








生物技术通报 ›› 2024, Vol. 40 ›› Issue (11): 125-141.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0360
收稿日期:2024-04-15
出版日期:2024-11-26
发布日期:2024-12-19
通讯作者:
李辰,女,博士,研究员,研究方向:蛋白质组学;E-mail: cli@shsmu.edu.cn作者简介:顾蕾,女,硕士,实验师,研究方向:蛋白质组学;E-mail: leigu@shsmu.edu.cn张誉露、唐尚睿、于浩月同为本文第一作者
基金资助:
GU Lei( ), ZHANG Yu-lu, TANG Shang-rui, YU Hao-yue, LI Chen(
), ZHANG Yu-lu, TANG Shang-rui, YU Hao-yue, LI Chen( )
)
Received:2024-04-15
Published:2024-11-26
Online:2024-12-19
摘要:
单细胞多组学技术为生命科学和医学注入新的活力,能够从更微观的细胞尺度、更高维的时空角度、更交叉的组学视野来探索生命的未知、拓展医学的边界。单细胞蛋白质组学是从时间和空间上研究单个或单一类型细胞的蛋白质表达异质性。尽管存在通量、灵敏度、分辨率等技术难点,基于质谱的细胞尺度的时空蛋白质组学技术凭借无偏检测、鉴定种类多、定量准确等特点而备受关注。本文综述了近年来单细胞蛋白质组学的发展,从单细胞分选、样品前处理、质谱检测和数据分析四个方面介绍相关技术进展,并探讨了其在生命科学与医学等研究中的应用,展望了未来面临的挑战和发展前景。在单细胞水平上进行时空蛋白质组学研究的挑战与机遇并存,而新兴的单细胞蛋白质组学技术无疑将为相关研究领域提供新的思路方法、认知线索和多模态数据等宝贵资源。
顾蕾, 张誉露, 唐尚睿, 于浩月, 李辰. 基于质谱的单细胞蛋白质组学技术的发展及应用[J]. 生物技术通报, 2024, 40(11): 125-141.
GU Lei, ZHANG Yu-lu, TANG Shang-rui, YU Hao-yue, LI Chen. Development and Application of Mass Spectrometry-based Single-cell Proteomics Technologies[J]. Biotechnology Bulletin, 2024, 40(11): 125-141.
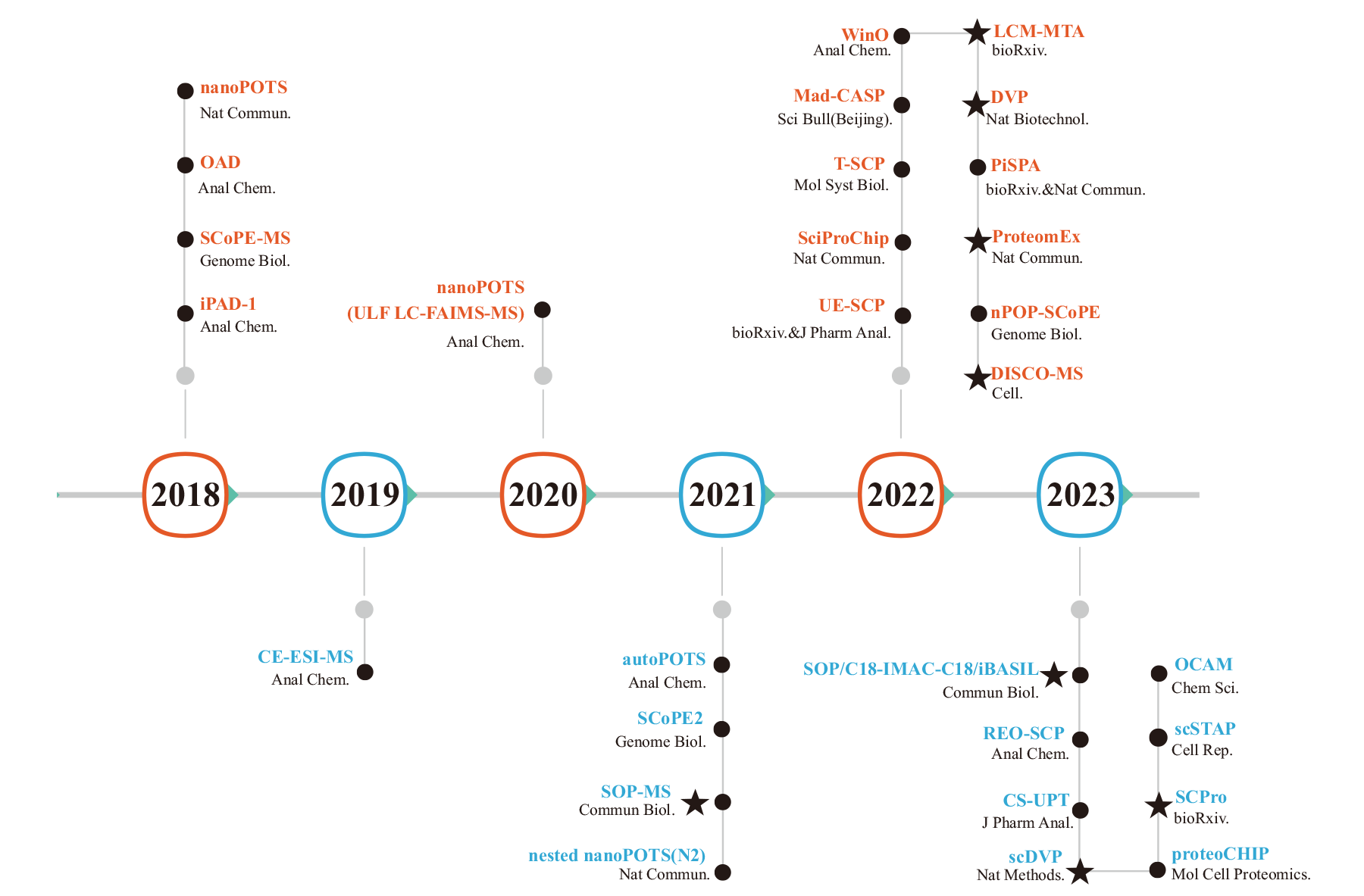
图1 基于质谱的单细胞蛋白质组学技术发展图 图中按照时间顺序列举2018-2023年代表性的单细胞蛋白质组学技术;●细胞悬液的单细胞蛋白质组学技术,★空间蛋白质组学技术
Fig. 1 Development of mass spectrometry-based single-cell proteomics technologies The figure lists representative single-cell proteomics techniques from 2018 to 2023 in chronological order; ● indicates single-cell proteomics techniques for cell suspensions, ★ indicates spatial proteomics techniques
| 单细胞蛋白质组学工具 Single-cell proteomics tools | 专用设备 Customized equipment | 标记 Label | 细胞类型 Cell types | 细胞分选分离方法 Cell isolation methods | 质谱仪 Mass spectrometers | 样品前处理通量 Pretreatment throughput(Cells per run) | 质谱检测通量 MS throughput(Cells per day) | 单个细胞中平均鉴定的蛋白质数目 Average identified protein number per cell | 单细胞蛋白质组覆盖总深度Depth of proteome coverage(n = number of cells) | 参考文献 References | 技术应用 Technical applications | 应用相关文献Related references |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| iPAD-1 | 需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Fusion Tribrid MS | 1 | 24 | 271 | 406(n=10) | [ | 图谱研究 | [ |
| OAD | 需要 | 非标记 | HeLa | 液滴微流控技术 | Orbitrap Elite MS | 1 | 4 | 51 | / | [ | 发育研究 | [ |
| nanoPOTS | 需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Fusion Lumos Tribrid MS | 27 | / | 669 | / | [ | 疾病精准分析 | [ |
| CE-ESI-MS | 需要 | 非标记 | Xenopus laevis(D11), zebrafish embryos | 毛细管电泳 | Orbitrap Q-Exactive Plus | / | / | 450-800 | / | [ | 发育研究 | [ |
| ULF LC-FAIMS-MS | 需要 | 非标记 | HeLa, U-937 | 纳升液相色谱毛细管 | Orbitrap Fusion Lumos Tribrid MS | 5 | / | 2 348 | / | [ | 癌症研究 | [ |
| nested nanoPOTS | 需要 | TMT16 | C10, RAW, SVEC | 全自动单细胞分选技术 | Orbitrap Eclipse Tribrid MS | 243 | 108 | 1 716 | 2 457(n=108) | [ | 免疫研究 | [ |
| iProChip | 需要 | 非标记 | MEC-1 | 双层聚二甲基硅氧烷(PDMS)装置 | Orbitrap Eclipse Tribrid MS | 9 | 9 | 455 | / | [ | 癌症研究 | [ |
| SciProChip | 需要 | 非标记 | PC-9 | 20 | 16 | 1 500 | 1 995(n=10) | |||||
| PiSPA | 需要 | 非标记 | A549 | 液滴微流控技术 | timsTOF Pro | 1 | / | 3 008 | 5 093(n=37) | [ | 癌症研究 | [ |
| nPOP | 需要 | TMT18 | U-937, WM989 | 全自动单细胞分选技术 | Orbitrap Q-Exactive | 2 016 | 212 | 997 | 2 844(n=1543) | [ | 癌症研究 | [ |
| proteoCHIP | 需要 | TMT16 | HeLa, HEK-293 | 全自动单细胞分选技术 | Orbitrap Exploris 480 MS | 592 | 384 | 1 940(20× carrier) | 3 674(n=276) | [ | 癌症研究 | [ |
