








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 111-132.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1450
赵艳侠1( ), 李倩2, 孙家波1, 梁红敏1, 李冰冰2(
), 李倩2, 孙家波1, 梁红敏1, 李冰冰2( )
)
收稿日期:2025-12-30
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
李冰冰,女,博士,教授,研究方向 :果实品质调控与生物育种;E-mail: libingbing@cau.edu.cn作者简介:赵艳侠,女,博士,助理研究员,研究方向 :草莓种质创新与绿色高效生产;E-mail: zhaoyanxia2368@sina.com
基金资助:
ZHAO Yan-xia1( ), LI Qian2, SUN Jia-bo1, LIANG Hong-min1, LI Bing-bing2(
), LI Qian2, SUN Jia-bo1, LIANG Hong-min1, LI Bing-bing2( )
)
Received:2025-12-30
Published:2026-03-26
Online:2026-04-23
摘要:
草莓(Fragaria × ananassa)作为全球重要的经济浆果,其果实品质由外观、质地、风味与营养等多个性状共同决定,直接影响商品价值与产业竞争力。为突破传统育种瓶颈,系统解析果实品质形成的分子调控机制已成为分子设计育种的核心基础。本综述系统总结草莓果实颜色、硬度、大小、糖、酸、香气物质及抗氧化成分等关键品质性状的遗传调控基础,揭示了MYB、NAC、WRKY等转录因子在品质形成多层级调控网络的核心作用;进一步阐明了以脱落酸为核心、生长素和赤霉素等其他激素通过协同或拮抗互作来调控果实成熟与品质代谢的分子通路;并整合了温度、光照等环境因子,通过影响激素信号与转录因子活性,进而影响果实品质形成的机制。当前研究多基于二倍体野生草莓,对八倍体栽培种中多等位基因互作、复杂调控网络及基因型‒环境互作方面的研究仍存在明显不足。未来研究需结合多组学技术、CRISPR/Cas9基因编辑及人工智能预测模型,深入解析栽培草莓关键调控网络的等位变异功能,开发实用分子标记,构建智能设计育种体系,从而实现抗逆高产、轻简优质、种子繁殖型等符合未来产业需求的草莓新品种的定向选育。本综述为草莓品质的遗传改良提供理论依据,也为其他园艺作物的品质调控研究提供参考。
赵艳侠, 李倩, 孙家波, 梁红敏, 李冰冰. 草莓果实品质形成的关键调控基因及分子网络解析[J]. 生物技术通报, 2026, 42(3): 111-132.
ZHAO Yan-xia, LI Qian, SUN Jia-bo, LIANG Hong-min, LI Bing-bing. Key Regulatory Genes and Molecular Networks Dissection Underlying Strawberry Fruit Quality Formation[J]. Biotechnology Bulletin, 2026, 42(3): 111-132.
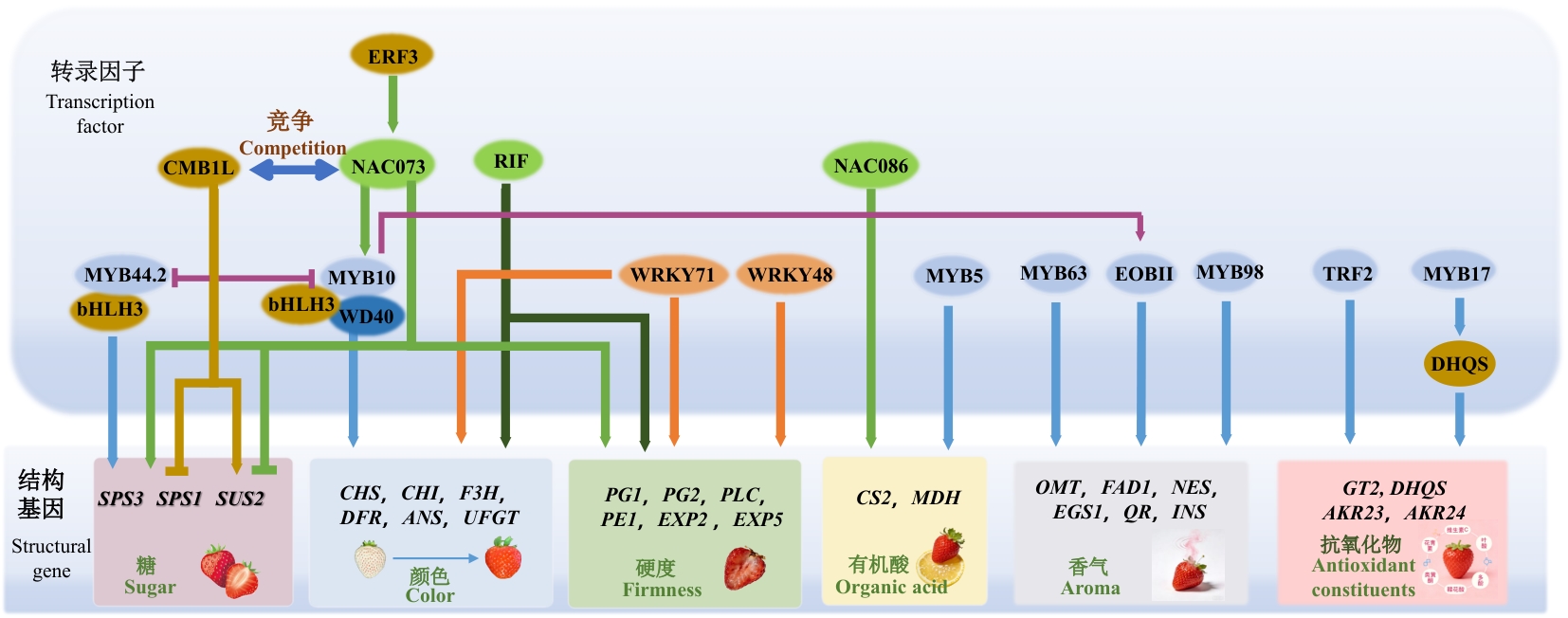
图1 草莓果实品质形成的转录调控网络不同颜色的椭圆及线条样式具有特定含义。绿色代表NAC家族转录因子,蓝色代表MYB家族转录因子,橙色代表WRKY家族转录因子,黄色代表其他转录因子;实线箭头表示“促进”作用,短横线加竖线表示“抑制”作用
Fig. 1 Transcriptional regulatory network underlying strawberry fruit quality formationDifferent colors of ellipses and line styles have specific meanings. Green indicates NAC family transcription factors, blue indicates MYB family transcription factors, orange indicates WRKY family transcription factors, and yellow indicates other transcription factors; solid arrows indicate the “promotion” effect, and a short horizontal line with a vertical line indicates the “inhibition” effect
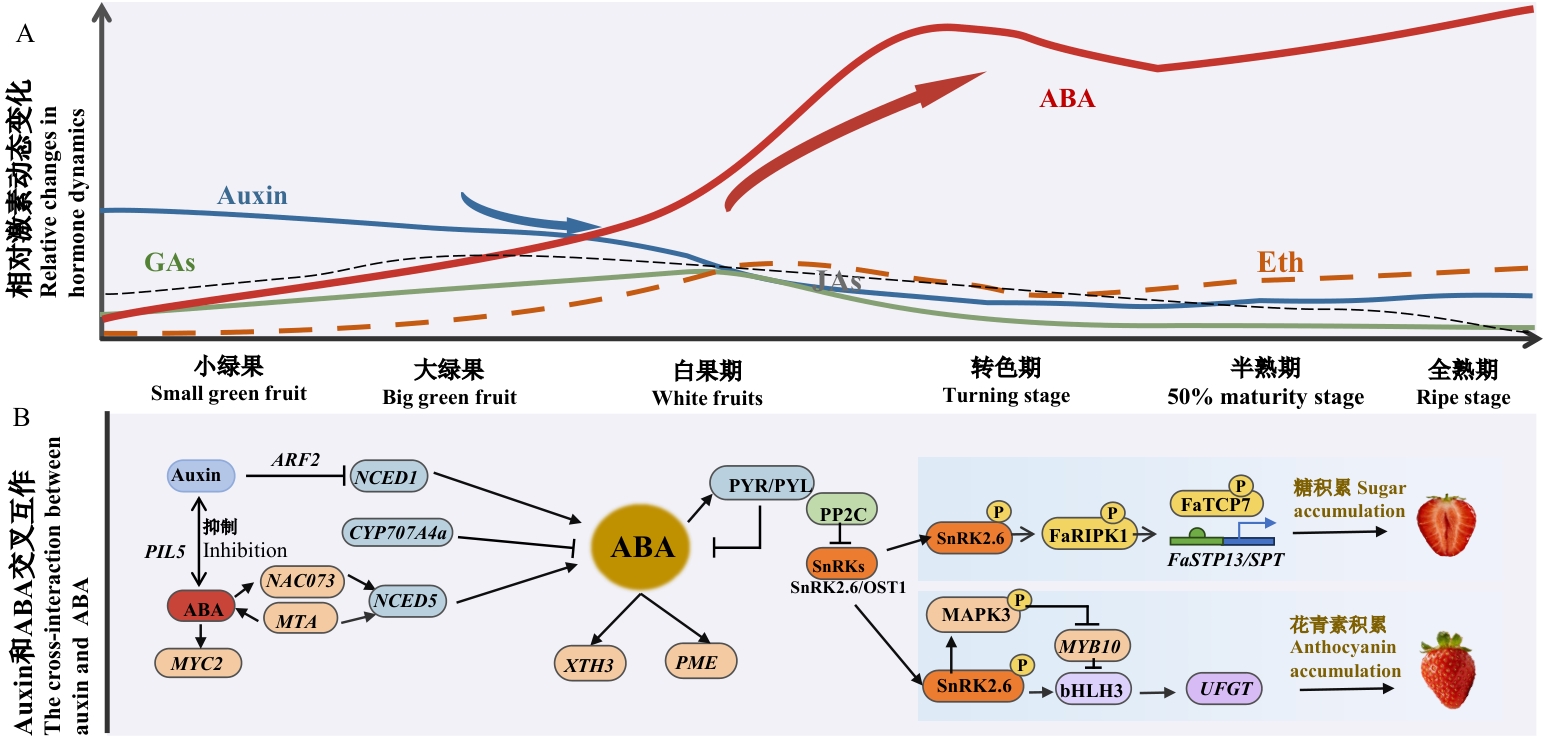
图2 草莓果实生长和成熟调控的激素网络A图为草莓果实发育进程中激素的动态变化。脱落酸(abscisic acid, ABA,红实线):在转色期含量骤升,促成熟;生长素(auxin,蓝实线):小绿至大绿果阶段占比高,抑制早期成熟、促进细胞分裂;赤霉素(gibberellins, GAs,绿实线):协同auxin促进果实膨大、延迟成熟;乙烯(ethylene, Eth,橙色虚线):在转色期上升,与ABA协同触发成熟;茉莉酸(jasmonic acids, JAs,黑虚线):全程低丰度且稳定。B图为auxin与ABA的交叉互作网络。Auxin通过ARF2/IAA等促进ABA合成基因(NCED1/5)表达、抑制降解基因(CYP707A)活性,提升ABA水平;ABA结合PYR/PYL受体后,激活SnRK2.6等核心激酶,磷酸化FaTCP7解除其对糖转运基因FaSTP13和FaSPT的抑制,促进糖积累;并启动MYB10/bHLH3级联,结合UFGT基因启动子促进花青素积累
Fig. 2 Hormonal regulatory network governing strawberry fruit growth and ripeningFigure A shows the dynamic changes of hormones during the development of strawberry fruits. Abscisic acid (ABA, red solid line): Its content surges sharply during the color-turning period, promoting ripening. Auxin (blue solid line): It accounts for a high proportion from the small green fruit stage to the large green fruit stage; inhibiting early ripening, promotes cell division. Gibberellins (GAs, green solid line): It synergizes with auxin to promote fruit enlargement and delay ripening. Ethylene (Eth, orange dashed line): It rises during the color-turning period, and cooperates with ABA to trigger ripening. Jasmonic acids (JAs, black dashed line): It maintains a low abundance and stability throughout the development process. Figure B shows the cross-interaction network between auxin and ABA. Auxin promotes the expressions of ABA synthesis genes (NCED1/5) and inhibits the activities of degradation genes (CYP707A) through ARF2/IAA, increasing the ABA level. After ABA binds to PYR/PYL receptors, it activates core kinases such as SnRK2.6, phosphorylates FaTCP7 to relieve its inhibition on sugar transporter genes FaSTP13 and FaSPT, and promotes sugar accumulation. It also initiates MYB10/bHLH3 cascade and binds to the promoter of the UFGT gene to promote anthocyanin accumulation
| [1] | Qiao Q, Edger PP, Xue L, et al. Evolutionary history and pan-genome dynamics of strawberry (Fragaria spp.) [J]. Proc Natl Acad Sci U S A, 2021, 118(45): e2105431118. |
| [2] | Liston A, Cronn R, Ashman TL. Fragaria: a genus with deep historical roots and ripe for evolutionary and ecological insights [J]. American J Botany, 2014, 101(10): 1686-1699. |
