








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 92-100.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0903
Previous Articles Next Articles
XU Meng-ge( ), SONG Huo-yan, LUO Jia, SU Yi, ZHOU Hui-wen, WANG Can, KONG Ke-ke(
), SONG Huo-yan, LUO Jia, SU Yi, ZHOU Hui-wen, WANG Can, KONG Ke-ke( )
)
Received:2025-08-20
Online:2026-04-26
Published:2026-04-30
Contact:
KONG Ke-ke
E-mail:943723370@qq.com;kekekong2011011034@163.com
XU Meng-ge, SONG Huo-yan, LUO Jia, SU Yi, ZHOU Hui-wen, WANG Can, KONG Ke-ke. Expression Analysis and Interaction Protein Screening of GmRLK19 in Soybean[J]. Biotechnology Bulletin, 2026, 42(4): 92-100.
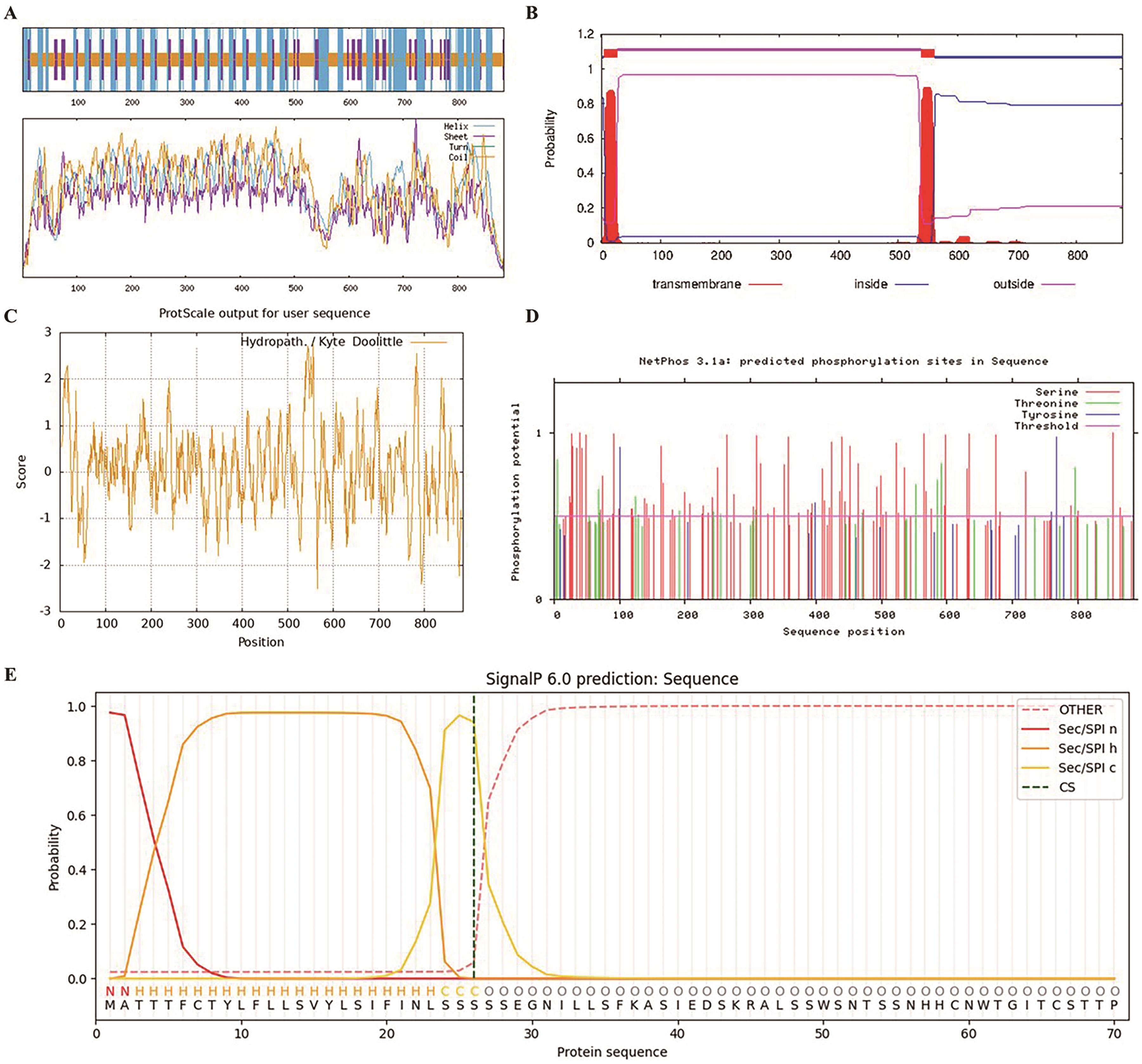
Fig. 1 Bioinformatics analysis of GmRLK19A: Secondary structure prediction; B: transmembrane structure prediction; C: hydrophilic/hydrophobic prediction; D: phosphorylation site prediction; E: signal peptide prediction
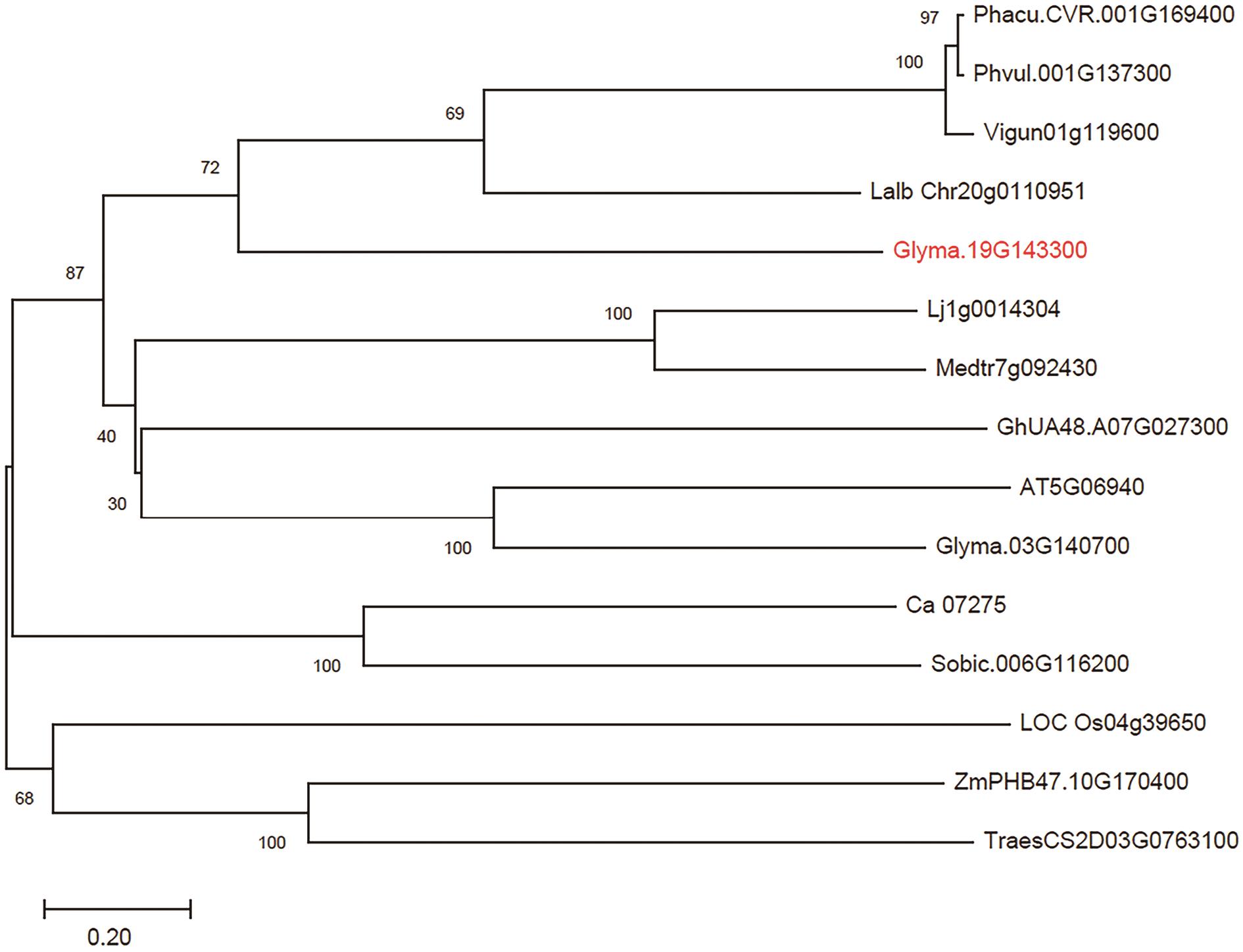
Fig. 2 Phylogenetic tree analysis of GmRLK19 and its homologous proteinPhylogenetic tree analysis of GmRLK19 and homologous proteins. The numbers in the nodes indicate 1 000-time Bootstrap. It is depicted using the full-length amino acid sequences of the 14 proteins that are homologous with GmRLK19, including Phacu.CVR.001G169400 (Phaseolus vulgaris); Phvul.001G137300 (Phaseolus acutifolius); Vigun01g119600 (Vigna unguiculata); Lalb Chr20g0110951 (Lupinus albus); Glyma.19G143300 (Glycine max); Lj1g0014304 (Lotus japonicus); Medtr7g092430 (Medicago truncatula); GhUA48.A07G027300 (Gossypium hirsutum); AT5G06940 (Arabidopsis thaliana); Glyma.03G140700 (Glycine max); Ca_07275 (Cicer arietinum); Sobic.006G116200 (Sorghum bicolor); LOC Os04g39650 (Oryza sativa); ZmPHB47.10G170400 (Zea mays); TraesCS2D03G0763100 (Triticum aestivum)
