








Biotechnology Bulletin ›› 2024, Vol. 40 ›› Issue (4): 148-158.doi: 10.13560/j.cnki.biotech.bull.1985.2023-1045
Previous Articles Next Articles
LI Hui1( ), WEN Yu-fang1, WANG Yue1, JI Chao1, SHI Guo-you1, LUO Ying2, ZHOU Yong1, LI Zhi-min1, WU Xiao-yu1, YANG You-xin2(
), WEN Yu-fang1, WANG Yue1, JI Chao1, SHI Guo-you1, LUO Ying2, ZHOU Yong1, LI Zhi-min1, WU Xiao-yu1, YANG You-xin2( ), LIU Jian-ping1(
), LIU Jian-ping1( )
)
Received:2023-11-07
Online:2024-04-26
Published:2024-04-30
Contact:
YANG You-xin, LIU Jian-ping
E-mail:lh15736630394@163.com;yangyouxin@jxau.edu.cn;JianpingLiu@jxau.edu.cn
LI Hui, WEN Yu-fang, WANG Yue, JI Chao, SHI Guo-you, LUO Ying, ZHOU Yong, LI Zhi-min, WU Xiao-yu, YANG You-xin, LIU Jian-ping. Expression Characteristics and Functions of CaPIF4 in Capsicum annuum Under Salt Stress[J]. Biotechnology Bulletin, 2024, 40(4): 148-158.
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') |
|---|---|
| CaPIF4-F | ATGAATCCTTGTCTTCCTG |
| CaPIF4-R | TCAAACCTGAGAAGCACC |
| CaPIF4-qPCR-F | TGGTTGATCCATCAATCCCTT |
| CaPIF4-qPCR-R | CAAACCTGAGAAGCACCCTGA |
| CaPIF4-VIGS-F | TGAGTAAGGTTACCGAATTCATGAATCCTTGTC- TTCCTGAAT |
| CaPIF4-VIGS-R | GTGAGCTCGGTACCGGATCCACAAAATACTTTG- TCAAATGAAT |
| pGBKT7-CaPIF4-F | ATGGCCATGGAGGCCGAATTCATGAATCCTTGT- CTTCCTGAATGG |
| pGBKT7-CaPIF4-R | CCGCTGCAGGTCGACGGATCCAACCTGAGAAGCACCCTG |
| p3301-CaPIF4-F | ACGGGGGACTCTTGACCATGGATGAATCCTTG- TCTTCCTGAATGG |
| p3301-CaPIF4-F | GCCCTTGCTCACCATCCATGGAACCTGAGAAGCACCCTG |
| CaACTIN-F | AGGGATGGGTCAAAAGGATGC |
| CaACTIN-F | GAGACAACACCGCCTGAATAGC |
Table1 List of primers
| 引物名称 Primer name | 引物序列 Primer sequence(5'-3') |
|---|---|
| CaPIF4-F | ATGAATCCTTGTCTTCCTG |
| CaPIF4-R | TCAAACCTGAGAAGCACC |
