








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 17-25.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0590
Previous Articles Next Articles
YAN Qi-qi( ), BU Yu-fen, ZHANG Xiao-xin, MA Xiao-cen, JING Yan-ping(
), BU Yu-fen, ZHANG Xiao-xin, MA Xiao-cen, JING Yan-ping( )
)
Received:2025-06-07
Online:2026-04-26
Published:2026-04-30
Contact:
JING Yan-ping
E-mail:18514235629@163.com;ypjing@bjfu.edu.cn
YAN Qi-qi, BU Yu-fen, ZHANG Xiao-xin, MA Xiao-cen, JING Yan-ping. Advances in the Studies of Plant C2 Domain Abscisic Acid-related Protein[J]. Biotechnology Bulletin, 2026, 42(4): 17-25.
| 蛋白名称 Protein name | 物种 Species | 主要功能 Major functions | 参考文献 Reference |
|---|---|---|---|
| AtCAR1 | 拟南芥 | 参与ABA信号,增强ABA敏感性;负调控碱胁迫响应 | [ |
| AtCAR4 | 拟南芥 | 参与ABA信号;正调控盐胁迫与生物胁迫抗性 | [ |
| AtCAR5 | 拟南芥 | 参与ABA信号 | [ |
| AtCAR6/EHB1 | 拟南芥 | 负调控向光性与向重性;负调控铁吸收;负调控碱胁迫响应 | [ |
| AtCAR9 | 拟南芥 | 参与ABA信号;正调控干旱胁迫耐受性 | [ |
| AtCAR10 | 拟南芥 | 负调控碱胁迫响应 | [ |
| IbCAR1 | 甘薯 | 增强盐胁迫下细胞完整性,激活ROS清除系统 | [ |
| OsGAP1/CAR4 | 水稻 | 增强盐胁迫与生物胁迫抗性 | [ |
Table 1 Main functions of CAR proteins in different plants
| 蛋白名称 Protein name | 物种 Species | 主要功能 Major functions | 参考文献 Reference |
|---|---|---|---|
| AtCAR1 | 拟南芥 | 参与ABA信号,增强ABA敏感性;负调控碱胁迫响应 | [ |
| AtCAR4 | 拟南芥 | 参与ABA信号;正调控盐胁迫与生物胁迫抗性 | [ |
| AtCAR5 | 拟南芥 | 参与ABA信号 | [ |
| AtCAR6/EHB1 | 拟南芥 | 负调控向光性与向重性;负调控铁吸收;负调控碱胁迫响应 | [ |
| AtCAR9 | 拟南芥 | 参与ABA信号;正调控干旱胁迫耐受性 | [ |
| AtCAR10 | 拟南芥 | 负调控碱胁迫响应 | [ |
| IbCAR1 | 甘薯 | 增强盐胁迫下细胞完整性,激活ROS清除系统 | [ |
| OsGAP1/CAR4 | 水稻 | 增强盐胁迫与生物胁迫抗性 | [ |
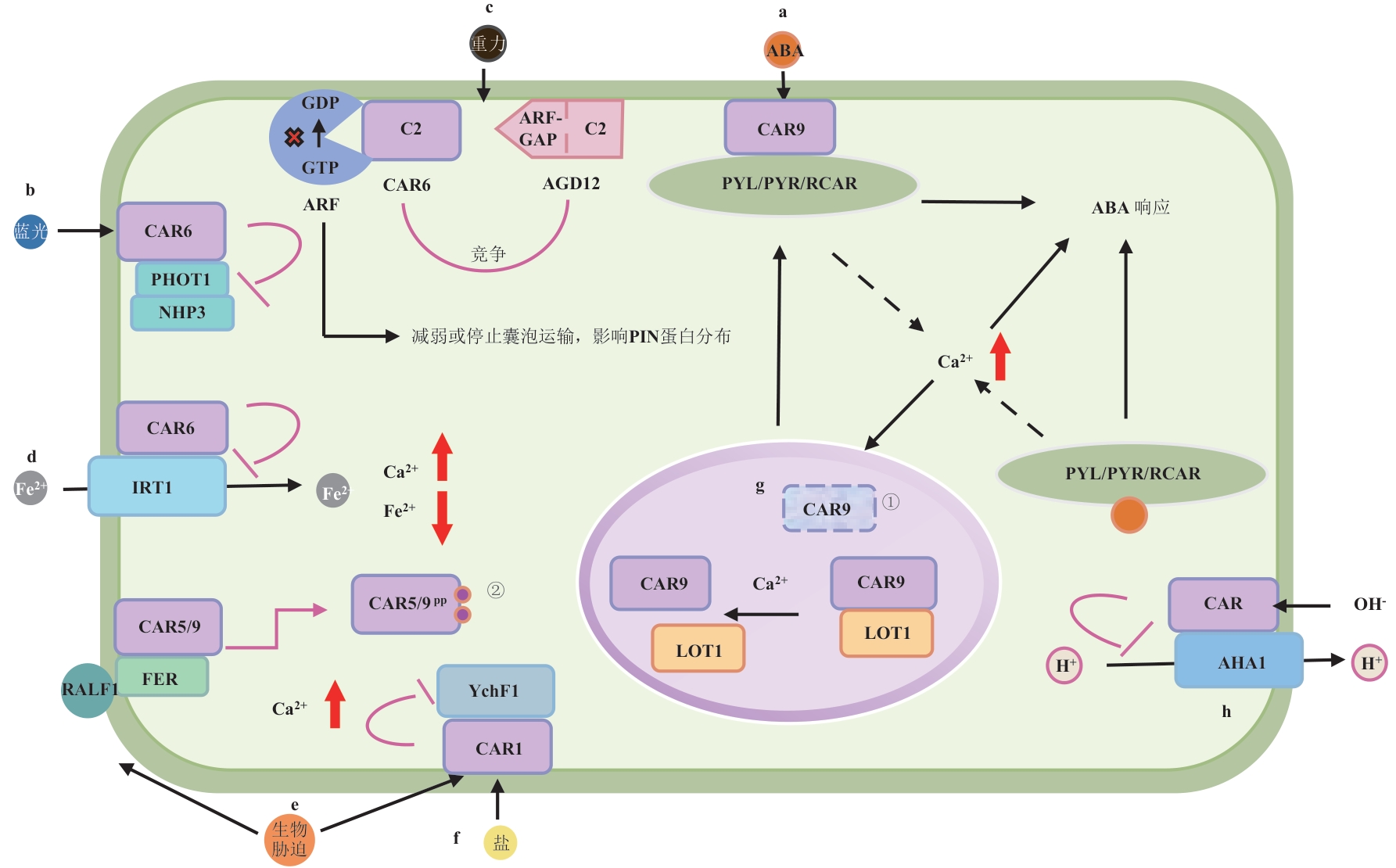
Fig. 2 Regulatory network and biological functions associated with CAR proteina & g: The LOT1 protein interacts with the CAR protein in the nucleus, inhibiting the ubiquitination modification of the CAR protein and thereby maintaining its stability. When subjected to drought stress or in the presence of ABA signaling, ABA is recognized by cytosolic or nuclear (not shown) PYR/PYL/RCAR receptors (abbreviated as PYLs). This recognition triggers an increase in cytosolic or nuclear Ca²⁺ concentration, which reduces the affinity between the CAR protein and the LOT1 protein. This promotes the relocation of the CAR protein to the plasma membrane, where it mediates the membrane recruitment of ABA receptor PYLs. This process enables the cell to respond to ABA signaling and enhances the plant’s drought resistance[19-20]; b: CAR binds to the BTB/POZ domain of NPH3, thereby impairing CUL3-NPH3 assembly, inhibiting ubiquitin-dependent degradation of PHOT1, and consequently attenuating plant phototropism[26]; c: CAR competitively inhibits ARF-GAP family members (e.g., AGD12) by binding to ADP-ribosylation factor (ARF) GTPases. This disrupts ARF GTP/GDP cycling, impairs PIN protein polar localization via vesicle trafficking defects, and compromises root gravitropism[27]; d: CAR protein negatively regulates the plant iron transporter IRT1, thereby affecting plant iron uptake[23]; e: CAR proteins negatively regulate YchF1-a negative regulator of pathogen infection-thereby relieving its inhibitory effect and enhancing plant resistance to biotic stresses[29]. Additionally, CAR proteins stabilize lipid raft structures, promoting the enrichment of immune-related proteins and strengthening immune signaling. Furthermore, CARs that interact with FER undergo phosphorylation, dissociate from the membrane, maintain the dynamic balance of membrane protein content, and prevent excessive activation of immune signaling[25]; f: CAR inhibits the salt stress negative regulator YchF1, conferring enhanced salinity tolerance[32]; h: CAR downregulates the plasma membrane H⁺-ATPase AHA1, thereby compromising plant tolerance to alkaline stress[28]. Red arrows indicate ion concentration increase (↑) or decrease (↓). Label ① marks ubiquitination of CAR, and ② indicates phosphorylation of CAR
| [1] | Ono Y, Kurokawa T, Kawahara K, et al. Cloning of rat brain protein kinase C complementary DNA [J]. FEBS Lett, 1986, 203(2): 111-115. |
| [2] | Parker PJ, Coussens L, Totty N, et al. The complete primary structure of protein kinase C-the major phorbol ester receptor [J]. Science, 1986, 233(4766): 853-859. |
| [3] | Ono Y, Kurokawa T, Fujii T, et al. Two types of complementary DNAs of rat brain protein kinase C Heterogeneity determined by alternative splicing [J]. FEBS Lett, 1986, 206(2): 347-352. |
| [4] | Geribaldi-Doldán N, Gómez-Oliva R, Domínguez-García S, et al. Protein kinase C: targets to regenerate brain injuries [J]. Front Cell Dev Biol, 2019, 7: 39. |
| [5] | Nishizuka Y. The molecular heterogeneity of protein kinase C and its implications for cellular regulation [J]. Nature, 1988, 334(6184): 661-665. |
| [6] | Bazzi MD, Nelsestuen GL. Protein kinase C interaction with calcium: a phospholipid-dependent process [J]. Biochemistry, 1990, 29(33): 7624-7630. |
| [7] | Bazzi MD, Nelsestuen GL. Association of protein kinase C with phospholipid vesicles [J]. Biochemistry, 1987, 26(1): 115-122. |
| [8] | Zhang DP, Aravind L. Identification of novel families and classification of the C2 domain superfamily elucidate the origin and evolution of membrane targeting activities in eukaryotes [J]. Gene, 2010, 469(1/2): 18-30. |
| [9] | 陈锦华. 拟南芥C2结构域蛋白的进化分析及其功能研究 [D]. 长沙: 湖南农业大学, 2019. |
| Chen JH. Analysis of evolution and function of C2 domain proteins in Arabidopsis [D]. Changsha: Hunan Agricultural University, 2019. | |
| [10] | Zhang HJ, Zeng YT, Seo J, et al. Global identification and characterization of C2 domain-containing proteins associated with abiotic stress response in rice (Oryza sativa L.) [J]. Int J Mol Sci, 2022, 23(4): 2221. |
