








生物技术通报 ›› 2025, Vol. 41 ›› Issue (1): 240-251.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0469
寇焙森1,2,3( ), 程萌萌1,2,3, 郭雪琴1,2,3, 葛彬1,2,3, 刘迪1,2,3, 陆海1,2,3, 李慧1,2,3(
), 程萌萌1,2,3, 郭雪琴1,2,3, 葛彬1,2,3, 刘迪1,2,3, 陆海1,2,3, 李慧1,2,3( )
)
收稿日期:2024-05-20
出版日期:2025-01-26
发布日期:2025-01-22
通讯作者:
李慧,女,副教授,研究方向:树木分子生物学;E-mail: lihui830@bjfu.edu.cn作者简介:寇焙森,男,硕士研究生,研究方向:树木分子生物学;E-mail: koubeisen@163.com
基金资助:
KOU Bei-sen1,2,3( ), CHENG Meng-meng1,2,3, GUO Xue-qin1,2,3, GE Bin1,2,3, LIU Di1,2,3, LU Hai1,2,3, LI Hui1,2,3(
), CHENG Meng-meng1,2,3, GUO Xue-qin1,2,3, GE Bin1,2,3, LIU Di1,2,3, LU Hai1,2,3, LI Hui1,2,3( )
)
Received:2024-05-20
Published:2025-01-26
Online:2025-01-22
摘要:
【目的】探究组蛋白乙酰化修饰在杨树茎生长发育中的分子机制。【方法】利用组蛋白去乙酰化酶抑制剂曲古抑菌素A(trichostatin A, TSA)对84K杨树(Populus alba × P. glandulosa)进行不同时间处理,采用Western blot检测杨树茎中组蛋白乙酰化水平变化情况,然后用RNA-Seq技术对其茎中基因表达谱进行转录组分析,最后利用石蜡切片和扫描电镜观察木质部的表型变化。【结果】Western Blot 结果表明2 μmol/L TSA处理2 h能明显提高茎中组蛋白H3的乙酰化水平,随着TSA处理时间延长,组蛋白H3的乙酰化水平进一步升高。转录组结果显示,TSA处理2 h、12 h共得到5 625个差异基因表达,其中2 h上调基因2 158个,下调基因1 556个;12 h上调基因905个、下调基因1 006个。GO功能分析发现,差异上调基因主要富集在细胞壁组分,DNA结合转录因子活性;差异下调基因主要富集在光合作用、响应非生物刺激等词条;KEGG通路分析发现差异上调基因显著富集到木质素合成等通路。表型分析结果显示,相较于未处理组,TSA处理导致植株高度下降9.03%,但是茎直径和木质部厚度未见显著性差异。【结论】组蛋白H3乙酰化水平上升通过促进转录活性或细胞壁相关基因的表达来参与杨树茎的发育,从而影响植株的高度。
寇焙森, 程萌萌, 郭雪琴, 葛彬, 刘迪, 陆海, 李慧. 组蛋白去乙酰化酶抑制剂TSA处理对杨树茎生长发育的影响[J]. 生物技术通报, 2025, 41(1): 240-251.
KOU Bei-sen, CHENG Meng-meng, GUO Xue-qin, GE Bin, LIU Di, LU Hai, LI Hui. Effects of Histone Deacetylase Inhibitor TSA Treatment on the Stem Development of Poplar[J]. Biotechnology Bulletin, 2025, 41(1): 240-251.
| 样品 Simple | 原始数据Raw reads(×106) | 高质量数据Clean reads(×106) | Q20/% | Q30/% | 比对到参考基因组 Alignment to reference genome /% | 唯一参考序列数 Number of unique reference sequences/% | 比对到参考基因组 Alignment to reference genome/% | 唯一参考序列数 Number of unique reference sequences/% |
|---|---|---|---|---|---|---|---|---|
| WT0 | 47.77 | 42.32 | 97.2 | 91.3 | 96.69 | 84.51 | 96.69 | 84.51 |
| CL2 | 48.93 | 42.84 | 97.3 | 91.8 | 96.44 | 84.38 | 96.44 | 84.38 |
| CL12 | 46.02 | 42.46 | 97.1 | 91.4 | 96.98 | 85.22 | 96.98 | 85.22 |
表1 转录组测序数据基本情况
Table 1 Details of transcriptome sequencing data
| 样品 Simple | 原始数据Raw reads(×106) | 高质量数据Clean reads(×106) | Q20/% | Q30/% | 比对到参考基因组 Alignment to reference genome /% | 唯一参考序列数 Number of unique reference sequences/% | 比对到参考基因组 Alignment to reference genome/% | 唯一参考序列数 Number of unique reference sequences/% |
|---|---|---|---|---|---|---|---|---|
| WT0 | 47.77 | 42.32 | 97.2 | 91.3 | 96.69 | 84.51 | 96.69 | 84.51 |
| CL2 | 48.93 | 42.84 | 97.3 | 91.8 | 96.44 | 84.38 | 96.44 | 84.38 |
