








生物技术通报 ›› 2026, Vol. 42 ›› Issue (4): 227-238.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0740
倪飞飞1,2( ), 陈益存1,2, 高暝1,2, 张盛剿3, 彭方有3, 陈涛梅3, 赵耘霄1,2(
), 陈益存1,2, 高暝1,2, 张盛剿3, 彭方有3, 陈涛梅3, 赵耘霄1,2( ), 汪阳东1,2(
), 汪阳东1,2( )
)
收稿日期:2025-07-09
出版日期:2026-04-26
发布日期:2026-04-30
通讯作者:
赵耘霄,女,副研究员,研究方向 :化学工业原料林育种;E-mail: zyx_yunxiao@caf.ac.cn作者简介:倪飞飞,男,硕士研究生,研究方向 :特色林木资源育种与培育;E-mail: nifeifei2042634075@163.com
基金资助:
NI Fei-fei1,2( ), CHEN Yi-cun1,2, GAO Ming1,2, ZHANG Sheng-jiao3, PENG Fang-you3, CHEN Tao-mei3, ZHAO Yun-xiao1,2(
), CHEN Yi-cun1,2, GAO Ming1,2, ZHANG Sheng-jiao3, PENG Fang-you3, CHEN Tao-mei3, ZHAO Yun-xiao1,2( ), WANG Yang-dong1,2(
), WANG Yang-dong1,2( )
)
Received:2025-07-09
Published:2026-04-26
Online:2026-04-30
摘要:
目的 筛选并验证可能参与调控山苍子萜类合成的TCP转录因子。 方法 基于山苍子全基因组及果实不同发育时期的转录组数据,采用生物信息学方法对TCP转录因子的理化性质、染色体定位、基因结构、保守基序、顺式作用元件及物种内共线性关系进行系统分析,并利用RT-qPCR检测CIN亚家族成员在不同发育时期及组织中的表达模式,对关键转录因子进行亚细胞定位,并结合酵母单杂和瞬时转化技术验证其功能。 结果 共鉴定出30个TCP基因家族成员(LcTCPs),分布于9条染色体,命名为LcTCP1-LcTCP30,开放读码框长度504-1 755 bp,分子量17 711.98-63 421.45 Da。系统进化将其划分为PCF(17个)、CIN(7个)和CYC/TB1(6个)3个亚家族,保守基序分析结果与分组一致。基因扩张主要受全基因组复制事件驱动,各亚家族间成员高度保守。RT-qPCR结果显示,CIN亚家族中LcTCP6与LcTCP11在精油合成高峰期及果实中的表达量较高,提示其与萜类合成相关。其中LcTCP11与关键基因LcDXS5在果实发育过程中呈现相似的表达模式,并定位于细胞核。功能验证表明LcTCP11能够调控LcDXS5的表达。 结论 从山苍子基因组中共鉴定30个TCP转录因子,其中LcTCP11与萜类合成关键基因LcDXS5呈协同表达,且在果实中表达显著高于其他组织,推测其可能通过调控LcDXS5参与山苍子萜类合成。
倪飞飞, 陈益存, 高暝, 张盛剿, 彭方有, 陈涛梅, 赵耘霄, 汪阳东. 山苍子TCP基因鉴定及其调控萜类合成的研究[J]. 生物技术通报, 2026, 42(4): 227-238.
NI Fei-fei, CHEN Yi-cun, GAO Ming, ZHANG Sheng-jiao, PENG Fang-you, CHEN Tao-mei, ZHAO Yun-xiao, WANG Yang-dong. Identification of Litsea cubebaTCP Genes and Their Roles in the Regulation of Terpenoid Biosynthesis[J]. Biotechnology Bulletin, 2026, 42(4): 227-238.
