








生物技术通报 ›› 2025, Vol. 41 ›› Issue (10): 303-312.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0488
梁欣敏1( ), 崔玉琴1,2, 雷梦婷1,3, 韩婧1, 贾定洪1, 王波1, 彭卫红1,2, 何晓兰1,3, 刘询1(
), 崔玉琴1,2, 雷梦婷1,3, 韩婧1, 贾定洪1, 王波1, 彭卫红1,2, 何晓兰1,3, 刘询1( )
)
收稿日期:2025-05-13
出版日期:2025-10-26
发布日期:2025-10-28
通讯作者:
刘询,男,博士,助理研究员,研究方向 :食用菌遗传育种;E-mail: liuxun0712@163.com作者简介:梁欣敏,女,硕士,研究方向 :食用菌遗传育种;E-mail: liangxinmin_lxm@163.com
基金资助:
LIANG Xin-min1( ), CUI Yu-qin1,2, LEI Meng-ting1,3, HAN Jing1, JIA Ding-hong1, WANG Bo1, PENG Wei-hong1,2, HE Xiao-lan1,3, LIU Xun1(
), CUI Yu-qin1,2, LEI Meng-ting1,3, HAN Jing1, JIA Ding-hong1, WANG Bo1, PENG Wei-hong1,2, HE Xiao-lan1,3, LIU Xun1( )
)
Received:2025-05-13
Published:2025-10-26
Online:2025-10-28
摘要:
目的 在金针菇(Flammulina filiformis)基因组中鉴定酪氨酸酶(TYR)和漆酶(Lac)基因家族成员,分析TYR和Lac家族基因的组织表达模式及在不同颜色金针菇菌盖表皮中的相对表达水平,为研究TYR和Lac基因在金针菇颜色差异调控中的功能提供理论基础。 方法 基于基因组数据系统鉴定金针菇TYR和Lac基因家族成员,利用实时荧光定量PCR(RT-qPCR)技术分析基因在不同颜色菌株(白色、黄色和褐色)菌柄和菌盖表皮组织的表达水平,并对酶活性进行测定。 结果 在金针菇基因组中共鉴定到3个TYR基因和11个Lac基因,编码蛋白均具有该家族典型保守结构域和保守基序,并与其他食用菌的相同基因家族成员亲缘关系较近。RT-qPCR分析显示,TYR3、Lac6和Lac11基因则在不同颜色菌株菌盖表皮的表达水平均高于菌柄表皮组织;Lac2在颜色较深的褐色菌盖表皮中显著高表达,而在白色和黄色菌株相对较低。进一步测定酶活性显示,白色菌盖表皮组织中的TYR酶活性显著高于褐色和黄色菌株;而Lac酶活性则在褐色菌盖表皮组织更高,其次是黄色,白色菌株中Lac酶活最低。 结论 在金针菇中鉴定出3个TYR和11个Lac家族基因;其中,漆酶家族成员可能通过催化酚类物质氧化聚合参与金针菇黑色素合成,导致金针菇菌盖呈现褐色表型。
梁欣敏, 崔玉琴, 雷梦婷, 韩婧, 贾定洪, 王波, 彭卫红, 何晓兰, 刘询. 酪氨酸酶和漆酶基因在不同颜色金针菇中的表达模式[J]. 生物技术通报, 2025, 41(10): 303-312.
LIANG Xin-min, CUI Yu-qin, LEI Meng-ting, HAN Jing, JIA Ding-hong, WANG Bo, PENG Wei-hong, HE Xiao-lan, LIU Xun. Expression Patterns of Tyrosinase and Laccase Genes in Flammulina filiformis with Different Colors[J]. Biotechnology Bulletin, 2025, 41(10): 303-312.