| CL12 | 46.02 | 42.46 | 97.1 | 91.4 | 96.98 | 85.22 | 96.98 | 85.22 |
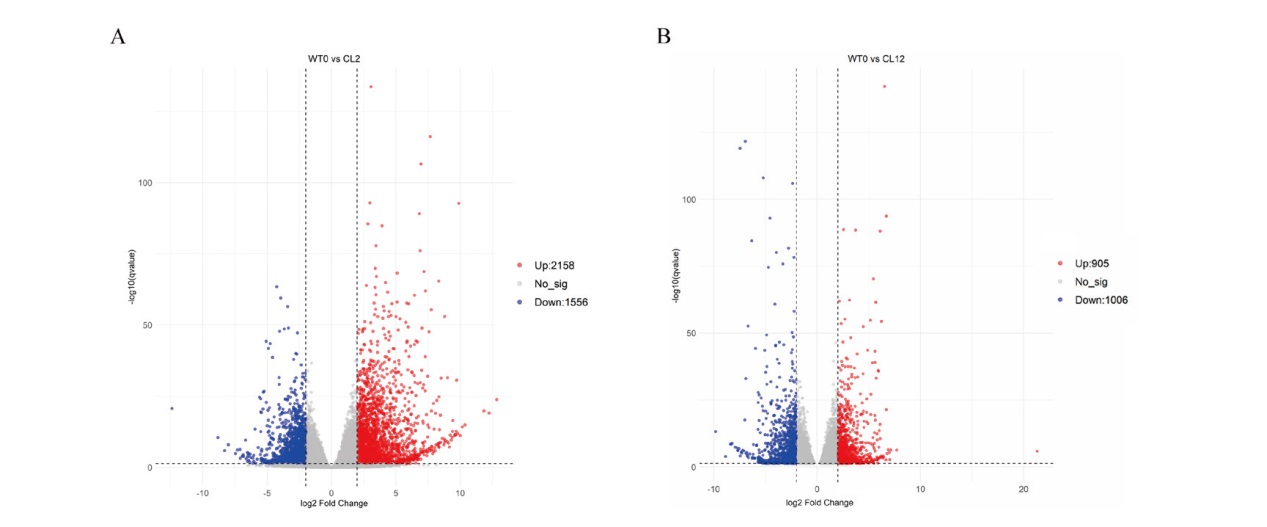
图2 差异基因数目统计 A:WT0 vs CL2差异基因火山图;B:WT0 vs CL12差异基因火山图;红点:上调差异基因;蓝点:下调差异基因;灰点:无差异基因
Fig. 2 Statistical data of DEGs A: Volcano plot of DEGs between WT0 and CL2. B: Volcano plot of DEGs between WT0 and CL12. Red dots: Up-regulated DEGs. Blue dots: Down-regulated DEGs. Gray dots: Non- DEGs
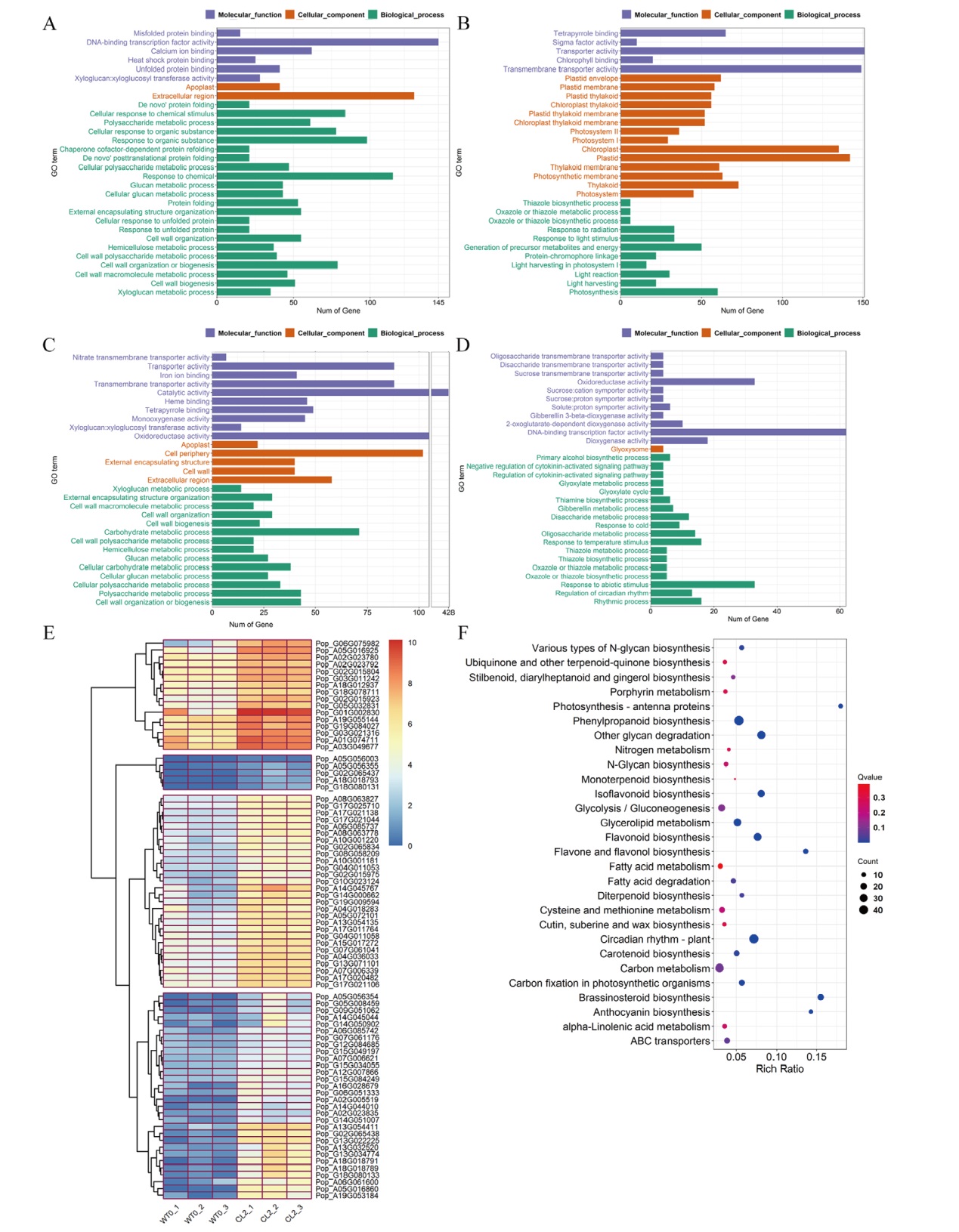
图3 差异基因GO功能富集柱状图、KEGG富集柱状图及表达量聚类热图 A-B:2 h上调(A)、下调(B)差异基因GO功能富集柱状图;C-D:12 h上调(C)、下调(D)差异基因GO功能富集柱状图;E:细胞壁发育相关差异基因表达量聚类热图;F:12 h差异上调基因KEGG富集柱状图; Go term:Go条目;Rich Ratio:富集度比率
Fig. 3 GO functional enrichment histogram, KEGG pathway enrichment histogram and expression clustering heat map of DEGs A-B: GO enrichment histogram of 2 h up-regulated(A)and down-regulated(B)DEGs. C-D: GO enrichment histogram of 12 h up-regulated(C)and down-regulated(D)DEGs. E: Cluster heat map of DEGS associated with cell wall development. F: KEGG enrichment histogram of 12 h up-regulated DEGs
| 基因ID Gene ID | log2 (CL2/WT0) | Q value (WT0-vs-CL2) | 描述 Description |
|---|---|---|---|
| Pop_A01G003964 | 5.769956725 | 1.32E-41 | ERF1A |
| Pop_A01G032260 | 5.649644852 | 1.26E-05 | ERF21 |
| Pop_A01G059811 | 5.539774145 | 5.91E-07 | ERF61 |
| Pop_A01G060003 | 7.0861912 | 1.99E-09 | DREB1E |
| Pop_A04G018519 | 3.029867059 | 3.13E-06 | bHLH162 |
| Pop_A06G061940 | 2.433933971 | 0.032353534 | ORG2 |
| Pop_A14G044066 | 2.36624488 | 0.001496264 | bZip53 |
| Pop_A08G063663 | 2.588359391 | 4.82E-13 | CDF2 |
| Pop_G19G009429 | 2.356323627 | 6.09E-16 | CDF3 |
| Pop_G09G015087 | 2.033526993 | 0.003029387 | ZAT10 |
| Pop_A16G090138 | 3.342912788 | 3.50E-13 | PHL7 |
| Pop_G10G047973 | 6.429548834 | 0.001027136 | ORR25 |
| Pop_A09G014447 | 2.372666749 | 6.59E-06 | SCR3 |
| Pop_A06G062082 | 6.325653816 | 0.001158459 | HSF4b |
| Pop_A16G055308 | 2.667183627 | 5.18E-06 | HSF3 |
| Pop_A02G066179 | 5.110284639 | 8.26E-05 | MYB73 |
| Pop_A07G011878 | 2.013444428 | 1.37E-12 | MYB44 |
| Pop_A08G063813 | 2.04957107 | 0.013470953 | MYB108 |
| Pop_A14G000372 | 3.81186488 | 1.23E-08 | MYB44 |
| Pop_A15G064557 | 4.679506153 | 1.94E-16 | MYB62 |
| Pop_A04G019588 | 2.206753814 | 6.93E-06 | NAC73 |
| Pop_A04G026457 | 3.711825186 | 0.000111049 | TCP13 |
| Pop_A01G059075 | 3.408636654 | 6.29E-05 | WRKY28 |
| Pop_A01G080041 | 3.413796772 | 6.86E-30 | WRKY40 |
| Pop_A02G012299 | 2.225001206 | 0.013563496 | WRKY22 |
| Pop_A03G019779 | 2.390109471 | 0.001165373 | WRKY75 |
表2 转录因子相关部分差异基因功能分析
Table 2 Functional analysis of transcription factor-related DEGs