| [11] | Zhao LY, Wang JZ, Zhou YT, et al. Genome-wide identification of plant C2 domain-containing protein family and the role of OsNTMC2T2.2 under chilling stress in rice [J]. Environ Exp Bot, 2025, 237: 106202. |
| [12] | Wang NN, Shi YY, Jiang Q, et al. A 14-3-3 protein positively regulates rice salt tolerance by stabilizing phospholipase C1 [J]. Plant Cell Environ, 2023, 46(4): 1232-1248. |
| [13] | Han SC, Wang YL, Li YX, et al. The OsNAC41-RoLe1-OsAGAP module promotes root development and drought resistance in upland rice [J]. Mol Plant, 2024, 17(10): 1573-1593. |
| [14] | Schneider R, Tang L, Lampugnani ER, et al. Two complementary mechanisms underpin cell wall patterning during xylem vessel development [J]. Plant Cell, 2017, 29(10): 2433-2449. |
| [15] | Sun Y, Zhao JY, Li YT, et al. Genome-wide analysis of the C2 domain family in soybean and identification of a putative abiotic stress response gene GmC2-148 [J]. Front Plant Sci, 2021, 12: 620544. |
| [16] | Niu JS, Li ZJ, Zhu JR, et al. Genome-wide identification and characterization of the C2 domain family in Sorghum bicolor (L.) and expression profiles in response to saline-alkali stress [J]. Physiol Mol Biol Plants, 2022, 28(9): 1695-1711. |
| [17] | 张兰军, 姬飞腾, 王丽丽, 等. 复苏植物旋蒴苣苔C2结构域小蛋白BhC2DP1参与植物对ABA的反应 [J]. 植物学报, 2012, 47(1): 11-27. |
| Zhang LJ, Ji FT, Wang LL, et al. A small C2-domain protein from the resurrection plant Boea hygrometrica promotes plant responses to abscisic acid [J]. Chin Bull Bot, 2012, 47(1): 11-27. | |
| [18] | Guo AY, Wu WQ, Bai D, et al. Recruitment of HAB1 and SnRK2.2 by C2-domain protein CAR1 in plasma membrane ABA signaling [J]. Plant J, 2024, 119(1): 237-251. |
| [19] | Rodriguez L, Gonzalez-Guzman M, Diaz M, et al. C2-domain abscisic acid-related proteins mediate the interaction of PYR/PYL/RCAR abscisic acid receptors with the plasma membrane and regulate abscisic acid sensitivity in Arabidopsis [J]. Plant Cell, 2014, 26(12): 4802-4820. |
| [20] | Qin T, Tian QZ, Wang GF, et al. LOWER TEMPERATURE 1 enhances ABA responses and plant drought tolerance by modulating the stability and localization of C2-domain ABA-related proteins in Arabidopsis [J]. Mol Plant, 2019, 12(9): 1243-1258. |
| [21] | Diaz M, Sanchez-Barrena MJ, Gonzalez-Rubio JM, et al. Calcium-dependent oligomerization of CAR proteins at cell membrane modulates ABA signaling [J]. Proc Natl Acad Sci USA, 2016, 113(3): E396-E405. |
| [22] | Rizo J, Südhof TC. C2-domains, structure and function of a universal Ca2+-binding domain [J]. J Biol Chem, 1998, 273(26): 15879-15882. |
| [23] | Khan I, Gratz R, Denezhkin P, et al. Calcium-promoted interaction between the C2-domain protein EHB1 and metal transporter IRT1 inhibits Arabidopsis iron acquisition [J]. Plant Physiol, 2019, 180(3): 1564-1581. |
| [24] | Cui MM, Gupta SK, Bauer P. Role of the plant-specific calcium-binding C2-domain abscisic acid-related (car) protein family in environmental signaling [J]. Eur J Cell Biol, 2023, 102(2): 151322. |