| [3] | Edger PP, Poorten TJ, VanBuren R, et al. Origin and evolution of the octoploid strawberry genome [J]. Nat Genet, 2019, 51(3): 541-547. |
| [4] | Hardigan MA, Lorant A, Pincot DDA, et al. Unraveling the complex hybrid ancestry and domestication history of cultivated strawberry [J]. Mol Biol Evol, 2021, 38(6): 2285-2305. |
| [5] | Pincot DDA, Ledda M, Feldmann MJ, et al. Social network analysis of the genealogy of strawberry: retracing the wild roots of heirloom and modern cultivars [J]. G3, 2021, 11(3): jkab015. |
| [6] | Castillejo C, Waurich V, Wagner H, et al. Allelic variation of MYB10 is the major force controlling natural variation in skin and flesh color in strawberry (Fragaria spp.) fruit [J]. Plant Cell, 2020, 32(12): 3723-3749. |
| [7] | Medina-Puche L, Cumplido-Laso G, Amil-Ruiz F, et al. MYB10 plays a major role in the regulation of flavonoid/phenylpropanoid metabolism during ripening of Fragaria × ananassa fruits [J]. J Exp Bot, 2014, 65(2): 401-417. |
| [8] | Hawkins C, Caruana J, Schiksnis E, et al. Genome-scale DNA variant analysis and functional validation of a SNP underlying yellow fruit color in wild strawberry [J]. Sci Rep, 2016, 6: 29017. |
| [9] | Zhang JX, Lei YY, Wang BT, et al. The high-quality genome of diploid strawberry (Fragaria nilgerrensis) provides new insights into anthocyanin accumulation [J]. Plant Biotechnol J, 2020, 18(9): 1908-1924. |
| [10] | Wang H, Zhang H, Yang Y, et al. The control of red colour by a family of MYB transcription factors in octoploid strawberry (Fragaria × ananassa) fruits [J]. Plant Biotechnol J, 2020, 18(5): 1169-1184. |
| [11] | Zhang JX, Liu S, Zhao S, et al. A telomere-to-telomere haplotype-resolved genome of white-fruited strawberry reveals the complexity of fruit colour formation of cultivated strawberry [J]. Plant Biotechnol J, 2025, 23(1): 78-80. |
| [12] | Yuan HZ, Cai WJ, Chen XD, et al. Heterozygous frameshift mutation in FaMYB10 is responsible for the natural formation of red and white-fleshed strawberry (Fragaria × ananassa Duch) [J]. Front Plant Sci, 2022, 13: 1027567. |
| [13] | Yuan HZ, Wang C, Quan FY, et al. Allelic variation and light-responsive regulation of FaMYB10-2 underlie tissue-specific anthocyanin accumulation in strawberry [J]. Plant Physiol, 2025. DOI: 10.1093/plphys/kiaf665 . |
| [14] | Duan XY, Wang KR, Tang RK, et al. Recent advances in biosynthesis and regulation of strawberry anthocyanins [J]. Hortic Res, 2025, 12(8): uhaf135. |
| [15] | Schaart JG, Dubos C, Romero De La Fuente I, et al. Identification and characterization of MYB-bHLH-WD40 regulatory complexes controlling proanthocyanidin biosynthesis in strawberry (Fragaria × ananassa) fruits [J]. New Phytol, 2013, 197(2): 454-467. |
| [16] | Jiang LY, Yue ML, Liu YQ, et al. A novel R2R3-MYB transcription factor FaMYB5 positively regulates anthocyanin and proanthocyanidin biosynthesis in cultivated strawberries (Fragaria × ananassa) [J]. Plant Biotechnol J, 2023, 21(6): 1140-1158. |
| [17] | Zhang ZY, Shi YN, Ma YC, et al. The strawberry transcription factor FaRAV1 positively regulates anthocyanin accumulation by activation of FaMYB10 and anthocyanin pathway genes [J]. Plant Biotechnol J, 2020, 18(11): 2267-2279. |
| [18] | Fan JM, Cao MH, Bi XY, et al. A FvERF3-FvNAC073 module regulates strawberry fruit size and ripening [J]. Plant J, 2025, 122(5): e70262. |
| [19] | Kui LW, McGhie TK, Wang M, et al. Engineering the anthocyanin regulatory complex of strawberry (Fragaria vesca) [J]. Front Plant Sci, 2014, 5: 651. |
| [20] | Martín-Pizarro C, Vallarino JG, Osorio S, et al. The NAC transcription factor FaRIF controls fruit ripening in strawberry [J]. Plant Cell, 2021, 33(5): 1574-1593. |
| [21] | Chen YT, Liu LP, Feng QQ, et al. FvWRKY50 is an important gene that regulates both vegetative growth and reproductive growth in strawberry [J]. Hortic Res, 2023, 10(7): uhad115. |
| [22] | Yue ML, Jiang LY, Zhang NT, et al. Importance of FaWRKY71 in strawberry (Fragaria × ananassa) fruit ripening [J]. Int J Mol Sci, 2022, 23(20): 12483. |
| [23] | Zheng GH, Hu SQ, Cheng SM, et al. Factor of DNA methylation 1 affects woodland strawberry plant stature and organ size via DNA methylation [J]. Plant Physiol, 2023, 191(1): 335-351. |
| [24] | Zhang ZH, Tian RR, Wan JY, et al. Combined analysis of metabolomics and transcriptomics reveals the effects of sugar treatment on postharvest strawberry fruit quality [J]. BMC Plant Biol, 2025, 25: 882. |