| 1 598(no carrier) | ||||||||||||
| scSTAP | 需要 | 非标记 | 小鼠oocytes | 序控液滴微流控技术 | timsTOF Pro | / | 15-20 | 2 663 | 3 363(n=36) | [ | 发育研究 | [ |
| OCAM | 需要 | 非标记 | HeLa, 小鼠oocytes | 毛细管烷基化微反应 | Orbitrap Q-Exactive, Orbitrap Exploris 480 MS | 100 | / | 3 457 (single mouse oocyte) 1 509 (single HeLa cell) | / | [ | 发育研究 | [ |
| SCoPE | 不需要 | TMT10 | Jurkat, U-937 | 人工挑选 | LTQ Orbitrap Elite | 8 | 48 | / | 767(n=24) | [ | 图谱研究 | [ |
| SCoPE2 | 不需要 | TMT16 | Monocyte, macrophage | 流式细胞荧光分选术 | Orbitrap Q-Exactive | / | 200 | 1 000 | 3 042(n=1490) | [ | 免疫研究 | [ |
| SOP-MS | 不需要 | 非标记 | MCF10A, MCF7 | 流式细胞荧光分选术,激光捕获显微切割 | Orbitrap Q-Exactive Plus | / | / | 146 | / | [ | 癌症研究 | [ |
| SOP/C18-IMAC-C18/iBASIL | 不需要 | TMT标记 | AML | 流式细胞荧光分选术 | Orbitrap Q-Exactive Plus | / | / | 1 926 | 2 622(n=104) | [ | 免疫研究 | [ |
| autoPOTS | 不需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Exploris 480 MS | / | 10 | 301 | / | [ | 免疫研究 | [ |
| T-SCP | 不需要 | 非标记 | HeLa | 流式细胞荧光分选术 | timsTOF Pro | 308 | 41 | 2 083 | 2 501(n=231) | [ | 图谱研究 | [ |
| Mad-CASP | 不需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Eclipse Tribrid MS | / | 16 | 1 240 | / | [ | 疾病精准分析 | [ |
| WinO | 不需要 | TMT10 | RPMI8226 | 细胞分选仪 | Orbitrap Fusion Tribrid MS | / | 144 | 845 | / | [ | 免疫研究 | [ |
| UE-SCP | 不需要 | TMT6 | HeLa, HEK-293T | 全自动单细胞分选技术 | timsTOF Pro | 308 | 96 | 2 249 | 4 230(n=128) | [ | 癌症研究 | [ |
| CS-UPT | 不需要 | 非标记, TMT6, TMT8 | 小鼠MII oocytes, zygotes | 人工挑选 | timsTOF Pro | / | 18 非标记,(70, 50×car rier,100×carrier),(140, no carrier) | 2 665非标记,(2 182, 50× carrier),(2 371, 100× carrier),(1 568, no carrier) | / | [ | 发育研究 | [ |
| REO-SCP | 不需要 | 非标记 | HeLa | 全自动单细胞分选技术 | Orbitrap Exploris 480 MS | 384 | / | 1 208 | / | [ | 图谱研究 | [ |
| DVP | 不需要 | 非标记 | U2OS | 激光显微切割技术 | timsTOF Pro | / | / | 4 500-4 800(每个样品平均267个细胞核) | 5 085(8个重复) | [ | 癌症研究 | [ |
| scDVP | 不需要 | 非标记 | 小鼠肝细胞 | 激光显微切割技术 | timsTOF SCP | / | 80 | 1 700-2 700(1/3-1/2个细胞的体积) | / | [ | 图谱研究 | [ |
| LCM-MTA | 不需要 | 非标记 | 人结直肠癌癌组织与癌旁组织 | 激光显微切割技术 | timsTOF Pro | / | / | 536(15个细胞的体积) | / | [ | 癌症研究 | [ |
| ProteomEx | 不需要 | 非标记 | 小鼠脑组织 | 激光显微切割技术 | timsTOF Pro | / | / | 928(160 μm横向分辨率) | / | [ | 图谱研究 | [ |
| DISCO-MS | 不需要 | 非标记 | 小鼠脑、心脏和肺组织 | 激光显微切割技术 | timsTOF Pro | / | / | 1 400(每个ROI) | / | [ | 癌症研究 | [ |
| SCPro | 不需要 | 非标记 | HEK 293T | 激光显微切割技术 | timsTOF Pro | / | / | 800(2.4个细胞的体积) | / | [ | 疾病精准分析 | [ |
表1 代表性的单细胞蛋白质组学技术及其应用
Table 1 Representative single-cell proteomics techniques and their applications
| 单细胞蛋白质组学工具 Single-cell proteomics tools | 专用设备 Customized equipment | 标记 Label | 细胞类型 Cell types | 细胞分选分离方法 Cell isolation methods | 质谱仪 Mass spectrometers | 样品前处理通量 Pretreatment throughput(Cells per run) | 质谱检测通量 MS throughput(Cells per day) | 单个细胞中平均鉴定的蛋白质数目 Average identified protein number per cell | 单细胞蛋白质组覆盖总深度Depth of proteome coverage(n = number of cells) | 参考文献 References | 技术应用 Technical applications | 应用相关文献Related references |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| iPAD-1 | 需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Fusion Tribrid MS | 1 | 24 | 271 | 406(n=10) | [ | 图谱研究 | [ |
| OAD | 需要 | 非标记 | HeLa | 液滴微流控技术 | Orbitrap Elite MS | 1 | 4 | 51 | / | [ | 发育研究 | [ |