| 基因ID Gene ID | log2 (CL2/WT0) | Q value (WT0-vs-CL2) | 描述 Description |
|---|---|---|---|
| Pop_A01G003964 | 5.769956725 | 1.32E-41 | ERF1A |
| Pop_A01G032260 | 5.649644852 | 1.26E-05 | ERF21 |
| Pop_A01G059811 | 5.539774145 | 5.91E-07 | ERF61 |
| Pop_A01G060003 | 7.0861912 | 1.99E-09 | DREB1E |
| Pop_A04G018519 | 3.029867059 | 3.13E-06 | bHLH162 |
| Pop_A06G061940 | 2.433933971 | 0.032353534 | ORG2 |
| Pop_A14G044066 | 2.36624488 | 0.001496264 | bZip53 |
| Pop_A08G063663 | 2.588359391 | 4.82E-13 | CDF2 |
| Pop_G19G009429 | 2.356323627 | 6.09E-16 | CDF3 |
| Pop_G09G015087 | 2.033526993 | 0.003029387 | ZAT10 |
| Pop_A16G090138 | 3.342912788 | 3.50E-13 | PHL7 |
| Pop_G10G047973 | 6.429548834 | 0.001027136 | ORR25 |
| Pop_A09G014447 | 2.372666749 | 6.59E-06 | SCR3 |
| Pop_A06G062082 | 6.325653816 | 0.001158459 | HSF4b |
| Pop_A16G055308 | 2.667183627 | 5.18E-06 | HSF3 |
| Pop_A02G066179 | 5.110284639 | 8.26E-05 | MYB73 |
| Pop_A07G011878 | 2.013444428 | 1.37E-12 | MYB44 |
| Pop_A08G063813 | 2.04957107 | 0.013470953 | MYB108 |
| Pop_A14G000372 | 3.81186488 | 1.23E-08 | MYB44 |
| Pop_A15G064557 | 4.679506153 | 1.94E-16 | MYB62 |
| Pop_A04G019588 | 2.206753814 | 6.93E-06 | NAC73 |
| Pop_A04G026457 | 3.711825186 | 0.000111049 | TCP13 |
| Pop_A01G059075 | 3.408636654 | 6.29E-05 | WRKY28 |
| Pop_A01G080041 | 3.413796772 | 6.86E-30 | WRKY40 |
| Pop_A02G012299 | 2.225001206 | 0.013563496 | WRKY22 |
| Pop_A03G019779 | 2.390109471 | 0.001165373 | WRKY75 |
| 基因ID Gene ID | log2(CL12/WT0) | Q value | 描述 Description |
|---|---|---|---|
| Pop_A02G023792 | 1.224720934 | 1.09E-10 | GAUT8 |
| Pop_G02G015804 | 1.084322434 | 1.96E-08 | GAUT8 |
| Pop_G18G078711 | 1.659192116 | 3.08E-11 | CesA1 |
| Pop_A18G012937 | 1.547823248 | 7.46E-12 | CesA1 |
| Pop_A03G049677 | 2.0312488 | 1.97E-08 | PAE7 |
| Pop_A05G016925 | 1.624518546 | 3.92E-06 | PAE12 |
| Pop_G03G021316 | 1.771860445 | 3.93E-08 | PAE7 |
| Pop_G05G032831 | 1.404050162 | 4.41E-07 | PAE12 |
| Pop_G03G011242 | 1.373107821 | 4.15E-14 | PT |
| Pop_A02G023780 | 2.640491502 | 1.58E-21 | XTH2 |
| Pop_A19G055144 | 0.555458286 | 0.19872077 | XTH9 |
| Pop_G01G002830 | 2.869514357 | 0.070436159 | XTH27 |
| Pop_G02G015923 | 3.12618234 | 4.66E-63 | XTH21 |
| Pop_A01G074711 | 2.335379968 | 0.044535379 | XTHA |
| Pop_G19G084027 | 0.768927412 | 0.025571895 | XTH9 |
| Pop_G06G075982 | 1.258093365 | 0.194655673 | XTH6/TCH4 |
表3 细胞壁发育相关部分差异基因功能分析
Table 3 Functional analysis of cell wall development-related DEGs
| 基因ID Gene ID | log2(CL12/WT0) | Q value | 描述 Description |
|---|---|---|---|
