








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (10): 41-52.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0638
Previous Articles Next Articles
ZHANG Shuo1,2( ), KAN Jun-hu1, ZHOU Jia-wei1, WU Zhi-qiang1(
), KAN Jun-hu1, ZHOU Jia-wei1, WU Zhi-qiang1( )
)
Received:2024-07-05
Online:2024-10-26
Published:2024-11-20
Contact:
WU Zhi-qiang
E-mail:szhang@webmail.hzau.edu.cn;wuzhiqiang@caas.cn
ZHANG Shuo, KAN Jun-hu, ZHOU Jia-wei, WU Zhi-qiang. Advance in Plant Mitochondrial Genome Editing[J]. Biotechnology Bulletin, 2024, 40(10): 41-52.
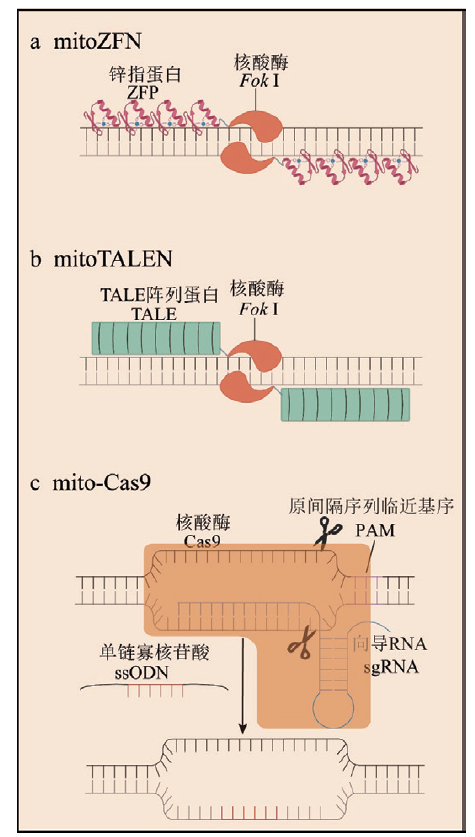
Fig. 1 Schematic diagram of mitochondrial genome editing technology a: In mitoZFN, tandem ZFP zinc finger proteins specifically recognize target sequences, and the Fok I dimer exhibits nuclease activity. b: In mitoTALEN, tandem TALE proteins specifically recognize target sequences, and the Fok I dimer exhibits nuclease activity. c: In mito-Cas9, the Cas9 protein can enter the mitochondria with the help of MTS, while sgRNA and single-stranded oligonucleotides can only enter the mitochondria to a limited extent, restricting the editing efficiency of mito-Cas9

Fig. 2 Schematic representation of mitochondrial genome editing techniques applied in plants a: Codon-optimized mitoTALEN has been applied to plant mitochondrial research. b: mt-DdCBEs/mitoTALECD utilizes tandem TALE proteins to specifically recognize target sequences. The DddA deaminase converts cytosine to uracil, and UGI acts as a uracil glycosylase inhibitor to protect uracil, achieving C-to-T editing through DNA replication. c: In TALED, DddA deaminase opens double-stranded DNA, and the single-strand-specific TadA8e adenine deaminase converts adenine to inosine, achieving A-to-G editing through DNA replication. d: In mitoCRISPR/Cas9, the Cas9 protein, aided by MTS, enters the mitochondria and binds with sgRNA to achieve target sequence cleavage
| 编辑工具 Editing tool | 物种 Species | 靶基因 Target gene | 编辑类型 Editing type | 表型 Phenotype | 文献 Reference |
|---|---|---|---|---|---|
| mitoTALEN | 水稻Oryza sativa | orf79 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 油菜 Brassica napus | orf125 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 拟南芥Arabidopsis thaliana | atp6-1, atp6-2 | 敲除 | 功能冗余;双突致死 | [ |
| mitoTALEN | 水稻Oryza sativa | orf352 | 敲除 | 恢复花粉育性;不结实 | [ |
| mitoTALEN | 水稻Oryza sativa | orf312 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 番茄Solanum lycopersicum | orf137 | 敲除 | 恢复花粉育性 | [ |
| TALEN-GDM | 烟草 Nicotiana tabacum | nad9 | 点突变 | 无明显表型 | [ |
| TALEN-GDM | 烟草Nicotiana tabacum | nad9 | 敲除 | 生长迟缓;叶片和花发育缺陷;雄性不育 | [ |