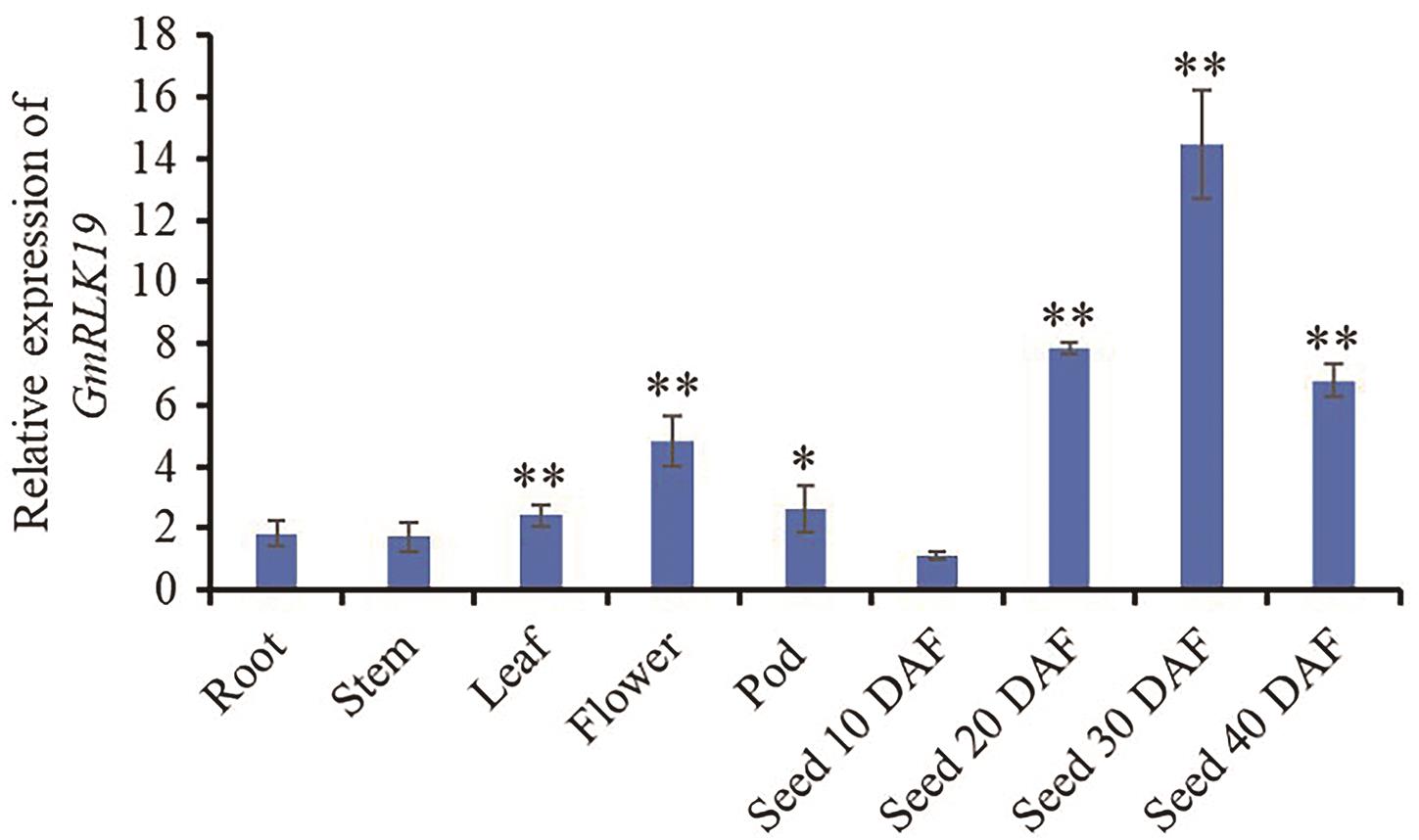
Fig. 3 Relative expressions of GmRLK19 gene in different soybean tissuesThe tissue samples were selected from the root, stem, leaf and 2 cm pod of the PI595843 of 14-day old seedlings, and the seeds at different development stages at 10-40 d after flowering were also collected. The relative expressions were normalized in the seed at 10DAF from PI595843, as the control (relative expression = 1). The error bars indicate the standard error (n = 3). Seed 10 DAF-Seed 40 DAF: the seed of the 10, 20, 30, and 40 d after flowering, respectively. * and ** indicate significant difference between tissue samples and seed at 10 DAF at 0.01 and 0.05 level (Student’s t-test, two-tail), respectively
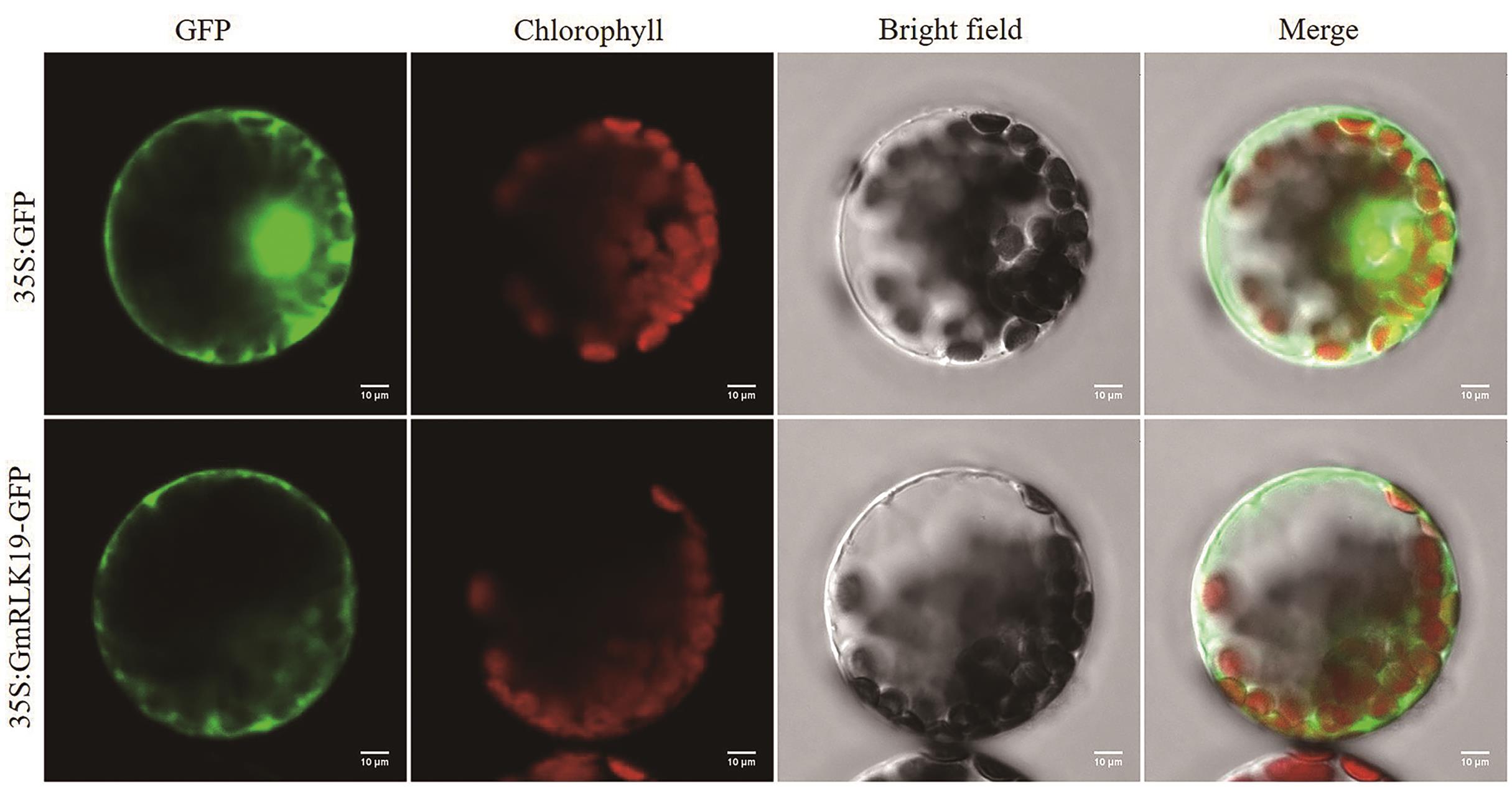
Fig. 4 Subcellular localization of GmRLK19 proteinTransient expression of GFP protein or GmRLK19-GFP fusion protein under the control of CaMV 35S promoter in tobacco cells. GFP, green fluorescence protein. Chlorophyll, chloroplast spontaneous fluorescence. Bright field, brighten field. Merged, merged images of the GFP, chlorophyll, and bright field images. Scale bar = 10 μm
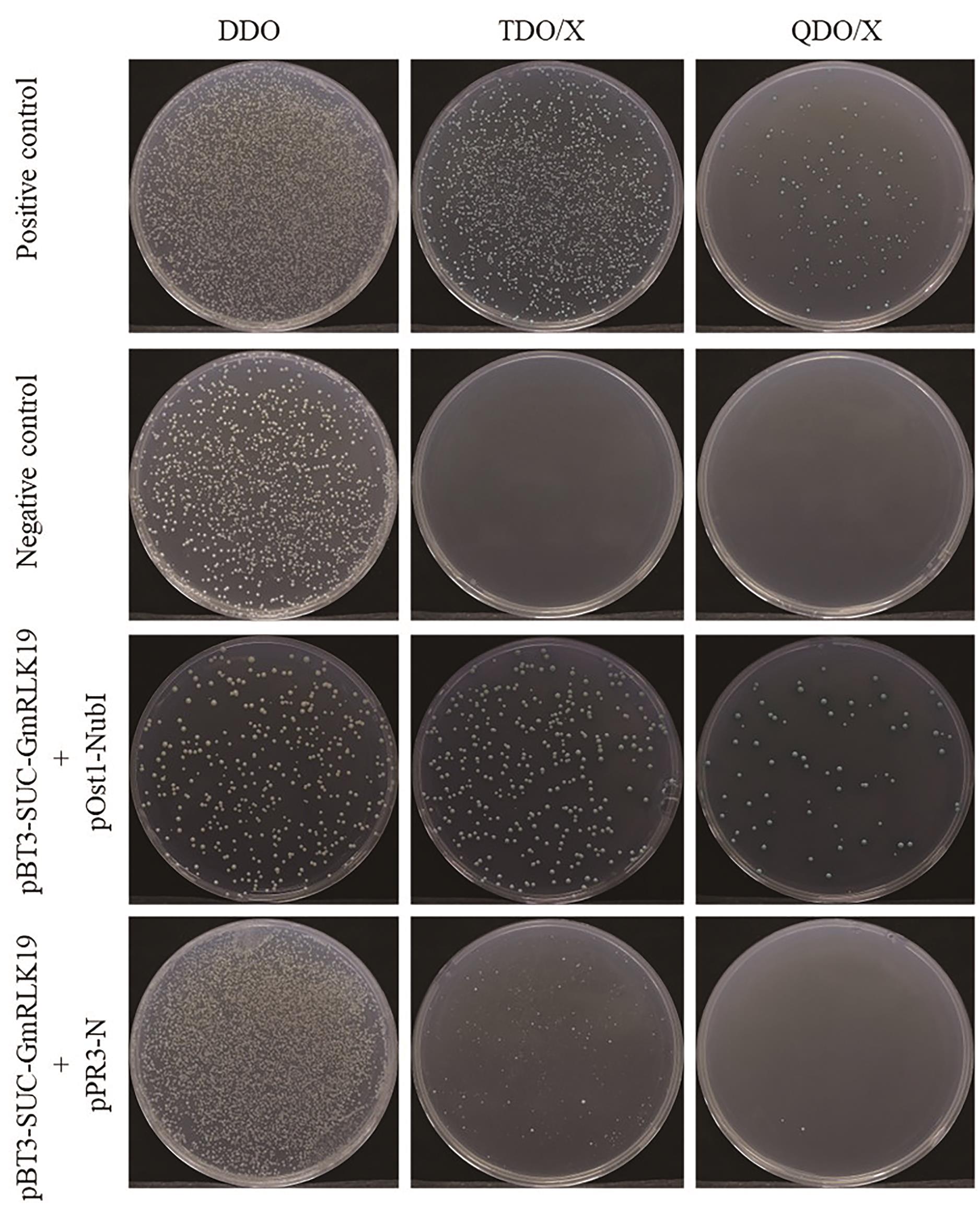

Fig. 5 Results of self-activation detectionFrom top to bottom, the groups are: positive control, negative control, functional validation group, and self-activation group. From left to right, the plates are: DDO, TDO/X, and QDO/X plate
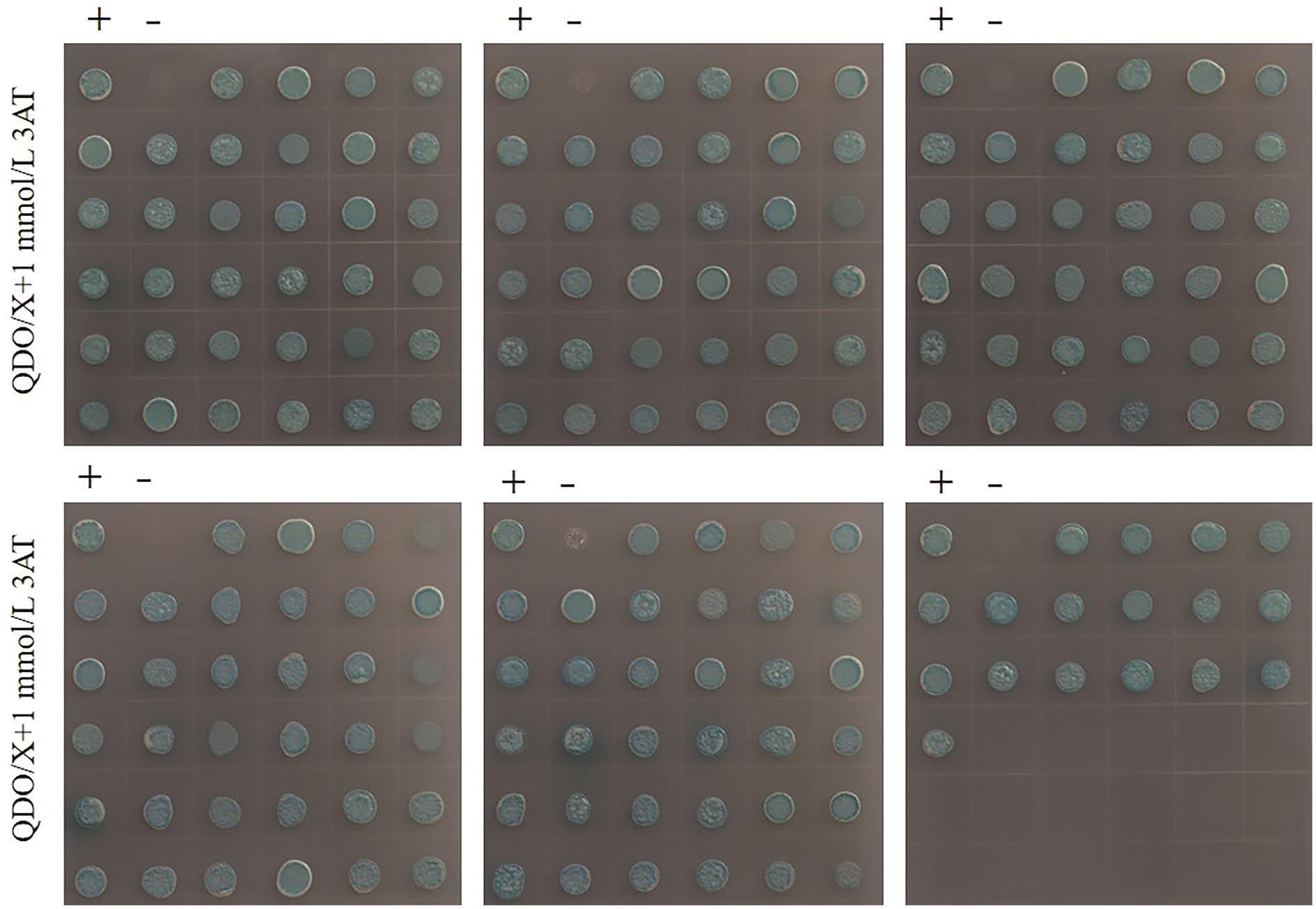