| CaPIF4-qPCR-F | TGGTTGATCCATCAATCCCTT |
| CaPIF4-qPCR-R | CAAACCTGAGAAGCACCCTGA |
| CaPIF4-VIGS-F | TGAGTAAGGTTACCGAATTCATGAATCCTTGTC- TTCCTGAAT |
| CaPIF4-VIGS-R | GTGAGCTCGGTACCGGATCCACAAAATACTTTG- TCAAATGAAT |
| pGBKT7-CaPIF4-F | ATGGCCATGGAGGCCGAATTCATGAATCCTTGT- CTTCCTGAATGG |
| pGBKT7-CaPIF4-R | CCGCTGCAGGTCGACGGATCCAACCTGAGAAGCACCCTG |
| p3301-CaPIF4-F | ACGGGGGACTCTTGACCATGGATGAATCCTTG- TCTTCCTGAATGG |
| p3301-CaPIF4-F | GCCCTTGCTCACCATCCATGGAACCTGAGAAGCACCCTG |
| CaACTIN-F | AGGGATGGGTCAAAAGGATGC |
| CaACTIN-F | GAGACAACACCGCCTGAATAGC |
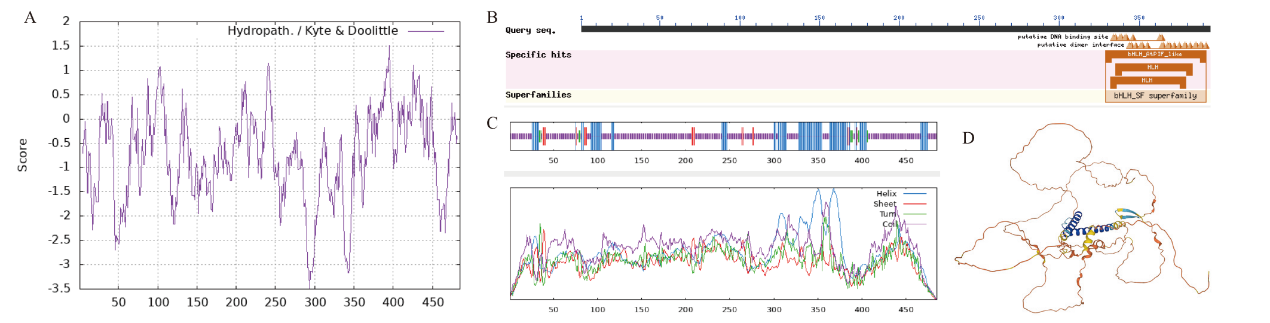
Fig. 2 Sequence analysis of CaPIF4 A: Hydrophilicity analysis of CaPIF4 protein. B: Conserved domain analysis of CaPIF4. C: Secondary structure of CaPIF4 protein. D: Tertiary structure of CaPIF4 protein
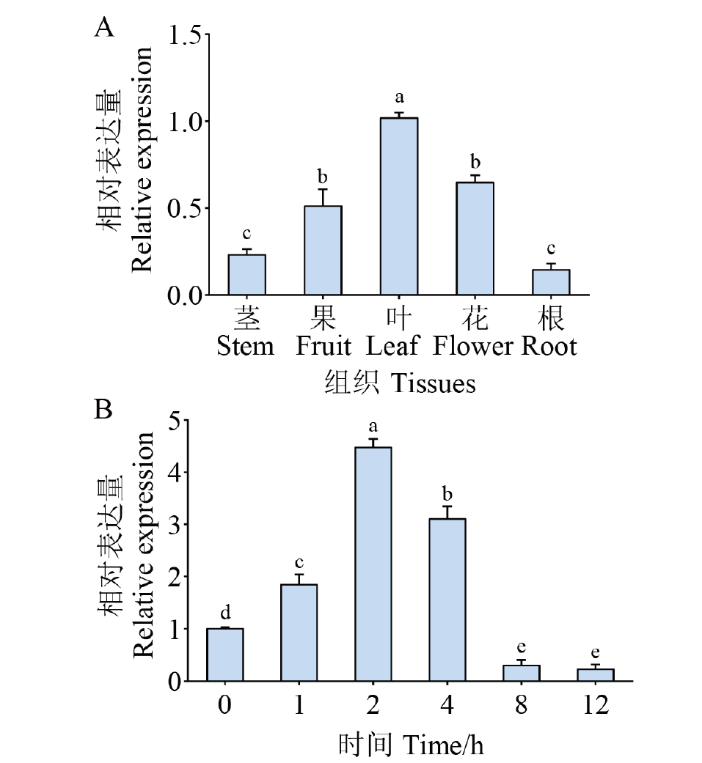
Fig. 3 Expression pattern analysis of CaPIF4 A: Expression analysis of CaPIF4 in different tissues. B: Expression analysis of CaPIF4 in the leaves under salt stress. Mean and SD values were obtained from three independent experiments, with 3 plants per experiments. The different letters indicate significant differences at P ≤0.05

Fig. 4 Construction of pTRV2-CaPIF4 vector A: Enzyme digesting verification of pTRV2-CaPIF4 vector; B: pTRV2-CaPIF4 vector sequence alignment; M: BM2000 DNA marker; 1-3: three repeats
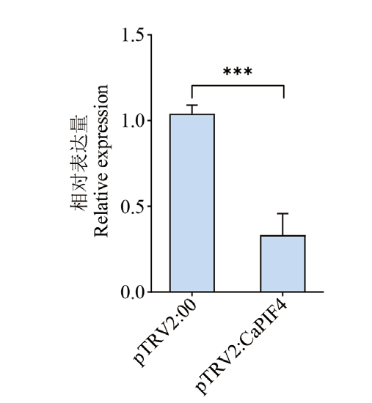
Fig. 6 Detection of CaPIF4 gene silencing efficiency The error line in the figure refers to the standard deviation. * indicate statistical significance (*P<0.05,**P<0.01,***P<0.001 ) compared to control. The same below
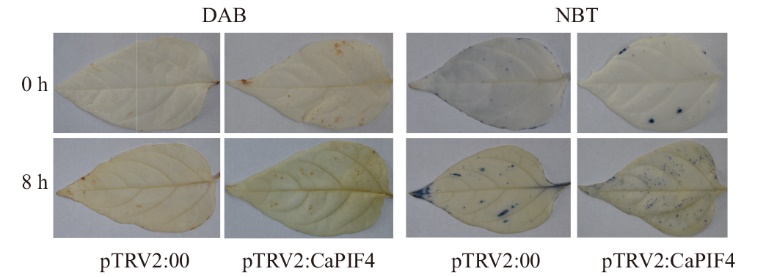