| mitoTALEN | 拟南芥Arabidopsis thaliana | nad7 | 敲除 | 严重的生长迟缓;致死 | [ |
| mitoTALEN | 水稻Oryza sativa | WA352 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 西兰花Brassica oleracea | orf138 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 马铃薯Solanum tuberosum | orf125 | 敲除 | N/A | [ |
| mt-DdCBEs | 油菜Brassica napus | atp6, rps14 | C-to-T | N/A | [ |
| mt-DdCBEs | 莴苣Lactuca sativa | atp6 | C-to-T | N/A | [ |
| mitoTALECD | 拟南芥Arabidopsis thaliana | atp1 | C-to-T | 恢复正常生长 | [ |
| mitoTALECD | 马铃薯Solanum tuberosum | orf125 | C-to-T | N/A | [ |
| mTALED | 水稻Oryza sativa | atp6-2 | A-to-G | N/A | [ |
| mitoCRISPR/Cas9 | 烟草Nicotiana tabacum | atp9 | 敲除 | 雄性不育 | [ |
Table 1 Application of mitochondrial genome editing technology in the study of plant mitochondrial gene functions
| 编辑工具 Editing tool | 物种 Species | 靶基因 Target gene | 编辑类型 Editing type | 表型 Phenotype | 文献 Reference |
|---|---|---|---|---|---|
| mitoTALEN | 水稻Oryza sativa | orf79 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 油菜 Brassica napus | orf125 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 拟南芥Arabidopsis thaliana | atp6-1, atp6-2 | 敲除 | 功能冗余;双突致死 | [ |
| mitoTALEN | 水稻Oryza sativa | orf352 | 敲除 | 恢复花粉育性;不结实 | [ |
| mitoTALEN | 水稻Oryza sativa | orf312 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 番茄Solanum lycopersicum | orf137 | 敲除 | 恢复花粉育性 | [ |
| TALEN-GDM | 烟草 Nicotiana tabacum | nad9 | 点突变 | 无明显表型 | [ |
| TALEN-GDM | 烟草Nicotiana tabacum | nad9 | 敲除 | 生长迟缓;叶片和花发育缺陷;雄性不育 | [ |
| mitoTALEN | 拟南芥Arabidopsis thaliana | nad7 | 敲除 | 严重的生长迟缓;致死 | [ |
| mitoTALEN | 水稻Oryza sativa | WA352 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 西兰花Brassica oleracea | orf138 | 敲除 | 恢复花粉育性 | [ |
| mitoTALEN | 马铃薯Solanum tuberosum | orf125 | 敲除 | N/A | [ |
| mt-DdCBEs | 油菜Brassica napus | atp6, rps14 | C-to-T | N/A | [ |
| mt-DdCBEs | 莴苣Lactuca sativa | atp6 | C-to-T | N/A | [ |
| mitoTALECD | 拟南芥Arabidopsis thaliana | atp1 | C-to-T | 恢复正常生长 | [ |
| mitoTALECD | 马铃薯Solanum tuberosum | orf125 | C-to-T | N/A | [ |
| mTALED | 水稻Oryza sativa | atp6-2 | A-to-G | N/A | [ |
| mitoCRISPR/Cas9 | 烟草Nicotiana tabacum | atp9 | 敲除 | 雄性不育 | [ |
| [1] |
Sloan DB, Warren JM, Williams AM, et al. Cytonuclear integration and co-evolution[J]. Nat Rev Genet, 2018, 19(10): 635-648.
doi: 10.1038/s41576-018-0035-9 pmid: 30018367 |
| [2] | Wang J, Kan SL, Liao XZ, et al. Plant organellar genomes: much done, much more to do[J]. Trends Plant Sci, 2024, 29(7): 754-769. |
| [3] |
Lynch M, Koskella B, Schaack S. Mutation pressure and the evolution of organelle genomic architecture[J]. Science, 2006, 311(5768): 1727-1730.