基因 Gene | 正向引物 Forward primer (5′-3′) | 反向引物 Reverse primer (5′-3′) |
|---|---|---|
| GAPDH | CCTCTGCTCACTTGAAGGGT | GCGTTGGAGATGACTTTGAA |
| TYR1 | AGGTGTGGTGCATCCAGAAG | GAACCAGAAGGACTTCGCCA |
| TYR2 | CGACCGCATGCTCTCACTAT | GGTGTCAACGGGGTATTGGT |
| TYR3 | GTACTGGAACGAACTCCCCG | CTTCCCCTGGGCCTAAGTTG |
| Lac1 | GGCCATTATGTCGACGGTCT | ACCAATCGCCAAGAACGACT |
| Lac2 | TCTACGATCCCGACGATCCA | GGCCGTTGATCAGGATAGCA |
| Lac3 | GTTTGCGAAGACTTTGGCGT | GACATGCTGACCAGACGGAA |
| Lac4 | ACCTGCGACGTTGAGAATGT | TGGGGTGCTCATCAATGGAC |
| Lac5 | CATCCATCGGCCTAACCGAA | TTTTCCCGCGACTACGTTGA |
| Lac6 | AGTGCAAGGAACTCGCTACC | TGAGTGTTGATCCCATCGGC |
| Lac7 | TTGTCCATGCCAATCAGCCT | GCGTCAGGAGCACCTTCATA |
| Lac8 | CGCTACCGCTTCCGGATTAT | TGGCTGCACGTTGATACCAT |
| Lac9 | CCAACGCTACTCCCTCATCC | ACATAATGCAGCACACCCGA |
| Lac10 | ATCTCCTGCGACCCAAACTG | ACTGAATCGACGAGAAGCGG |
| Lac11 | GGACCGACATCGAAGAGTCC | GCCCAGTTTGCGTTGGTATC |
表1 本研究使用的引物列表
Table 1 Primers used in this study
基因 Gene | 正向引物 Forward primer (5′-3′) | 反向引物 Reverse primer (5′-3′) |
|---|---|---|
| GAPDH | CCTCTGCTCACTTGAAGGGT | GCGTTGGAGATGACTTTGAA |
| TYR1 | AGGTGTGGTGCATCCAGAAG | GAACCAGAAGGACTTCGCCA |
| TYR2 | CGACCGCATGCTCTCACTAT | GGTGTCAACGGGGTATTGGT |
| TYR3 | GTACTGGAACGAACTCCCCG | CTTCCCCTGGGCCTAAGTTG |
| Lac1 | GGCCATTATGTCGACGGTCT | ACCAATCGCCAAGAACGACT |
| Lac2 | TCTACGATCCCGACGATCCA | GGCCGTTGATCAGGATAGCA |
| Lac3 | GTTTGCGAAGACTTTGGCGT | GACATGCTGACCAGACGGAA |
| Lac4 | ACCTGCGACGTTGAGAATGT | TGGGGTGCTCATCAATGGAC |
| Lac5 | CATCCATCGGCCTAACCGAA | TTTTCCCGCGACTACGTTGA |
| Lac6 | AGTGCAAGGAACTCGCTACC | TGAGTGTTGATCCCATCGGC |
| Lac7 | TTGTCCATGCCAATCAGCCT | GCGTCAGGAGCACCTTCATA |
| Lac8 | CGCTACCGCTTCCGGATTAT | TGGCTGCACGTTGATACCAT |
| Lac9 | CCAACGCTACTCCCTCATCC | ACATAATGCAGCACACCCGA |
| Lac10 | ATCTCCTGCGACCCAAACTG | ACTGAATCGACGAGAAGCGG |
| Lac11 | GGACCGACATCGAAGAGTCC | GCCCAGTTTGCGTTGGTATC |
基因名 Gene name | 蛋白大小Protein size (aa) | 分子量 Molecular weight (kD) | 等电点 pI | 不稳定指数Instability index | 亲水性平均值 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|