| [25] | Chen Q, Long Y, Yang M, et al. FaPKc2.2 negatively modulates strawberry fruit ripening by reprograming the carbon metabolic pathway [J]. Sci Hortic, 2022, 301: 111114. |
| [26] | Gu TT, Jia SF, Huang XR, et al. Transcriptome and hormone analyses provide insights into hormonal regulation in strawberry ripening [J]. Planta, 2019, 250(1): 145-162. |
| [27] | Guo L, Luo X, Li MZ, et al. Mechanism of fertilization-induced auxin synthesis in the endosperm for seed and fruit development [J]. Nat Commun, 2022, 13: 3985. |
| [28] | Kou XH, Zhou JQ, Wu CE, et al. The interplay between ABA/ethylene and NAC TFs in tomato fruit ripening: a review [J]. Plant Mol Biol, 2021, 106(3): 223-238. |
| [29] | Rosli HG, Civello PM, Martínez GA. Changes in cell wall composition of three Fragaria × ananassa cultivars with different softening rate during ripening [J]. Plant Physiol Biochem, 2004, 42(10): 823-831. |
| [30] | Peng ZZ, Liu GS, Li HL, et al. Molecular and genetic events determining the softening of fleshy fruits: a comprehensive review [J]. Int J Mol Sci, 2022, 23(20): 12482. |
| [31] | Zhang YH, Wang S, Li X, et al. FaERF6 activates the polygalacturonase gene FaPG1 to promote strawberry fruit softening [J]. Postharvest Biol Technol, 2026, 234: 114129. |
| [32] | Zhou HC, Li G, Zhao X, et al. Comparative analysis of polygalacturonase in the fruit of strawberry cultivars [J]. Genet Mol Res, 2015, 14(4): 12776-12787. |
| [33] | García JM, Posé S, Muñoz-Blanco J, et al. The polygalacturonase FaPG1 gene plays a key role in strawberry fruit softening [J]. Plant Signal Behav, 2009, 4(8): 766-768. |
| [34] | López-Casado G, Sánchez-Raya C, Ric-Varas PD, et al. CRISPR/Cas9 editing of the polygalacturonase FaPG1 gene improves strawberry fruit firmness [J]. Hortic Res, 2023, 10(3): uhad011. |
| [35] | Muñoz P, Roldán-Guerra FJ, Verma S, et al. Genome-wide association studies in a diverse strawberry collection unveil loci controlling agronomic and fruit quality traits [J]. Plant Genome, 2024, 17(4): e20509. |
| [36] | Lin YX, He H, Wen YL, et al. Comprehensive analysis of the pectate lyase gene family and the role of FaPL1 in strawberry softening [J]. Int J Mol Sci, 2023, 24(17): 13217. |
| [37] | Medina-Escobar N, Cárdenas J, Moyano E, et al. Cloning, molecular characterization and expression pattern of a strawberry ripening-specific cDNA with sequence homology to pectate lyase from higher plants [J]. Plant Mol Biol, 1997, 34(6): 867-877. |
| [38] | Jiménez-Bermúdez S, Redondo-Nevado J, Muñoz-Blanco J, et al. Manipulation of strawberry fruit softening by antisense expression of a pectate lyase gene [J]. Plant Physiol, 2002, 128(2): 751-759. |
| [39] | Benítez‐Burraco A, Blanco‐Portales R, Redondo-Nevado J, et al. Cloning and characterization of two ripening-related strawberry (Fragaria × ananassa cv. Chandler) pectate lyase genes [J]. J Exp Bot, 2003, 54(383): 633-645. |
| [40] | Zhang WW, Zhao SQ, Gu S, et al. FvWRKY48 binds to the pectate lyase FvPLA promoter to control fruit softening in Fragaria vesca [J]. Plant Physiol, 2022, 189(2): 1037-1049. |
| [41] | Harrison EP, McQueen‐Mason SJ, Manning K. Expression of six expansin genes in relation to extension activity in developing strawberry fruit [J]. J Exp Bot, 2001, 52(360): 1437-1446. |
| [42] | Dotto MC, Martínez GA, Civello PM. Expression of expansin genes in strawberry varieties with contrasting fruit firmness [J]. Plant Physiol Biochem, 2006, 44(5/6): 301-307. |
| [43] | Mu Q, Li XY, Luo JH, et al. Characterization of expansin genes and their transcriptional regulation by histone modifications in strawberry [J]. Planta, 2021, 254(2): 21. |
| [44] | Molina-Hidalgo FJ, Franco AR, Villatoro C, et al. The strawberry (Fragaria × ananassa) fruit-specific rhamnogalacturonate lyase 1 (FaRGLyase1) gene encodes an enzyme involved in the degradation of cell-wall middle lamellae [J]. J Exp Bot, 2013, 64(6): 1471-1483. |
| [45] | Paniagua C, Blanco-Portales R, Barceló-Muñoz M, et al. Antisense down-regulation of the strawberry β-galactosidase gene FaβGal4 increases cell wall galactose levels and reduces fruit softening [J]. J Exp Bot, 2016, 67(3): 619-631. |