doi: 10.1126/science.1118884 pmid: 16556832 |
| [4] | Hu ZJ, Yang L, Zhang ML, et al. A novel protein CYTB-187AA encoded by the mitochondrial gene CYTB modulates mammalian early development[J]. Cell Metab, 2024, 36(7): 1586-1597.e7. |
| [5] |
Wallace DC, Singh G, Lott MT, et al. Mitochondrial DNA mutation associated with Leber's hereditary optic neuropathy[J]. Science, 1988, 242(4884): 1427-1430.
doi: 10.1126/science.3201231 pmid: 3201231 |
| [6] |
Kauppila TES, Kauppila JHK, Larsson NG. Mammalian mitochondria and aging: an update[J]. Cell Metab, 2017, 25(1): 57-71.
doi: S1550-4131(16)30502-2 pmid: 28094012 |
| [7] | Yang L, Lin XB, Tang HT, et al. Mitochondrial DNA mutation exacerbates female reproductive aging via impairment of the NADH/NAD+ redox[J]. Aging Cell, 2020, 19(9): e13206. |
| [8] |
DiMauro S, Hirano M, Schon EA. Approaches to the treatment of mitochondrial diseases[J]. Muscle Nerve, 2006, 34(3): 265-283.
pmid: 16810684 |
| [9] |
Zhang J, Liu H, Luo SY, et al. Live birth derived from oocyte spindle transfer to prevent mitochondrial disease[J]. Reprod Biomed Online, 2017, 34(4): 361-368.
doi: S1472-6483(17)30041-X pmid: 28385334 |
| [10] |
Minczuk M, Papworth MA, Miller JC, et al. Development of a single-chain, quasi-dimeric zinc-finger nuclease for the selective degradation of mutated human mitochondrial DNA[J]. Nucleic Acids Res, 2008, 36(12): 3926-3938.
doi: 10.1093/nar/gkn313 pmid: 18511461 |
| [11] | Kim JS, Chen J. Base editing of organellar DNA with programmable deaminases[J]. Nat Rev Mol Cell Biol, 2024, 25(1): 34-45. |
| [12] | Kozik A, Rowan BA, Lavelle D, et al. The alternative reality of plant mitochondrial DNA: one ring does not rule them all[J]. PLoS Genet, 2019, 15(8): e1008373. |
| [13] | Palmer JD, Herbon LA. Plant mitochondrial DNA evolves rapidly in structure, but slowly in sequence[J]. J Mol Evol, 1988, 28(1/2): 87-97. |
| [14] |
Christensen AC. Plant mitochondria are a riddle wrapped in a mystery inside an Enigma[J]. J Mol Evol, 2021, 89(3): 151-156.
doi: 10.1007/s00239-020-09980-y pmid: 33486550 |
| [15] |
Chen LT, Liu YG. Male sterility and fertility restoration in crops[J]. Annu Rev Plant Biol, 2014, 65: 579-606.
doi: 10.1146/annurev-arplant-050213-040119 pmid: 24313845 |
| [16] |
Luo DP, Xu H, Liu ZL, et al. A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice[J]. Nat Genet, 2013, 45(5): 573-577.
doi: 10.1038/ng.2570 pmid: 23502780 |
| [17] |
Minczuk M, Papworth MA, Kolasinska P, et al. Sequence-specific modification of mitochondrial DNA using a chimeric zinc finger methylase[J]. Proc Natl Acad Sci USA, 2006, 103(52): 19689-19694.
doi: 10.1073/pnas.0609502103 pmid: 17170133 |
| [18] |
Bacman SR, Williams SL, Pinto M, et al. Specific elimination of mutant mitochondrial genomes in patient-derived cells by mitoTALENs[J]. Nat Med, 2013, 19(9): 1111-1113.
doi: 10.1038/nm.3261 pmid: 23913125 |
| [19] | Zhu HC, Li C, Gao CX. Applications of CRISPR-Cas in agriculture and plant biotechnology[J]. Nat Rev Mol Cell Biol, 2020, 21(11): 661-677. |
| [20] |
Pacesa M, Pelea O, Jinek M. Past, present, and future of CRISPR genome editing technologies[J]. Cell, 2024, 187(5): 1076-1100.
doi: 10.1016/j.cell.2024.01.042 pmid: 38428389 |
| [21] | Bi R, Li Y, Xu M, et al. Direct evidence of CRISPR-Cas9-mediated mitochondrial genome editing[J]. Innovation(Camb), 2022, 3(6): 100329. |
| [22] |
Gammage PA, Moraes CT, Minczuk M. Mitochondrial genome engineering: the revolution may not be CRISPR-ized[J]. Trends Genet, 2018, 34(2): 101-110.