| TYR1 | 318 | 35.4 | 4.86 | 41.54 | -0.361 | Extracellular space |
| TYR2 | 336 | 37.2 | 6.22 | 37.63 | -0.369 | Endomembrane system |
| TYR3 | 311 | 35.2 | 5.11 | 31.54 | -0.292 | Extracellular space |
| Lac1 | 387 | 42.5 | 4.53 | 33.57 | -0.033 | Extracellular space |
| Lac2 | 441 | 47.3 | 4.57 | 33.28 | 0.135 | Plasma membrane |
| Lac3 | 516 | 55.8 | 4.18 | 30.58 | -0.024 | Extracellular space |
| Lac4 | 642 | 68.9 | 4.75 | 35.55 | 0.018 | Extracellular space |
| Lac5 | 398 | 44.4 | 4.91 | 30.93 | -0.299 | Extracellular space |
| Lac6 | 788 | 86.5 | 5.36 | 34.82 | -0.122 | Extracellular space |
| Lac7 | 446 | 49.3 | 5.02 | 25.42 | -0.204 | Extracellular space |
| Lac8 | 523 | 57.1 | 5.17 | 29.37 | -0.020 | Extracellular space |
| Lac9 | 371 | 40.5 | 5.11 | 38.78 | -0.029 | Cytoplasm |
| Lac10 | 391 | 43.2 | 5.82 | 37.78 | 0.096 | Extracellular space |
| Lac11 | 196 | 21.4 | 4.17 | 27.25 | 0.051 | Extracellular space |
表2 金针菇酪氨酸酶和漆酶家族成员的理化性质
Table 2 Physicochemical properties of tyrosinase and laccase family members in F. filiformis
基因名 Gene name | 蛋白大小Protein size (aa) | 分子量 Molecular weight (kD) | 等电点 pI | 不稳定指数Instability index | 亲水性平均值 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|
| TYR1 | 318 | 35.4 | 4.86 | 41.54 | -0.361 | Extracellular space |
| TYR2 | 336 | 37.2 | 6.22 | 37.63 | -0.369 | Endomembrane system |
| TYR3 | 311 | 35.2 | 5.11 | 31.54 | -0.292 | Extracellular space |
| Lac1 | 387 | 42.5 | 4.53 | 33.57 | -0.033 | Extracellular space |
| Lac2 | 441 | 47.3 | 4.57 | 33.28 | 0.135 | Plasma membrane |
| Lac3 | 516 | 55.8 | 4.18 | 30.58 | -0.024 | Extracellular space |
| Lac4 | 642 | 68.9 | 4.75 | 35.55 | 0.018 | Extracellular space |
| Lac5 | 398 | 44.4 | 4.91 | 30.93 | -0.299 | Extracellular space |
| Lac6 | 788 | 86.5 | 5.36 | 34.82 | -0.122 | Extracellular space |
| Lac7 | 446 | 49.3 | 5.02 | 25.42 | -0.204 | Extracellular space |
| Lac8 | 523 | 57.1 | 5.17 | 29.37 | -0.020 | Extracellular space |
| Lac9 | 371 | 40.5 | 5.11 | 38.78 | -0.029 | Cytoplasm |