| Pop_A02G023792 | 1.224720934 | 1.09E-10 | GAUT8 |
| Pop_G02G015804 | 1.084322434 | 1.96E-08 | GAUT8 |
| Pop_G18G078711 | 1.659192116 | 3.08E-11 | CesA1 |
| Pop_A18G012937 | 1.547823248 | 7.46E-12 | CesA1 |
| Pop_A03G049677 | 2.0312488 | 1.97E-08 | PAE7 |
| Pop_A05G016925 | 1.624518546 | 3.92E-06 | PAE12 |
| Pop_G03G021316 | 1.771860445 | 3.93E-08 | PAE7 |
| Pop_G05G032831 | 1.404050162 | 4.41E-07 | PAE12 |
| Pop_G03G011242 | 1.373107821 | 4.15E-14 | PT |
| Pop_A02G023780 | 2.640491502 | 1.58E-21 | XTH2 |
| Pop_A19G055144 | 0.555458286 | 0.19872077 | XTH9 |
| Pop_G01G002830 | 2.869514357 | 0.070436159 | XTH27 |
| Pop_G02G015923 | 3.12618234 | 4.66E-63 | XTH21 |
| Pop_A01G074711 | 2.335379968 | 0.044535379 | XTHA |
| Pop_G19G084027 | 0.768927412 | 0.025571895 | XTH9 |
| Pop_G06G075982 | 1.258093365 | 0.194655673 | XTH6/TCH4 |
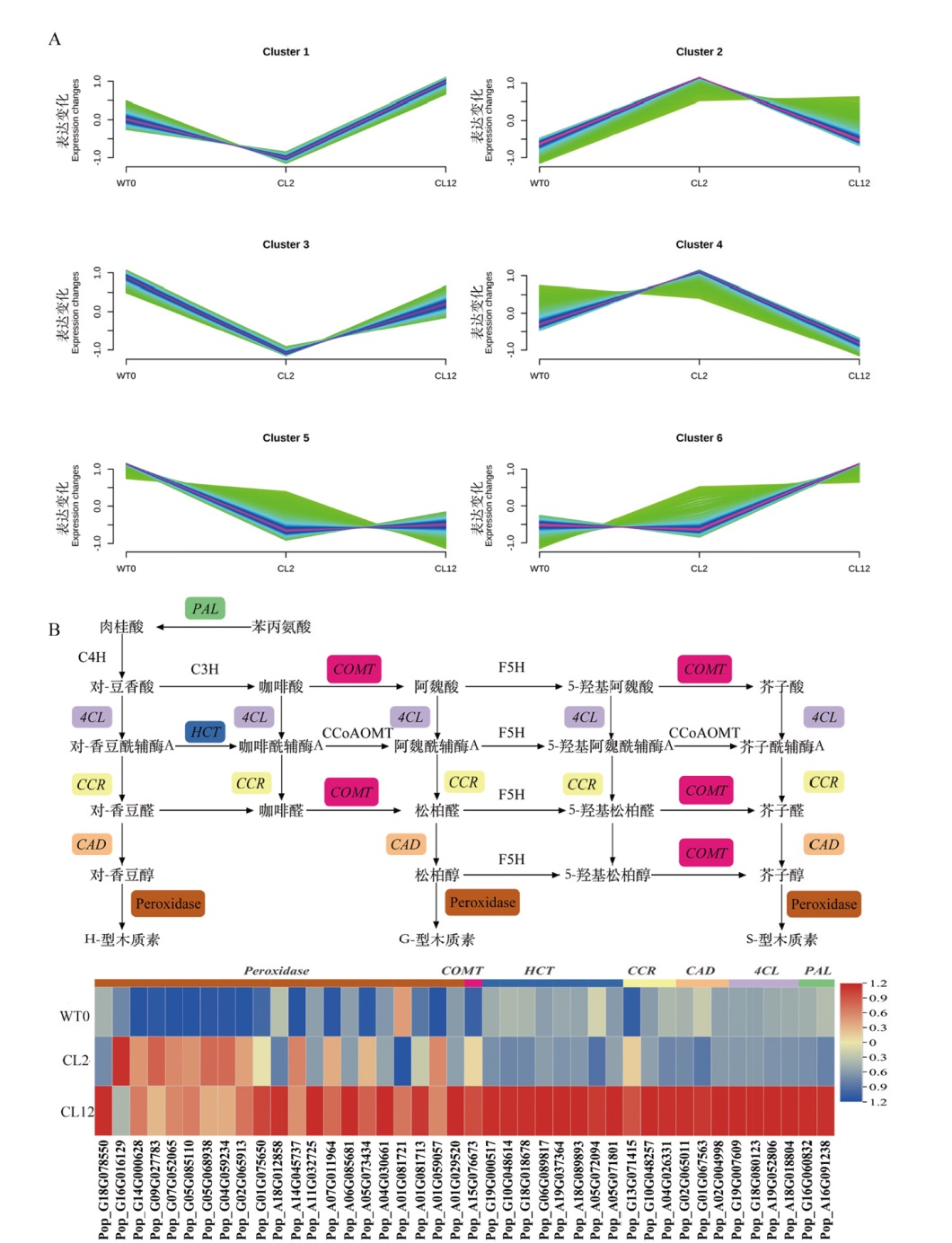
图4 差异基因时间聚类及苯丙烷通路简图 A:时间聚类分析;B:苯丙烷通路图(框体标注代表差异表达基因);WT0:对照组;CL2:TSA处理2 h;CL12:TSA处理12 h
Fig. 4 DEGs temporal clustering and schematic representation of the phenylpropanoid pathway A: DEGs temporal clustering. B: Schematic representation of the phenylpropanoid pathway(Black boxes indicate genes encoding enzymes that are differentially expressed). WT0: Control. CL2:TSA treatment for 2 h. CL12:TSA treatment for 12 h
| 基因ID Gene ID | log2(CL12/WT0) | Q value | 描述 Description |
|---|---|---|---|
| Pop_A19G052806 | 2.403845576 | 4.60E-20 | 4CL2 |
| Pop_A18G018804 | 2.667476561 | 2.32E-10 | 4CL4 |
| Pop_G19G007609 | 3.667350303 | 2.16E-23 | 4CL2 |
| Pop_G18G080123 | 2.367966412 | 1.40E-07 | 4CL4 |
| Pop_A02G004998 | 3.561801582 | 7.11E-17 | CAD3 |
| Pop_G01G067563 | 1.220128387 | 0.02149788 | CAD9 |
| Pop_G02G065011 | 2.056960752 | 2.93E-12 | CAD3 |
| Pop_G10G048257 | 2.444394374 | 5.10E-14 | CCR3 |
| Pop_A04G026331 | 1.602900842 | 0.002526309 | CCR10 |
| Pop_G13G071415 | 3.465894044 | 6.89E-07 | CCR2 |
| Pop_A15G076673 | 3.006325753 | 1.29E-06 | COMT3 |
| Pop_A18G089893 | 2.322128861 | 0.000122309 | HCT |
| Pop_G06G089817 | 4.974228282 | 6.97E-05 | HCT |
| Pop_A19G037364 | 2.298057613 | 0.000168854 | HCT |
| Pop_G10G048614 | 2.130717747 | 1.18E-08 | HCT |
| Pop_G18G018678 | 2.236704931 | 0.078108456 | HCT |
| Pop_A05G071801 | 3.065678506 | 3.14E-07 | HCT1 |
| Pop_G19G000517 | 1.836359474 | 2.75E-05 | HCT |
| Pop_A05G072094 | 1.301262613 | 0.407703359 | HCT |
| Pop_A16G091238 | 1.580246846 | 6.22E-05 | PAL |
| Pop_G16G060832 | 2.402572772 | 4.33E-12 | PAL |
| Pop_A01G029520 | 3.361672474 | 0.005884321 | Peroxidase |
| Pop_A14G045737 | 3.712025804 | 5.26E-06 | Peroxidase |