Fig. 6 The 187 positive clones obtained double hybridization library screening“+” indicates positive control, pTSU2-APP+NubG-Fe65; “-” indicates negative control, pTSU2-APP+pPR3-N
| [1] | Kato S, Sayama T, Fujii K, et al. A major and stable QTL associated with seed weight in soybean across multiple environments and genetic backgrounds [J]. Theor Appl Genet, 2014, 127(6): 1365-1374. |
| [2] | Gao CX. Genome engineering for crop improvement and future agriculture [J]. Cell, 2021, 184(6): 1621-1635. |
| [3] | Li J, Zhang YH, Ma RR, et al. Identification of ST1 reveals a selection involving hitchhiking of seed morphology and oil content during soybean domestication [J]. Plant Biotechnol J, 2022, 20(6): 1110-1121. |
| [4] | Duan ZB, Zhang M, Zhang ZF, et al. Natural allelic variation of GmST05 controlling seed size and quality in soybean [J]. Plant Biotechnol J, 2022, 20(9): 1807-1818. |
| [5] | Liang S, Duan ZB, He XM, et al. Natural variation in GmSW17 controls seed size in soybean [J]. Nat Commun, 2024, 15: 7417. |
| [6] | Goettel W, Zhang HY, Li Y, et al. POWR1 is a domestication gene pleiotropically regulating seed quality and yield in soybean [J]. Nat Commun, 2022, 13: 3051. |
| [7] | Di Q, Dong LD, Jiang L, et al. Genome-wide association study and RNA-seq identifies GmWRI1-like transcription factor related to the seed weight in soybean [J]. Front Plant Sci, 2023, 14: 1268511. |
| [8] | Wang SD, Yokosho K, Guo RZ, et al. The soybean sugar transporter GmSWEET15 mediates sucrose export from endosperm to early embryo [J]. Plant Physiol, 2019, 180(4): 2133-2141. |