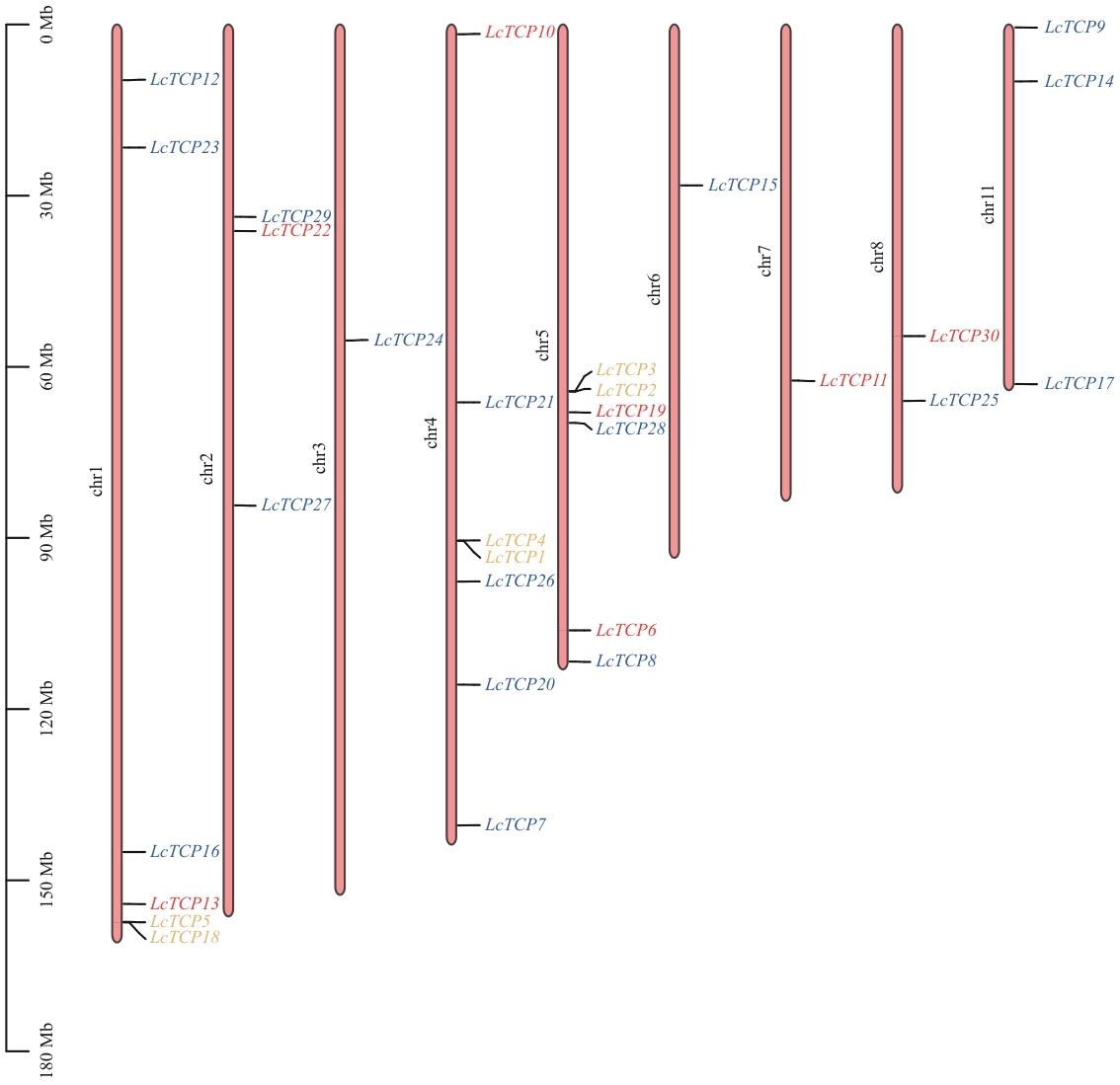
图1 山苍子LcTCPs基因的染色体定位蓝色表示PCF亚家族,黄色表示CYC/TB1亚家族,红色表示CIN亚家族
Fig. 1 Chromosomal localization of LcTCP genesBlue indicates the PCF subfamily, yellow indicates the CYC/TB1 subfamily, and red indicates the CIN subfamily
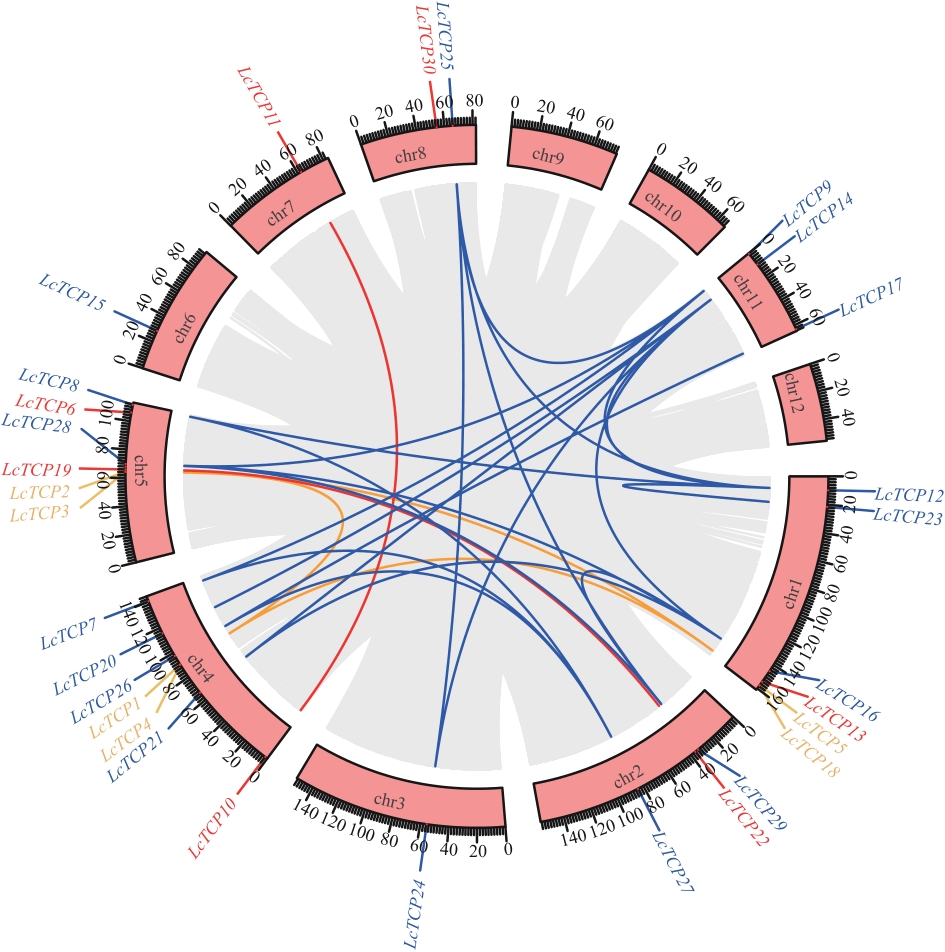
图4 LcTCPs基因的共线性关系粉色框表示染色体,蓝色线表示PCF亚家族基因共线性,红色线表示CIN亚家族基因共线性,黄色线表示CYC/TB1亚家族基因共线性,灰色背景是山苍子基因组所有基因中存在的片段重复基因
Fig. 4 Synteny relationship of LcTCP genesPink boxes indicate chromosomes, blue lines indicate syntenic relationships among PCF subfamily genes, red lines indicate those of the CIN subfamily, and yellow lines indicate those of the CYC/TB1 subfamily. The gray background indicates segmentally duplicated genes across the L. cubeba genome
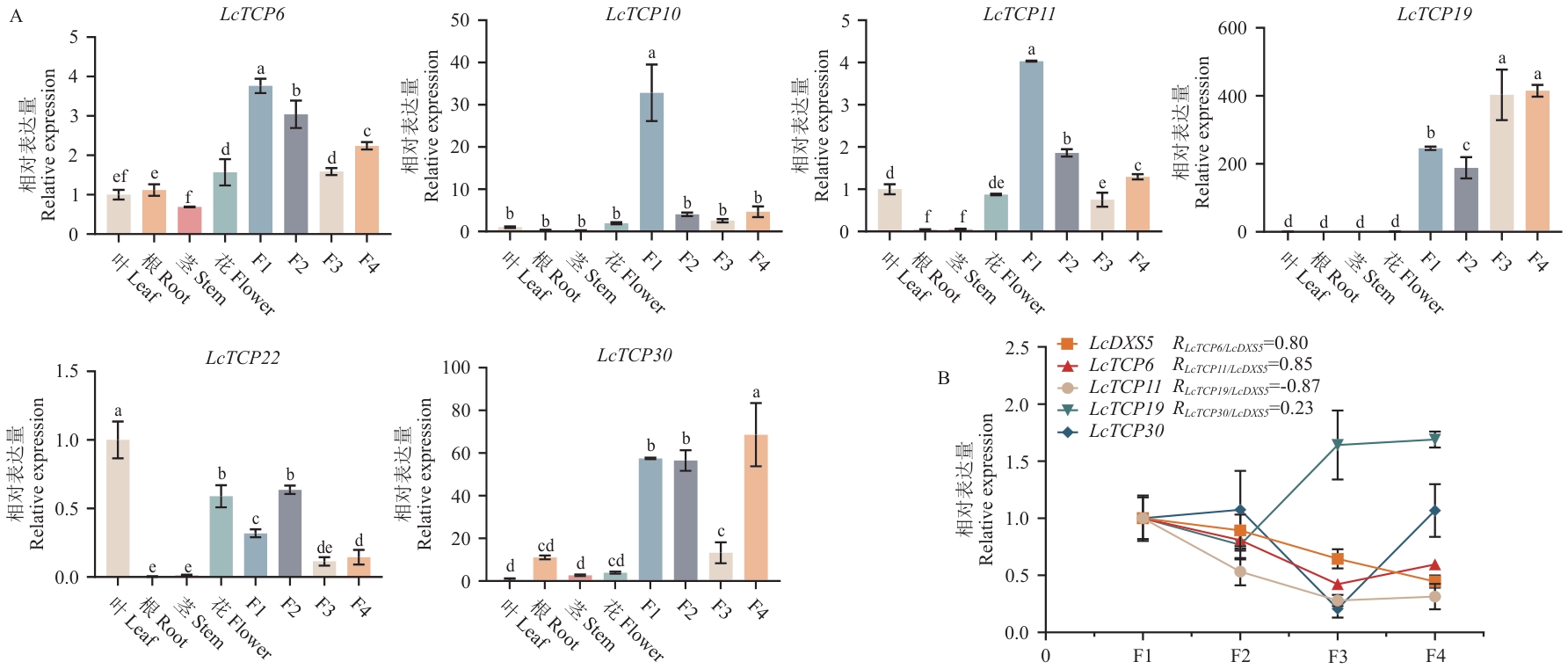
图6 不同组织的表达模式及果实不同发育时期表达相关性A:6个山苍子CIN亚家族成员在果实不同发育时期及不同组织的表达模式;B:LcTCP6、LcTCP11、LcTCP19、LcTCP30与LcDXS5在果实不同发育阶段的表达相关性;F1:花后30 d;F2:花后60 d,精油合成高峰期;F3:花后90 d;F4:花后120 d。不同字母表示差异显著(P<0.05)
Fig. 6 Expression patterns in different tissues and correlation of expression at different fruit developmental stagesA: Expression patterns of 6 CIN subfamily members from L. cubeba at different fruit developmental stages and in various tissues. B: Expression correlation among LcTCP6, LcTCP11, LcTCP19, LcTCP30, and LcDXS5 at different fruit developmental stages. F1: 30 d after flowering; F2: 60 d after flowering, the peak period of essential oil synthesis; F3: 90 d after flowering; F4: 120 d after flowering. Different lower letters indicate significant differences (P<0.05)
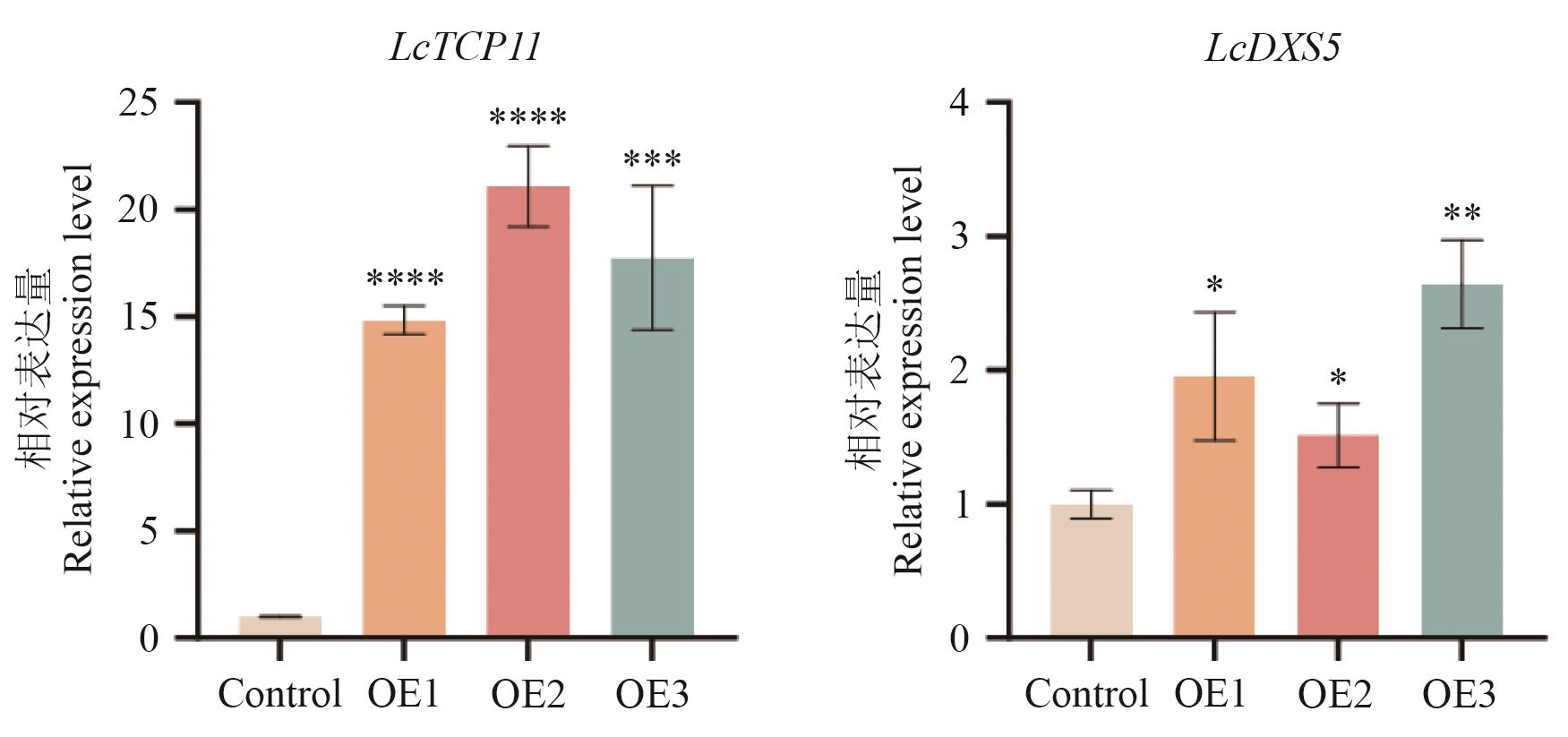
图7 瞬时转化山苍子叶片LcTCP11及LcDXS5的表达水平OE1、OE2、OE3代表3个过表达株系,数据以均值±标准差(SD)表示,来自3次重复实验(*P<0.05,**P<0.01,***P<0.001,****P<0.000 1)
Fig. 7 Expression analysis of LcTCP11 and LcDXS5 in transiently transformed L. cubeba leavesOE1, OE2, and OE3 refer to three overexpression lines. Data are expressed as mean±SD from three replicates (*P<0.05, **P<0.01, ***P<0.001,and ****P<0.000 1)