| Pop_G01G075650 | 2.826658953 | 0.006729353 | Peroxidase |
| Pop_G04G059234 | 1.493633494 | 5.56E-06 | Peroxidase |
| Pop_A07G011964 | 3.400742219 | 5.98E-09 | Peroxidase |
| Pop_G02G065913 | 2.501334426 | 0.013145367 | Peroxidase |
| Pop_A06G085681 | 2.902707699 | 0.003229047 | Peroxidase |
| Pop_A04G030661 | 2.73324656 | 1.12E-05 | Peroxidase |
| Pop_G07G052065 | 3.735543904 | 1.65E-06 | Peroxidase |
| Pop_G14G000628 | 2.779869353 | 0.006974024 | Peroxidase |
| Pop_G16G016129 | 0.785917676 | 0.479463762 | Peroxidase |
| Pop_G05G068938 | 1.80032676 | 1.21E-06 | Peroxidase |
| Pop_A01G059057 | 1.923239289 | 5.70E-28 | Peroxidase |
| Pop_G09G027783 | 1.931046589 | 6.19E-07 | Peroxidase |
| Pop_G05G085110 | 2.311933723 | 1.03E-11 | Peroxidase |
| Pop_A05G073434 | 2.266091449 | 1.44E-08 | Peroxidase |
| Pop_G18G078550 | 2.34410638 | 1.22E-13 | Peroxidase |
| Pop_A18G012858 | 1.673704695 | 3.66E-08 | Peroxidase |
| Pop_A01G081713 | 6.43779021 | 0.000215695 | Peroxidase |
| Pop_A01G081721 | 0.235613668 | 0.858752785 | Peroxidase |
| Pop_A11G032725 | 3.074623654 | 4.13E-15 | Peroxidase |
表4 苯丙烷合成相关差异基因功能分析
Table 4 Function analysis of DEGs involved in phenylpropanoid biosynthesis
| 基因ID Gene ID | log2(CL12/WT0) | Q value | 描述 Description |
|---|---|---|---|
| Pop_A19G052806 | 2.403845576 | 4.60E-20 | 4CL2 |
| Pop_A18G018804 | 2.667476561 | 2.32E-10 | 4CL4 |
| Pop_G19G007609 | 3.667350303 | 2.16E-23 | 4CL2 |
| Pop_G18G080123 | 2.367966412 | 1.40E-07 | 4CL4 |
| Pop_A02G004998 | 3.561801582 | 7.11E-17 | CAD3 |
| Pop_G01G067563 | 1.220128387 | 0.02149788 | CAD9 |
| Pop_G02G065011 | 2.056960752 | 2.93E-12 | CAD3 |
| Pop_G10G048257 | 2.444394374 | 5.10E-14 | CCR3 |
| Pop_A04G026331 | 1.602900842 | 0.002526309 | CCR10 |
| Pop_G13G071415 | 3.465894044 | 6.89E-07 | CCR2 |
| Pop_A15G076673 | 3.006325753 | 1.29E-06 | COMT3 |
| Pop_A18G089893 | 2.322128861 | 0.000122309 | HCT |
| Pop_G06G089817 | 4.974228282 | 6.97E-05 | HCT |
| Pop_A19G037364 | 2.298057613 | 0.000168854 | HCT |
| Pop_G10G048614 | 2.130717747 | 1.18E-08 | HCT |
| Pop_G18G018678 | 2.236704931 | 0.078108456 | HCT |
| Pop_A05G071801 | 3.065678506 | 3.14E-07 | HCT1 |
| Pop_G19G000517 | 1.836359474 | 2.75E-05 | HCT |
| Pop_A05G072094 | 1.301262613 | 0.407703359 | HCT |
| Pop_A16G091238 | 1.580246846 | 6.22E-05 | PAL |
| Pop_G16G060832 | 2.402572772 | 4.33E-12 | PAL |
| Pop_A01G029520 | 3.361672474 | 0.005884321 | Peroxidase |
| Pop_A14G045737 | 3.712025804 | 5.26E-06 | Peroxidase |
| Pop_G01G075650 | 2.826658953 | 0.006729353 | Peroxidase |
| Pop_G04G059234 | 1.493633494 | 5.56E-06 | Peroxidase |