doi: S0168-9525(17)30191-9 pmid: 29179920 |
| [23] |
Beerli RR, Barbas CF III. Engineering polydactyl zinc-finger transcription factors[J]. Nat Biotechnol, 2002, 20(2): 135-141.
pmid: 11821858 |
| [24] |
Li L, Wu LP, Chandrasegaran S. Functional domains in Fok I restriction endonuclease[J]. Proc Natl Acad Sci USA, 1992, 89(10): 4275-4279.
pmid: 1584761 |
| [25] |
Gammage PA, Rorbach J, Vincent AI, et al. Mitochondrially targeted ZFNs for selective degradation of pathogenic mitochondrial genomes bearing large-scale deletions or point mutations[J]. EMBO Mol Med, 2014, 6(4): 458-466.
doi: 10.1002/emmm.201303672 pmid: 24567072 |
| [26] |
Lim K, Cho SI, Kim JS. Nuclear and mitochondrial DNA editing in human cells with zinc finger deaminases[J]. Nat Commun, 2022, 13(1): 366.
doi: 10.1038/s41467-022-27962-0 pmid: 35042880 |
| [27] |
Urnov FD, Rebar EJ, Holmes MC, et al. Genome editing with engineered zinc finger nucleases[J]. Nat Rev Genet, 2010, 11(9): 636-646.
doi: 10.1038/nrg2842 pmid: 20717154 |
| [28] |
Moscou MJ, Bogdanove AJ. A simple cipher governs DNA recognition by TAL effectors[J]. Science, 2009, 326(5959): 1501.
doi: 10.1126/science.1178817 pmid: 19933106 |
| [29] | Boch J, Bonas U. Xanthomonas AvrBs3 family-type III effectors: discovery and function[J]. Annu Rev Phytopathol, 2010, 48: 419-436. |
| [30] |
Sakuma T, Ochiai H, Kaneko T, et al. Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity[J]. Sci Rep, 2013, 3: 3379.
doi: 10.1038/srep03379 pmid: 24287550 |
| [31] |
Boch J, Scholze H, Schornack S, et al. Breaking the code of DNA binding specificity of TAL-type III effectors[J]. Science, 2009, 326(5959): 1509-1512.
doi: 10.1126/science.1178811 pmid: 19933107 |
| [32] | Bacman SR, Barrera-Paez JD, Pinto M, et al. mitoTALEN reduces the mutant mtDNA load in neurons[J]. Mol Ther Nucleic Acids, 2024, 35(1): 102132. |
| [33] | Bacman SR, Kauppila JHK, Pereira CV, et al. MitoTALEN reduces mutant mtDNA load and restores tRNAAla levels in a mouse model of heteroplasmic mtDNA mutation[J]. Nat Med, 2018, 24(11): 1696-1700. |
| [34] | Komor AC, Kim YB, Packer MS, et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage[J]. Nature, 2016, 533(7603): 420-424. |
| [35] | Gaudelli NM, Komor AC, Rees HA, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage[J]. Nature, 2017, 551(7681): 464-471. |
| [36] | Mok BY, de Moraes MH, Zeng J, et al. A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing[J]. Nature, 2020, 583(7817): 631-637. |
| [37] |
Qi XL, Chen XX, Guo JY, et al. Precision modeling of mitochondrial disease in rats via DdCBE-mediated mtDNA editing[J]. Cell Discov, 2021, 7(1): 95.
doi: 10.1038/s41421-021-00325-7 pmid: 34663794 |
| [38] |
Wei YH, Xu CL, Feng H, et al. Human cleaving embryos enable efficient mitochondrial base-editing with DdCBE[J]. Cell Discov, 2022, 8(1): 7.
doi: 10.1038/s41421-021-00372-0 pmid: 35102133 |
| [39] |
Lee H, Lee S, Baek G, et al. Mitochondrial DNA editing in mice with DddA-TALE fusion deaminases[J]. Nat Commun, 2021, 12(1): 1190.