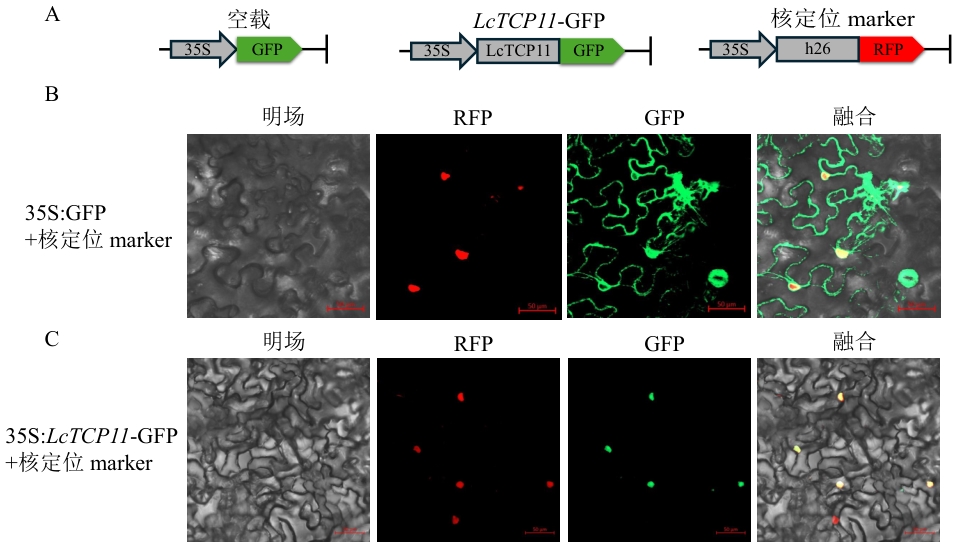
图8 LcTCP11亚细胞定位A:载体示意图;B:空载亚细胞定位;C:LcTCP11-GFP融合表达定位
Fig. 8 Subcellular localization of LcTCP11A: Schematic diagram of the vector. B: Subcellular localization of the empty vector. C: Localization of LcTCP11-GFP fusion expression
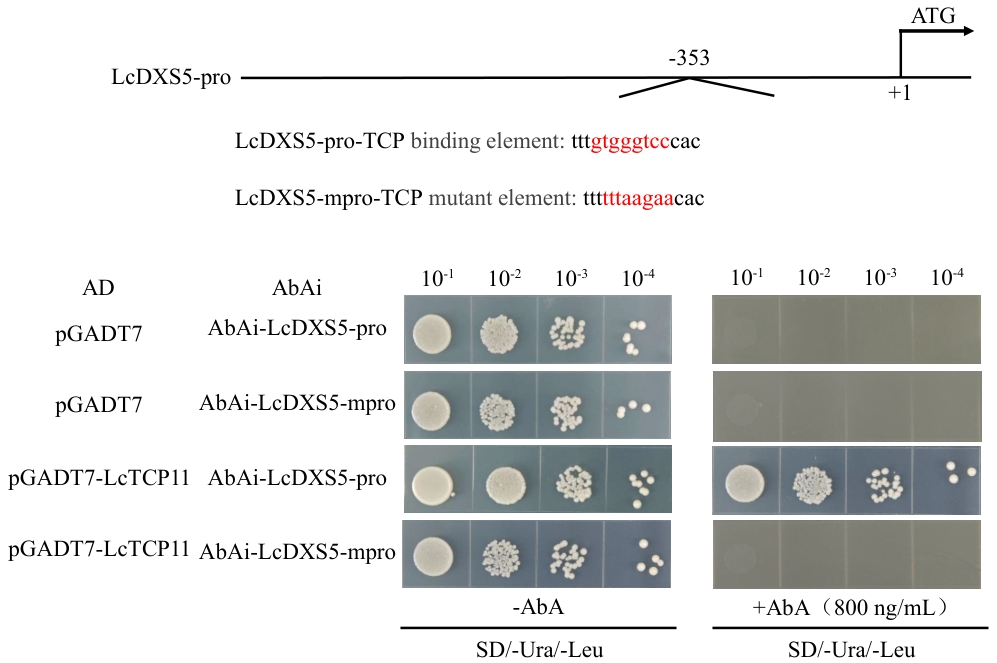
图9 酵母单杂交试验验证LcTCP11对LcDXS5启动子TCP元件的识别作用图中红色字体分别代表启动子上的结合元件序列以及突变之后的元件序列。将同时表达pGADT7-LcTCP11和pAbAi-3×gtgggtcc的酵母细胞在含有AbA的选择培养基(SD/-Trp/-Ura)上于30 ℃培养48 h。待长出单个菌落后,将单个菌落用水按10¹‒10⁴倍稀释后,每个稀释度取10 μL点种到新的含有AbA的选择培养基(SD/-Trp/-Ura)上,继续培养48 h
Fig. 9 Y1H assays verifying the recognition of LcTCP11 to the TCP element in LcDXS5 promoterThe red text in the figure indicates the binding element sequences on the promoter and the corresponding mutated element sequences, respectively. Yeast cells co-expressing pGADT7-LcTCP11 and pAbAi-3×gtgggtcc were cultured on selective medium containing AbA (SD/-Trp/-Ura) at 30 ℃ for 48 h. Once single colonies had grown, they were diluted in water to 10¹-10⁴ fold, and 10 μL of each dilution was spotted onto fresh selective medium containing AbA (SD/-Trp/-Ura) for a further 48 h of cultivation
| [1] | Lan JQ, Qin GJ. The regulation of CIN-like TCP transcription factors [J]. Int J Mol Sci, 2020, 21(12): 4498. |
| [2] | Cubas P, Lauter N, Doebley J, et al. The TCP domain: a motif found in proteins regulating plant growth and development [J]. Plant J, 1999, 18(2): 215-222. |
| [3] | Doebley J, Stec A, Hubbard L. The evolution of apical dominance in maize [J]. Nature, 1997, 386(6624): 485-488. |
| [4] | Luo D, Carpenter R, Vincent C, et al. Origin of floral asymmetry in Antirrhinum [J]. Nature, 1996, 383(6603): 794-799. |
| [5] | Shunichi Kosugi YO. PCF1 and PCF2 specifically bind to cis elements in the rice proliferating cell nuclear antigen gene [J]. Plant Cell, 1997, 9(9): 1607-1619. |