| [46] | Bustamante CA, Rosli HG, Añón MC, et al. β-Xylosidase in strawberry fruit: Isolation of a full-length gene and analysis of its expression and enzymatic activity in cultivars with contrasting firmness [J]. Plant Sci, 2006, 171(4): 497-504. |
| [47] | Lee HE, Manivannan A, Lee SY, et al. Chromosome level assembly of homozygous inbred line ‘Wongyo 3115’ facilitates the construction of a high-density linkage map and identification of QTLs associated with fruit firmness in octoploid strawberry (Fragaria × ananassa) [J]. Front Plant Sci, 2021, 12: 696229. |
| [48] | Witasari LD, Huang FC, Hoffmann T, et al. Higher expression of the strawberry xyloglucan endotransglucosylase/hydrolase genes FvXTH9 and FvXTH6 accelerates fruit ripening [J]. Plant J, 2019, 100(6): 1237-1253. |
| [49] | Alleva K, Marquez M, Villarreal N, et al. Cloning, functional characterization, and co-expression studies of a novel aquaporin (FaPIP2;1) of strawberry fruit [J]. J Exp Bot, 2010, 61(14): 3935-3945. |
| [50] | Carrasco-Orellana C, Stappung Y, Mendez-Yañez A, et al. Characterization of a ripening-related transcription factor FcNAC1 from Fragaria chiloensis fruit [J]. Sci Rep, 2018, 8: 10524. |
| [51] | Li XJ, Martín-Pizarro C, Zhou LL, et al. Deciphering the regulatory network of the NAC transcription factor FvRIF, a key regulator of strawberry (Fragaria vesca) fruit ripening [J]. Plant Cell, 2023, 35(11): 4020-4045. |
| [52] | Cai JF, Mo XL, Wen CJ, et al. FvMYB79 positively regulates strawberry fruit softening via transcriptional activation of FvPME38 [J]. Int J Mol Sci, 2022, 23(1): 101. |
| [53] | Xiao K, Fan JM, Bi XY, et al. A NAC transcription factor and a MADS-box protein antagonistically regulate sucrose accumulation in strawberry receptacles [J]. Plant Physiol, 2025, 197(3): kiaf043. |
| [54] | Rashid A, Ruan HX, Wang YS. The gene FvTST1 from strawberry modulates endogenous sugars enhancing plant growth and fruit ripening [J]. Front Plant Sci, 2022, 12: 774582. |
| [55] | Li X, Wang Y, Zhang C, et al. FvPHR1 improves the quality of woodland strawberry fruit by up-regulating the expression of FvPHT1;7 and FvSWEET9 [J]. Plant Cell Environ, 2025, 48(4): 2821-2834. |
| [56] | Zhang C, Liu YX, Wang BT, et al. CRISPR/Cas9 targeted knockout FvPHO2 can increase phosphorus content and improve fruit quality of woodland strawberry [J]. Sci Hortic, 2023, 317: 112078. |
| [57] | Choi HG, Moon BY, Kang NJ, et al. Yield loss and quality degradation of strawberry fruits cultivated under the deficient insolation conditions by shading [J]. Hortic Environ Biotechnol, 2014, 55(4): 263-270. |
| [58] | Wei LZ, Mao WW, Jia MR, et al. FaMYB44.2, a transcriptional repressor, negatively regulates sucrose accumulation in strawberry receptacles through interplay with FaMYB10 [J]. J Exp Bot, 2018, 69(20): 4805-4820. |
| [59] | Vallarino JG, Merchante C, Sánchez-Sevilla JF, et al. Characterizing the involvement of FaMADS9 in the regulation of strawberry fruit receptacle development [J]. Plant Biotechnol J, 2020, 18(4): 929-943. |
| [60] | Xu Y, Liu S, Sun HY, et al. Sucrose transport gene FaSWEET9a regulated by FaDOF2 transcription factor promotes sucrose accumulation in strawberry [J]. Plant Cell Rep, 2025, 44(6): 138. |
| [61] | Huang Y, Xu PH, Hou BZ, et al. Strawberry tonoplast transporter, FaVPT1, controls phosphate accumulation and fruit quality [J]. Plant Cell Environ, 2019, 42(9): 2715-2729. |
| [62] | Wang YY, Kong LX, Wang WH, et al. Global ubiquitinome analysis reveals the role of E3 ubiquitin ligase FaBRIZ in strawberry fruit ripening [J]. J Exp Bot, 2023, 74(1): 214-232. |
| [63] | Xing SN, Chen KL, Zhu HC, et al. Fine-tuning sugar content in strawberry [J]. Genome Biol, 2020, 21: 230. |
| [64] | Milosavljević D, Maksimović V, Milivojević J, et al. Sugars and organic acids in 25 strawberry cultivars: qualitative and quantitative evaluation [J]. Plants, 2023, 12(12): 2238. |
| [65] | Kallio H, Hakala M, Pelkkikangas AM, et al. Sugars and acids of strawberry varieties [J]. Eur Food Res Technol, 2000, 212(1): 81-85. |
| [66] | Luo Y, Ge C, Yang M, et al. Cytosolic/plastid glyceraldehyde-3-phosphate dehydrogenase is a negative regulator of strawberry fruit ripening [J]. Genes, 2020, 11(5): 580. |
| [67] | Yang M, Hou GY, Peng YT, et al. FaGAPC2/FaPKc2.2 and FaPEPCK reveal differential citric acid metabolism regulation in late development of strawberry fruit [J]. Front Plant Sci, 2023, 14: 1138865. |