| Lac10 | 391 | 43.2 | 5.82 | 37.78 | 0.096 | Extracellular space |
| Lac11 | 196 | 21.4 | 4.17 | 27.25 | 0.051 | Extracellular space |
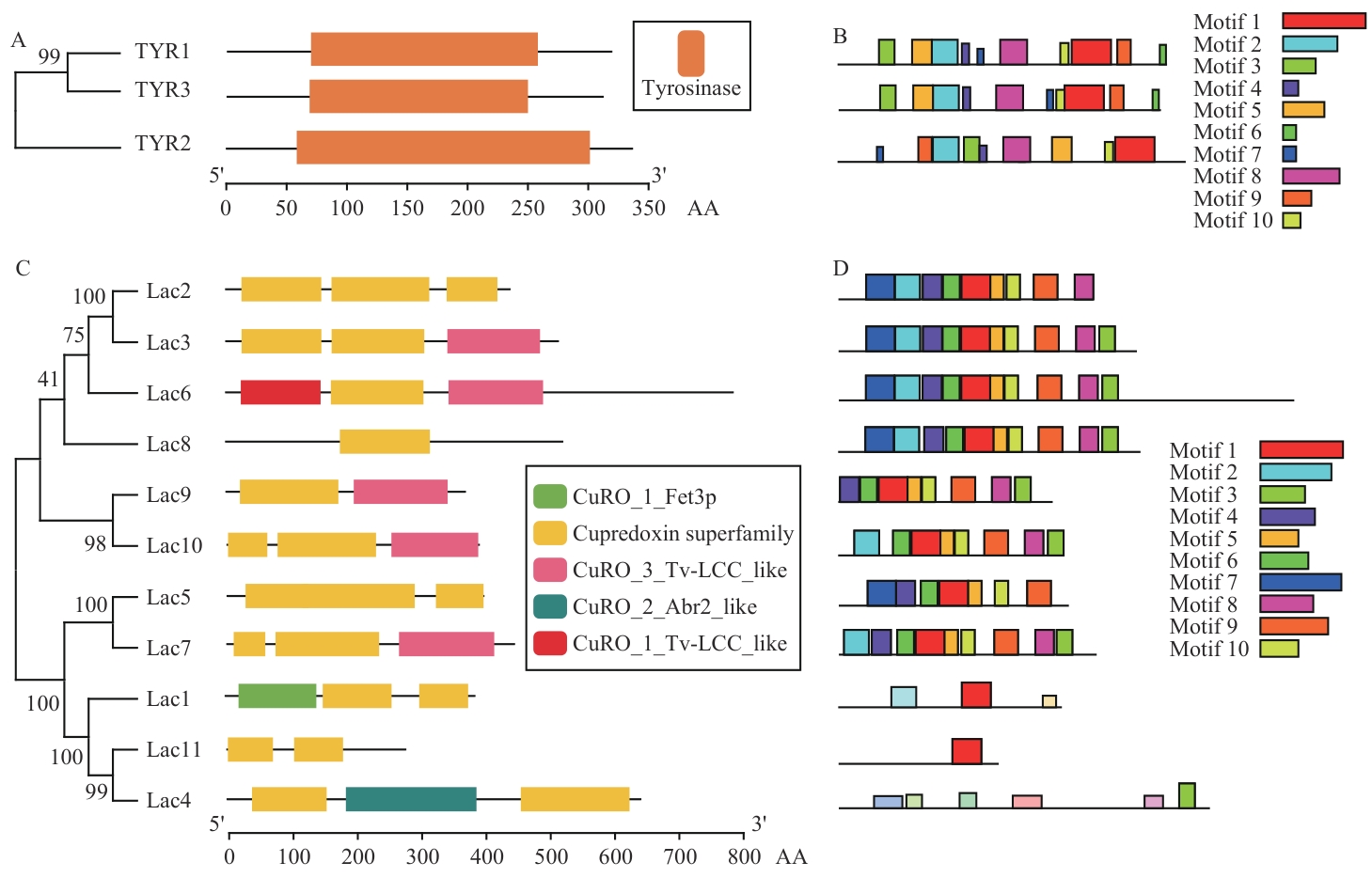
图3 TYR和Lac家族蛋白的保守结构域和保守基序A:TYR蛋白保守结构域;B:TYR蛋白保守基序;C:Lac蛋白保守结构域;D:Lac蛋白保守基序
Fig. 3 Conserved domains and conserved motifs of the TYR and Lac family proteinsA: Conserved domain of TYR protein. B: Conserved motif of TYR protein. C: Conserved domain of Lac protein. D: Conserved motif of Lac protein
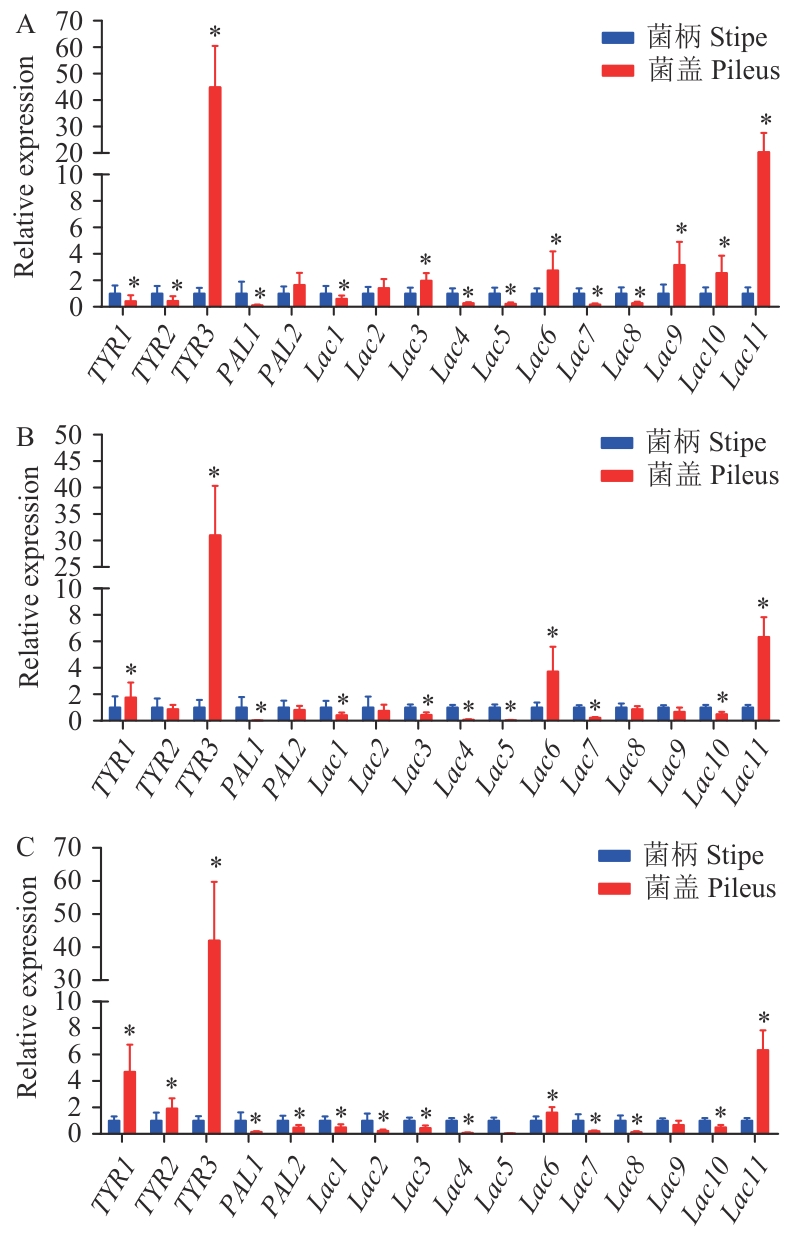
图4 酪氨酸酶和漆酶基因在不同颜色金针菇菌柄和菌盖表皮的表达模式A:白色金针菇;B:黄色金针菇;C:褐色金针菇
Fig. 4 Expression patterns of TYR and Lac genes in the epidermis of different-color F. filiformis stipes and pileusA: White F. filiformis. B: Yellow F. filiformis. C: Brown F. filiformis. *P<0.05
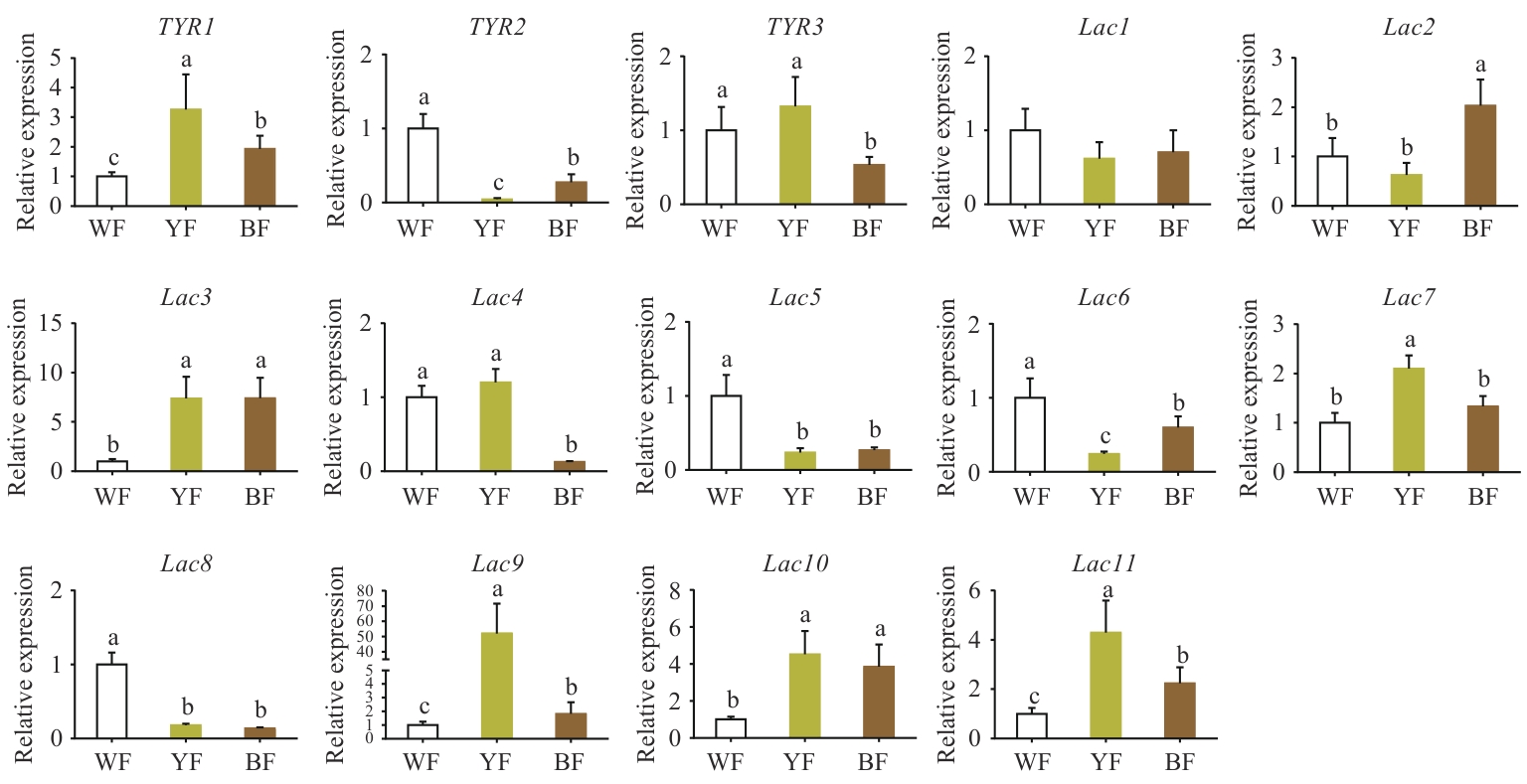
图5 不同颜色菌盖表皮中酪氨酸酶和漆酶基因的相对表达水平比较WF:白色菌盖表皮组织;YF:黄色菌盖表皮组织;BF:褐色菌盖表皮组织。不同字母代表差异显著P<0.05,下同
Fig. 5 Comparison of the relative expressions of TYR and Lac genes in the epidermis of pileus with different colorsWF: White pileus epidermal tissue. YF: Yellow pileus epidermal tissue. BF: Brown pileus epidermal tissue. Different letters refer to significant difference (P<0.05). The same below
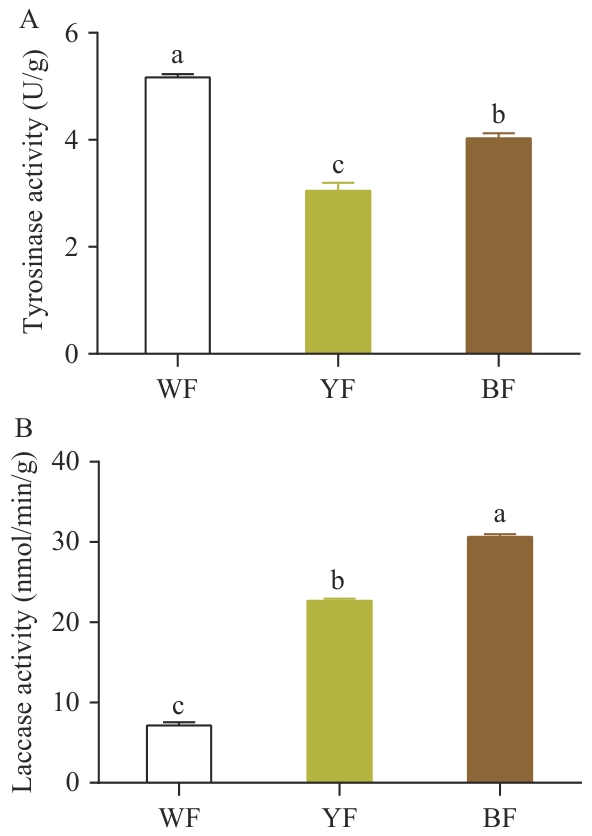
图6 不同颜色菌盖表皮中酪氨酸酶和漆酶活性对比分析A:菌盖表皮组织酪氨酸酶活性;B:菌盖表皮组织漆酶活性
Fig. 6 Comparative analysis of tyrosinase and laccase activities in the epidermis of pileus with different colorsA: Tyrosinase activity in the epidermal tissue of the pileus. B: Laccase activity in the epidermal tissue of the pileus
| [1] | Wang PM, Liu XB, Dai YC, et al. Phylogeny and species delimitation of Flammulina: taxonomic status of winter mushroom in east Asia and a new European species identified using an integrated approach [J]. Mycol Prog, 2018, 17(9): 1013-1030. |
| [2] | 戴玉成, 杨祝良. 中国五种重要食用菌学名新注 [J]. 菌物学报, 2018, 37(12): 1572-1577. |
| Dai YC, Yang ZL. Notes on the nomenclature of five important edible fungi in China [J]. Mycosystema, 2018, 37(12): 1572-1577. | |