| nanoPOTS | 需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Fusion Lumos Tribrid MS | 27 | / | 669 | / | [ | 疾病精准分析 | [ |
| CE-ESI-MS | 需要 | 非标记 | Xenopus laevis(D11), zebrafish embryos | 毛细管电泳 | Orbitrap Q-Exactive Plus | / | / | 450-800 | / | [ | 发育研究 | [ |
| ULF LC-FAIMS-MS | 需要 | 非标记 | HeLa, U-937 | 纳升液相色谱毛细管 | Orbitrap Fusion Lumos Tribrid MS | 5 | / | 2 348 | / | [ | 癌症研究 | [ |
| nested nanoPOTS | 需要 | TMT16 | C10, RAW, SVEC | 全自动单细胞分选技术 | Orbitrap Eclipse Tribrid MS | 243 | 108 | 1 716 | 2 457(n=108) | [ | 免疫研究 | [ |
| iProChip | 需要 | 非标记 | MEC-1 | 双层聚二甲基硅氧烷(PDMS)装置 | Orbitrap Eclipse Tribrid MS | 9 | 9 | 455 | / | [ | 癌症研究 | [ |
| SciProChip | 需要 | 非标记 | PC-9 | 20 | 16 | 1 500 | 1 995(n=10) | |||||
| PiSPA | 需要 | 非标记 | A549 | 液滴微流控技术 | timsTOF Pro | 1 | / | 3 008 | 5 093(n=37) | [ | 癌症研究 | [ |
| nPOP | 需要 | TMT18 | U-937, WM989 | 全自动单细胞分选技术 | Orbitrap Q-Exactive | 2 016 | 212 | 997 | 2 844(n=1543) | [ | 癌症研究 | [ |
| proteoCHIP | 需要 | TMT16 | HeLa, HEK-293 | 全自动单细胞分选技术 | Orbitrap Exploris 480 MS | 592 | 384 | 1 940(20× carrier) | 3 674(n=276) | [ | 癌症研究 | [ |
| 1 598(no carrier) | ||||||||||||
| scSTAP | 需要 | 非标记 | 小鼠oocytes | 序控液滴微流控技术 | timsTOF Pro | / | 15-20 | 2 663 | 3 363(n=36) | [ | 发育研究 | [ |
| OCAM | 需要 | 非标记 | HeLa, 小鼠oocytes | 毛细管烷基化微反应 | Orbitrap Q-Exactive, Orbitrap Exploris 480 MS | 100 | / | 3 457 (single mouse oocyte) 1 509 (single HeLa cell) | / | [ | 发育研究 | [ |
| SCoPE | 不需要 | TMT10 | Jurkat, U-937 | 人工挑选 | LTQ Orbitrap Elite | 8 | 48 | / | 767(n=24) | [ | 图谱研究 | [ |
| SCoPE2 | 不需要 | TMT16 | Monocyte, macrophage | 流式细胞荧光分选术 | Orbitrap Q-Exactive | / | 200 | 1 000 | 3 042(n=1490) | [ | 免疫研究 | [ |
| SOP-MS | 不需要 | 非标记 | MCF10A, MCF7 | 流式细胞荧光分选术,激光捕获显微切割 | Orbitrap Q-Exactive Plus | / | / | 146 | / | [ | 癌症研究 | [ |
| SOP/C18-IMAC-C18/iBASIL | 不需要 | TMT标记 | AML | 流式细胞荧光分选术 | Orbitrap Q-Exactive Plus | / | / | 1 926 | 2 622(n=104) | [ | 免疫研究 | [ |
| autoPOTS | 不需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Exploris 480 MS | / | 10 | 301 | / | [ | 免疫研究 | [ |
| T-SCP | 不需要 | 非标记 | HeLa | 流式细胞荧光分选术 | timsTOF Pro | 308 | 41 | 2 083 | 2 501(n=231) | [ | 图谱研究 | [ |
| Mad-CASP | 不需要 | 非标记 | HeLa | 流式细胞荧光分选术 | Orbitrap Eclipse Tribrid MS | / | 16 | 1 240 | / | [ | 疾病精准分析 | [ |
| WinO | 不需要 | TMT10 | RPMI8226 | 细胞分选仪 | Orbitrap Fusion Tribrid MS | / | 144 | 845 | / | [ | 免疫研究 | [ |
| UE-SCP | 不需要 | TMT6 | HeLa, HEK-293T | 全自动单细胞分选技术 | timsTOF Pro | 308 | 96 | 2 249 | 4 230(n=128) | [ | 癌症研究 | [ |
| CS-UPT | 不需要 | 非标记, TMT6, TMT8 | 小鼠MII oocytes, zygotes | 人工挑选 | timsTOF Pro | / | 18 非标记,(70, 50×car rier,100×carrier),(140, no carrier) | 2 665非标记,(2 182, 50× carrier),(2 371, 100× carrier),(1 568, no carrier) | / | [ | 发育研究 | [ |
| REO-SCP | 不需要 | 非标记 | HeLa | 全自动单细胞分选技术 | Orbitrap Exploris 480 MS | 384 | / | 1 208 | / | [ | 图谱研究 | [ |
| DVP | 不需要 | 非标记 | U2OS | 激光显微切割技术 | timsTOF Pro | / | / | 4 500-4 800(每个样品平均267个细胞核) | 5 085(8个重复) | [ | 癌症研究 | [ |
| scDVP | 不需要 | 非标记 | 小鼠肝细胞 | 激光显微切割技术 | timsTOF SCP | / | 80 | 1 700-2 700(1/3-1/2个细胞的体积) | / | [ | 图谱研究 | [ |
| LCM-MTA | 不需要 | 非标记 | 人结直肠癌癌组织与癌旁组织 | 激光显微切割技术 | timsTOF Pro | / | / | 536(15个细胞的体积) | / | [ | 癌症研究 | [ |
| ProteomEx | 不需要 | 非标记 | 小鼠脑组织 | 激光显微切割技术 | timsTOF Pro | / | / | 928(160 μm横向分辨率) | / | [ | 图谱研究 | [ |