| [6] | Navaud O, Dabos P, Carnus E, et al. TCP transcription factors predate the emergence of land plants [J]. J Mol Evol, 2007, 65(1): 23-33. |
| [7] | 马亚男. TCP类转录因子调控青蒿素生物合成的分子机制研究 [D]. 上海: 上海交通大学, 2019. |
| Ma YN. The studies on molecular mechanism of TCP transcription factors regulating artemisinin biosynthesis in Artemisia annua L. [D]. Shanghai: Shanghai Jiao Tong University, 2019. | |
| [8] | Martín-Trillo M, Cubas P. TCP genes: a family snapshot ten years later [J]. Trends Plant Sci, 2010, 15(1): 31-39. |
| [9] | O’Connor C, Varshosaz P, Moise AR. Mechanisms of feedback regulation of vitamin a metabolism [J]. Nutrients, 2022, 14(6): 1312. |
| [10] | Ma N, Zhang ZY, Liao FL, et al. The birth of artemisinin [J]. Pharmacol Ther, 2020, 216: 107658. |
| [11] | Costa EF, Magalhães WV, Di Stasi LC. Recent advances in herbal-derived products with skin anti-aging properties and cosmetic applications [J]. Molecules, 2022, 27(21): 7518. |
| [12] | Thoppil RJ, Bishayee A. Terpenoids as potential chemopreventive and therapeutic agents in liver cancer [J]. World J Hepatol, 2011, 3(9): 228-249. |
| [13] | Bakkali F, Averbeck S, Averbeck D, et al. Biological effects of essential oils-A review [J]. Food Chem Toxicol, 2008, 46(2): 446-475. |
| [14] | Liu XM, Zhang WW, Tang N, et al. Genomic-wide identification and expression analysis of AP2/ERF transcription factors in Zanthoxylum armatum reveals the candidate genes for the biosynthesis of terpenoids [J]. Plant Genome, 2024, 17(1): e20422. |
| [15] | Guo YH, Guo ZY, Zhong J, et al. Positive regulatory role of R2R3 MYBs in terpene biosynthesis in Lilium ‘Siberia’ [J]. Hortic Plant J, 2023, 9(5): 1024-1038. |
| [16] | Li HW, Chen NN, Zhang HB, et al. Multidimensional regulation of transcription factors: decoding the comprehensive signals of plant secondary metabolism [J]. Front Plant Sci, 2025, 16: 1522278. |