| [9] | Wang SD, Liu SL, Wang J, et al. Simultaneous changes in seed size, oil content and protein content driven by selection of SWEET homologues during soybean domestication [J]. Natl Sci Rev, 2020, 7(11): 1776-1786. |
| [10] | Hu Y, Liu Y, Tao JJ, et al. GmJAZ3 interacts with GmRR18a and GmMYC2a to regulate seed traits in soybean [J]. J Integr Plant Biol, 2023, 65(8): 1983-2000. |
| [11] | Hu Y, Liu Y, Lu L, et al. Global analysis of seed transcriptomes reveals a novel PLATZ regulator for seed size and weight control in soybean [J]. New Phytol, 2023, 240(6): 2436-2454. |
| [12] | Soltabayeva A, Dauletova N, Serik S, et al. Receptor-like kinases (LRR-RLKs) in response of plants to biotic and abiotic stresses [J]. Plants, 2022, 11(19): 2660. |
| [13] | Morinaka Y, Sakamoto T, Inukai Y, et al. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice [J]. Plant Physiol, 2006, 141(3): 924-931. |
| [14] | Jiang YH, Bao L, Jeong SY, et al. XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice [J]. Plant J, 2012, 70(3): 398-408. |
| [15] | Garcia D, Saingery V, Chambrier P, et al. Arabidopsis haiku mutants reveal new controls of seed size by endosperm [J]. Plant Physiol, 2003, 131(4): 1661-1670. |
| [16] | Luo M, Dennis ES, Berger F, et al. MINISEED3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR)KINASE gene, are regulators of seed size in Arabidopsis [J]. Proc Natl Acad Sci USA, 2005, 102(48): 17531-17536. |
| [17] | He CM, Wang J, Dong R, et al. Overexpression of an antisense RNA of maize receptor-like kinase gene ZmRLK7 enlarges the organ and seed size of transgenic Arabidopsis plants [J]. Front Plant Sci, 2020, 11: 579120. |