| DISCO-MS | 不需要 | 非标记 | 小鼠脑、心脏和肺组织 | 激光显微切割技术 | timsTOF Pro | / | / | 1 400(每个ROI) | / | [ | 癌症研究 | [ |
| SCPro | 不需要 | 非标记 | HEK 293T | 激光显微切割技术 | timsTOF Pro | / | / | 800(2.4个细胞的体积) | / | [ | 疾病精准分析 | [ |
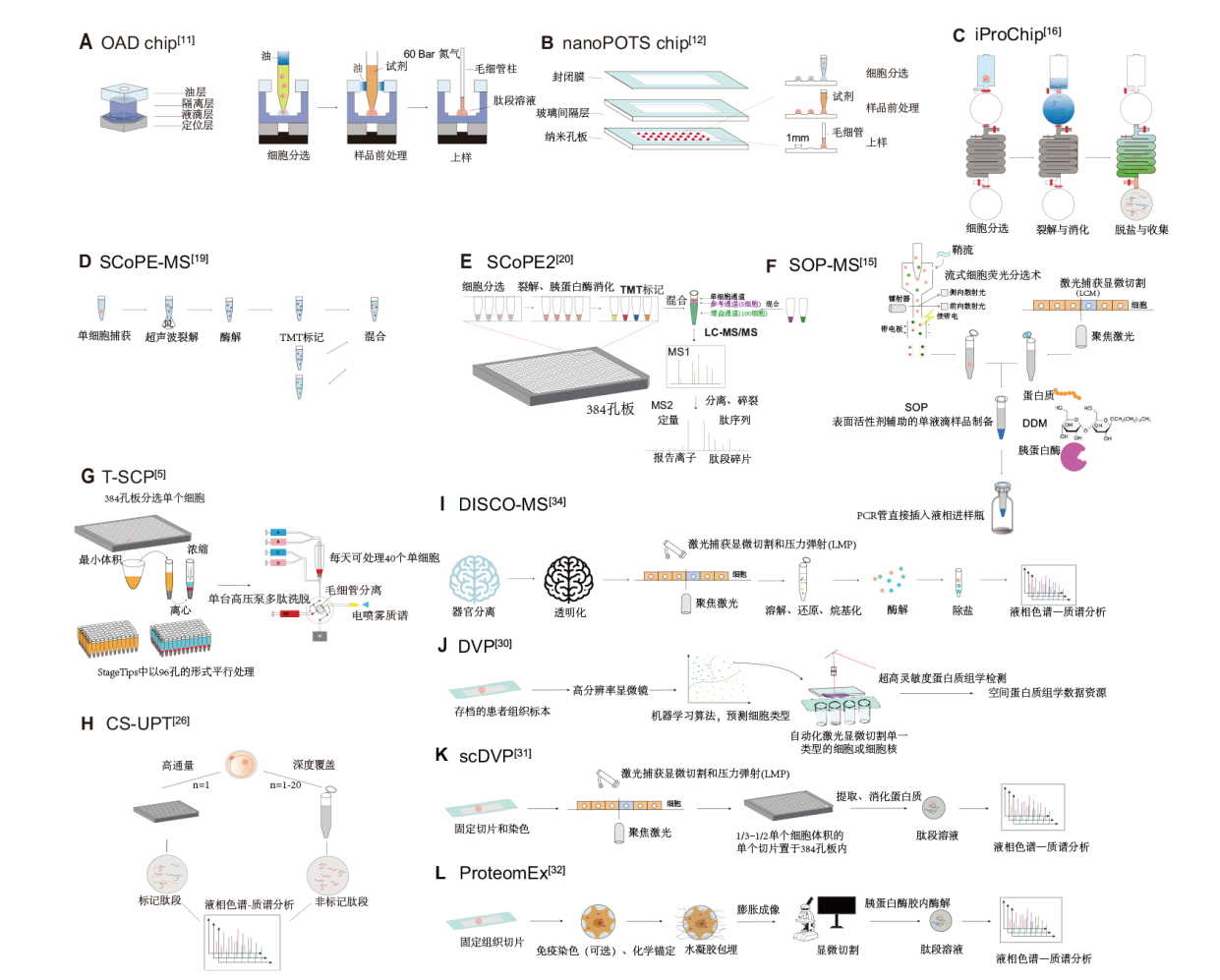
图2 代表性的单细胞蛋白质组学技术简化原理图 A:OAD chip,一种用于单细胞蛋白质组学分析的微流控设备,能够实现单细胞裂解与蛋白质消化、脱盐及样品收集等一体化操作;该芯片设计旨在减少样品损失并提高处理效率,尤其适用于极小体积样品的分析;B:nanoPOTS chip,一种专门用于处理极小样本量的微流控芯片,适用于纳升级和皮升级体积的蛋白质组学样品处理;该芯片能够将单个细胞样品进行高效裂解和消化,并极大减少样品损失;C:iProChip,一种集成化的蛋白质组学芯片,用于在微流控环境下进行细胞分选、样品前处理和质谱分析的自动化操作;该芯片设计旨在将多个步骤集成到一体,以提高操作效率和数据质量;D:SCoPE-MS,一种专门用于单细胞蛋白质组学分析的技术;它利用同位素标记的方法对多个单细胞样品进行高效分析;E:SCoPE2,是SCoPE-MS的升级版,具有更高的灵敏度和分辨率,适用于分析低丰度蛋白质;F:SOP-MS,一种标准操作规程下的质谱分析技术,常用于分析蛋白质样品;G:T-SCP,一种单细胞检测分辨率的高通量蛋白质组学技术;H:iProChip,一种用于单细胞分析的微流控芯片,能够实现从细胞分选到样品前处理的一体化操作;I:DISCO-MS,一种结合组织透明化和质谱分析的技术,适用于从组织样本中提取和分析蛋白质;J:DVP,结合显微成像和超高灵敏度质谱法的空间蛋白质组学分析技术;K:scDVP,DVP的单细胞版本;L:ProteomEx,一种简单、便捷、稳定、可重复的空间蛋白质组学分析方法,采用了水凝胶放大实际样品,再结合4D蛋白质组学技术助力微量样本的蛋白质组学分析;LC-MS/MS:液相色谱-串联质谱;TMT:同位素标记标签;SCoPE-MS:单细胞蛋白质组质谱技术;LMP:激光捕获显微切割和压力弹射;SCP:单细胞蛋白质组学
Fig. 2 Simplified schematics of representative single-cell proteomics techniques A: OAD chip is a microfluidic device used for single-cell proteomics analysis, capable of performing single-cell lysis, protein digestion, desalting, and sample collection in an integrated workflow. This chip is designed to minimize sample loss and enhance processing efficiency, especially suitable for small-volume samples. B: NanoPOTS chip is a microfluidic chip designed specifically for handling ultra-small sample volumes, suitable for proteomics sample preparation at the nanoliter and picoliter scale. This chip efficiently lyses and digests single-cell samples, significantly reducing sample loss. C: iProChip is an integrated proteomics chip used for automated operations of cell sorting, sample preparation, and mass spectrometry analysis in a microfluidic environment. The chip is designed to integrate multiple steps to improve operational efficiency and data quality. D: SCoPE-MS is a specialized technology for single-cell proteomics analysis. It uses isotope labeling to efficiently analyze multiple single-cell samples. E: SCoPE2 is an upgraded version of SCoPE-MS, featuring higher sensitivity and resolution, making it suitable for analyzing low-abundance proteins. F: SOP-MS is a mass spectrometry technique under standard operating procedures, commonly used for protein sample analysis. G: T-SCP is a high-throughput technique for single-cell proteomics, capable of processing large numbers of single-cell samples. H: iProChip is a microfluidic chip used for single-cell analysis, capable of integrating cell sorting and sample preparation in one platform. I: DISCO-MS is a technique combining tissue clearing and mass spectrometry analysis, suitable for extracting and analyzing proteins from tissue samples. J: DVP is a technique used for proteomics analysis, combining microscopy imaging with mass spectrometry. K: scDVP is the single-cell version of DVP, specifically used to analyze protein expression in single cells. L: ProteomEx is a high-resolution mass spectrometry technology used for analyzing archived tissue samples, capable of extracting single-cell information from complex tissue samples. LC-MS/MS: Liquid chromatography-tandem mass spectrometry. TMT: Tandem mass tag. SCoPE-MS: Single-cell proteomics by mass spectrometry. LMP: Laser capture microdissection and pressure catapulting. SCP: Single-cell proteomics
