








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 170-181.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0873
Previous Articles Next Articles
GUO Dao-xiang( ), SU Quan, LI Ao-ting, WANG Yan, CHEN Qing, WANG Xiao-rong, HE Wen(
), SU Quan, LI Ao-ting, WANG Yan, CHEN Qing, WANG Xiao-rong, HE Wen( )
)
Received:2025-08-11
Online:2026-04-26
Published:2026-04-30
Contact:
HE Wen
E-mail:guodaoxiang@stu.sicau.edu.cn;hewen0724@sicau.edu.cn
GUO Dao-xiang, SU Quan, LI Ao-ting, WANG Yan, CHEN Qing, WANG Xiao-rong, HE Wen. Identification of the MYB Gene Family in Pomelo (Citrus grandis) and Functional Analysis in Chlorosis of Graft Incompatibility[J]. Biotechnology Bulletin, 2026, 42(4): 170-181.
基因 Gene | 正向引物序列 Forward primer sequence (5′‒3′) | 反向引物序列 Reverse primer sequence (5′‒3′) | 退火温度 Annealing temperature (℃) | 产物长度 Primer length (bp) |
|---|---|---|---|---|
| β-Tubulin | ACATCCCGCCTAAGGGTCTG | TTCCTCCGAAACATAGCCGTA | 58 | 150 |
| q-CgAPL8 | ATTGGAGCCCTTCTCTCAGC | AACTGCTCTGTCTCAGCCAG | 58 | 150 |
| q-CgISA2 | CCATTACTACACGGCTCTTCT | GCGATTGAGGCTTCTTGATAA | 58 | 150 |
| q-CgISA3 | GGCATTGGGTGACTGAGTTT | CCCATCTCTTCCAATGAGGA | 58 | 150 |
| q-CgMYB53 | GAGGCACATTCCCAAAGCTG | TGTTCGTCCTGGAAGTCTCC | 58 | 150 |
| q-Cg2g041650 | TGGGAGCTCCCAAGTAAGGC | GGCTTATCAGTCCCCGTTGC | 58 | 150 |
| q-CgGBSS2 | TAGCCGCTTCAAGCTTTGTCT | CAATCATTAAGGATGGCCCGTTCT | 58 | 150 |
| q-MYB45 | TCATGGTGAGGGCAATTGGG | CAATGACCATCGGTTGCCAA | 58 | 150 |
| q-MYB75 | ATTCCCAGCAGAGGCACTAG | CCACGCTCTCAAACACAACG | 58 | 150 |
| q-MYB40 | ACTCGAAGAGCGACCCATAA | TATCGTTCGACAAGCCTGGT | 58 | 150 |
| q-MYB113 | AGAGGACCAGCTATTGCTGA | ATGATCTCGTCTTCGTCGGG | 58 | 150 |
| q-MYB136 | AAGCACAGAAAGGGCTTGTG | AATCCATCTCAGTCTGCAGC | 58 | 150 |
| q-MYB129 | CGCCCACATCAAAAAACACG | ATCCATCGGAGTCTGCAACT | 58 | 150 |
| q-MYB2 | TCGAGTTGGACAGCCAAGAA | CAACGAGCATCTCATACCGC | 58 | 150 |
| q-MYB89 | GAATCAAGTTCTGGCCTGCA | TTGGCTTGGGAGCAAAATCG | 58 | 150 |
| q-MYB14 | CTTCGAAAGGGCTTGTGGTC | ACAGCTCTTGCCACATCTCT | 58 | 150 |
| q-CgBE3 | GACCGTCAACTCCCCTCATA | GCTGATCAACCCTTGGAAAA | 58 | 150 |
| q-CgBE1 | ACTTCGCTTCCTTCTGTCCA | CCAGAAACATCTTCGGCAAT | 58 | 150 |