Fig. 10 ROS accumulation in the control group and VIGS silenced plants in the 3rd and 4th leaves before and after salt stress A: 3,3'-Diaminobenzidine (DAB) staining. B: Nitro-blue tetrazolium (NBT) staining
| [1] |
Leivar P, Quail PH. PIFs: pivotal components in a cellular signaling hub[J]. Trends Plant Sci, 2011, 16(1): 19-28.
doi: 10.1016/j.tplants.2010.08.003 pmid: 20833098 |
| [2] |
Lee N, Choi G. Phytochrome-interacting factor from Arabidopsis to liverwort[J]. Curr Opin Plant Biol, 2017, 35: 54-60.
doi: 10.1016/j.pbi.2016.11.004 URL |
| [3] |
Pham VN, Kathare PK, Huq E. Phytochromes and phytochrome interacting factors[J]. Plant Physiol, 2018, 176(2): 1025-1038.
doi: 10.1104/pp.17.01384 pmid: 29138351 |
| [4] |
Inoue K, Nishihama R, Kataoka H, et al. Phytochrome signaling is mediated by phytochrome interacting factor in the liverwort Marchantia polymorpha[J]. Plant Cell, 2016, 28(6): 1406-1421.
doi: 10.1105/tpc.15.01063 URL |
| [5] |
Possart A, Xu TF, Paik I, et al. Characterization of phytochrome interacting factors from the moss Physcomitrella patens illustrates conservation of phytochrome signaling modules in land plants[J]. Plant Cell, 2017, 29(2): 310-330.
doi: 10.1105/tpc.16.00388 URL |
| [6] |
Nakamura Y, Kato T, Yamashino T, et al. Characterization of a set of phytochrome-interacting factor-like bHLH proteins in Oryza sativa[J]. Biosci Biotechnol Biochem, 2007, 71(5): 1183-1191.
doi: 10.1271/bbb.60643 URL |
| [7] |
Shi QB, Zhang HS, Song XY, et al. Functional characterization of the maize phytochrome-interacting factors PIF4 and PIF5[J]. Front Plant Sci, 2018, 8: 2273.
doi: 10.3389/fpls.2017.02273 URL |
| [8] |
Wu GX, Zhao YP, Shen RX, et al. Characterization of maize phytochrome-interacting factors in light signaling and photomorphogenesis[J]. Plant Physiol, 2019, 181(2): 789-803.