| 单细胞蛋白质组学数据分析软件Software for single-cell proteomics data analysis | 构建方式Construction method | 输出方式 Output method | 适配的数据采集模式 Compatible data acquisition modes | 测试数据集 Test datasets | 特点 Features | 参考文献 References |
|---|---|---|---|---|---|---|
| DO-MS | R | HTML报告 | DDA、DIA | SCoPE、SCoPE2 | 交互式可视化分析 | [ |
| SCPCompanion | C# | XML文件 | DDA | SCoPE、N2 | 推荐仪器和数据分析参数 | [ |
| SCeptre | Python | YAML文件 | DDA | AML细胞群 | 数据归一化处理 | [ |
| DART-ID | Python和R | HTML报告 | DDA | SCoPE、SCoPE2 | 对齐肽段保留时间 | [ |
| IceR | R | TSV文件 | DDA(PIP)、DIA | DeMix-Q、IonStar、HRM-DIA等 | 使用离子电流信息进行混合肽段鉴定 | [ |
| DeepSCP | Python(深度学习) | CSV文件 | DDA(TDC) | SCoPE2、N2 | 首次应用深度学习技术进行DDA分析 | [ |
表2 代表性的单细胞蛋白质组学数据分析软件
Table 2 Representative software for single-cell proteomics data analysis
| 单细胞蛋白质组学数据分析软件Software for single-cell proteomics data analysis | 构建方式Construction method | 输出方式 Output method | 适配的数据采集模式 Compatible data acquisition modes | 测试数据集 Test datasets | 特点 Features | 参考文献 References |
|---|---|---|---|---|---|---|
| DO-MS | R | HTML报告 | DDA、DIA | SCoPE、SCoPE2 | 交互式可视化分析 | [ |
| SCPCompanion | C# | XML文件 | DDA | SCoPE、N2 | 推荐仪器和数据分析参数 | [ |
| SCeptre | Python | YAML文件 | DDA | AML细胞群 | 数据归一化处理 | [ |
| DART-ID | Python和R | HTML报告 | DDA | SCoPE、SCoPE2 | 对齐肽段保留时间 | [ |
| IceR | R | TSV文件 | DDA(PIP)、DIA | DeMix-Q、IonStar、HRM-DIA等 | 使用离子电流信息进行混合肽段鉴定 | [ |
| DeepSCP | Python(深度学习) | CSV文件 | DDA(TDC) | SCoPE2、N2 | 首次应用深度学习技术进行DDA分析 | [ |
| [1] |
Aslam B, Basit M, Nisar MA, et al. Proteomics: technologies and their applications[J]. J Chromatogr Sci, 2017, 55(2): 182-196.
doi: 10.1093/chromsci/bmw167 pmid: 28087761 |
| [2] |
Kelly RT. Single-cell proteomics: progress and prospects[J]. Mol Cell Proteomics, 2020, 19(11): 1739-1748.
doi: 10.1074/mcp.R120.002234 pmid: 32847821 |
| [3] |
Vistain LF, Tay S. Single-cell proteomics[J]. Trends Biochem Sci, 2021, 46(8): 661-672.
doi: 10.1016/j.tibs.2021.01.013 pmid: 33653632 |
| [4] |
Woo J, Williams SM, Markillie LM, et al. High-throughput and high-efficiency sample preparation for single-cell proteomics using a nested nanowell chip[J]. Nat Commun, 2021, 12(1): 6246.
doi: 10.1038/s41467-021-26514-2 pmid: 34716329 |
| [5] | Brunner AD, Thielert M, Vasilopoulou C, et al. Ultra-high sensitivity mass spectrometry quantifies single-cell proteome changes upon perturbation[J]. Mol Syst Biol, 2022, 18(3): e10798. |
| [6] | Jiang YR, Zhu L, Cao LR, et al. Simultaneous deep transcriptome and proteome profiling in a single mouse oocyte[J]. Cell Rep, 2023, 42(11): 113455. |
| [7] |
Levy E, Slavov N. Single cell protein analysis for systems biology[J]. Essays Biochem, 2018, 62(4): 595-605.
doi: 10.1042/EBC20180014 pmid: 30072488 |
| [8] | Lundberg E, Borner GHH. Spatial proteomics: a powerful discovery tool for cell biology[J]. Nat Rev Mol Cell Biol, 2019, 20(5): 285-302. |
| [9] | Cheung TK, Lee CY, Bayer FP, et al. Defining the carrier proteome limit for single-cell proteomics[J]. Nat Meth, 2021, 18: 76-83. |
| [10] |
Shao X, Wang XT, Guan S, et al. Integrated proteome analysis device for fast single-cell protein profiling[J]. Anal Chem, 2018, 90(23): 14003-14010.