| q-CgAPL3 | TCTTAAGAGCGGGGACTTGA | CCACATTTTTGGGATCTGC | 58 | 150 |
| q-CgBAM6 | GCTGTTGCAGAGATGGTTGA | GAAGATCATCAACGGCCAAT | 58 | 150 |
| q-CgBAM4 | TCTCCGGGCTAAGAGTTCAA | CCAGTGGCAACATCACAAAC | 58 | 150 |
| CgMYB53 | ATGGCGGGTAAGCGCAAGA | CAAAATCCCAAATCCGAT | 60 | 753 |
| OE-CgMYB53 | GAACACGGGGGACGAGCTCGGTACCATGGCGGGTAAGCGCAAGA | GTGGTGGTGGTCGACGGATCCTCACCAAAATCCCAAATCCGAT | 62 | 799 |
Table 1 Primer sequences for RT-qPCR, gene cloning, and plasmid construction
基因 Gene | 正向引物序列 Forward primer sequence (5′‒3′) | 反向引物序列 Reverse primer sequence (5′‒3′) | 退火温度 Annealing temperature (℃) | 产物长度 Primer length (bp) |
|---|---|---|---|---|
| β-Tubulin | ACATCCCGCCTAAGGGTCTG | TTCCTCCGAAACATAGCCGTA | 58 | 150 |
| q-CgAPL8 | ATTGGAGCCCTTCTCTCAGC | AACTGCTCTGTCTCAGCCAG | 58 | 150 |
| q-CgISA2 | CCATTACTACACGGCTCTTCT | GCGATTGAGGCTTCTTGATAA | 58 | 150 |
| q-CgISA3 | GGCATTGGGTGACTGAGTTT | CCCATCTCTTCCAATGAGGA | 58 | 150 |
| q-CgMYB53 | GAGGCACATTCCCAAAGCTG | TGTTCGTCCTGGAAGTCTCC | 58 | 150 |
| q-Cg2g041650 | TGGGAGCTCCCAAGTAAGGC | GGCTTATCAGTCCCCGTTGC | 58 | 150 |
| q-CgGBSS2 | TAGCCGCTTCAAGCTTTGTCT | CAATCATTAAGGATGGCCCGTTCT | 58 | 150 |
| q-MYB45 | TCATGGTGAGGGCAATTGGG | CAATGACCATCGGTTGCCAA | 58 | 150 |
| q-MYB75 | ATTCCCAGCAGAGGCACTAG | CCACGCTCTCAAACACAACG | 58 | 150 |
| q-MYB40 | ACTCGAAGAGCGACCCATAA | TATCGTTCGACAAGCCTGGT | 58 | 150 |
| q-MYB113 | AGAGGACCAGCTATTGCTGA | ATGATCTCGTCTTCGTCGGG | 58 | 150 |
| q-MYB136 | AAGCACAGAAAGGGCTTGTG | AATCCATCTCAGTCTGCAGC | 58 | 150 |
| q-MYB129 | CGCCCACATCAAAAAACACG | ATCCATCGGAGTCTGCAACT | 58 | 150 |
| q-MYB2 | TCGAGTTGGACAGCCAAGAA | CAACGAGCATCTCATACCGC | 58 | 150 |
| q-MYB89 | GAATCAAGTTCTGGCCTGCA | TTGGCTTGGGAGCAAAATCG | 58 | 150 |
| q-MYB14 | CTTCGAAAGGGCTTGTGGTC | ACAGCTCTTGCCACATCTCT | 58 | 150 |
| q-CgBE3 | GACCGTCAACTCCCCTCATA | GCTGATCAACCCTTGGAAAA | 58 | 150 |
| q-CgBE1 | ACTTCGCTTCCTTCTGTCCA | CCAGAAACATCTTCGGCAAT | 58 | 150 |
| q-CgAPL3 | TCTTAAGAGCGGGGACTTGA | CCACATTTTTGGGATCTGC | 58 | 150 |
| q-CgBAM6 | GCTGTTGCAGAGATGGTTGA | GAAGATCATCAACGGCCAAT | 58 | 150 |
| q-CgBAM4 | TCTCCGGGCTAAGAGTTCAA | CCAGTGGCAACATCACAAAC | 58 | 150 |