| [17] | Liu C, Lv TT, Shen YH, et al. Genome-wide identification and integrated analysis of TCP genes controlling ginsenoside biosynthesis in Panax ginseng [J]. BMC Plant Biol, 2024, 24(1): 47. |
| [18] | Ma YN, Xu DB, Yan X, et al. Jasmonate- and abscisic acid-activated AaGSW1-AaTCP15/AaORA transcriptional cascade promotes artemisinin biosynthesis in Artemisia annua [J]. Plant Biotechnol J, 2021, 19(7): 1412-1428. |
| [19] | Ma YN, Xu DB, Li L, et al. Jasmonate promotes artemisinin biosynthesis by activating the TCP14-ORA complex in Artemisia annua [J]. Sci Adv, 2018, 4(11): eaas9357. |
| [20] | 詹忠根. 转录因子调控青蒿素生物合成的作用机制研究进展 [J]. 中草药, 2022, 53(19): 6258-6272. |
| Zhan ZG. Research progress on transcriptional regulation mechanism of artemisinin biosynthesis [J]. Chin Tradit Herb Drugs, 2022, 53(19): 6258-6272. | |
| [21] | Gao J, Chen YC, Gao M, et al. LcWRKY17, a WRKY transcription factor from Litsea cubeba, effectively promotes monoterpene synthesis [J]. Int J Mol Sci, 2023, 24(8): 7210. |
| [22] | Si LL, Chen YC, Han XJ, et al. Chemical composition of essential oils of Litsea cubeba harvested from its distribution areas in China [J]. Molecules, 2012, 17(6): 7057-7066. |
| [23] | Gao M, Lin LY, Chen YC, et al. Digital gene expression profiling to explore differentially expressed genes associated with terpenoid biosynthesis during fruit development in Litsea cubeba [J]. Molecules, 2016, 21(9): 1251. |
| [24] | Wang MY, Gao M, Zhao YX, et al. LcERF19, an AP2/ERF transcription factor from Litsea cubeba, positively regulates geranial and neral biosynthesis [J]. Hortic Res, 2022, 9: uhac093. |
| [25] | Zhao YX, Wang MY, Chen YC, et al. LcERF134 increases the production of monoterpenes by activating the terpene biosynthesis pathway in Litsea cubeba [J]. Int J Biol Macromol, 2023, 232: 123378. |