| [25] | Chen WJ, Zhou HN, Xu F, et al. CAR modulates plasma membrane nano-organization and immune signaling downstream of RALF1-FERONIA signaling pathway [J]. New Phytol, 2023, 237(6): 2148-2162. |
| [26] | Knauer T, Dümmer M, Landgraf F, et al. A negative effector of blue light-induced and gravitropic bending in Arabidopsis [J]. Plant Physiol, 2011, 156(1): 439-447. |
| [27] | Dümmer M, Michalski C, Essen LO, et al. EHB1 and AGD12, two calcium-dependent proteins affect gravitropism antagonistically in Arabidopsis thaliana [J]. J Plant Physiol, 2016, 206: 114-124. |
| [28] | Guo AY, Wu WQ, Liu WC, et al. C2-domain abscisic acid-related proteins regulate the dynamics of a plasma membrane H+-ATPase in response to alkali stress [J]. Plant Physiol, 2024, 196(4): 2784-2794. |
| [29] | Cheng SS, Ku YS, Cheung MY, et al. AtGAP1 promotes the resistance to Pseudomonas syringae pv. tomato DC3000 by regulating cell-wall thickness and stomatal aperture in Arabidopsis [J]. Int J Mol Sci, 2022, 23(14): 7540. |
| [30] | Cheung MY, Li MW, Yung YL, et al. The unconventional P-loop NTPase OsYchF1 and its regulator OsGAP1 play opposite roles in salinity stress tolerance [J]. Plant Cell Environ, 2013, 36(11): 2008-2020. |
| [31] | Cheung MY, Zeng NY, Tong SW, et al. Constitutive expression of a rice GTPase-activating protein induces defense responses [J]. New Phytol, 2008, 179(2): 530-545. |
| [32] | You C, Li C, Ma M, et al. A C2-domain abscisic acid-related gene, IbCAR1, positively enhances salt tolerance in sweet potato (Ipomoea batatas (L.) lam.) [J]. Int J Mol Sci, 2022, 23(17): 9680. |
| [33] | Cheung MY, Ngo JC, Chen ZZ, et al. A structure model explaining the binding between a ubiquitous unconventional G-protein (OsYchF1) and a plant-specific C2-domain protein (OsGAP1) from rice [J]. Biochem J, 2020, 477(20): 3935-3949. |
| [34] | Soon FF, Ng LM, Zhou XE, et al. Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases [J]. Science, 2012, 335(6064): 85-88. |
| [35] | Ma Y, Szostkiewicz I, Korte A, et al. Regulators of PP2C phosphatase activity function as abscisic acid sensors [J]. Science, 2009, 324(5930): 1064-1068. |
| [36] | Park SY, Fung P, Nishimura N, et al. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins [J]. Science, 2009, 324(5930): 1068-1071. |
| [37] | Nishimura N, Sarkeshik A, Nito K, et al. PYR/PYL/RCAR family members are major in-vivo ABI1 protein phosphatase 2C-interacting proteins in Arabidopsis [J]. Plant J, 2010, 61(2): 290-299. |
| [38] | Christie JM, Suetsugu N, Sullivan S, et al. Shining light on the function of NPH3/RPT2-like proteins in phototropin signaling [J]. Plant Physiol, 2018, 176(2): 1015-1024. |
| [39] | Christie JM. Phototropin blue-light receptors [J]. Annu Rev Plant Biol, 2007, 58: 21-45. |