| CgMYB53 | ATGGCGGGTAAGCGCAAGA | CAAAATCCCAAATCCGAT | 60 | 753 |
| OE-CgMYB53 | GAACACGGGGGACGAGCTCGGTACCATGGCGGGTAAGCGCAAGA | GTGGTGGTGGTCGACGGATCCTCACCAAAATCCCAAATCCGAT | 62 | 799 |
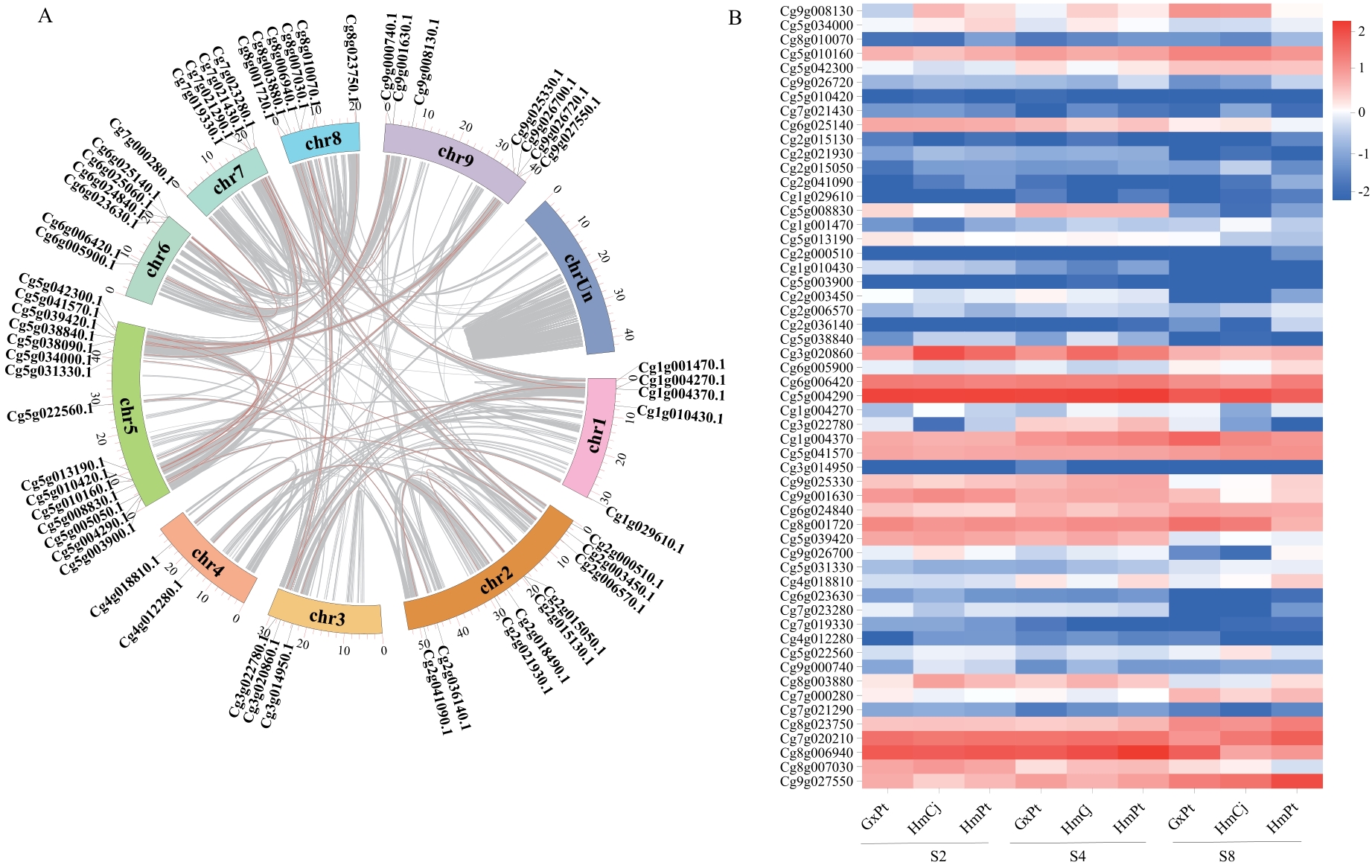
Fig. 2 Co-linearity analysis of CgMYBs genesA: Co-linearity analysis of CgMYBs genes. B: Expression heatmap of genes involved in co-linearity CgMYBs in different graft combinations. Compatible graft combinations: GxPt (‘Guanxi Miyou’ grafted onto trifoliate orange) and HmCj (‘Hongmian Miyou’ grafted onto ‘Shuzhen No.1’). Incompatible graft combination: HmPt (‘Hongmian Miyou’ grafted onto trifoliate orange). S2: Relative chlorophyll content [Soil and Plant Analyzer Development (SPAD) value] between 70-80; S4: SPAD value between 50-60; S8: SPAD value between 10-20. The same below
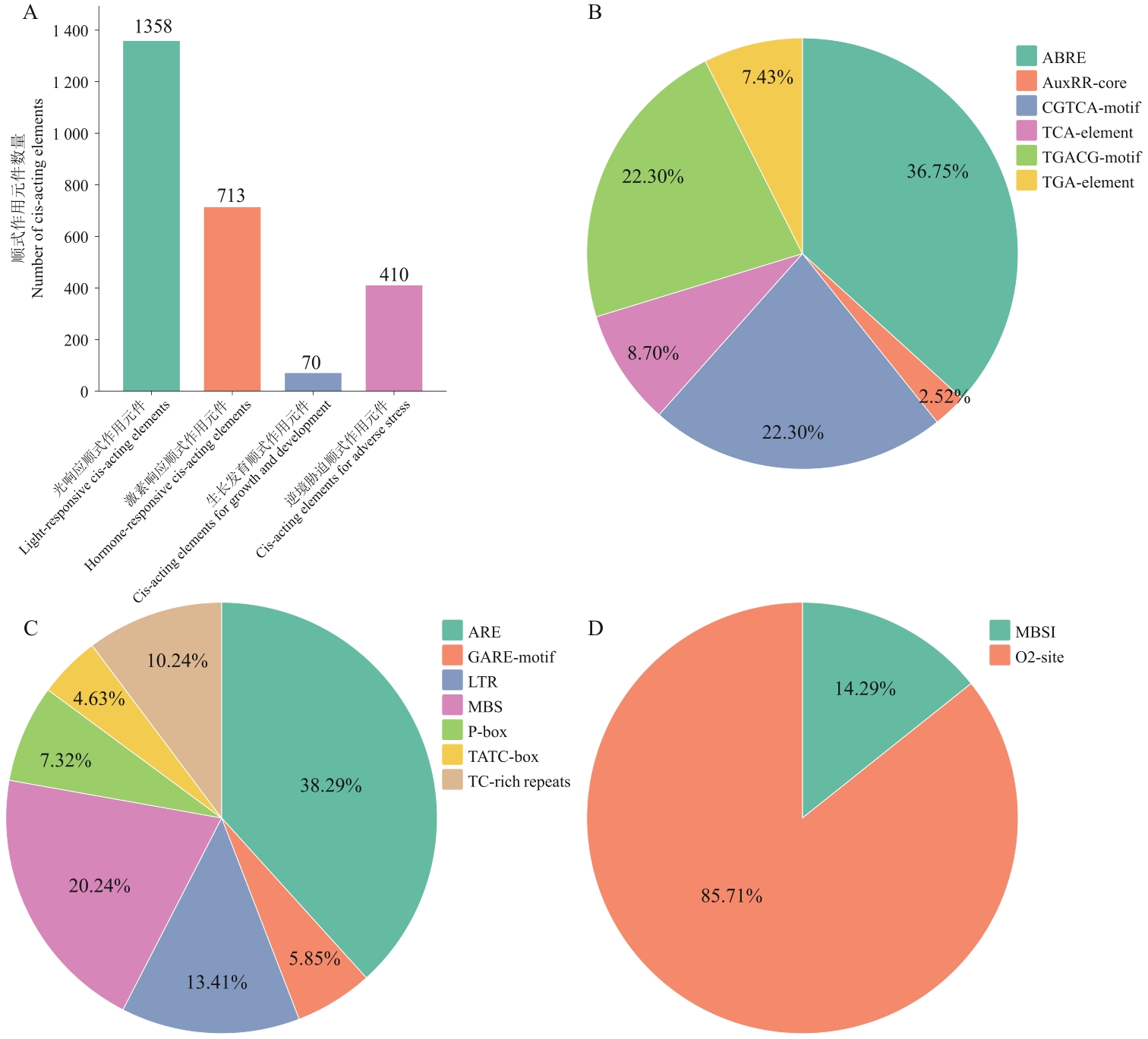
Fig. 3 Assay result of promoter elements of the R2R3-CgMYBs genesA: Numbers of various promoter elements of the R2R3-CgMYBs. B: Promotor elements in hormone responsive elements of the R2R3-CgMYBs. C: Promotor elements in stress responsive elements of the R2R3-CgMYBs. D: Promotor elements in plant growth-related responsive elements of the R2R3-CgMYBs
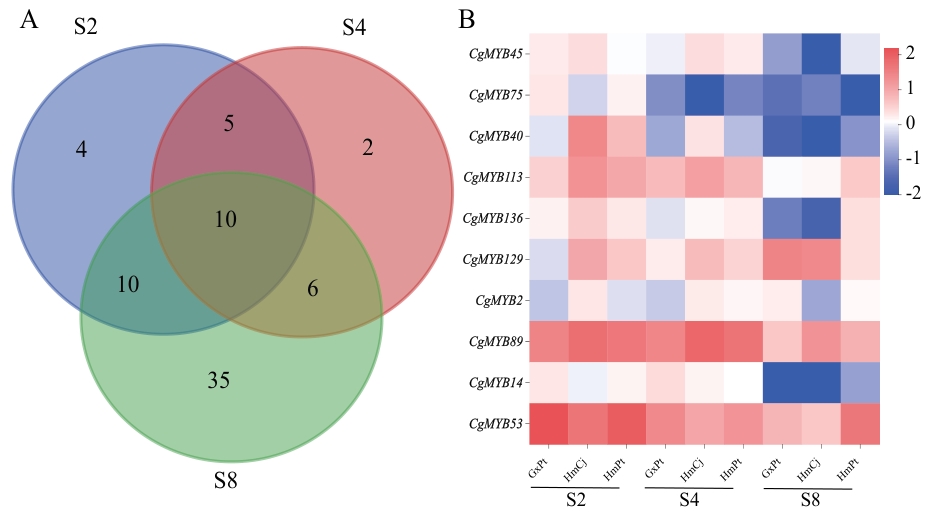
Fig. 6 Expression profile analysis of genes involved in CgMYBs in different graft combinationsA: Venn analysis of MYB genes in chlorotic phase of HmPt (incompatible graft). B: Expression heatmap of the ten CgMYBs in different graft combinations [Log10(FPKM+0.01)]
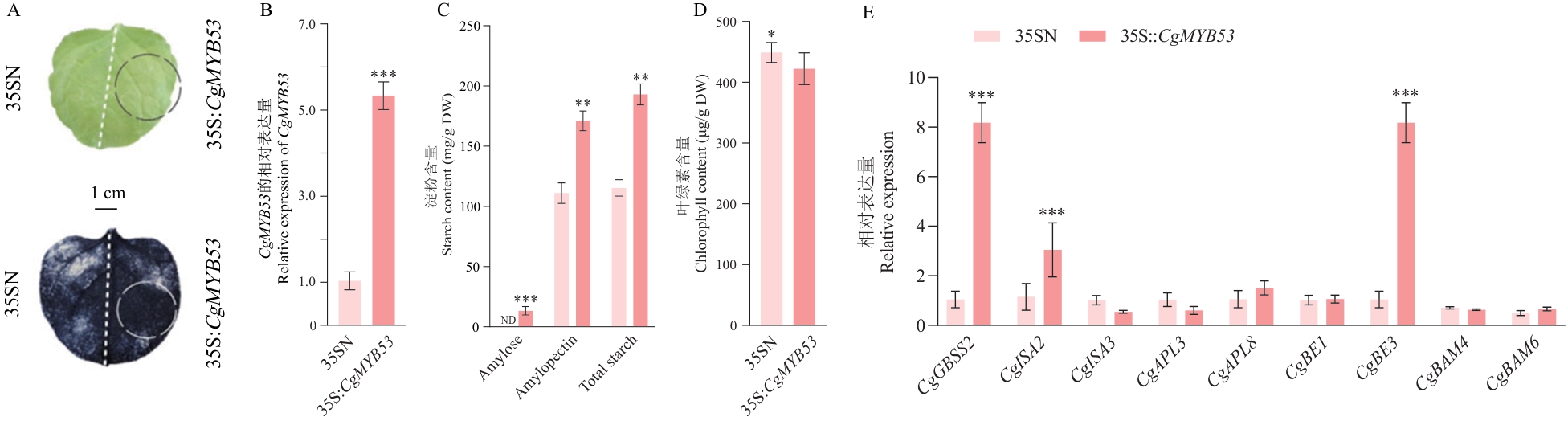
Fig. 8 Transient overexpressions of CgMYB53 in tobacco leavesA: Phenotype and iodine-potassium iodide staining of CgMYB53-transient overexpression tobacco leaves. B: Relative expressions of CgMYB53. C: Starch content in tobacco leaves. D: Chlorophyll and carotenoid content in tobacco leaves (ND stands for not detected). E: Relative expressions of starch metabolism-related genes. * P<0.05; ** P<0.01; *** P<0.001
| [1] | 邓秀新. 中国柑橘育种60年回顾与展望 [J]. 园艺学报, 2022, 49(10): 2063-2074. |
| Deng XX. A review and perspective for citrus breeding in China during the last six decades [J]. Acta Hortic Sin, 2022, 49(10): 2063-2074. | |