| [3] | 中国食用菌协会. 2023年度全国食用菌统计调查结果分析[J]. 中国食用菌, 2025, 44(1): 120-129. |
| China Association of Edible Fungi. Analysis of the results of the national edible fungi statistical survey in 2023[J]. Edible Fungi of China, 2025, 44(1): 120-129. | |
| [4] | 王波. 金针菇新品种'川金33'的选育和栽培应用 [J]. 食药用菌, 2018, 26(1): 38-39. |
| Wang B. Breeding and cultivation application of a new Flammulina velutipes variety ‘Chuanjin 33’ [J]. Edible Med Mushrooms, 2018, 26(1): 38-39. | |
| [5] | 刘询, 何晓兰, 贾定洪, 等. 基于非靶向代谢组学分析黄色和白色金针菇代谢物差异[J]. 菌物学报, 2024, 43(8): 73-85. |
| Liu X, He XL, Jia DH, et al. Analysis of metabolite difference between yellow and white strains of Flammulina filiformis based on untargeted metabolomics [J]. Mycosystema, 2024. 43(8): 73-85. | |
| [6] | Jolivet S, Arpin N, Wichers HJ, et al. Agaricus bisporus browning: a review [J]. Mycol Res, 1998, 102(12): 1459-1483. |
| [7] | Claus H, Decker H. Bacterial tyrosinases [J]. Syst Appl Microbiol, 2006, 29(1): 3-14. |
| [8] | 韩建荣. 木耳子实体阶段酪氨酸酶活性与温度光照和子实体颜色的关系 [J]. 中国食用菌, 1993, 12(2): 11-13. |
| Han JR. Relationship of activities of tyrosinase with the cultivation temperature and licht as well as the fruitbody colqur during fruiting of au ricul Aria au Ricula [J]. Edible Fungi China, 1993, 12(2): 11-13. | |
| [9] | Zhang Y, Wu XL, Huang CY, et al. Isolation and identification of pigments from oyster mushrooms with black, yellow and pink caps [J]. Food Chem, 2022, 372: 131171. |
| [10] | Zhang Y, Huang CY, van Peer AF, et al. Fine mapping and functional analysis of the gene PcTYR, involved in control of cap color of Pleurotus cornucopiae [J]. Appl Environ Microbiol, 2022, 88(7): e0217321. |
| [11] | 刘丽娜. 金针菇颜色突变型的遗传属性研究 [D]. 长沙: 湖南师范大学, 2016. |
| Liu LN. Study on genetic properties of color mutant of Flammulina velutipes [D]. Changsha: Hunan Normal University, 2016. | |
| [12] | Zeng JY, Shi DD, Chen Y, et al. FvbHLH1 regulates the accumulation of phenolic compounds in the yellow cap of Flammulina velutipes [J]. J Fungi, 2023, 9(11): 1063. |