doi: 10.1038/s41467-021-21464-1 pmid: 33608520 |
| [40] | Guo JY, Chen XX, Liu ZW, et al. DdCBE mediates efficient and inheritable modifications in mouse mitochondrial genome[J]. Mol Ther Nucleic Acids, 2021, 27: 73-80. |
| [41] | Lei ZX, Meng HW, Liu LL, et al. Mitochondrial base editor induces substantial nuclear off-target mutations[J]. Nature, 2022, 606(7915): 804-811. |
| [42] |
Wei YH, Li ZF, Xu K, et al. Mitochondrial base editor DdCBE causes substantial DNA off-target editing in nuclear genome of embryos[J]. Cell Discov, 2022, 8(1): 27.
doi: 10.1038/s41421-022-00391-5 pmid: 35304438 |
| [43] |
Mok BY, Kotrys AV, Raguram A, et al. CRISPR-free base editors with enhanced activity and expanded targeting scope in mitochondrial and nuclear DNA[J]. Nat Biotechnol, 2022, 40(9): 1378-1387.
doi: 10.1038/s41587-022-01256-8 pmid: 35379961 |
| [44] | Lee S, Lee H, Baek G, et al. Precision mitochondrial DNA editing with high-fidelity DddA-derived base editors[J]. Nat Biotechnol, 2023, 41(3): 378-386. |
| [45] |
Sun HF, Wang ZJ, Shen L, et al. Developing mitochondrial base editors with diverse context compatibility and high fidelity via saturated spacer library[J]. Nat Commun, 2023, 14(1): 6625.
doi: 10.1038/s41467-023-42359-3 pmid: 37857619 |
| [46] | Cho SI, Lee S, Mok YG, et al. Targeted A-to-G base editing in human mitochondrial DNA with programmable deaminases[J]. Cell, 2022, 185(10): 1764-1776.e12. |
| [47] | Cho SI, Lim K, Hong S, et al. Engineering TALE-linked deaminases to facilitate precision adenine base editing in mitochondrial DNA[J]. Cell, 2024, 187(1): 95-109.e26. |
| [48] | Hu JC, Sun Y, Li BS, et al. Strand-preferred base editing of organellar and nuclear genomes using CyDENT[J]. Nat Biotechnol, 2024, 42(6): 936-945. |
| [49] | Yi ZY, Zhang XX, Tang W, et al. Strand-selective base editing of human mitochondrial DNA using mitoBEs[J]. Nat Biotechnol, 2024, 42(3): 498-509. |
| [50] | Li SY, Xia LQ. Precise gene replacement in plants through CRISPR/Cas genome editing technology: current status and future perspectives[J]. aBIOTECH, 2019, 1(1): 58-73. |
| [51] |
Gao CX. Genome engineering for crop improvement and future agriculture[J]. Cell, 2021, 184(6): 1621-1635.
doi: 10.1016/j.cell.2021.01.005 pmid: 33581057 |
| [52] | Ma GG, Kuang YJ, Lu ZW, et al. CRISPR/Sc++-mediated genome editing in rice[J]. J Integr Plant Biol, 2021, 63(9): 1606-1610. |
| [53] |
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering[J]. Cell, 2014, 157(6): 1262-1278.
doi: S0092-8674(14)00604-7 pmid: 24906146 |
| [54] | Jo A, Ham S, Lee GH, et al. Efficient mitochondrial genome editing by CRISPR/Cas9[J]. Biomed Res Int, 2015, 2015: 305716. |
| [55] | Bian WP, Chen YL, Luo JJ, et al. Knock-In strategy for editing human and zebrafish mitochondrial DNA using mito-CRISPR/Cas9 system[J]. ACS Synth Biol, 2019, 8(4): 621-632. |
| [56] | Yoo BC, Yadav NS, Orozco EM Jr, et al. Cas9/gRNA-mediated genome editing of yeast mitochondria and Chlamydomonas chloroplasts[J]. PeerJ, 2020, 8: e8362. |
| [57] |
Johnston SA, Anziano PQ, Shark K, et al. Mitochondrial transformation in yeast by bombardment with microprojectiles[J]. Science, 1988, 240(4858): 1538-1541.
pmid: 2836954 |
| [58] |
Remacle C, Cardol P, Coosemans N, et al. High-efficiency biolistic transformation of Chlamydomonas mitochondria can be used to insert mutations in complex I genes[J]. Proc Natl Acad Sci USA, 2006, 103(12): 4771-4776.