doi: 10.1021/acs.analchem.8b03692 pmid: 30375851 |
| [11] | Li ZY, Huang M, Wang XK, et al. Nanoliter-scale oil-air-droplet chip-based single cell proteomic analysis[J]. Anal Chem, 2018, 90(8): 5430-5438. |
| [12] |
Zhu Y, Piehowski PD, Zhao R, et al. Nanodroplet processing platform for deep and quantitative proteome profiling of 10-100 mammalian cells[J]. Nat Commun, 2018, 9(1): 882.
doi: 10.1038/s41467-018-03367-w pmid: 29491378 |
| [13] |
Lombard-Banek C, Moody SA, Manzini MC, et al. Microsampling capillary electrophoresis mass spectrometry enables single-cell proteomics in complex tissues: developing cell clones in live Xenopus laevis and zebrafish embryos[J]. Anal Chem, 2019, 91(7): 4797-4805.
doi: 10.1021/acs.analchem.9b00345 pmid: 30827088 |
| [14] |
Greguš M, Kostas JC, Ray S, et al. Improved sensitivity of ultralow flow LC-MS-based proteomic profiling of limited samples using monolithic capillary columns and FAIMS technology[J]. Anal Chem, 2020, 92(21): 14702-14712.
doi: 10.1021/acs.analchem.0c03262 pmid: 33054160 |
| [15] | Tsai CF, Zhang PF, Scholten D, et al. Surfactant-assisted one-pot sample preparation for label-free single-cell proteomics[J]. Commun Biol, 2021, 4(1): 265. |
| [16] |
Gebreyesus ST, Siyal AA, Kitata RB, et al. Streamlined single-cell proteomics by an integrated microfluidic chip and data-independent acquisition mass spectrometry[J]. Nat Commun, 2022, 13(1): 37.
doi: 10.1038/s41467-021-27778-4 pmid: 35013269 |
| [17] |
Wang Y, Guan ZY, Shi SW, et al. Pick-up single-cell proteomic analysis for quantifying up to 3000 proteins in a Mammalian cell[J]. Nat Commun, 2024, 15: 1279.
doi: 10.1038/s41467-024-45659-4 pmid: 38341466 |
| [18] | Tsai CF, Wang YT, Hsu CC, et al. A streamlined tandem tip-based workflow for sensitive nanoscale phosphoproteomics[J]. Commun Biol, 2023, 6: 70. |
| [19] |
Budnik B, Levy E, Harmange G, et al. SCoPE-MS: mass spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation[J]. Genome Biol, 2018, 19(1): 161.
doi: 10.1186/s13059-018-1547-5 pmid: 30343672 |
| [20] |
Specht H, Emmott E, Petelski AA, et al. Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2[J]. Genome Biol, 2021, 22(1): 50.
doi: 10.1186/s13059-021-02267-5 pmid: 33504367 |
| [21] |
Liang YR, Acor H, McCown MA, et al. Fully automated sample processing and analysis workflow for low-input proteome profiling[J]. Anal Chem, 2021, 93(3): 1658-1666.
doi: 10.1021/acs.analchem.0c04240 pmid: 33352054 |
| [22] |
Li SQ, Su KC, Zhuang ZK, et al. A simple, rapid, and practical method for single-cell proteomics based on mass-adaptive coating of synthetic peptides[J]. Sci Bull, 2022, 67(6): 581-584.
doi: 10.1016/j.scib.2021.12.022 pmid: 36546118 |
| [23] |
Masuda T, Inamori Y, Furukawa A, et al. Water droplet-in-oil digestion method for single-cell proteomics[J]. Anal Chem, 2022, 94(29): 10329-10336.
doi: 10.1021/acs.analchem.1c05487 pmid: 35817413 |
| [24] |
Leduc A, Huffman RG, Cantlon J, et al. Exploring functional protein covariation across single cells using nPOP[J]. Genome Biol, 2022, 23(1): 261.
doi: 10.1186/s13059-022-02817-5 pmid: 36527135 |
| [25] | Gu L, Li Z, Wang Q, et al. An ultra-sensitive and easy-to-use multiplexed single-cell proteomic analysis[J]. bioRxiv, 2022. DOI: https://doi.org/10.1101/2022.01.02.474723. |
| [26] |
Gu L, Li XM, Zhu WC, et al. Ultrasensitive proteomics depicted an in-depth landscape for the very early stage of mouse maternal-to-zygotic transition[J]. J Pharm Anal, 2023, 13(8): 942-954.
doi: 10.1016/j.jpha.2023.05.003 pmid: 37719194 |
| [27] |
Matzinger M, Müller E, Dürnberger G, et al. Robust and easy-to-use one-pot workflow for label-free single-cell proteomics[J]. Anal Chem, 2023, 95(9): 4435-4445.
doi: 10.1021/acs.analchem.2c05022 pmid: 36802514 |
| [28] | Ctortecka C, Hartlmayr D, Seth A, et al. An automated nanowell-array workflow for quantitative multiplexed single-cell proteomics sample preparation at high sensitivity[J]. Mol Cell Proteomics, 2023, 22(12): 100665. |
| [29] | Gu L, Li X, Li Z, et al. Increasing the sensitivity, recovery, and integrality of spatially resolved proteomics by LCM-MTA[J]. bioRxiv, 2022. DOI: https://doi.org/10.1101/2022.08.21.504675. |
| [30] |
Mund A, Coscia F, Kriston A, et al. Deep Visual Proteomics defines single-cell identity and heterogeneity[J]. Nat Biotechnol, 2022, 40(8): 1231-1240.
doi: 10.1038/s41587-022-01302-5 pmid: 35590073 |
| [31] |
Rosenberger FA, Thielert M, Strauss MT, et al. Spatial single-cell mass spectrometry defines zonation of the hepatocyte proteome[J]. Nat Methods, 2023, 20(10): 1530-1536.
doi: 10.1038/s41592-023-02007-6 pmid: 37783884 |
| [32] |
Li L, Sun CJ, Sun YT, et al. Spatially resolved proteomics via tissue expansion[J]. Nat Commun, 2022, 13(1): 7242.
doi: 10.1038/s41467-022-34824-2 pmid: 36450705 |
| [33] | Tsai CF, Zhao R, Williams SM, et al. An improved boosting to amplify signal with isobaric labeling(iBASIL)strategy for precise quantitative single-cell proteomics[J]. Mol Cell Proteomics, 2020, 19(5): 828-838. |
| [34] |
Bhatia HS, Brunner AD, Öztürk F, et al. Spatial proteomics in three-dimensional intact specimens[J]. Cell, 2022, 185(26): 5040-5058.e19.