| Pop_A07G011964 | 3.400742219 | 5.98E-09 | Peroxidase |
| Pop_G02G065913 | 2.501334426 | 0.013145367 | Peroxidase |
| Pop_A06G085681 | 2.902707699 | 0.003229047 | Peroxidase |
| Pop_A04G030661 | 2.73324656 | 1.12E-05 | Peroxidase |
| Pop_G07G052065 | 3.735543904 | 1.65E-06 | Peroxidase |
| Pop_G14G000628 | 2.779869353 | 0.006974024 | Peroxidase |
| Pop_G16G016129 | 0.785917676 | 0.479463762 | Peroxidase |
| Pop_G05G068938 | 1.80032676 | 1.21E-06 | Peroxidase |
| Pop_A01G059057 | 1.923239289 | 5.70E-28 | Peroxidase |
| Pop_G09G027783 | 1.931046589 | 6.19E-07 | Peroxidase |
| Pop_G05G085110 | 2.311933723 | 1.03E-11 | Peroxidase |
| Pop_A05G073434 | 2.266091449 | 1.44E-08 | Peroxidase |
| Pop_G18G078550 | 2.34410638 | 1.22E-13 | Peroxidase |
| Pop_A18G012858 | 1.673704695 | 3.66E-08 | Peroxidase |
| Pop_A01G081713 | 6.43779021 | 0.000215695 | Peroxidase |
| Pop_A01G081721 | 0.235613668 | 0.858752785 | Peroxidase |
| Pop_A11G032725 | 3.074623654 | 4.13E-15 | Peroxidase |

图6 TSA处理杨树植株表型分析 A:未处理(左)及TSA处理(右)杨树植株;B:植株高度统计;C-N:石蜡切片分析杨树茎的横切面,C、D、E为对照植株第18茎节,F、G、H为TSA处理植株第18茎节,I、J、K为对照植株第16茎节,L、M、N为TSA处理植株第16茎节,C、F、I、L标尺为200 μm,D、G、J、M标尺为100 μm,E、H、K、N标尺为50 μm;O:植株茎直径统计;P:植株木质部厚度统计;*P<0.05;**P<0.01
Fig. 6 Phenotypic analysis of poplar plants treated with TSA A: Untreated(left)and TSA-treated(right)poplar plants. B: Statistics of plant heights. C-N: Paraffin section analysis of poplar stem cross-sections. C-E: The 18th stem segment of control plants. F-H: The 18th stem segment of TSA-treated plants. I-K: The 16th stem segment of control plants. L-N: The 16th stem segment of TSA-treated plants; C, F, I, L scale bar = 200 μm; D, G, J, M scale bar=100 μm; E, H, K, N scale bar=50 μm. O: Statistics of plant stem diameter. P: Statistics of plant xylem thickness. *P<0.05, **P<0.01

图7 扫描电镜分析杨树木质部表型 A-D:扫描电镜下杨树木质部结构,A、B为未处理组,C、D为TSA处理组,A、C标尺为500 μm,B、D标尺为100 μm;E:导管面积统计;F:导管数量统计
Fig. 7 SEM analysis of the xylem phenotype in poplar A-D: SEM analysis of the xylem structure in poplar. A, B: Untreated group; C, D: TSA-treated group. A, C scale bars = 500 μm; B, D scale bars =100 μm. E: Statistics of vessel area. F: Statistics of vessel number
| [1] | 张冰, 夏德安, 马旭俊. 组蛋白去乙酰化酶在杨树根再生和生长中的功能[J]. 江苏农业科学, 2017, 45(5): 40-43. |
| Zhang B, Xia DA, Ma XJ. Function of histone deacetylase in regeneration and growth of Yang Shugen[J]. Jiangsu Agric Sci, 2017, 45(5): 40-43. | |
| [2] |
Yang XJ, Seto E. HATs and HDACs: from structure, function and regulation to novel strategies for therapy and prevention[J]. Oncogene, 2007, 26(37): 5310-5318.