pmid: 16537419 |
| [59] |
Li WM, Ruf S, Bock R. Chloramphenicol acetyltransferase as selectable marker for plastid transformation[J]. Plant Mol Biol, 2011, 76(3-5): 443-451.
doi: 10.1007/s11103-010-9678-4 pmid: 20721602 |
| [60] | Lin JY, Liu YC, Tseng YH, et al. TALE-based organellar genome editing and gene expression in plants[J]. Plant Cell Rep, 2024, 43(3): 61. |
| [61] |
Kazama T, Okuno M, Watari Y, et al. Curing cytoplasmic male sterility via TALEN-mediated mitochondrial genome editing[J]. Nat Plants, 2019, 5(7): 722-730.
doi: 10.1038/s41477-019-0459-z pmid: 31285556 |
| [62] |
Omukai S, Arimura SI, Toriyama K, et al. Disruption of mitochondrial open reading frame 352 partially restores pollen development in cytoplasmic male sterile rice[J]. Plant Physiol, 2021, 187(1): 236-246.
doi: 10.1093/plphys/kiab236 pmid: 34015134 |
| [63] | Takatsuka A, Kazama T, Arimura SI, et al. TALEN-mediated depletion of the mitochondrial gene orf312 proves that it is a Tadukan-type cytoplasmic male sterility-causative gene in rice[J]. Plant J, 2022, 110(4): 994-1004. |
| [64] | Zhou JW, Nie LY, Zhang S, et al. Mitochondrial genome editing of WA352 via mitoTALENs restore fertility in cytoplasmic male sterile rice[J]. Plant Biotechnol J, 2024, 22(7): 1960-1962. |
| [65] |
Kuwabara K, Arimura SI, Shirasawa K, et al. orf137 triggers cytoplasmic male sterility in tomato[J]. Plant Physiol, 2022, 189(2): 465-468.
doi: 10.1093/plphys/kiac082 pmid: 35212743 |
| [66] |
Xu FY, Su TB, Zhang XC, et al. Editing of ORF138 restores fertility of Ogura cytoplasmic male sterile broccoli via mitoTALENs[J]. Plant Biotechnol J, 2024, 22(5): 1325-1334.
doi: 10.1111/pbi.14268 pmid: 38213067 |
| [67] | Arimura SI, Nakazato I. Genome editing of plant mitochondrial and chloroplast genomes[J]. Plant Cell Physiol, 2024, 65(4): 477-483. |
| [68] | Møller IM, Rasmusson AG, Van Aken O. Plant mitochondria - past, present and future[J]. Plant J, 2021, 108(4): 912-959. |
| [69] | Arimura SI, Ayabe H, Sugaya H, et al. Targeted gene disruption of ATP synthases 6-1 and 6-2 in the mitochondrial genome of Arabi-dopsis thaliana by mitoTALENs[J]. Plant J, 2020, 104(6): 1459-1471. |
| [70] | Ayabe H, Toyoda A, Iwamoto A, et al. Mitochondrial gene defects in Arabidopsis can broadly affect mitochondrial gene expression through copy number[J]. Plant Physiol, 2023, 191(4): 2256-2275. |
| [71] |
Forner J, Kleinschmidt D, Meyer EH, et al. Targeted introduction of heritable point mutations into the plant mitochondrial genome[J]. Nat Plants, 2022, 8(3): 245-256.
doi: 10.1038/s41477-022-01108-y pmid: 35301443 |
| [72] |
Forner J, Kleinschmidt D, Meyer EH, et al. Targeted knockout of a conserved plant mitochondrial gene by genome editing[J]. Nat Plants, 2023, 9(11): 1818-1831.
doi: 10.1038/s41477-023-01538-2 pmid: 37814021 |
| [73] | Kang BC, Bae SJ, Lee S, et al. Chloroplast and mitochondrial DNA editing in plants[J]. Nat Plants, 2021, 7(7): 899-905. |
| [74] | Nakazato I, Okuno M, Zhou C, et al. Targeted base editing in the mitochondrial genome of Arabidopsis thaliana[J]. Proc Natl Acad Sci USA, 2022, 119(20): e2121177119. |
| [75] |
Small ID, Schallenberg-Rüdinger M, Takenaka M, et al. Plant organellar RNA editing: what 30 years of research has revealed[J]. Plant J, 2020, 101(5): 1040-1056.