doi: 10.1104/pp.19.00239 pmid: 31350363 |
| [9] | Zheng PF, Wang X, Yang YY, et al. Identification of phytochrome-interacting factor family members and functional analysis of MdPIF4 in Malus domestica[J]. Int J Mol Sci, 2020, 21(19): 7350. |
| [10] | Rosado D, Gramegna G, Cruz A, et al. Phytochrome interacting factors(PIFs)in Solanum lycopersicum: Diversity, evolutionary history and expression profiling during different developmental processes[J]. PLoS One, 2016, 11(11): e0165929. |
| [11] |
Ding JH, Zhang B, Li Y, et al. Phytochrome B and PHYTOCHROME INTERACTING FACTOR8 modulate seasonal growth in trees[J]. New Phytol, 2021, 232(6): 2339-2352.
doi: 10.1111/nph.17350 pmid: 33735450 |
| [12] |
Li WQ, Liu Y, Wang WP, et al. Phytochrome-interacting factor(PIF)in rapeseed(Brassica napus L.): Genome-wide identification, evolution and expression analyses during abiotic stress, light quality and vernalization[J]. Int J Biol Macromol, 2021, 180: 14-27.
doi: 10.1016/j.ijbiomac.2021.03.055 URL |
| [13] |
Leivar P, Monte E. PIFs: systems integrators in plant development[J]. Plant Cell, 2014, 26(1): 56-78.
doi: 10.1105/tpc.113.120857 URL |
| [14] |
Sakuraba Y, Bülbül S, Piao WL, et al. Arabidopsis early flowering3 increases salt tolerance by suppressing salt stress response pathways[J]. Plant J, 2017, 92(6): 1106-1120.
doi: 10.1111/tpj.2017.92.issue-6 URL |
| [15] |
Gao Y, Wu MQ, Zhang MJ, et al. Roles of a maize phytochrome-interacting factors protein ZmPIF3 in regulation of drought stress responses by controlling stomatal closure in transgenic rice without yield penalty[J]. Plant Mol Biol, 2018, 97(4/5): 311-323.
doi: 10.1007/s11103-018-0739-4 |
| [16] |
Ma L, Han R, Yang YQ, et al. Phytochromes enhance SOS2-mediated PIF1 and PIF3 phosphorylation and degradation to promote Arabidopsis salt tolerance[J]. Plant Cell, 2023, 35(8): 2997-3020.
doi: 10.1093/plcell/koad117 URL |
| [17] |
Mo WP, Tang WJ, Du YX, et al. PHYTOCHROME-INTERACTING FACTOR-LIKE14 and SLENDER RICE1 interaction controls seedling growth under salt stress[J]. Plant Physiol, 2020, 184(1): 506-517.
doi: 10.1104/pp.20.00024 pmid: 32581115 |
| [18] |
Yang YX, Guang YL, Wang F, et al. Characterization of phytochrome-interacting factor genes in pepper and functional analysis of CaPIF8 in cold and salt stress[J]. Front Plant Sci, 2021, 12: 746517.
doi: 10.3389/fpls.2021.746517 URL |
| [19] |
Paik I, Kathare PK, Kim JI, et al. Expanding roles of PIFs in signal integration from multiple processes[J]. Mol Plant, 2017, 10(8): 1035-1046.
doi: S1674-2052(17)30198-3 pmid: 28711729 |
| [20] |
Bu QY, Zhu L, Huq E. Multiple kinases promote light-induced degradation of PIF1[J]. Plant Signal Behav, 2011, 6(8): 1119-1121.
doi: 10.4161/psb.6.8.16049 pmid: 21758014 |
| [21] |
Al-Sady B, Ni WM, Kircher S, et al. Photoactivated phytochrome induces rapid PIF3 phosphorylation prior to proteasome-mediated degradation[J]. Mol Cell, 2006, 23(3): 439-446.