| [68] | Liu YX, Zhu L, Yang MJ, et al. R2R3-MYB transcription factor FaMYB5 is involved in citric acid metabolism in strawberry fruits [J]. J Plant Physiol, 2022, 277: 153789. |
| [69] | Li CL, Dougherty L, Coluccio AE, et al. Apple ALMT9 requires a conserved C-terminal domain for malate transport underlying fruit acidity [J]. Plant Physiol, 2020, 182(2): 992-1006. |
| [70] | Peng YJ, Yuan YY, Chang WJ, et al. Transcriptional repression of MdMa1 by MdMYB21 in Ma locus decreases malic acid content in apple fruit [J]. Plant J, 2023, 115(5): 1231-1242. |
| [71] | He JX, Xu YT, Huang D, et al. TRIPTYCHON-LIKE regulates aspects of both fruit flavor and color in citrus [J]. J Exp Bot, 2022, 73(11): 3610-3624. |
| [72] | Butelli E, Licciardello C, Ramadugu C, et al. Noemi controls production of flavonoid pigments and fruit acidity and illustrates the domestication routes of modern citrus varieties [J]. Curr Biol, 2019, 29(1): 158-164.e2. |
| [73] | Ye J, Wang X, Hu TX, et al. An InDel in the promoter of Al-ACTIVATED MALATE TRANSPORTER9 selected during tomato domestication determines fruit malate contents and aluminum tolerance [J]. Plant Cell, 2017, 29(9): 2249-2268. |
| [74] | Rey-Serra P, Mnejja M, Monfort A. Inheritance of esters and other volatile compounds responsible for the fruity aroma in strawberry [J]. Front Plant Sci, 2022, 13: 959155. |
| [75] | Lyzhin AS, Luk’yanchuk IV, Zhbanova EV. Polymorphism of the FaOMT and FaFAD1 genes for fruit flavor volatiles in strawberry varieties and wild species from the genetic collection of the Michurin Federal Research Center [J]. Vestn VOGiS, 2020, 24(1): 5-11. |
| [76] | Oh Y, Barbey CR, Chandra S, et al. Genomic characterization of the fruity aroma gene, FaFAD1, reveals a gene dosage effect on γ-decalactone production in strawberry (Fragaria × ananassa) [J]. Front Plant Sci, 2021, 12: 639345. |
| [77] | Fan Z, Tieman DM, Knapp SJ, et al. A multi-omics framework reveals strawberry flavor genes and their regulatory elements [J]. New Phytol, 2022, 236(3): 1089-1107. |
| [78] | Wang SS, Shi MY, Zhang Y, et al. The R2R3-MYB transcription factor FaMYB63 participates in regulation of eugenol production in strawberry [J]. Plant Physiol, 2022, 188(4): 2146-2165. |
| [79] | Medina-Puche L, Molina-Hidalgo FJ, Boersma M, et al. An R2R3-MYB transcription factor regulates eugenol production in ripe strawberry fruit receptacles [J]. Plant Physiol, 2015, 168(2): 598-614. |
| [80] | Molina-Hidalgo FJ, Medina-Puche L, Cañete-Gómez C, et al. The fruit-specific transcription factor FaDOF2 regulates the production of eugenol in ripe fruit receptacles [J]. J Exp Bot, 2017, 68(16): 4529-4543. |
| [81] | Pan ZF, Jiang RY, Xie XB, et al. The DOF transcription factor, FaDOF1 affects eugenol accumulation in strawberry [J]. Plant Growth Regul, 2024, 104(2): 991-1002. |
| [82] | Zhang YY, Yin XR, Xiao YW, et al. An ETHYLENE RESPONSE FACTOR-MYB transcription complex regulates furaneol biosynthesis by activating QUINONE OXIDOREDUCTASE expression in strawberry [J]. Plant Physiol, 2018, 178(1): 189-201. |
| [83] | Song CK, Hong XT, Zhao S, et al. Glucosylation of 4-hydroxy-2,5-dimethyl-3(2H)-furanone, the key strawberry flavor compound in strawberry fruit [J]. Plant Physiol, 2016, 171(1): 139-151. |
| [84] | Fang XP, Shen JS, Zhang LQ, et al. Metabolomic and transcriptomic integration reveals the mechanism of aroma formation as strawberries naturally turn colors while ripening [J]. Food Chem, 2024, 460: 140765. |
| [85] | Urrutia M, Schwab W, Hoffmann T, et al. Genetic dissection of the (poly) phenol profile of diploid strawberry (Fragaria vesca) fruits using a NIL collection [J]. Plant Sci, 2016, 242: 151-168. |
| [86] | Lei YY, Nie YX, Wang J, et al. The FvMYB17 targets FvDHQS to promote ellagic acid biosynthesis in strawberry [J]. Plant J, 2025, 122(4): e70224. |
| [87] | Schulenburg K, Feller A, Hoffmann T, et al. Formation of β- glucogallin, the precursor of ellagic acid in strawberry and raspberry [J]. J Exp Bot, 2016, 67(8): 2299-2308. |
| [88] | Miller K, Feucht W, Schmid M. Bioactive compounds of strawberry and blueberry and their potential health effects based on human intervention studies: a brief overview [J]. Nutrients, 2019, 11(7): 1510. |