| [2] | 朱世平, 陈传武, 刘兆俊, 等. 卡里佐枳橙在中国的生产适应性调查与比较研究 [J]. 中国南方果树, 2020, 49(3): 1-8. |
| Zhu SP, Chen CW, Liu ZJ, et al. Investigation and comparison of the performance of carrizo citrange in China [J]. South China Fruits, 2020, 49(3): 1-8. | |
| [3] | He W, Xie R, Wang Y, et al. Comparative transcriptomic analysis on compatible/incompatible grafts in citrus [J]. Hortic Res, 2022, 9: uhab072. |
| [4] | He W, Wang Y, Chen Q, et al. Dissection of the mechanism for compatible and incompatible graft combinations of Citrus grandis (L.) Osbeck (‘Hongmian Miyou’) [J]. Int J Mol Sci, 2018, 19(2): 505. |
| [5] | He W, Xie R, Guo DX, et al. The starch excess and key genes underlying citrus leaf chlorosis by rootstock-scion incompatibility [J]. Int J Biol Macromol, 2024, 282(Pt 4): 137111. |
| [6] | He W, Xie R, Luo L, et al. Comparative transcriptomic analysis of inarching invigorating rootstock onto incompatible grafts in citrus [J]. Int J Mol Sci, 2022, 23(23): 14523. |
| [7] | Zhang LC, Zhao GY, Jia JZ, et al. Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress [J]. J Exp Bot, 2012, 63(1): 203-214. |
| [8] | Karpinska B, Karlsson M, Srivastava M, et al. MYB transcription factors are differentially expressed and regulated during secondary vascular tissue development in hybrid aspen [J]. Plant Mol Biol, 2004, 56(2): 255-270. |
| [9] | Lim PO, Kim HJ, Nam HG. Leaf senescence [J]. Annu Rev Plant Biol, 2007, 58: 115-136. |
| [10] | Guo YF, Ren GD, Zhang KW, et al. Leaf senescence: progression, regulation, and application [J]. Mol Hortic, 2021, 1(1): 5. |
| [11] | Zhang YM, Guo PR, Xia XL, et al. Multiple layers of regulation on leaf senescence: new advances and perspectives [J]. Front Plant Sci, 2021, 12: 788996. |
| [12] | Yongfeng Guo SG. AtMYB2 regulates whole plant senescence by inhibiting cytokinin-mediated branching at late stages of development in Arabidopsis [J]. Plant Physiol, 2011, 156(3): 1612-1619. |
| [13] | Zhang X, Ju HW, Chung MS, et al. The R-R-type MYB-like transcription factor, AtMYBL, is involved in promoting leaf senescence and modulates an abiotic stress response in Arabidopsis [J]. Plant Cell Physiol, 2011, 52(1): 138-148. |