| [40] | Briggs WR, Christie JM. Phototropins 1 and 2: versatile plant blue-light receptors [J]. Trends Plant Sci, 2002, 7(5): 204-210. |
| [41] | Pedmale UV, Liscum E. Regulation of phototropic signaling in Arabidopsis via phosphorylation state changes in the phototropin 1-interacting protein NPH3 [J]. J Biol Chem, 2007, 282(27): 19992-20001. |
| [42] | Sakai T, Ken HG. Molecular genetic analysis of phototropism in Arabidopsis [J]. Plant Cell Physiol, 2012, 53(9): 1517-1534. |
| [43] | Vernoud V, Horton AC, Yang ZB, et al. Analysis of the small GTPase gene superfamily of Arabidopsis [J]. Plant Physiol, 2003, 131(3): 1191-1208. |
| [44] | Dümmer M, Forreiter C, Galland P. Gravitropism in Arabidopsis thaliana: Root-specific action of the EHB gene and violation of the resultant law [J]. J Plant Physiol, 2015, 189: 24-33. |
| [45] | Cointry V, Vert G. The bifunctional transporter-receptor IRT1 at the heart of metal sensing and signalling [J]. New Phytol, 2019, 223(3): 1173-1178. |
| [46] | Reyt G, Boudouf S, Boucherez J, et al. Iron- and ferritin-dependent reactive oxygen species distribution: impact on Arabidopsis root system architecture [J]. Mol Plant, 2015, 8(3): 439-453. |
| [47] | Vert G, Grotz N, Dédaldéchamp F, et al. IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth [J]. Plant Cell, 2002, 14(6): 1223-1233. |
| [48] | Yung YL, Cheung MY, Miao R, et al. Site-directed mutagenesis shows the significance of interactions with phospholipids and the G-protein OsYchF1 for the physiological functions of the rice GTPase-activating protein 1 (OsGAP1) [J]. J Biol Chem, 2015, 290(39): 23984-23996. |
| [49] | 强晓楠, 李鑫, 陈佳, 等. 拟南芥RALF多肽家族的功能多样性初步分析 [J]. 生物技术通报, 2019, 35(1): 2-10. |
| Qiang XN, Li X, Chen J, et al. Preliminary analysis of functional diversity of RALF peptide family in Arabidopsis thaliana [J]. Biotechnol Bull, 2019, 35(1): 2-10. | |
| [50] | 谌伟军. 拟南芥RALF-FER信号调控CAR家族介导膜筏形成及其初步功能研究 [D]. 长沙: 湖南大学, 2022. |
| Chen WJ. The receptor kinase FERONIA-CAR axis regulates proper lipid order formation and its preliminary function to response RALF signaling [D]. Changsha: Hunan University, 2022. | |
| [51] | Fu Q, Li HB, Wang BQ, et al. The RALF1 peptide-FERONIA complex phosphorylates the endosomal sorting protein FREE1 to attenuate abscisic acid signaling [J]. Plant Physiol, 2024, 197(1): kiae625. |
| [1] | LIU Lin-ya, LIU Huan-yan, LIANG Xin-yu, SONG Shu-yi, HE Bin, WANG Xu-ying, HUANG Ya-cheng. Genome-wide Identification and Expression Analysis of BGAL Gene Family in Actinidiachinensis var. Hongyang [J]. Biotechnology Bulletin, 2026, 42(3): 312-323. |
| [2] | MA Ying-ying, YOU Hui-wan, ZHENG Ji-rong, WANG Qiao-mei, LIU Li-hong. Advances in the Quality Formation Mechanism of Horticultural Crops Based on Multi-level Regulation of PSY [J]. Biotechnology Bulletin, 2026, 42(3): 96-110. |