| [89] | Agius F, González-Lamothe R, Caballero JL, et al. Engineering increased vitamin C levels in plants by overexpression of a D-galacturonic acid reductase [J]. Nat Biotechnol, 2003, 21(2): 177-181. |
| [90] | Broad RC, Bonneau JP, Hellens RP, et al. Manipulation of ascorbate biosynthetic, recycling, and regulatory pathways for improved abiotic stress tolerance in plants [J]. Int J Mol Sci, 2020, 21(5): 1790. |
| [91] | Liu HB, Wei LZ, Ni Y, et al. Genome-wide analysis of ascorbic acid metabolism related genes in Fragaria × ananassa and its expression pattern analysis in strawberry fruits [J]. Front Plant Sci, 2022, 13: 954505. |
| [92] | Wei LZ, Liu HB, Ni Y, et al. FaAKR23 modulates ascorbic acid and anthocyanin accumulation in strawberry (Fragaria × ananassa) fruits [J]. Antioxidants, 2022, 11(9): 1828. |
| [93] | Hou GY, Yang M, He CX, et al. Genome-wide identification and comparative transcriptome methods reveal FaMDHAR50 regulating ascorbic acid regeneration and quality formation of strawberry fruits [J]. Int J Mol Sci, 2023, 24(11): 9510. |
| [94] | Li BJ, Shi YN, Jia HR, et al. Abscisic acid mediated strawberry receptacle ripening involves the interplay of multiple phytohormone signaling networks [J]. Front Plant Sci, 2023, 14: 1117156. |
| [95] | Liao X, Li MS, Liu B, et al. Interlinked regulatory loops of ABA catabolism and biosynthesis coordinate fruit growth and ripening in woodland strawberry [J]. Proc Natl Acad Sci U S A, 2018, 115(49): E11542-E11550. |
| [96] | Yu JQ, Yang XK, Gao HS, et al. Transcriptome atlas provides novel insights into ABA-mediated modulation of strawberry fruit ripening and quality [J]. Hortic Adv, 2025, 3: 16. |
| [97] | Chen XX, Gao JH, Shen YY. Abscisic acid controls sugar accumulation essential to strawberry fruit ripening via the FaRIPK1-FaTCP7-FaSTP13/FaSPT module [J]. Plant J, 2024, 119(3): 1400-1417. |
| [98] | Huang FL, Sun MM, Yao ZJ, et al. Protein kinase SnRK2.6 phosphorylates transcription factor bHLH3 to regulate anthocyanin homeostasis during strawberry fruit ripening [J]. J Exp Bot, 2024, 75(18): 5627-5640. |
| [99] | Wang W, Ouyang J, Li YT, et al. A signaling cascade mediating fruit trait development via phosphorylation-modulated nuclear accumulation of JAZ repressor [J]. J Integr Plant Biol, 2024, 66(6): 1106-1125. |
| [100] | Zhou LL, Tang RK, Li XJ, et al. N6-methyladenosine RNA modification regulates strawberry fruit ripening in an ABA-dependent manner [J]. Genome Biol, 2021, 22: 168. |
| [101] | Zhang YT, Long Y, Liu YT, et al. MAPK5 and MAPK10 overexpression influences strawberry fruit ripening, antioxidant capacity and resistance to Botrytis cinerea [J]. Planta, 2022, 255: 19. |
| [102] | Kim J, Lee JG, Hong Y, et al. Analysis of eight phytohormone concentrations, expression levels of ABA biosynthesis genes, and ripening-related transcription factors during fruit development in strawberry [J]. J Plant Physiol, 2019, 239: 52-60. |
| [103] | Feng J, Dai C, Luo HF, et al. Reporter gene expression reveals precise auxin synthesis sites during fruit and root development in wild strawberry [J]. J Exp Bot, 2019, 70(2): 563-574. |
| [104] | Jang YJ, Kim T, Lin MK, et al. Genome-wide gene network uncover temporal and spatial changes of genes in auxin homeostasis during fruit development in strawberry (F. × ananassa) [J]. BMC Plant Biol, 2024, 24: 876. |
| [105] | Liu H, Ying YY, Zhang L, et al. Isolation and characterization of two YUCCA flavin monooxygenase genes from cultivated strawberry (Fragaria × ananassa Duch.) [J]. Plant Cell Rep, 2012, 31(8): 1425-1435. |
| [106] | Liu H, Xie WF, Zhang L, et al. Auxin biosynthesis by the YUCCA6 flavin monooxygenase gene in woodland strawberry [J]. J Integr Plant Biol, 2014, 56(4): 350-363. |
| [107] | Pi MT, Hu SQ, Cheng LC, et al. The MADS-box gene FveSEP3 plays essential roles in flower organogenesis and fruit development in woodland strawberry [J]. Hortic Res, 2021, 8: 247. |
| [108] | Zhou JH, Sittmann J, Guo L, et al. Gibberellin and auxin signaling genes RGA1 and ARF8 repress accessory fruit initiation in diploid strawberry [J]. Plant Physiol, 2021, 185(3): 1059-1075. |
| [109] | Fait A, Hanhineva K, Beleggia R, et al. Reconfiguration of the achene and receptacle metabolic networks during strawberry fruit development[J]. Plant Physiol, 2008, 148(2): 730-750. |