doi: 10.1111/tpj.14578 |
| [76] | Small I, Melonek J, Bohne AV, et al. Plant organellar RNA maturation[J]. Plant Cell, 2023, 35(6): 1727-1751. |
| [77] | Zhou C, Okuno M, Nakazato I, et al. Targeted A-to-G base editing in the organellar genomes of Arabidopsis with monomeric programmable deaminases[J]. Plant Physiol, 2024, 194(4): 2278-2287. |
| [78] | Chang YZ, Liu BL, Jiang YY, et al. Induce male sterility by CRISPR/Cas9-mediated mitochondrial genome editing in tobacco[J]. Funct Integr Genomics, 2023, 23(3): 205. |
| [79] |
Nicolia A, Scotti N, D'Agostino N, et al. Mitochondrial DNA editing in potato through mitoTALEN and mitoTALECD: molecular characterization and stability of editing events[J]. Plant Methods, 2024, 20(1): 4.
doi: 10.1186/s13007-023-01124-9 pmid: 38183104 |
| [80] |
Srivastava S, Moraes CT. Manipulating mitochondrial DNA heteroplasmy by a mitochondrially targeted restriction endonuclease[J]. Hum Mol Genet, 2001, 10(26): 3093-3099.
pmid: 11751691 |
| [81] |
Tanaka M, Borgeld HJ, Zhang J, et al. Gene therapy for mitochondrial disease by delivering restriction endonuclease SmaI into mitochondria[J]. J Biomed Sci, 2002, 9(6 Pt 1):534-541.
doi: 10.1159/000064726 pmid: 12372991 |
| [82] |
Bayona-Bafaluy MP, Blits B, Battersby BJ, et al. Rapid directional shift of mitochondrial DNA heteroplasmy in animal tissues by a mitochondrially targeted restriction endonuclease[J]. Proc Natl Acad Sci USA, 2005, 102(40): 14392-14397.
pmid: 16179392 |
| [83] |
Lee S, Lee H, Baek G, et al. Enhanced mitochondrial DNA editing in mice using nuclear-exported TALE-linked deaminases and nucleases[J]. Genome Biol, 2022, 23(1): 211.
doi: 10.1186/s13059-022-02782-z pmid: 36224582 |
| [84] | Xu K, Feng H, Zhang HH, et al. Structure-guided discovery of highly efficient cytidine deaminases with sequence-context independence[J]. Nat Biomed Eng, 2024. |
| [85] |
Wang G, Chen HW, Oktay Y, et al. PNPASE regulates RNA import into mitochondria[J]. Cell, 2010, 142(3): 456-467.
doi: 10.1016/j.cell.2010.06.035 pmid: 20691904 |
| [86] | Wang PC, Zhang LX, Chen SY, et al. ANT2 functions as a translocon for mitochondrial cross-membrane translocation of RNAs[J]. Cell Res, 2024, 34(7): 504-521. |
| [87] | 王熠晨, 王颖, 陈妤, 等. 线粒体基因编辑技术研究进展[J]. 浙江大学学报: 医学版, 2023, 52(4): 460-472. |
| Wang YC, Wang Y, Chen Y, et al. Research progress in mitochondrial gene editing technology[J]. J Zhejiang Univ Med Sci, 2023, 52(4): 460-472. | |
| [88] |
Kolesnikova O, Kazakova H, Comte C, et al. Selection of RNA aptamers imported into yeast and human mitochondria[J]. RNA, 2010, 16(5): 926-941.