doi: 10.1038/sj.onc.1210599 pmid: 17694074 |
| [3] |
Dangl M, Brosch G, Haas H, et al. Comparative analysis of HD2 type histone deacetylases in higher plants[J]. Planta, 2001, 213(2): 280-285.
pmid: 11469594 |
| [4] | Servet C, Silva NCE, et al. Histone acetyltransferase AtGCN5/HAG1 is a versatile regulator of developmental and inducible gene expression in Arabidopsis[J]. Mol Plant, 2010, 3(4): 670-677. |
| [5] | 李淑娟, 张超, 张彦妮, 等. 毛果杨HDAC基因家族序列及其表达分析[J]. 西北农林科技大学学报: 自然科学版, 2015, 43(3): 63-76. |
| Li SJ, Zhang C, Zhang YN, et al. Sequence and gene expression of histone deacetylases(HDAC)gene family of Populus trichocar-pa[J]. J Northwest A F Univ Nat Sci Ed, 2015, 43(3): 63-76. | |
| [6] |
Chen XS, Ding AB, et al. Functions and mechanisms of plant histone deacetylases[J]. Sci China Life Sci, 2020, 63(2): 206-216.
doi: 10.1007/s11427-019-1587-x pmid: 31879846 |
| [7] |
Krogan NT, Hogan K, Long JA. APETALA2 negatively regulates multiple floral organ identity genes in Arabidopsis by recruiting the co-repressor TOPLESS and the histone deacetylase HDA19[J]. Development, 2012, 139(22): 4180-4190.
doi: 10.1242/dev.085407 pmid: 23034631 |
| [8] | Yu CW, Liu XC, Luo M, et al. HISTONE DEACETYLASE6 interacts with FLOWERING LOCUS D and regulates flowering in Arabidop-sis[J]. Plant Physiol, 2011, 156(1): 173-184. |
| [9] | Yu CW, Chang KY, Wu KQ. Genome-wide analysis of gene regulatory networks of the FVE-HDA6-FLD complex in Arabidopsis[J]. Front Plant Sci, 2016, 7: 555. |
| [10] | Kang MJ, Jin HS, Noh YS, et al. Repression of flowering under a noninductive photoperiod by the HDA9-AGL19-FT module in Arabidopsis[J]. New Phytol, 2015, 206(1): 281-294. |
| [11] | Luo M, Yu CW, Chen FF, et al. Histone deacetylase HDA6 is functionally associated with AS1 in repression of KNOX genes in Arabidopsis[J]. PLoS Genet, 2012, 8(12): e1003114. |
| [12] | Chen XS, Lu L, Mayer KS, et al. POWERDRESS interacts with HISTONE DEACETYLASE 9 to promote aging in Arabidop-sis[J]. eLife, 2016, 5: e17214. |
| [13] | Li HC, Torres-Garcia J, Latrasse D, et al. Plant-specific histone deacetylases HDT1/2 regulate GIBBERELLIN 2-OXIDASE2 expression to control Arabidopsis root meristem cell number[J]. Plant Cell, 2017, 29(9): 2183-2196. |
| [14] | Chung PJ, Kim YS, et al. The histone deacetylase OsHDAC1 epigenetically regulates the OsNAC6 gene that controls seedling root growth in rice[J]. Plant J, 2009, 59(5): 764-776. |
| [15] |
Luo M, Cheng K, et al. Plant responses to abiotic stress regulated by histone deacetylases[J]. Front Plant Sci, 2017, 8: 2147.
doi: 10.3389/fpls.2017.02147 pmid: 29326743 |
| [16] | Lee HG, Seo PJ. MYB96 recruits the HDA15 protein to suppress negative regulators of ABA signaling in Arabidopsis[J]. Nat Commun, 2019, 10(1): 1713. |
| [17] | Shen Y, Lei TT, Cui XY, et al. Arabidopsis histone deacetylase HDA15 directly represses plant response to elevated ambient temperature[J]. Plant J, 2019, 100(5): 991-1006. |
| [18] | To TK, Nakaminami K, Kim JM, et al. Arabidopsis HDA6 is required for freezing tolerance[J]. Biochem Biophys Res Commun, 2011, 406(3): 414-419. |
| [19] | Dai XF, Zhai R, Lin JJ, et al. Cell-type-specific PtrWOX4a and PtrVCS2 form a regulatory nexus with a histone modification system for stem cambium development in Populus trichocarpa[J]. Nat Plants, 2023, 9(1): 96-111. |
| [20] | Zhang YZ, Yin B, Zhang JX, et al. Histone deacetylase HDT1 is involved in stem vascular development in Arabidopsis[J]. Int J Mol Sci, 2019, 20(14): 3452. |
| [21] | Hirai R, Wang SM, Demura T, et al. Histone deacetylation controls xylem vessel cell differentiation via transcriptional regulation of a transcription repressor complex OFP1/4-MYB75-KNAT7-BLH6[J]. Front Plant Sci, 2022, 12: 825810. |
| [22] | Li S, Lin YCJ, et al. The AREB1 transcription factor influences histone acetylation to regulate drought responses and tolerance in Populus trichocarpa[J]. Plant Cell, 2019, 31(3): 663-686. |
| [23] | Nowak K, Morończyk J, Grzyb M, et al. miR172 regulates WUS during somatic embryogenesis in Arabidopsis via AP2[J]. Cells, 2022, 11(4): 718. |
| [24] |
Zheng Y, Ge JY, Bao C, et al. Histone deacetylase HDA9 and WRKY53 transcription factor are mutual antagonists in regulation of plant stress response[J]. Mol Plant, 2020, 13(4): 598-611.