| [26] | Ni FF, Zhao YX, Gao M, et al. LcMYB43 enhances monoterpene biosynthesis by activating 1-deoxy-D-xylulose-5-phosphate synthase gene expression in Litsea cubeba [J]. Int J Biol Macromol, 2025, 304(Pt 1): 140860. |
| [27] | Zhao YX, Chen YC, Gao M, et al. Alcohol dehydrogenases regulated by a MYB44 transcription factor underlie Lauraceae citral biosynthesis [J]. Plant Physiol, 2024, 194(3): 1674-1691. |
| [28] | Yang JH, Chen YC, Gao M, et al. Comprehensive identification of bHLH transcription factors in Litsea cubeba reveals candidate gene involved in the monoterpene biosynthesis pathway [J]. Front Plant Sci, 2022, 13: 1081335. |
| [29] | Li ST, Zachgo S. TCP3 interacts with R2R3-MYB proteins, promotes flavonoid biosynthesis and negatively regulates the auxin response in Arabidopsis thaliana [J]. Plant J, 2013, 76(6): 901-913. |
| [30] | 宋姝熠, 蒋开秀, 刘欢艳, 等. ‘红阳’猕猴桃TCP基因家族鉴定及其在果实中的表达分析 [J]. 生物技术通报, 2025, 41(3): 190-201. |
| Song SY, Jiang KX, Liu HY, et al. Identification of the TCP gene family in Actinidia chinensis var. Hongyang and their expression analysis in fruit [J]. Biotechnol Bull, 2025, 41(3): 190-201. | |
| [31] | Chen YC, Li Z, Zhao YX, et al. The Litsea genome and the evolution of the laurel family [J]. Nat Commun, 2020, 11(1): 1675. |
| [32] | Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets [J]. Mol Biol Evol, 2016, 33(7): 1870-1874. |
| [33] | Viola IL, Camoirano A, Gonzalez DH. Redox-dependent modulation of anthocyanin biosynthesis by the TCP transcription factor TCP15 during exposure to high light intensity conditions in Arabidopsis [J]. Plant Physiol, 2016, 170(1): 74-85. |
| [34] | Liu DH, Luo Y, Han H, et al. Genome-wide analysis of citrus TCP transcription factors and their responses to abiotic stresses [J]. BMC Plant Biol, 2022, 22(1): 325. |