doi: 10.1016/j.molcel.2006.06.011 pmid: 16885032 |
| [22] |
Shen H, Moon J, Huq E. PIF1 is regulated by light-mediated degradation through the ubiquitin-26S proteasome pathway to optimize photomorphogenesis of seedlings in Arabidopsis[J]. Plant J, 2005, 44(6): 1023-1035.
doi: 10.1111/j.1365-313X.2005.02606.x pmid: 16359394 |
| [23] |
Ni WM, Xu SL, Chalkley RJ, et al. Multisite light-induced phosphorylation of the transcription factor PIF3 is necessary for both its rapid degradation and concomitant negative feedback modulation of photoreceptor phyB levels in Arabidopsis[J]. Plant Cell, 2013, 25(7): 2679-2698.
doi: 10.1105/tpc.113.112342 URL |
| [24] | Paik I, Huq E. Rapid examination of phytochrome-phytochrome interacting factor(PIF)interaction by in vitro coimmunoprecipitation assay[J]. Methods Mol Biol, 2019, 2026: 21-28. |
| [25] |
Ni WM, Xu SL, González-Grandío E, et al. PPKs mediate direct signal transfer from phytochrome photoreceptors to transcription factor PIF3[J]. Nat Commun, 2017, 8: 15236.
doi: 10.1038/ncomms15236 pmid: 28492231 |
| [26] |
Bernardo-García S, de Lucas M, Martínez C, et al. BR-dependent phosphorylation modulates PIF4 transcriptional activity and shapes diurnal hypocotyl growth[J]. Genes Dev, 2014, 28(15): 1681-1694.
doi: 10.1101/gad.243675.114 URL |
| [27] |
Nozue K, Harmer SL, Maloof JN. Genomic analysis of circadian clock-, light-, and growth-correlated genes reveals PHYTOCHROME-INTERACTING FACTOR5 as a modulator of auxin signaling in Arabidopsis[J]. Plant Physiol, 2011, 156(1): 357-372.
doi: 10.1104/pp.111.172684 URL |
| [28] |
Lorrain S, Allen T, Duek PD, et al. Phytochrome-mediated inhibition of shade avoidance involves degradation of growth-promoting bHLH transcription factors[J]. Plant J, 2008, 53(2): 312-323.
doi: 10.1111/j.1365-313X.2007.03341.x pmid: 18047474 |
| [29] |
Huang GT, Ma SL, Bai LP, et al. Signal transduction during cold, salt, and drought stresses in plants[J]. Mol Biol Rep, 2012, 39(2): 969-987.
doi: 10.1007/s11033-011-0823-1 URL |
| [30] | Ma YL, Cao J, He JH, et al. Molecular mechanism for the regulation of ABA homeostasis during plant development and stress responses[J]. Int J Mol Sci, 2018, 19(11): 3643. |
| [31] |
Zhu JK. Salt and drought stress signal transduction in plants[J]. Annu Rev Plant Biol, 2002, 53: 247-273.
doi: 10.1146/arplant.2002.53.issue-1 URL |
| [32] | 李俊贞, 何乐祖, 赵春梅, 等. 盐胁迫对黄果厚壳桂幼苗荧光和生理特性的影响[J]. 山西农业科学, 2021, 49(8): 919-923. |
| Li JZ, He LZ, Zhao CM, et al. Effects of salt stress on the photosynthetic and physiological characteristics of cryptocarya concinna seedlings[J]. J Shanxi Agric Sci, 2021, 49(8): 919-923. | |
| [33] | 付丽, 刘加珍, 陶宝先, 等. 盐生植物对盐渍土壤环境的适应机制研究综述[J]. 江苏农业科学, 2021, 49(15): 32-39. |
| Fu L, Liu JZ, Tao BX, et al. A review on the adaptationmechanism of halophytes to saline soil environment[J]. J Jiangsu Agric Sci, 2021, 49(15): 32-39. | |
| [34] |
Huang RD. Research progress on plant tolerance to soil salinity and alkalinity in sorghum[J]. J Integr Agric, 2018, 17(4): 739-746.