| [18] | Zhou FL, Guo Y, Qiu LJ. Genome-wide identification and evolutionary analysis of leucine-rich repeat receptor-like protein kinase genes in soybean [J]. BMC Plant Biol, 2016, 16(1): 58. |
| [19] | Kim S, Kim SJ, Shin YJ, et al. An atypical soybean leucine-rich repeat receptor-like kinase, GmLRK1, may be involved in the regulation of cell elongation [J]. Planta, 2009, 229(4): 811-821. |
| [20] | Li XP, Gan R, Li PL, et al. Identification and functional characterization of a leucine-rich repeat receptor-like kinase gene that is involved in regulation of soybean leaf senescence [J]. Plant Mol Biol, 2006, 61(6): 829-844. |
| [21] | Yang L, Wu KC, Gao P, et al. GsLRPK, a novel cold-activated leucine-rich repeat receptor-like protein kinase from Glycine soja, is a positive regulator to cold stress tolerance [J]. Plant Sci, 2014, 215/216: 19-28. |
| [22] | Xu MG, Kong KK, Miao L, et al. Identification of major quantitative trait loci and candidate genes for seed weight in soybean [J]. Theor Appl Genet, 2023, 136(1): 22. |
| [23] | Larkin MA, Blackshields G, Brown NP, et al. Clustal W and clustal X version 2.0 [J]. Bioinformatics, 2007, 23(21): 2947-2948. |
| [24] | Tamura K, Stecher G, Peterson D, et al. MEGA6: molecular evolutionary genetics analysis version 6.0 [J]. Mol Biol Evol, 2013, 30(12): 2725-2729. |
| [25] | Yoo SD, Cho YH, Sheen J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis [J]. Nat Protoc, 2007, 2(7): 1565-1572. |
| [26] | Dievart A, Gottin C, Périn C, et al. Origin and diversity of plant receptor-like kinases [J]. Annu Rev Plant Biol, 2020, 71: 131-156. |
| [27] | Gampala SS, Kim TW, He JX, et al. An essential role for 14-3-3 proteins in brassinosteroid signal transduction in Arabidopsis [J]. Dev Cell, 2007, 13(2): 177-189. |