| [14] | Buelbuel S, Sakuraba Y, Sedaghatmehr M, et al. Arabidopsis BBX14 negatively regulates nitrogen starvation- and dark-induced leaf senescence [J]. Plant J, 2023, 116(1): 251-268. |
| [15] | Wang WB, He XF, Yan XM, et al. Chromosome-scale genome assembly and insights into the metabolome and gene regulation of leaf color transition in an important oak species, Quercus dentata [J]. New Phytol, 2023, 238(5): 2016-2032. |
| [16] | Cao J, Yang Q, Zhao YN, et al. MYB47 delays leaf senescence by modulating jasmonate pathway via direct regulation of CYP94B3/CYP94C1 expression in Arabidopsis [J]. New Phytol, 2025, 246(5): 2192-2206. |
| [17] | He SC, Zhi F, Min YC, et al. The MYB59 transcription factor negatively regulates salicylic acid- and jasmonic acid-mediated leaf senescence [J]. Plant Physiol, 2023, 192(1): 488-503. |
| [18] | Yang JW, Huang JZ, Wu X, et al. NtMYB1 and NtNCED1/2 control abscisic acid biosynthesis and tepal senescence in Chinese Narcissus (Narcissus tazetta) [J]. J Exp Bot, 2023, 74(21): 6505-6521. |
| [19] | Zhu ZF, Lu LL, Chen SY, et al. Transcription factor CitMYB16 positively regulates cutin and wax biosynthesis in citrus by directly activating CitDCR and CitKCS2 [J]. Plant J, 2025, 123(3): e70324. |
| [20] | Liu CY, Long JM, Zhu KJ, et al. Characterization of a citrus R2R3-MYB transcription factor that regulates the flavonol and hydroxycinnamic acid biosynthesis [J]. Sci Rep, 2016, 6: 25352. |
| [21] | Wang TT, Song X, Zhang M, et al. CsCPC, an R3-MYB transcription factor, acts as a negative regulator of citric acid accumulation in Citrus [J]. Plant J, 2025, 121(1): e17189. |
| [22] | Liu H, Wang X, Liu SJ, et al. Citrus Pan-Genome to Breeding Database (CPBD): a comprehensive genome database for citrus breeding [J]. Mol Plant, 2022, 15(10): 1503-1505. |
| [23] | Mistry J, Chuguransky S, Williams L, et al. Pfam: The protein families database in 2021 [J]. Nucleic Acids Res, 2021, 49(D1): D412-D419. |
| [24] | Lescot M, Déhais P, Thijs G, et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences [J]. Nucleic Acids Res, 2002, 30(1): 325-327. |