doi: 10.1016/S2095-3119(17)61728-3 URL |
| [35] | 秦峰梅, 张红香, 武祎, 等. 盐胁迫对黄花苜蓿发芽及幼苗生长的影响[J]. 草业学报, 2010, 19(4): 71-78. |
| Qin FM, Zhang HX, Wu Y, et al. Effects of salt stress on germination and seedling growth of Medicago falcata[J]. Acta Prataculturae Sin, 2010, 19(4): 71-78. | |
| [36] |
时振振, 李胜, 杨柯, 等. 盐胁迫下豌豆幼苗对内外源NO的生理生化响应[J]. 草业学报, 2014, 23(5): 193-200.
doi: 10.11686/cyxb20140522 |
| Shi ZZ, Li S, Yang K, et al. Physiological and biochemical response of pea seedlings to endogenous and exogenous NO under salt stress[J]. Acta Prataculturae Sin, 2014, 23(5): 193-200. | |
| [37] | 孙璐, 黄瑞冬. 高粱幼苗保护酶系统对盐胁迫的初期响应[J]. 沈阳农业大学学报, 2014, 45(2): 134-137. |
| Sun L, Huang RD. Responses to salt stress of protective enzyme system in sorghum seedlings[J]. J Shenyang Agric Univ, 2014, 45(2): 134-137. | |
| [38] | 田晓艳, 刘延吉, 郭迎春. 盐胁迫对NHC牧草Na+、K+、Pro、可溶性糖及可溶性蛋白的影响[J]. 草业科学, 2008, 25(10): 34-38. |
| Tian XY, Liu YJ, Guo YC. Effect of salt stress on Na+, K+, Pro, soluble sugar and soluble protein of NHC[J]. Pratacultural Sci, 2008, 25(10): 34-38. | |
| [39] | Feng XJ, Li JR, Qi SL, et al. Light affects salt stress-induced transcriptional memory of P5CS1 in Arabidopsis[J]. Proc Natl Acad Sci USA. 2016, 113(51): E8335-E8343. |
| [40] | Seong ES, Cho HS, Choi D, et al. Tomato plants overexpressing CaKR1 enhanced tolerance to salt and oxidative stress[J]. Biochem Biophys Res Commun, 2007 ; 363(4): 983-988. |
| [41] |
宋阳, 崔晓晗, 张明, 等. 盐胁迫对番茄幼苗生理特性及离子分布的影响[J]. 北方农业学报, 2019, 47(4): 115-121.
doi: 10.3969/j.issn.2096-1197.2019.04.20 |
| Song Y, Cui XH, Zhang M, et al. Effects of salt stress on physiological characteristics and ion distribution of tomato seedlings[J]. J North Agric, 2019, 47(4): 115-121. |
| [1] | XIAO Ya-ru, JIA Ting-ting, LUO Dan, WU Zhe, LI Li-xia. Cloning and Expression Analysis of CsERF025L Transcription Factor in Cucumber [J]. Biotechnology Bulletin, 2024, 40(4): 159-166. |
| [2] | GUO Chun, SONG Gui-mei, YAN Yan, DI Peng, WANG Ying-ping. Genome Wide Identification and Expression Analysis of the bZIP Gene Family in Panax quinquefolius [J]. Biotechnology Bulletin, 2024, 40(4): 167-178. |
| [3] | GAO Yu-kun, ZHANG Jian-dong, YANG Pu-yuan, CHEN Dong-ming, WANG Zhi-bo, TIAN Yi-jin, Zakey Eldinn. E. A. Khlid, CUI Jiang-hui, CHANG Jin-hua. Responses of Sorghum Rhizosphere Soil Bacterial Communities to Salt Stress [J]. Biotechnology Bulletin, 2024, 40(4): 203-216. |
| [4] | GAO Zhi-wei, WEI Ming, YU Zu-long, WU Guo-qiang, WEI Jun-long. Identification of Salt-tolerant Plant Growth-promoting Bacterium W-1 and Its Effect on the Salt-tolerance of Sainfoin(Onobrychis viciaefolia) [J]. Biotechnology Bulletin, 2024, 40(4): 217-227. |