doi: S1674-2052(19)30408-3 pmid: 31891777 |
| [25] | Gao SY, Yin MX, Xu MY, et al. Transcription factors PuPRE6/PuMYB12 and histone deacetylase PuHDAC9-like regulate sucrose levels in pear[J]. Plant Physiol, 2024, 194(3): 1577-1592. |
| [26] | 李慧, 郭晓蕊, 刘雅琳, 等. 木材形成过程中次生壁沉积和细胞程序性死亡的分子调控机制[J]. 中国科学: 生命科学, 2020, 50(2): 123-135. |
| Li H, Guo XR, Liu YL, et al. The molecular mechanism in secondary wall deposition and programmed cell death of wood formation[J]. Sci Sin Vitae, 2020, 50(2): 123-135. | |
| [27] | Shahin L, Zhang L, Mohnen D, et al. Insights into pectin O-acetylation in the plant cell wall: structure, synthesis, and modification[J]. Cell Surf, 2023, 9: 100099. |
| [28] |
Fan NN, Su LT, Lv AM, et al. PECTIN ACETYLESTERASE12 regulates shoot branching via acetic acid and auxin accumulation in alfalfa shoots[J]. Plant Physiol, 2024, 195(1): 518-533.
doi: 10.1093/plphys/kiae071 pmid: 38365203 |
| [29] | Hrmova M, Stratilová B, et al. Broad specific xyloglucan: xyloglucosyl transferases are formidable players in the re-modelling of plant cell wall structures[J]. Int J Mol Sci, 2022, 23(3): 1656. |
| [30] |
Zhong RQ, Cui DT, Ye ZH. Secondary cell wall biosynthesis[J]. New Phytol, 2019, 221(4): 1703-1723.
doi: 10.1111/nph.15537 pmid: 30312479 |
| [31] | 李金花, 张绮纹, 牛正田, 等. 木质素生物合成及其基因调控的研究进展[J]. 世界林业研究, 2007, 20(1): 29-37. |
| Li JH, Zhang QW, Niu ZT, et al. Advances in study of lignin biosynthesis and genetic engineering modification[J]. World For Res, 2007, 20(1): 29-37. |
| [1] | 裴旭娟, 狄靖宜, 刘浩, 高伟霞. 基于转录组分析挖掘兽疫链球菌透明质酸分子量调控元件[J]. 生物技术通报, 2025, 41(1): 347-356. |
| [2] | 岳丽昕, 王清华, 刘泽洲, 孔素萍, 高莉敏. 基于转录组和WGCNA筛选大葱雄性不育相关基因[J]. 生物技术通报, 2024, 40(9): 212-224. |
| [3] | 聂祝欣, 郭瑾, 乔子洋, 李微薇, 张学燕, 刘春阳, 王静. 黑果枸杞不同发育时期果实花色苷合成的转录组分析[J]. 生物技术通报, 2024, 40(8): 106-117. |
| [4] | 周麟, 黄顺满, 苏文坤, 姚响, 屈燕. 滇山茶bHLH基因家族鉴定及花色形成相关基因筛选[J]. 生物技术通报, 2024, 40(8): 142-151. |
| [5] | 王睿, 戚继. 整合组织学图像信息增强空间转录组细胞聚类的分辨率[J]. 生物技术通报, 2024, 40(8): 39-46. |
| [6] | 廖杨梅, 赵国春, 翁学煌, 贾黎明, 陈仲. 无患子雄性不育品种‘琦蕊’不同发育时期雄花转录组分析[J]. 生物技术通报, 2024, 40(7): 197-206. |
| [7] | 高萌萌, 赵天宇, 焦馨悦, 林春晶, 关哲允, 丁孝羊, 孙妍妍, 张春宝. 大豆细胞质雄性不育系及其恢复系的比较转录组分析[J]. 生物技术通报, 2024, 40(7): 137-149. |
| [8] | 白志元, 徐菲, 杨午, 王明贵, 杨玉花, 张海平, 张瑞军. 大豆细胞质雄性不育弱恢复型杂种F1育性转变的转录组分析[J]. 生物技术通报, 2024, 40(6): 134-142. |
| [9] | 秦健, 李振月, 何浪, 李俊玲, 张昊, 杜荣. 肌源性细胞分化的单细胞转录谱变化及细胞间通讯分析[J]. 生物技术通报, 2024, 40(6): 330-342. |
| [10] | 吴迪, 游小凤, 郑亦铮, 林楠, 张燕燕, 魏艺聪. 草珊瑚中类胡萝卜素合成的内源激素调控机制分析[J]. 生物技术通报, 2024, 40(5): 203-214. |
| [11] | 郭纯, 宋桂梅, 闫艳, 邸鹏, 王英平. 西洋参bZIP基因家族全基因组鉴定和表达分析[J]. 生物技术通报, 2024, 40(4): 167-178. |
| [12] | 钟匀, 林春, 刘正杰, 董陈文华, 毛自朝, 李兴玉. 芦笋皂苷合成相关糖基转移酶基因克隆及原核表达分析[J]. 生物技术通报, 2024, 40(4): 255-263. |
| [13] | 杨淇, 魏子迪, 宋娟, 童堃, 杨柳, 王佳涵, 刘海燕, 栾维江, 马轩. 水稻组蛋白H1三突变体的创建和转录组学分析[J]. 生物技术通报, 2024, 40(4): 85-96. |
| [14] | 谢倩, 江来, 贺进, 刘玲玲, 丁明月, 陈清西. 不同鲜食品质橄榄果实转录组测序及酚类代谢途径相关调控基因挖掘[J]. 生物技术通报, 2024, 40(3): 215-228. |
| [15] | 梁佳林, 赵爽, 李幸儿, 赵成周, 李萍. 尼泊尔黄堇Aux/IAA基因家族的鉴定与UVB处理下表达模式分析[J]. 生物技术通报, 2024, 40(12): 182-192. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||