| [35] | Zhou Y, Zhang DZ, An JX, et al. TCP transcription factors regulate shade avoidance via directly mediating the expression of both PHYTOCHROME INTERACTING FACTORs and auxin biosynthetic genes [J]. Plant Physiol, 2018, 176(2): 1850-1861. |
| [36] | 唐羽翔, 高旭, 崔亚宁, 等. 植物TCP转录因子研究进展 [J]. 科学通报, 2022, 67(33): 3964-3975. |
| Tang YX, Gao X, Cui YN, et al. Research progress of plant TCP transcription factors [J]. Chin Sci Bull, 2022, 67(33): 3964-3975. | |
| [37] | Viola IL, Gonzalez DH. TCP transcription factors in plant reproductive development: juggling multiple roles [J]. Biomolecules, 2023, 13(5): 750. |
| [38] | Aguilar-Martínez JA, Poza-Carrión C, Cubas P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds [J]. Plant Cell, 2007, 19(2): 458-472. |
| [39] | Koyama T, Mitsuda N, Seki M, et al. TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis [J]. Plant Cell, 2010, 22(11): 3574-3588. |
| [40] | Miao-Ying Wang PZ. The cotton transcription factor TCP14 functions in auxin-mediated epidermal cell differentiation and elongation[J]. Plant Physiol, 2013, 162(3): 1669-1680. |
| [41] | Tian C, Zhai LS, Zhu WJ, et al. Characterization of the TCP gene family in Chrysanthemum nankingense and the role of CnTCP4 in cold tolerance [J]. Plants, 2022, 11(7): 936. |
| [42] | Chen SY, Chen B, Xu XN. Genome-wide identification and expression analysis of TCP transcription factors in Chrysanthemum indicum reveals their critical role in the response to various abiotic stresses [J]. BMC Plant Biol, 2025, 25(1): 631. |
| [43] | Xu YX, Ma L, Zeng XC, et al. Genome-wide identification and analysis of BrTCP transcription factor family genes involved in cold stress tolerance in winter rapeseed (Brassica rapa L.) [J]. Int J Mol Sci, 2024, 25(24): 13592. |
| [44] | Sprenger GA, Schörken U, Wiegert T, et al. Identification of a thiamin-dependent synthase in Escherichia coli required for the formation of the 1-deoxy-D-xylulose 5-phosphate precursor to isoprenoids, thiamin, and pyridoxol [J]. Proc Natl Acad Sci USA, 1997, 94(24): 12857-12862. |