| [5] | ZHANG Yu, SHI Lei, GONG Lei, NIE Feng-jie, YANG Jiang-wei, LIU Xuan, YANG Wen-jing, ZHANG Guo-hui, XIE Rui-xia, ZHANG Li. Genome-wide Identification of Potato WOX Gene Family and Its Expression Analysis in in vitro Regeneration and Abiotic Stress [J]. Biotechnology Bulletin, 2024, 40(3): 170-180. |
| [6] | WU Xing-xing, HONG Hai-bo, GAN Zhi-cheng, LI Rui-ning, HUANG Xian-zhong. Cloning and Preliminary Functional Analysis of CaPI Gene in Capsicum annuum L. [J]. Biotechnology Bulletin, 2024, 40(3): 193-201. |
| [7] | XIE Qian, JIANG Lai, HE Jin, LIU Ling-ling, DING Ming-yue, CHEN Qing-xi. Regulatory Genes Mining Related to Transcriptome Sequencing and Phenolic Metabolism Pathway of Canarium album Fruit with Different Fresh Food Quality [J]. Biotechnology Bulletin, 2024, 40(3): 215-228. |
| [8] | SHEN Tian-hong, QI Xiao-bo, ZHAO Rui-feng, MA Xin-rong. Research Progress in the Molecular Mechanisms of Microalgae Responding to Salt Stress [J]. Biotechnology Bulletin, 2024, 40(3): 89-99. |
| [9] | CHEN Yan-mei. Crosstalk Between Different Post-translational Modifications and Its Regulatory Mechanisms in Plant Growth and Development [J]. Biotechnology Bulletin, 2024, 40(2): 1-8. |
| [10] | YANG Yan, HU Yang, LIU Ni-ru, YIN Lu, YANG Rui, WANG Peng-fei, MU Xiao-peng, ZHANG Shuai, CHENG Chun-zhen, ZHANG Jian-cheng. Cloning and Functional Analysis of MbbZIP43 Gene in ‘Hongmantang’ Red-flesh Apple [J]. Biotechnology Bulletin, 2024, 40(2): 146-159. |
| [11] | LU Yu-dan, LIU Xiao-chi, FENG Xin, CHEN Gui-xin, CHEN Yi-ting. Identification of the Kiwifruit BBX Gene Family and Analysis of Their Transcriptional Characteristics [J]. Biotechnology Bulletin, 2024, 40(2): 172-182. |
| [12] | LI Hao, WU Guo-qiang, WEI Ming, HAN Yue-xin. Genome-wide Identification of the BvBADH Gene Family in Sugar Beet(Beta vulgaris)and Their Expression Analysis Under High Salt Stress [J]. Biotechnology Bulletin, 2024, 40(2): 233-244. |
| [13] | XU Yang, ZHANG Rui-ying, DAI Liang-xiang, ZHANG Guan-chu, DING Hong, ZHANG Zhi-meng. Regulation of Nitrogen Application on Peanut Seed Germination and Spermosphere Bacterial Community Structure Under Salt Stress [J]. Biotechnology Bulletin, 2024, 40(2): 253-265. |
| [14] | ZHANG Dan-dan, ZHAO Rui-xue, XIAN Guo-jian, XIONG He. Trait-regulated-genes Ontology Model Construction and Application by Integrating Cross-species Scientific Data [J]. Biotechnology Bulletin, 2024, 40(2): 313-324. |
| [15] | ZHANG Hong-min, LONG Wen, LAO Xiao-qing, CHEN Wen-yan, SHANG Xue-mei, WANG Hong-lian, WANG Li, SU Hong-wei, SHEN Hong-ping, SHEN Hong-chun. Construction of Pmepa1 Knockout TCMK1 Mouse Renal Tubular Epithelial Cell Line Using CRISPR/Cas9 Technology [J]. Biotechnology Bulletin, 2024, 40(2): 73-79. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||