| [25] | Kumar S, Stecher G, Li M, et al. MEGA X: Molecular evolutionary genetics analysis across computing platforms [J]. Mol Biol Evol, 2018, 35(6): 1547-1549. |
| [26] | 唐云, 闫海彦, 赵亚雄, 等. 碘比色法测定高粱中直链淀粉和支链淀粉的方法探讨 [J]. 食品工业科技, 2023, 44(13): 272-280. |
| Tang Y, Yan HY, Zhao YX, et al. Determination of amylose and amylopectin in sorghum by iodine colorimetric method [J]. Sci Technol Food Ind, 2023, 44(13): 272-280. | |
| [27] | Wellburn AR, Lichtenthaler H. Formulae and program to determine total carotenoids and chlorophylls a and b of leaf extracts in different solvents [M]//Advances in Photosynthesis Research. Dordrecht: Springer Netherlands, 1984: 9-12. |
| [28] | Dubos C, Stracke R, Grotewold E, et al. MYB transcription factors in Arabidopsis [J]. Trends Plant Sci, 2010, 15(10): 573-581. |
| [29] | Yang JH, Zhang BH, Gu G, et al. Genome-wide identification and expression analysis of the R2R3-MYB gene family in tobacco (Nicotiana tabacum L.) [J]. BMC Genomics, 2022, 23(1): 432. |
| [30] | Chen YH, Yang XY, He K, et al. The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family [J]. Plant Mol Biol, 2006, 60(1): 107-124. |
| [31] | Ding Y, Yang QH, Waheed A, et al. Genome-wide characterization and functional identification of MYB genes in Malus sieversii infected by Valsa mali [J]. Front Plant Sci, 2023, 14: 1112681. |
| [32] | Li XL, Xue C, Li JM, et al. Genome-wide identification, evolution and functional divergence of MYB transcription factors in Chinese white pear (Pyrus bretschneideri) [J]. Plant Cell Physiol, 2016, 57(4): 824-847. |
| [33] | Liu H, Xiong JS, Jiang YT, et al. Evolution of the R2R3-MYB gene family in six Rosaceae species and expression in woodland strawberry [J]. J Integr Agric, 2019, 18(12): 2753-2770. |
| [34] | Hou XJ, Li SB, Liu SR, et al. Genome-wide classification and evolutionary and expression analyses of citrus MYB transcription factor families in sweet orange [J]. PLoS One, 2014, 9(11): e112375. |
| [35] | Zhang DW, Zhou HP, Zhang Y, et al. Diverse roles of MYB transcription factors in plants [J]. J Integr Plant Biol, 2025, 67(3): 539-562. |