| [110] | Bustamante CA, Civello PM, Martínez GA. Cloning of the promoter region of β-xylosidase (FaXyl1) gene and effect of plant growth regulators on the expression of FaXyl1 in strawberry fruit [J]. Plant Sci, 2009, 177(1): 49-56. |
| [111] | Sun JH, Luo JJ, Tian L, et al. New evidence for the role of ethylene in strawberry fruit ripening [J]. J Plant Growth Regul, 2013, 32(3): 461-470. |
| [112] | Cherian S, Figueroa CR, Nair H. ‘Movers and shakers’ in the regulation of fruit ripening: a cross-dissection of climacteric versus non-climacteric fruit [J]. J Exp Bot, 2014, 65(17): 4705-4722. |
| [113] | Zhang YT, Deng MY, Gu XJ, et al. Ethylene signaling pathway genes in strawberry and their expression patterns during fruit ripening [J]. Agronomy, 2023, 13(7): 1930. |
| [114] | Sánchez-Sevilla JF, Vallarino JG, Osorio S, et al. Gene expression atlas of fruit ripening and transcriptome assembly from RNA-seq data in octoploid strawberry (Fragaria × ananassa) [J]. Sci Rep, 2017, 7: 13737. |
| [115] | Dong YH, Song MY, Liu XX, et al. Effects of exogenous KT and BA on fruit quality in strawberry (Fragaria vesca) [J]. J Hortic Sci Biotechnol, 2022, 97(2): 236-243. |
| [116] | Gan LJ, Wei MM, Cao SQ, et al. Overexpression of housekeeping gene FveIPT2 enhances anthocyanin and terpenoid accumulation in strawberry fruits with minimal impact on plant growth and development [J]. Hortic Res, 2025, 12(8): uhaf130. |
| [117] | Symons GM, Chua YJ, Ross JJ, et al. Hormonal changes during non-climacteric ripening in strawberry [J]. J Exp Bot, 2012, 63(13): 4741-4750. |
| [118] | Csukasi F, Osorio S, Gutierrez JR, et al. Gibberellin biosynthesis and signalling during development of the strawberry receptacle [J]. New Phytol, 2011, 191(2): 376-390. |
| [119] | 郭正兵, 冯英娜, 王全智. 茉莉酸类物质在草莓果实发育中的作用及其分子机理分析 [J]. 西北植物学报, 2015, 35(12): 2462-2468. |
| Guo ZB, Feng YN, Wang QZ. Molecular mechanism of jasmonates in strawberry fruit development [J]. Acta Bot Boreali Occidentalia Sin, 2015, 35(12): 2462-2468. | |
| [120] | Zhang ZB, Lu SW, Yu WB, et al. Jasmonate increases terpene synthase expression, leading to strawberry resistance to Botrytis cinerea infection [J]. Plant Cell Rep, 2022, 41(5): 1243-1260. |
| [121] | Vaezi S, Asghari M, Farokhzad A, et al. Exogenous methyl jasmonate enhances phytochemicals and delays senescence in harvested strawberries by modulating GABA shunt pathway [J]. Food Chem, 2022, 393: 133418. |
| [122] | Concha CM, Figueroa NE, Poblete LA, et al. Methyl jasmonate treatment induces changes in fruit ripening by modifying the expression of several ripening genes in Fragaria chiloensis fruit [J]. Plant Physiol Biochem, 2013, 70: 433-444. |
| [123] | Liu C, Liu Z, Li X, et al. Jasmonate modulates strawberry susceptibility to anthracnose by activating SnRK2.1 to regulate the WRKY50-JAZ5 module [J]. Plant Biotechnol J, 2025. DOI:10.1111/pbi.70492 . |
| [124] | Jia MR, Ding N, Zhang Q, et al. A FERONIA-like receptor kinase regulates strawberry (Fragaria × ananassa) fruit ripening and quality formation [J]. Front Plant Sci, 2017, 8: 1099. |
| [125] | Chen JX, Mao LC, Lu WJ, et al. Transcriptome profiling of postharvest strawberry fruit in response to exogenous auxin and abscisic acid [J]. Planta, 2016, 243(1): 183-197. |
| [126] | Li BJ, Shi YN, Xiao YN, et al. AUXIN RESPONSE FACTOR 2 mediates repression of strawberry receptacle ripening via auxin-ABA interplay [J]. Plant Physiol, 2024, 196(4): 2638-2653. |
| [127] | Christou A, Filippou P, Manganaris GA, et al. Sodium hydrosulfide induces systemic thermotolerance to strawberry plants through transcriptional regulation of heat shock proteins and aquaporin [J]. BMC Plant Biol, 2014, 14: 42. |
| [128] | Hu Y, Han YT, Wei W, et al. Identification, isolation, and expression analysis of heat shock transcription factors in the diploid woodland strawberry Fragaria vesca [J]. Front Plant Sci, 2015, 6: 736. |
| [129] | Mao WW, Han Y, Chen YT, et al. Low temperature inhibits anthocyanin accumulation in strawberry fruit by activating FvMAPK3-induced phosphorylation of FvMYB10 and degradation of Chalcone Synthase 1 [J]. Plant Cell, 2022, 34(4): 1226-1249. |
| [130] | Chen S, Wang XJ, Cheng Y, et al. Effects of supplemental lighting on flavonoid and anthocyanin biosynthesis in strawberry flesh revealed via metabolome and transcriptome co-analysis [J]. Plants, 2024, 13(8): 1070. |