| [13] | Im JH, Yu HW, Park CH, et al. Phenylalanine ammonia-lyase: a key gene for color discrimination of edible mushroom Flammulina velutipes [J]. J Fungi, 2023, 9(3): 339. |
| [14] | Liang YQ, Luo KM, Wang BL, et al. Inhibition of polyphenol oxidase for preventing browning in edible mushrooms: a review [J]. J Food Sci, 2024, 89(11): 6796-6817. |
| [15] | Martin E, Dubessay P, Record E, et al. Recent advances in laccase activity assays: a crucial challenge for applications on complex substrates [J]. Enzyme Microb Technol, 2024, 173: 110373. |
| [16] | Yan LL, Xu RP, Bian YB, et al. Expression profile of laccase gene family in white-rot basidiomycete Lentinula edodes under different environmental stresses [J]. Genes, 2019, 10(12): 1045. |
| [17] | Zhang JL, Tanaka Y, Ono A, et al. Gene expression analysis for stem browning in the mushroom Lentinula edodes [J]. Mycoscience, 2024, 65(5): 253-259. |
| [18] | 李子豪, 李小凤, 杜芳, 等. 刺芹侧耳不同生长发育阶段漆酶基因家族表达 [J]. 食用菌学报, 2021, 28(4): 1-6. |
| Li ZH, Li XF, Du F, et al. Expression of laccase gene family at different developmental stages of Pleurotus eryngii [J]. Acta Edulis Fungi, 2021, 28(4): 1-6. | |
| [19] | 刘冬梅, 刁文彤, 孙雪言, 等. 灵芝漆酶基因家族的克隆及其在不同发育阶段的表达分析 [J]. 微生物学杂志, 2025, 45(1): 34-42. |
| Liu DM, Diao WT, Sun XY, et al. Cloning and expression analysis of laccase genes family at developmental stages of Ganoderma lucidum [J]. Journal of Microbiology, 2025, 45(1): 34-42. | |
| [20] | He Y, Liu B, Ouyang XQ, et al. Whole-genome sequencing and fine map analysis of Pholiota nameko [J]. J Fungi, 2025, 11(2): 112. |
| [21] | 吉军, 张玉琎, 肖鑫, 等. 白假鬼伞漆酶基因家族鉴定及表达模式 [J]. 植物生理学报, 2023, 59(11): 2027-2038. |
| Ji J, Zhang YJ, Xiao X, et al. Identification and expression analysis on laccase gene family in Coprinellus disseminates [J]. Plant Physiology Journal, 2023, 59(11): 2027-2038. | |
| [22] | Liang XM, Han J, Cui YQ, et al. Whole-genome sequencing of Flammulina filiformis and multi-omics analysis in response to low temperature [J]. J Fungi, 2025, 11(3): 229. |