| [28] | Parry LIC G. Complex regulation of the TIR1/AFB family of auxin receptors [J]. Proc Natl Acad Sci U S A, 2009, 106(52): 22540-22545. |
| [29] | Qu J, Kang SG, Hah C, et al. Molecular and cellular characterization of GA-Stimulated Transcripts GASA4 and GASA6 in Arabidopsis thaliana [J]. Plant Sci, 2016, 246: 1-10. |
| [30] | Li N, Xu R, Li YH. Molecular networks of seed size control in plants [J]. Annu Rev Plant Biol, 2019, 70: 435-463. |
| [31] | Lu X, Xiong Q, Cheng T, et al. A PP2C-1 allele underlying a quantitative trait locus enhances soybean 100-seed weight [J]. Mol Plant, 2017, 10(5): 670-684. |
| [32] | Jia ML, Li YN, Wang ZY, et al. TaIAA21 represses TaARF25-mediated expression of TaERFs required for grain size and weight development in wheat [J]. Plant J, 2021, 108(6): 1754-1767. |
| [33] | Wang L, Yang YM, Yang ZY, et al. GmFtsH25 overexpression increases soybean seed yield by enhancing photosynthesis and photosynthates [J]. J Integr Plant Biol, 2023, 65(4): 1026-1040. |
| [34] | Hu DZ, Li X, Yang ZY, et al. Downregulation of a gibberellin 3β- hydroxylase enhances photosynthesis and increases seed yield in soybean [J]. New Phytol, 2022, 235(2): 502-517. |
| [1] | SU Yan-zhu, LI Da, ZHANG Ai-ai, LIU Yong-guang, ZHANG Xiu-rong, XUE Qi-qin. Identification and Expression Analysis of CAD Gene Family in Soybean(Glycine max (L.) Merr.) [J]. Biotechnology Bulletin, 2026, 42(4): 101-113. |
| [2] | LIU Qing-yuan, WU Hong-qi, CHEN Xiu-e, CHEN Jian, JIANG Yuan-ze, HE Yan-zi, YU Qi-wei, LIU Ren-xiang. Function of Transcription Factor NtMYB96a in Regulating the Tolerance of Tobacco to Drought [J]. Biotechnology Bulletin, 2026, 42(4): 239-250. |