| [131] | Liu YQ, Tang L, Wang YP, et al. The blue light signal transduction module FaCRY1-FaCOP1-FaHY5 regulates anthocyanin accumulation in cultivated strawberry [J]. Front Plant Sci, 2023, 14: 1144273. |
| [132] | Li Y, Xu PB, Chen GQ, et al. FvbHLH9 functions as a positive regulator of anthocyanin biosynthesis by forming a HY5-bHLH9 transcription complex in strawberry fruits [J]. Plant Cell Physiol, 2020, 61(4): 826-837. |
| [133] | Kadomura-Ishikawa Y, Miyawaki K, Noji S, et al. Phototropin 2 is involved in blue light-induced anthocyanin accumulation in Fragaria × ananassa fruits [J]. J Plant Res, 2013, 126(6): 847-857. |
| [134] | Li H, Larsen DH, Schouten RE, et al. Red, blue and far-red light affect strawberry plant development and fruit quality without changing the susceptibility to Botrytis cinerea infection [J]. Environ Exp Bot, 2025, 233: 106133. |
| [135] | Xu F, Cao SF, Shi LY, et al. Blue light irradiation affects anthocyanin content and enzyme activities involved in postharvest strawberry fruit [J]. J Agric Food Chem, 2014, 62(20): 4778-4783. |
| [136] | Sun YF, Yang XF, Wu RR, et al. DNA methylation controlling abscisic acid catabolism responds to light to mediate strawberry fruit ripening [J]. J Integr Plant Biol, 2024, 66(8): 1718-1734. |
| [137] | Liu CY, Jiang XB, Yuan ZH. Plant responses and adaptations to salt stress: a review [J]. Horticulturae, 2024, 10(11): 1221. |
| [138] | Geilfus CM. Chloride in soil: From nutrient to soil pollutant [J]. Environ Exp Bot, 2019, 157: 299-309. |
| [139] | Li WH, Zhong JL, Zhang LH, et al. Overexpression of a Fragaria vesca MYB transcription factor gene (FvMYB82) increases salt and cold tolerance in Arabidopsis thaliana [J]. Int J Mol Sci, 2022, 23(18): 10538. |
| [140] | Li WH, Li P, Chen HY, et al. Overexpression of a Fragaria vesca 1R-MYB transcription factor gene (FvMYB114) increases salt and cold tolerance in Arabidopsis thaliana [J]. Int J Mol Sci, 2023, 24(6): 5261. |
| [141] | He SS, Yang H, Cao RQ, et al. 5-Aminolevulinic acid-induced salt tolerance in strawberry (cv. ‘Benihoppe’): possible role of nitric oxide on interception of salt ions in roots [J]. Sci Hortic, 2022, 304: 111294. |
| [142] | Wei B, Zhang JT, Wang LJ. ALA improves salt tolerance of strawberry by alleviating the negative regulation of FaMYB44 on FaCLC expression [J]. J Plant Physiol, 2025, 315: 154633. |
| [143] | Galli V, da Silva Messias R, Perin EC, et al. Mild salt stress improves strawberry fruit quality [J]. LWT, 2016, 73: 693-699. |
| [144] | Yuan XF, Hong S, Xiong W, et al. Development of fungal-mediated soil suppressiveness against Fusarium wilt disease via plant residue manipulation [J]. Microbiome, 2021, 9: 200. |
| [145] | MacKenzie SJ, Peres NA. Use of leaf wetness and temperature to time fungicide applications to control anthracnose fruit rot of strawberry in Florida [J]. Plant Dis, 2012, 96(4): 522-528. |
| [1] | 秦子璐, 孙海燕, 陈赢男. 植物表皮毛发育分子调控机制研究进展[J]. 生物技术通报, 2026, 42(9): 1-13. |
| [2] | 刘青媛, 吴洪启, 陈秀娥, 陈剑, 姜远泽, 何燕子, 喻奇伟, 刘仁祥. 转录因子NtMYB96a调控烟草耐旱性的功能研究[J]. 生物技术通报, 2026, 42(4): 239-250. |
| [3] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [4] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [5] | 彭楚, 孙娟利, 郑蓓蓓, 张若西, 韩月彭, 赵云. 果实花色苷酶促降解机制研究进展[J]. 生物技术通报, 2026, 42(3): 133-144. |
| [6] | 罗龙鑫, 李智, 李彤, 冯资权, 翟欣悦, 梁成林, 张亚丽, 吴上, 李媛媛, 姜翰. 苹果果实糖分形成的分子基础[J]. 生物技术通报, 2026, 42(3): 156-171. |
| [7] | 殷诗情, 田泰, 黄凤庭, 冯珑强, 王浩, 张静, 何文, 陈清, 王小蓉, 王燕. 果树果实硬度的调控机制研究进展[J]. 生物技术通报, 2026, 42(3): 213-229. |
| [8] | 杜丹, 郭翔, 胡鑫, 潘宇. 质体发育调控果实成熟与品质的研究进展[J]. 生物技术通报, 2026, 42(3): 48-59. |
| [9] | 王鹤瑶, 孙红梅. 园艺植物表皮毛功能及其形成机制研究进展[J]. 生物技术通报, 2026, 42(3): 242-254. |
| [10] | 付万祥, 王涛, 谭澍, 余思远, 熊欢, 邹锋. 外源糖对锥栗果实品质、淀粉酶活性及其基因表达的影响[J]. 生物技术通报, 2026, 42(3): 374-382. |
| [11] | 孙焱森, 魏立翔, 李若冰, 张程志, 聂宇航, 李杰, 才学鹏, 乔军, 孟庆玲. AOL-113转录因子对少孢节丛孢菌菌丝生长、胁迫响应及捕食能力的调控作用[J]. 生物技术通报, 2026, 42(2): 338-350. |
| [12] | 董亚茹, 朱红, 王照红, 赵东晓, 刘惠芬. 桑树MnDREB6E的克隆及耐盐抗旱性分析[J]. 生物技术通报, 2026, 42(2): 306-316. |
| [13] | 张冬岭, 张寅生, 王建军, 叶飞宇, 卢子涵, 马晨晨, 柳华峰, 胡德升, 邓亚洲, 曹丽茹. 玉米HSFs转录因子家族在干旱胁迫下的表达特性及功能[J]. 生物技术通报, 2026, 42(2): 178-187. |
| [14] | 吴翠翠, 陈登科, 兰刚, 夏芝, 李朋波. 花生转录因子AhHDZ70的生物信息学分析及耐盐耐旱性研究[J]. 生物技术通报, 2026, 42(1): 198-207. |
| [15] | 张驰昊, 刘晋囡, 晁跃辉. 蒺藜苜蓿bZIP转录因子MtbZIP29的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 241-250. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||