| [45] | Yuan Y, Ren SY, Liu XF, et al. SlWRKY35 positively regulates carotenoid biosynthesis by activating the MEP pathway in tomato fruit [J]. New Phytol, 2022, 234(1): 164-178. |
| [46] | Tian L, Shi JW, Yang L, et al. Molecular cloning and functional analysis of DXS and FPS genes from Zanthoxylum bungeanum Maxim [J]. Foods, 2022, 11(12): 1746. |
| [1] | 孙婷, 张艳, 刘玉珊, 冯媛媛, 秦恒山, 张军, 何小岗, 张景荣. 黄秋葵AeF3H基因的克隆与功能分析[J]. 生物技术通报, 2026, 42(4): 153-160. |
| [2] | 冯成蒿, 党雨乐, 王志泽, 聂蔚丹, 杨中敏, 杜崇. 基于RNA-seq筛选番茄防御南方根结线虫病的差异基因与RPP13的表达特征分析[J]. 生物技术通报, 2026, 42(4): 141-152. |
| [3] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [4] | 殷诗情, 田泰, 黄凤庭, 冯珑强, 王浩, 张静, 何文, 陈清, 王小蓉, 王燕. 果树果实硬度的调控机制研究进展[J]. 生物技术通报, 2026, 42(3): 213-229. |
| [5] | 赵艳侠, 李倩, 孙家波, 梁红敏, 李冰冰. 草莓果实品质形成的关键调控基因及分子网络解析[J]. 生物技术通报, 2026, 42(3): 111-132. |
| [6] | 徐泽, 周陈平, 邝瑞彬, 吴夏明, 杨敏, 刘传和, 贺涵, 魏岳荣. 番木瓜PG基因家族鉴定及其在果实软化中的功能[J]. 生物技术通报, 2026, 42(3): 349-361. |
| [7] | 张冬岭, 张寅生, 王建军, 叶飞宇, 卢子涵, 马晨晨, 柳华峰, 胡德升, 邓亚洲, 曹丽茹. 玉米HSFs转录因子家族在干旱胁迫下的表达特性及功能[J]. 生物技术通报, 2026, 42(2): 178-187. |
| [8] | 农韦优, 赵昌祖, 钱禛锋, 丁倩, 王誉洁, 陈疏影, 何丽莲, 李富生. 蔗茅EfBBX基因家族鉴定及冷胁迫下表达模式分析[J]. 生物技术通报, 2026, 42(2): 267-277. |
| [9] | 杨娟, 冯慧, 吉乃喆, 孙丽萍, 王赟, 张佳楠, 赵世伟. 月季AP2/ERF转录因子RcERF4和RcRAP2-12的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 150-160. |
| [10] | 吴翠翠, 陈登科, 兰刚, 夏芝, 李朋波. 花生转录因子AhHDZ70的生物信息学分析及耐盐耐旱性研究[J]. 生物技术通报, 2026, 42(1): 198-207. |
| [11] | 龙林茜, 曾银萍, 王茜, 邓玉萍, 葛敏茜, 陈彦灼, 李鑫娟, 杨军, 邹建. 向日葵GH3基因家族鉴定及其在花发育中的功能分析[J]. 生物技术通报, 2026, 42(1): 125-138. |
| [12] | 陈强, 于璎霏, 张颖, 张冲. 茉莉酸甲酯对薄皮甜瓜‘绿宝石’采后冷害的调控[J]. 生物技术通报, 2025, 41(9): 105-114. |
| [13] | 张超超, 韩开元, 王彤, 陈仲. 毛白杨PtoYABBY2和PtoYABBY12的克隆及功能分析[J]. 生物技术通报, 2025, 41(9): 256-264. |
| [14] | 史发超, 姜永华, 刘海伦, 文英杰, 严倩. 荔枝LcTFL1基因的克隆与功能分析[J]. 生物技术通报, 2025, 41(9): 159-167. |
| [15] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||