| [3] | CHEN Deng-ke, LAN Gang, XIA Zhi, HOU Bao-guo, YANG Liu-liu, CAO Cai-rong, LI Peng-bo, WU Cui-cui. Identification of ZF-HD Gene Family in Arachis hypogaea and Analysis in Response to Abiotic Stress [J]. Biotechnology Bulletin, 2026, 42(4): 114-128. |
| [4] | JIANG Xin-hua, FANG Tian-yu, ZHANG Jing-jing, LI Xiang-yuan, ZHANG Bang-yue, LIAO Xiao-shan, RONG Duo-yan. Identification and Functional Analysis of the MpPP2A-C Gene in Marchantia polymorpha [J]. Biotechnology Bulletin, 2026, 42(4): 216-226. |
| [5] | ZHANG Gao-xiang, WU Yu-bi, GUO Ya-jing, JI Wei, YANG Zhong-yi. Identification and Expression Analysis of WD40 Gene Family in Grape [J]. Biotechnology Bulletin, 2026, 42(3): 324-337. |
| [6] | LIU Lin-ya, LIU Huan-yan, LIANG Xin-yu, SONG Shu-yi, HE Bin, WANG Xu-ying, HUANG Ya-cheng. Genome-wide Identification and Expression Analysis of BGAL Gene Family in Actinidiachinensis var. Hongyang [J]. Biotechnology Bulletin, 2026, 42(3): 312-323. |
| [7] | YANG Zi-han, LI Kui-xiu, LI Jun-liang, LYU Wen-hui, HE Wen-ting, LIU Xu-yan, LIU Guan-ze. Identification and Expression Analysis of the RALF Gene Family in Panax notoginseng [J]. Biotechnology Bulletin, 2026, 42(2): 278-292. |
| [8] | REN Yun-er, WU Guo-qiang, CHENG Bin, WEI Ming. Genome-wide Identification of the BvATGs Genes Family in Sugar Beet (Beta vulgaris L.) and Analysis of Their Expression Pattern under Salt Stress [J]. Biotechnology Bulletin, 2026, 42(1): 184-197. |
| [9] | YANG Yue-qin, XING Ying, ZHONG Zi-he, TIAN Wei-jun, YANG Xue-qing, WANG Jian-xu. Expression and Functional Analysis of OsMATE34 in Rice under Mercury Stress [J]. Biotechnology Bulletin, 2026, 42(1): 86-94. |