| [23] | 杜金茹, 王守现, 严冬, 等. 基于全基因组数据分析香菇黑色素合成途径及相关基因 [J]. 菌物学报, 2023, 42(5):1114-1128. |
| Du JR, Wang SX, Yan D, et al. Analyses of melanin synthesis pathway and the related genes in Lentinula edodes based on whole genome sequence comparison [J]. Mycosystema, 2023, 42(5): 1114-1128. | |
| [24] | Wilkins MR, Gasteiger E, Bairoch A, et al. Protein identification and analysis tools in the ExPASy server [J]. Methods Mol Biol, 1999, 112: 531-552. |
| [25] | Savojardo C, Martelli PL, Fariselli P, et al. BUSCA: an integrative web server to predict subcellular localization of proteins [J]. Nucleic Acids Res, 2018, 46(W1): W459-W466. |
| [26] | 王波, 刘询, 何晓兰, 等. 一株褐色金针菇及其应用:中国, CN202410393413.4 [P]. 2024-08-13. |
| Wang B, Liu X, He XL, et al. A brown Flammulina filiformis and its applications: China, CN202410393413.4 [P]. 2024-08-13. | |
| [27] | Yu CY. Molecular mechanism of manipulating seed coat coloration in oilseed Brassica species [J]. J Appl Genet, 2013, 54(2): 135-145. |
| [28] | Wang M, Song TT, Jin QL, et al. From white to reddish-brown: the anthocyanin journey in Stropharia rugosoannulata driven by auxin and genetic regulators [J]. J Agric Food Chem, 2025, 73(1): 954-966. |
| [29] | 吴林, 朱刚, 陈明杰, 等. 草菇基因组中11个漆酶同源基因的生物信息学分析及铜离子对其表达的影响 [J]. 菌物学报, 2014, 33(2): 323-333. |
| Wu L, Zhu G, Chen MJ, et al. The bioinformatic analyses and the gene expression induced by Cu2+ of 11 laccase homologous genes from Volvariella volvacea [J]. Mycosystema, 2014, 33(2): 323-333. | |
| [30] | 李雪松, 孙达锋, 华蓉, 等. 皱木耳漆酶生物信息学分析 [J]. 中国食用菌, 2023, 42(2): 52-59. |
| Li XS, Sun DF, Hua R, et al. Bioinformatics identification of laccase in Auricularia delicata [J]. Edible Fungi China, 2023, 42(2): 52-59. | |
| [31] | Fu NN, Li JX, Wang M, et al. Genes identification, molecular docking and dynamics simulation analysis of laccases from Amylostereum areolatum provides molecular basis of laccase bound to lignin [J]. Int J Mol Sci, 2020, 21(22): 8845. |