doi: 10.1261/rna.1914110 pmid: 20348443 |
| [1] | TONG Wei-jing, LUO Shu, LU Xin-lu, SHEN Jian-fu, LU Bai-yi, LI Kai-mian, MA Qiu-xiang, ZHANG Peng. CRISPR/Cas9 Editing MeHNL Gene to Generate Cassava Plants with Low Cyanogenic Glycoside [J]. Biotechnology Bulletin, 2024, 40(9): 11-19. |
| [2] | CUI Hai-yang, TAN Miao, QUAN Zhuang, CHEN Hong-li, DONG Yan-min, TANG Li-chun. Generation of Virus-free TRAC-knocked-in T Cells Using Cas9TX [J]. Biotechnology Bulletin, 2024, 40(9): 190-197. |
| [3] | HOU Wen-ting, SUN Lin, ZHANG Yan-jun, DONG He-zhong. Application of Gene-editing Technology for Germplasm Innovation and Genetic Improvement in Cotton [J]. Biotechnology Bulletin, 2024, 40(7): 68-77. |
| [4] | LONG Jing, CHEN Jing-min, LIU Xiao, ZHANG Yi-fan, ZHOU Li-bin, DU Yan. Repair Mechanisms of DNA Double-strand Breaks and Their Roles in Heavy Ion Mutagenesis and Gene Editing in Plants [J]. Biotechnology Bulletin, 2024, 40(7): 55-67. |
| [5] | CHEN Mo-yan, ZHU Cheng. Mechanism Study and Application of CRISPR/Cas12a-based Biosensing Platform [J]. Biotechnology Bulletin, 2024, 40(7): 90-98. |
| [6] | XIAO Yi-meng, YANG Wen, CHENG Yi-yi, LUO Gang. CRISPR-Cas9 Gene Editing Technology and Its Research Progress in Poultry [J]. Biotechnology Bulletin, 2024, 40(5): 38-47. |
| [7] | ZHANG Zhen, LI Qing, XU Jing, CHEN Kai-yuan, ZHANG Chun-zhi, ZHU Guang-tao. Construction and Application of Potato Mitochondrial Targeted Expression Vector [J]. Biotechnology Bulletin, 2024, 40(5): 66-73. |
| [8] | YANG Qi, WEI Zi-di, SONG Juan, TONG Kun, YANG Liu, WANG Jia-han, LIU Hai-yan, LUAN Wei-jiang, MA Xuan. Construction and Transcriptomic Analysis of Rice Histone H1 Triple Mutant [J]. Biotechnology Bulletin, 2024, 40(4): 85-96. |
| [9] | ZHU Tian-yi, KONG Gui-mei, JIAO Hong-mei, GUO Ting-ting, WU Ri-han, LIU Cui-cui, GAO Cheng-feng, LI Guo-cai. Establishment of A Bacterial Model of CRISPR/Cas9 Mediated adeG Gene Knockout in Escherichia coli [J]. Biotechnology Bulletin, 2024, 40(2): 55-64. |
| [10] | GAO Deng-ke, MA Bai-rong, GUO Yi-ying, LIU Wei, LIU Tian, JIN Ya-ping, JIANG Zhou, CHEN Hua-tao. Establishment of Quaking Knockout Mouse Embryonic Fibroblast Cell Line Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2024, 40(2): 65-72. |
| [11] | ZHANG Hong-min, LONG Wen, LAO Xiao-qing, CHEN Wen-yan, SHANG Xue-mei, WANG Hong-lian, WANG Li, SU Hong-wei, SHEN Hong-ping, SHEN Hong-chun. Construction of Pmepa1 Knockout TCMK1 Mouse Renal Tubular Epithelial Cell Line Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2024, 40(2): 73-79. |
| [12] | ZHOU Jia-wei, WU Zhi-qiang. Construction Method of mitoTALENs Mitochondrial Gene Editing Vector in Plants [J]. Biotechnology Bulletin, 2024, 40(10): 172-180. |
| [13] | PI Yi-fei, SONG Xin-hui, WANG Xi-lin, LI Jin-jin, SUN Chang-bin, XU Wei. High-throughput Screening System for Functional R-loop Loci Based on R-loop Targeted Editing Technology [J]. Biotechnology Bulletin, 2024, 40(10): 181-190. |
| [14] | LI Xin-ge, WANG Mei-xia, WANG Chen-yang, MA Gui-gen, ZHOU Chang-yong, WANG Ya-nan, ZHOU Huan-bin. Development and Optimization of Genome Editing in Rice with CRISPR/LanCas9 System [J]. Biotechnology Bulletin, 2024, 40(10): 233-242. |
| [15] | LI Ming-kun, BI Mei-ying, ZHANG Tian-hang, WU Xiang-yu, YANG Pei-ru, YING Ming. Restoration of Agricultural Function of Rhizobacteria by UgRNA/Cas9 Multi-gene Editing [J]. Biotechnology Bulletin, 2024, 40(10): 275-287. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||