. 生物技术通报, 2025, 41(6): 1-11. |
| [12] | 周倩, 唐梦君, 张小燕, 陆俊贤, 唐修君, 杨星星, 高玉时. 基于CRISPR-Cas系统的多重耐药菌防治技术研究进展[J]. 生物技术通报, 2025, 41(5): 42-51. |
| [13] | 高畅, 庄添驰, 李宁, 刘云, 顾鹏飞, 赵昕怡, 季明辉. RPA-CRISPR/Cas12a结合重力驱动微流控芯片的MTB快检方法的建立[J]. 生物技术通报, 2025, 41(5): 62-69. |
| [14] | 文博霖, 万敏, 胡建军, 王克秀, 景晟林, 王心悦, 朱博, 唐铭霞, 李兵, 何卫, 曾子贤. 马铃薯川芋50遗传转化及基因编辑体系的建立[J]. 生物技术通报, 2025, 41(4): 88-97. |
| [15] | 娄亚楠, 熊晶晶, 娄亚玲, 王福斌, 程文杰, 陈晓晶, 李加宇, 普少瑕, 魏红江, 赵红业. 多西环素长期高剂量诱导对滇南小耳猪肝肾功能的影响[J]. 生物技术通报, 2025, 41(4): 345-354. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||