| [32] | Litvintseva AP, Henson JM. Cloning, characterization, and transcription of three laccase genes from Gaeumannomyces graminis var. tritici, the take-all fungus [J]. Appl Environ Microbiol, 2002, 68(3): 1305-1311. |
| [33] | Jin WS, Li JH, Feng HC, et al. Importance of a laccase gene (Lcc1) in the development of Ganoderma tsugae [J]. Int J Mol Sci, 2018, 19(2): 471. |
| [34] | Chen SC, Ge W, Buswell JA. Molecular cloning of a new laccase from the edible straw mushroom Volvariella volvacea: possible involvement in fruit body development [J]. FEMS Microbiol Lett, 2004, 230(2): 171-176. |
| [1] | 张超超, 韩开元, 王彤, 陈仲. 毛白杨PtoYABBY2和PtoYABBY12的克隆及功能分析[J]. 生物技术通报, 2025, 41(9): 256-264. |
| [2] | 史发超, 姜永华, 刘海伦, 文英杰, 严倩. 荔枝LcTFL1基因的克隆与功能分析[J]. 生物技术通报, 2025, 41(9): 159-167. |
| [3] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [4] | 李玉珍, 李梦丹, 张蔚, 彭婷. 基于月季扩展蛋白基因家族鉴定的野蔷薇RmEXPB2基因功能研究[J]. 生物技术通报, 2025, 41(9): 182-194. |
| [5] | 黄诗宇, 田姗姗, 杨天为, 高曼熔, 张尚文. 赤苍藤WRI1基因家族的全基因组鉴定及表达模式分析[J]. 生物技术通报, 2025, 41(8): 242-254. |
| [6] | 康琴, 汪霞, 谌明洋, 徐静天, 陈诗兰, 廖平杨, 许文志, 吴卫, 徐东北. 薄荷UV-B受体基因McUVR8的克隆与表达分析[J]. 生物技术通报, 2025, 41(8): 255-266. |
| [7] | 段敏杰, 李怡斐, 王春萍, 黄任中, 黄启中, 张世才. 辣椒果实颜色性状与SSR分子标记的关联分析及指纹图谱构建[J]. 生物技术通报, 2025, 41(7): 81-94. |
| [8] | 王芳, 乔帅, 宋伟, 崔鹏娟, 廖安忠, 谭文芳, 杨松涛. 甘薯IbNRT2基因家族全基因组鉴定和表达分析[J]. 生物技术通报, 2025, 41(7): 193-204. |
| [9] | 许慧珍, SHANTWANA Ghimire, RAJU Kharel, 岳云, 司怀军, 唐勋. 马铃薯SUMO E3连接酶基因家族分析及StSIZ1基因的克隆与表达模式分析[J]. 生物技术通报, 2025, 41(6): 119-129. |
| [10] | 王天禧, 杨炳松, 潘荣君, 盖文贤, 梁美霞. 苹果PLATZ基因家族鉴定及MdPLATZ9基因功能研究[J]. 生物技术通报, 2025, 41(4): 176-187. |
| [11] | 宋佳怡, 苏芸丽, 郑兴艳, 夏文念, 杨冬梅, 胡慧贞. 金鱼草Expansin基因家族鉴定及其抗核盘菌相关基因筛选[J]. 生物技术通报, 2025, 41(4): 227-242. |
| [12] | 张益瑄, 马宇, 王童童, 盛苏奥, 宋家凤, 吕钊彦, 朱晓彪, 侯华兰. 马铃薯DIR家族全基因组鉴定及表达模式分析[J]. 生物技术通报, 2025, 41(3): 123-136. |
| [13] | 宋姝熠, 蒋开秀, 刘欢艳, 黄亚成, 刘林娅. ‘红阳’猕猴桃TCP基因家族鉴定及其在果实中的表达分析[J]. 生物技术通报, 2025, 41(3): 190-201. |
| [14] | 匡健华, 程志鹏, 赵永晶, 杨洁, 陈润乔, 陈龙清, 胡慧贞. 激素和非生物胁迫下荷花GH3基因家族的表达分析[J]. 生物技术通报, 2025, 41(2): 221-233. |
| [15] | 黄颖, 遇文婧, 刘雪峰, 刁桂萍. 山新杨谷胱甘肽转移酶基因的生物信息学与表达模式分析[J]. 生物技术通报, 2025, 41(2): 248-256. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||