doi: 10.1016/j.cell.2022.11.021 pmid: 36563667 |
| [35] | Xu Y, Wang X, Li Y, et al. Multimodal single cell-resolved spatial proteomics reveals pancreatic tumor heterogeneity[J]. bioRxiv, 2023. DOI: https://doi.org/10.1101/2023.11.04.565590. |
| [36] |
Dang YJ, Zhu L, Yuan P, et al. Functional profiling of stage-specific proteome and translational transition across human pre-implantation embryo development at a single-cell resolution[J]. Cell Discov, 2023, 9(1): 10.
doi: 10.1038/s41421-022-00491-2 pmid: 36693841 |
| [37] |
He YY, Yuan HM, Liang Y, et al. On-capillary alkylation micro-reactor: a facile strategy for proteo-metabolome profiling in the same single cells[J]. Chem Sci, 2023, 14(46): 13495-13502.
doi: 10.1039/d3sc05047e pmid: 38033888 |
| [38] | Zhu WC, Ding YF, Meng J, et al. Reading and writing of mRNA m6A modification orchestrate maternal-to-zygotic transition in mice[J]. Genome Biol, 2023, 24(1): 67. |
| [39] |
Gross A, Schoendube J, Zimmermann S, et al. Technologies for single-cell isolation[J]. Int J Mol Sci, 2015, 16(8): 16897-16919.
doi: 10.3390/ijms160816897 pmid: 26213926 |
| [40] | Yokoyama WM, Christensen M, Dos Santos G, et al. Production of monoclonal antibodies[J]. Curr Protoc Immunol, 2013, 102(1): 2.5.1-2.5.29. |
| [41] | Wright G, Tucker MJ, Morton PC, et al. Micromanipulation in assisted reproduction: a review of current technology[J]. Curr Opin Obstet Gynecol, 1998, 10(3): 221-226. |
| [42] |
van den Brink SC, Sage F, Vértesy Á, et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations[J]. Nat Methods, 2017, 14(10): 935-936.
doi: 10.1038/nmeth.4437 pmid: 28960196 |
| [43] |
Nakamura N, Ruebel K, Jin L, et al. Laser capture microdissection for analysis of single cells[J]. Methods Mol Med, 2007, 132: 11-18.
pmid: 17876072 |
| [44] | 宋扬, 林金明. 微流控芯片上单细胞操控与分析方法研究进展[J]. 中国科学: 化学, 2023, 53(8): 1472-1493. |
| Song Y, Lin JM. Recent advances in single-cell manipulation and analysis methods on microfluidic chips[J]. Sci Sin Chim, 2023, 53(8): 1472-1493. | |
| [45] |
Hosic S, Murthy SK, Koppes AN. Microfluidic sample preparation for single cell analysis[J]. Anal Chem, 2016, 88(1): 354-380.
doi: 10.1021/acs.analchem.5b04077 pmid: 26567589 |
| [46] |
Liu D, Sun ML, Zhang JW, et al. Single-cell droplet microfluidics for biomedical applications[J]. Analyst, 2022, 147(11): 2294-2316.
doi: 10.1039/d1an02321g pmid: 35506869 |
| [47] | Gao Y, Wu MR, Lin Y, et al. Acoustic microfluidic separation techniques and bioapplications: a review[J]. Micromachines, 2020, 11(10): 921. |
| [48] | 贺映云, 袁辉明, 梁振, 等. 单细胞蛋白质组学分析技术研究进展[J]. 分析试验室, 2022, 41(12): 1365-1378. |
| He YY, Yuan HM, Liang Z, et al. Progress in technologies for single-cell proteome analysis[J]. Chin J Anal Lab, 2022, 41(12): 1365-1378. | |
| [49] |
Shen Y, Tolicć N, Masselon C, et al. Nanoscale proteomics[J]. Anal Bioanal Chem, 2004, 378(4): 1037-1045.
pmid: 14647945 |
| [50] |
Shen YF, Zhao R, Berger SJ, et al. High-efficiency nanoscale liquid chromatography coupled on-line with mass spectrometry using nanoelectrospray ionization for proteomics[J]. Anal Chem, 2002, 74(16): 4235-4249.
pmid: 12199598 |
| [51] |
Shen YF, Tolić N, Masselon C, et al. Ultrasensitive proteomics using high-efficiency on-line micro-SPE-nanoLC-nanoESI MS and MS/MS[J]. Anal Chem, 2004, 76(1): 144-154.
pmid: 14697044 |
| [52] |
Zhu Y, Clair G, Chrisler WB, et al. Proteomic analysis of single mammalian cells enabled by microfluidic nanodroplet sample preparation and ultrasensitive NanoLC-MS[J]. Angew Chem Int Ed, 2018, 57(38): 12370-12374.
doi: 10.1002/anie.201802843 pmid: 29797682 |
| [53] |
Cong YZ, Liang YR, Motamedchaboki K, et al. Improved single-cell proteome coverage using narrow-bore packed NanoLC columns and ultrasensitive mass spectrometry[J]. Anal Chem, 2020, 92(3): 2665-2671.
doi: 10.1021/acs.analchem.9b04631 pmid: 31913019 |
| [54] | Orsburn BC. Acetic acid is a superior acidifier for sub-nanogram and single cell proteomic studies[J]. bioRxiv, 2023. DOI: https://doi.org/10.1101/2023.08.01.551522. |
| [55] | Ye ZL, Sabatier P, Van der Hoeven L, et al. High-throughput and scalable single cell proteomics identifies over 5000 proteins per cell[J]. bioRxiv, 2023. DOI: https://doi.org/10.1101/2023.11.27.568953. |
| [56] | Bubis JA, Arrey TN, Damoc E, et al. Challenging the AstralTM mass analyzer - up to 5300 proteins per single-cell at unseen quantitative accuracy to study cellular heterogeneity[J]. bioRxiv, 2024. DOI: https://doi.org/10.1101/2024.02.01.578358 |
| [57] | Ctortecka C, Clark NM, Boyle B, et al. Automated single-cell proteomics providing sufficient proteome depth to study complex biology beyond cell type classifications[J]. bioRxiv, 2024. DOI: https://doi.org/10.1101/2024.01.20.576369. |
| [58] |
Stadlmann J, Hudecz O, Krššáková G, et al. Improved sensitivity in low-input proteomics using micropillar array-based chromatography[J]. Anal Chem, 2019, 91(22): 14203-14207.
doi: 10.1021/acs.analchem.9b02899 pmid: 31612716 |
| [59] |
Amenson-Lamar EA, Sun LL, Zhang ZB, et al. Detection of 1 zmol injection of angiotensin using capillary zone electrophoresis coupled to a Q-Exactive HF mass spectrometer with an electrokinetically pumped sheath-flow electrospray interface[J]. Talanta, 2019, 204: 70-73.