| [10] | ZHANG Yue, DAI Yue-hua, ZHANG Ying-ying, LI Ao-hui, LI Chu-hui, XUE Jin-ai, QIN Hui-bin, CHEN Yan, NIE Meng-en, ZHANG Hai-ping. Cloning and Functional Analysis of the Soybean Enoyl-CoA Reductase ECR14 Gene [J]. Biotechnology Bulletin, 2026, 42(1): 95-104. |
| [11] | CHEN Jing-huan, FANG Guo-nan, ZHU Wen-hao, YE Guang-ji, SU Wang, HE Miao-miao, YANG Sheng-long, ZHOU Yun. Starch Characterization and Related Gene Expression Analysis of Potato Germplasm Resources [J]. Biotechnology Bulletin, 2026, 42(1): 170-183. |
| [12] | LI Ya-tao, ZHANG Zhi-peng, ZHAO Meng-yao, LYU Zhen, GAN Tian, WEI Hao, WU Shu-feng, MA Yu-chao. Whole Genome Analysis of Bradyrhizobium sp. Bd1 and the Negative Regulating Function of TetR3 during Cell Growth and Nodulation [J]. Biotechnology Bulletin, 2025, 41(9): 289-301. |
| [13] | LI Shan, MA Deng-hui, MA Hong-yi, YAO Wen-kong, YIN Xiao. Identification and Expression Analysis of SKP1 Gene Family in Grapevine (Vitis vinifera L.) [J]. Biotechnology Bulletin, 2025, 41(9): 147-158. |
| [14] | HUANG Guo-dong, DENG Yu-xing, CHENG Hong-wei, DAN Yan-nan, ZHOU Hui-wen, WU Lan-hua. Genome-wide Identification and Expression Analysis of the ZIP Gene Family in Soybean [J]. Biotechnology Bulletin, 2025, 41(9): 71-81. |
| [15] | GONG Hui-ling, XING Yu-jie, MA Jun-xian, CAI Xia, FENG Zai-ping. Identification of Laccase (LAC) Gene Family in Potato (Solanum tuberosum L.) and Its Expression Analysis under Salt Stresses [J]. Biotechnology Bulletin, 2025, 41(9): 82-93. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||