| [36] | Frangedakis E, Yelina NE, Billakurthi K, et al. MYB-related transcription factors control chloroplast biogenesis [J]. Cell, 2024, 187(18): 4859-4876.e22. |
| [37] | Li ZT, Gmitter FG, Grosser JW, et al. Isolation and characterization of a novel anthocyanin-promoting MYBA gene family in citrus [J]. Tree Genet Genomes, 2012, 8(4): 675-685. |
| [38] | Luo TT, Zhang H, Tan HK, et al. Two MYB transcription factors interact to inhibit the expression of cell wall metabolism and starch degradation genes in banana [J]. Plant Physiol, 2025, 198(4): kiaf239. |
| [39] | Xue J, Lu DB, Wang SG, et al. Integrated transcriptomic and metabolomic analysis provides insight into the regulation of leaf senescence in rice [J]. Sci Rep, 2021, 11(1): 14083. |
| [40] | Huang JY, Yan M, Zhu XY, et al. Gene mapping of starch accumulation and premature leaf senescence in the ossac3 mutant of rice [J]. Euphytica, 2018, 214(10): 177. |
| [41] | Huang KY, Chen L, Jiao GA, et al. OsMYBR1, a 1R-MYB family transcription factor regulates starch biosynthesis in rice endosperm [J]. Life, 2025, 15(6): 962. |
| [42] | Xiao QL, Wang YY, Du J, et al. ZmMYB14 is an important transcription factor involved in the regulation of the activity of the ZmBT1 promoter in starch biosynthesis in maize [J]. FEBS J, 2017, 284(18): 3079-3099. |
| [43] | Fan ZQ, Ba LJ, Shan W, et al. A banana R2R3-MYB transcription factor MaMYB3 is involved in fruit ripening through modulation of starch degradation by repressing starch degradation-related genes and MabHLH6 [J]. Plant J, 2018, 96(6): 1191-1205. |
| [1] | ZHANG Chi-hao, LIU Jin-nan, CHAO Yue-hui. Cloning and Functional Analysis of a bZIP Transcription Factor MtbZIP29 from Medicago truncatula [J]. Biotechnology Bulletin, 2026, 42(1): 241-250. |
| [2] | WANG Sai-di, ZHANG Gao-yang, LYU Huan-huan, SUN Zhong-ke, LI Cheng-wei. Research Advances in Amylose Biosynthesis and Strategies for Enhancing Its Content in Plants [J]. Biotechnology Bulletin, 2025, 41(8): 22-33. |
| [3] | LAI Shi-yu, LIANG Qiao-lan, WEI Lie-xin, NIU Er-bo, CHEN Ying-e, ZHOU Xin, YANG Si-zheng, WANG Bo. The Role of NbJAZ3 in the Infection of Nicotiana benthamiana by Alfalfa Mosaic Virus [J]. Biotechnology Bulletin, 2025, 41(8): 186-196. |