doi: S0039-9140(19)30573-9 pmid: 31357355 |
| [60] | Dou MW, Zhu Y, Liyu A, et al. Nanowell-mediated two-dimensional liquid chromatography enables deep proteome profiling of <1000 mammalian cells[J]. Chem Sci, 2018, 9(34): 6944-6951. |
| [61] |
Krieger JR, Wybenga-Groot LE, Tong JF, et al. Evosep one enables robust deep proteome coverage using tandem mass tags while significantly reducing instrument time[J]. J Proteome Res, 2019, 18(5): 2346-2353.
doi: 10.1021/acs.jproteome.9b00082 pmid: 30938160 |
| [62] |
Zheng RS, Stejskal K, Pynn C, et al. Deep single-shot NanoLC-MS proteome profiling with a 1500 bar UHPLC system, long fully porous columns, and HRAM MS[J]. J Proteome Res, 2022, 21(10): 2545-2551.
doi: 10.1021/acs.jproteome.2c00270 pmid: 36068014 |
| [63] |
Zhu Y, Zhao R, Piehowski PD, et al. Subnanogram proteomics: impact of LC column selection, MS instrumentation and data analysis strategy on proteome coverage for trace samples[J]. Int J Mass Spectrom, 2018, 427: 4-10.
doi: 10.1016/j.ijms.2017.08.016 pmid: 29576737 |
| [64] | Cox J. Prediction of peptide mass spectral libraries with machine learning[J]. Nat Biotechnol, 2023, 41(1): 33-43. |
| [65] |
Michalski A, Cox J, Mann M. More than 100, 000 detectable peptide species elute in single shotgun proteomics runs but the majority is inaccessible to data-dependent LC-MS/MS[J]. J Proteome Res, 2011, 10(4): 1785-1793.
doi: 10.1021/pr101060v pmid: 21309581 |
| [66] | Guan SH, Taylor PP, Han ZW, et al. Data dependent-independent acquisition(DDIA)proteomics[J]. J Proteome Res, 2020, 19(8): 3230-3237. |
| [67] |
Schoof EM, Furtwängler B, Üresin N, et al. Quantitative single-cell proteomics as a tool to characterize cellular hierarchies[J]. Nat Commun, 2021, 12(1): 3341.
doi: 10.1038/s41467-021-23667-y pmid: 34099695 |
| [68] |
Kalxdorf M, Müller T, Stegle O, et al. IceR improves proteome coverage and data completeness in global and single-cell proteomics[J]. Nat Commun, 2021, 12(1): 4787.
doi: 10.1038/s41467-021-25077-6 pmid: 34373457 |
| [69] | Wang B, Wang Y, Chen Y, et al. DeepSCP: utilizing deep learning to boost single-cell proteome coverage[J]. Brief Bioinform, 2022, 23(4): bbac214. |
| [70] |
Huang Z, Merrihew GE, Larson EB, et al. Brain proteomic analysis implicates actin filament processes and injury response in resilience to Alzheimer’s disease[J]. Nat Commun, 2023, 14(1): 2747.
doi: 10.1038/s41467-023-38376-x pmid: 37173305 |
| [71] | Sun FY, Li HY, Sun DQ, et al. Single-cell omics: experimental workflow, data analyses and applications[J]. Sci China Life Sci, 2024. DOI: 10.1007/s11427-023-2561-0. |
| [72] |
Malioutov D, Chen TC, Airoldi E, et al. Quantifying homologous proteins and proteoforms[J]. Mol Cell Proteomics, 2019, 18(1): 162-168.
doi: 10.1074/mcp.TIR118.000947 pmid: 30282776 |
| [1] | 杨伟杰, 杨周林, 朱浩东, 魏煜, 刘君, 刘训. 地衣素合成酶关键模块 LchAD 蛋白的性质和功能研究[J]. 生物技术通报, 2024, 40(3): 322-332. |
| [2] | 纪宏超, 李正艳. 基于质谱的未知次生代谢物结构解析研究进展与展望[J]. 生物技术通报, 2024, 40(10): 76-85. |
| [3] | 何诗瑜, 曾仲大, 李博岩. 空间分辨代谢组学在疾病诊断研究中的应用进展[J]. 生物技术通报, 2024, 40(1): 145-159. |
| [4] | 罗阳兰, 曹乃馨, 杨玉梅, 黄丽玲, 阎勇, 黄世旅. 三种桑黄子实体脂溶性成分的GC-MS分析及抗氧化活性[J]. 生物技术通报, 2023, 39(6): 149-157. |
| [5] | 苗玉娇, 朱龙佼, 许文涛. 质谱成像新基质及其在分析生物样本方面的研究进展[J]. 生物技术通报, 2022, 38(12): 156-167. |
| [6] | 张健, 才恒, 申振豪, 刘仲浩, 赵娟, 张巧珍, 高强. 烟酸促进短乳杆菌合成γ-氨基丁酸的机制[J]. 生物技术通报, 2022, 38(10): 235-242. |
| [7] | 蔡国磊, 陆小凯, 娄水珠, 杨海英, 杜刚. 芽孢杆菌LM基于全基因组的分类鉴定及抑菌原理的研究[J]. 生物技术通报, 2021, 37(8): 176-185. |
| [8] | 刘娟, 朱春晓, 肖雪琼, 莫陈汨, 王高峰, 肖炎农. 淡紫紫孢菌亲环蛋白PlCYP6 互作蛋白的筛选[J]. 生物技术通报, 2021, 37(7): 137-145. |
| [9] | 田鹤, 税光厚. 基于质谱技术的代谢组学分析方法研究进展[J]. 生物技术通报, 2021, 37(1): 24-32. |
| [10] | 殷志斌, 黄文洁, 伍欣宙, 晏石娟. 空间分辨代谢组学进展和挑战[J]. 生物技术通报, 2021, 37(1): 32-51. |
| [11] | 赖博文, 刘玢, 梁永康. 基于高分辨质谱的非靶向代谢组学在食品造假鉴定中的研究进展[J]. 生物技术通报, 2019, 35(2): 192-197. |
| [12] | 叶洲辰, 吴友根, 于靖, 张军锋, 杨东梅, 胡新文. 不同产地油茶籽油提取物的抗氧化活性比较分析及其营养评价[J]. 生物技术通报, 2019, 35(10): 80-88. |
| [13] | 黄放,林向民. 嗜水气单胞菌bamA、bamB、bamD突变株的构建及其对外膜蛋白转运的影响[J]. 生物技术通报, 2018, 34(5): 148-153. |
| [14] | 乔玉玲, 黄铮, 秦海艳, 宋兰兰, 陈继军, 安晨, 叶星, 毛晓燕. 抗CD52单克隆抗体HPLC-肽图分析方法的建立[J]. 生物技术通报, 2018, 34(11): 216-222. |
| [15] | 尚攀, 付元帅, 施志仪, 喻杰, 刘素萍. 甲状腺激素受体与Drosha蛋白互作鉴定分析[J]. 生物技术通报, 2017, 33(8): 206-212. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||