| [3] | CUI Zhi-han, WEI Qing-zhen, HU Na, BAO Chong-lai, WANG Hua-sen. Research Progress in IQD Genes in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 230-241. |
| [4] | FU Han, SUN Shu-hao, ZHANG Si-qing, AI Niu, YU Yang, YU Lian-wei, WANG Qiong-qiong, HAN Xiao-yu, SHI Yan, HAN Wei-li, YANG Xue. Genome-wide Identification and Expression Analysis of the BOI Gene Family in Nicotiana benthamiana [J]. Biotechnology Bulletin, 2026, 42(2): 207-217. |
| [5] | GAN Chen-lu, YOU Yu-ting, XIE Han-dan, ZENG Zi-xian, ZHU Bo. Research Progress in Flavin Monooxygenases in Plants [J]. Biotechnology Bulletin, 2026, 42(1): 1-12. |
| [6] | WANG Cong-huan, WU Guo-qiang, WEI Ming. Functional Mechanism of Plant CBL in Regulating the Responses to Abiotic and Biotic Stresses [J]. Biotechnology Bulletin, 2025, 41(7): 1-16. |
| [7] | WEI Yu-jia, LI Yan, KANG Yu-han, GONG Xiao-nan, DU Min, TU Lan, SHI Peng, YU Zi-han, SUN Yan, ZHANG Kun. Cloning and Expression Analysis of the CrMYB4 Gene in Carex rigescens [J]. Biotechnology Bulletin, 2025, 41(7): 248-260. |
| [8] | LI Rui, HU Ting, CHEN Shu-wei, WANG Yao, WANG Ji-ping. Positive Regulation of Anthocyanin Biosynthesis by PfMYB80 Transcription Factor in Perilla frutescens [J]. Biotechnology Bulletin, 2025, 41(6): 243-255. |
| [9] | ZAN Shu-wen, XIE Huan-huan, ZHANG Yu-qin, WANG Wen-Juan, ZHANG Peng-fei, LIANG Jin-jun, WEN Peng-fei. VvAGAMOUS Regulates Carpel Development through VvCRABS CLAW in Grape [J]. Biotechnology Bulletin, 2025, 41(5): 208-217. |
| [10] | WANG Bin, LIN Wei, XIAO Yan-hui, YUAN Xiao. Research Progress in the Roles of Plant Glycine-rich Protein Family [J]. Biotechnology Bulletin, 2025, 41(2): 1-17. |
| [11] | LI Jing-jing, HU Jin-hong, LIANG Wang-li, MA Yu-rong, LIANG Wen-yu, WANG Ling-xia. Differential Expression Analysis of Genes Related to NaCl Stress Response in Lycium barbarum 'Ningqi 1' [J]. Biotechnology Bulletin, 2025, 41(2): 202-209. |
| [12] | FANG Yu-jie, LIU Kuan, CUI Han-bing, WANG You-ping. Recent Advances in Plant SUMO E3 Ligases [J]. Biotechnology Bulletin, 2025, 41(12): 1-15. |
| [13] | WANG Bi-cheng, JING Hai-qing, WAN Kun, ZHANG Ying-ying, DING Jia-hao, LI Run-zhi, XUE Jin-ai, ZHANG Hai-ping. Identification of Soybean BCAT Gene Family and Functional Analysis of GmBCAT3 in Soybean Responses to Drought Stress [J]. Biotechnology Bulletin, 2025, 41(10): 196-209. |
| [14] | WU Ding-jie, CHEN Ying-ying, XU Jing, LIU Yuan, ZHANG Hang, LI Rui-li. Research Progress in Plant Gibberellin Oxidase and Its Functions [J]. Biotechnology Bulletin, 2024, 40(7): 43-54. |
| [15] | HAO Si-yi, ZHANG Jun-ke, WANG Bin, QU Peng-yan, LI Rui-de, CHENG Chun-zhen. Cloning and Expression Analysis of Banana EARLY FLOWERING 3(ELF3)Genes [J]. Biotechnology Bulletin, 2024, 40(5): 131-140. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||