| [4] | ZHANG Ze, YANG Xiu-li, NING Dong-xian. Identification of 4CL Gene Family in Arachis hypogaea L. and Expression Analysis in Response to Drought and Salt Stress [J]. Biotechnology Bulletin, 2025, 41(7): 117-127. |
| [5] | HU Yan-an, DAI Xin-lyu, ZHONG Jiao-yan, LI Rui-min, HUANG Gui-yan. Comparative Genomics-based Identification and Functional Characterization of Key Genes Resisting Huanglongbing in Citrus [J]. Biotechnology Bulletin, 2025, 41(11): 282-292. |
| [6] | PAN Ping-ping, XU Zhi-hao, ZHANG Yi-wen, LI Qing, WANG Zhong-hua. Prokaryotic Expression, Subcellular Localization and Expression Analysis of PcCHS Gene from Polygonatum cyrtonema Hua [J]. Biotechnology Bulletin, 2024, 40(5): 280-289. |
| [7] | YANG Yan, HU Yang, LIU Ni-ru, YIN Lu, YANG Rui, WANG Peng-fei, MU Xiao-peng, ZHANG Shuai, CHENG Chun-zhen, ZHANG Jian-cheng. Cloning and Functional Analysis of MbbZIP43 Gene in ‘Hongmantang’ Red-flesh Apple [J]. Biotechnology Bulletin, 2024, 40(2): 146-159. |
| [8] | LEI Mei-ling, RAO Wen-hua, HU Jin-feng, YUE Qi, WU Zu-jian, FAN Guo-cheng. Bacterial Diversity and Structure in Rhizosphere Soil of Citrus Infested with Huanglongbing [J]. Biotechnology Bulletin, 2024, 40(2): 266-276. |
| [9] | YE Liu-jian, HE Yu-lan, WANG Xiao-hu, WEI Sheng-bo, HE Shuang, ZHU Qi-xia, LU Jie, ZHOU Li-qin. Effects of Bacillus amyloliquefaciens YK3 on the Control of Citrus reticulata cv. Orah Canker and Its Influence on the Network of Phyllosphere Bacteria [J]. Biotechnology Bulletin, 2024, 40(11): 248-258. |
| [10] | LIU Jing-ju, ZHANG Yu-sen, CHEN Juan, SUN Bing-da, ZHAO Guo-zhu. Research Progress in Modern Taxonomy and Nomenclature of Aspergillus [J]. Biotechnology Bulletin, 2022, 38(7): 109-118. |
| [11] | ZHANG Jian-bo,JIN Yun-feng,WANG Sha-sha,YANG Hui-qin,PANG Tao,LI Jun-ying,CUI Ming-kun,GONG Ming,. Effects of Different Temperature on Starch Metabolism in Different Growth Stages of Tobacco(Nicotiana tabacum) [J]. Biotechnology Bulletin, 2016, 32(5): 200-211. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||