








生物技术通报 ›› 2026, Vol. 42 ›› Issue (4): 141-152.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0657
冯成蒿( ), 党雨乐, 王志泽, 聂蔚丹, 杨中敏, 杜崇(
), 党雨乐, 王志泽, 聂蔚丹, 杨中敏, 杜崇( )
)
收稿日期:2025-06-23
出版日期:2026-04-26
发布日期:2026-04-30
通讯作者:
杜崇,男,博士,副教授,研究方向 :番茄分子遗传育种;E-mail: godv2018@163.com作者简介:冯成蒿,男,硕士研究生,研究方向 :番茄分子遗传育种;E-mail: 1274849136@qq.com
基金资助:
FENG Cheng-hao( ), DANG Yu-le, WANG Zhi-ze, NIE Wei-dan, YANG Zhong-min, DU Chong(
), DANG Yu-le, WANG Zhi-ze, NIE Wei-dan, YANG Zhong-min, DU Chong( )
)
Received:2025-06-23
Published:2026-04-26
Online:2026-04-30
摘要:
目的 通过对抗、感番茄材料接种南方根结线虫后进行RNA-seq和加权基因共表达网络分析(weighted gene co-expression network analysis, WGCNA),关注和抗病相关的信号通路及基因,验证目标基因的根部表达模式,为后续上述基因分子功能研究提供理论依据。 方法 对抗病番茄材料‘18060’和感病材料‘Moneymaker’进行线虫接种,收集未接种、接种后2 d和4 d的根部进行RNA-seq和WGCNA协同分析,对可能参与抗病的主要信号通路及抗病相关基因进行筛选,并完成目标基因的组织特异性表达和根部表达水平测定。 结果 随着接种时间延长,不同阶段的差异表达基因(differential expressed genes, DEGs)数量逐渐增多,抗、感材料的组间比较在接种第2天筛选出1 900个DEGs,在第4天,DEGs数量增加至3 012个;KEGG分析显示,在抗、感材料组内和组间比较中,“植物‒病原菌互作”均被显著富集。在组间比较中,被富集到该通路的DEGs主要编码包括类受体蛋白/激酶、钙信号相关蛋白、热激蛋白以及转录因子等,其中还有编码RPP13蛋白的5个R基因亦被筛选到;聚焦接种时期的DEGs,WGCNA筛选出了具有较强显著性的模块MEbrown,依据in degree和out degree数量分别筛选出数值前10的nodes作为核心基因,发现其中有2个编码RPP13的DEGs;组织特异性和根部表达模式鉴定进一步表明,Solyc07g039410.3在番茄防卫南方根结线虫入侵中可能发挥关键作用。 结论 筛选出番茄抗南方根结线虫病进程中可能发挥作用的6个RPP13基因,其中,Solyc07g039420.1和Solyc07g039410.3在根部存在特异性表达特征,4 dpi时期,作为hub基因的Solyc07g039410.3被显著诱导并提升至6.23倍,该基因很可能在番茄防御南方根结线虫病的进程中发挥关键功能。
冯成蒿, 党雨乐, 王志泽, 聂蔚丹, 杨中敏, 杜崇. 基于RNA-seq筛选番茄防御南方根结线虫病的差异基因与RPP13的表达特征分析[J]. 生物技术通报, 2026, 42(4): 141-152.
FENG Cheng-hao, DANG Yu-le, WANG Zhi-ze, NIE Wei-dan, YANG Zhong-min, DU Chong. Screening of Differentially Expressed Genes in Tomato Defense against Meloidogyne incognita Based on RNA-seq and Expression Characteristics Analysis of RPP13 Gene[J]. Biotechnology Bulletin, 2026, 42(4): 141-152.
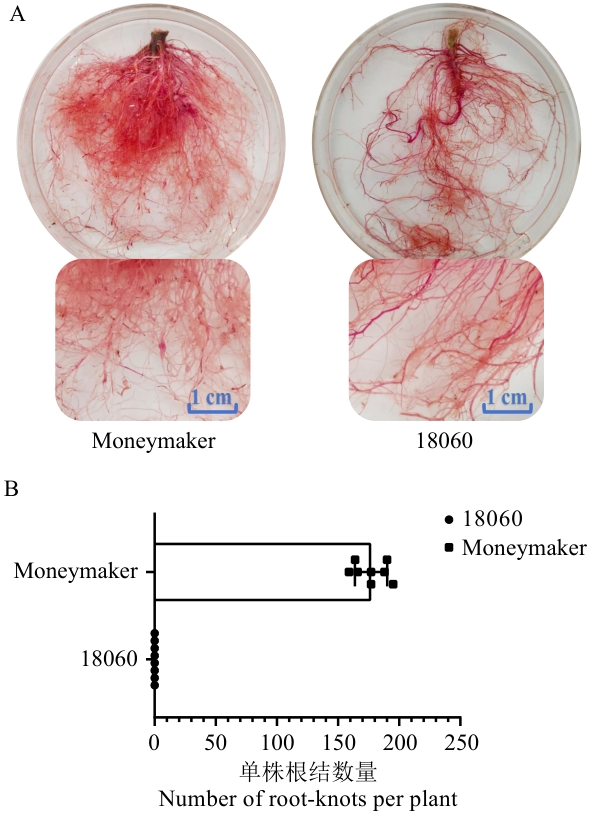
图1 抗病番茄‘18060’和感病番茄‘Moneymaker’接种后根结数量统计A:供试番茄材料接种后的根部酸性品红溶液染色结果;B:抗、感材料各10株供试番茄根部的根结数量统计
Fig. 1 Statistics of root galls numbers after inoculation of disease-resistant tomato '18060' and disease-susceptible tomato 'Moneymaker'A: The staining results of the root parts of the tested tomato materials after inoculation with acid fuchsin solution. B: Number statistics of root-knots on the roots of 10 tested tomato plants for both resistant and susceptible materials
| 样本Sample | 干净数据Clean data (Gb) | Q30 (%) | GC含量 GC content (%) | 总比对率Total mapped reads | 唯一映射的读段Unique mapped reads | 基因区间Intergenic (%) | 外显子Exon(%) |
|---|---|---|---|---|---|---|---|
| MM-0d-1 | 5.88 | 96.29 | 42.35 | 97.11 | 94.01 | 5.95 | 88.38 |
| MM-0d-2 | 6.16 | 95.50 | 42.19 | 97.14 | 94.26 | 6.21 | 87.17 |
| MM-0d-3 | 5.74 | 96.37 | 42.37 | 97.38 | 94.42 | 5.69 | 88.99 |
| MM-2d-1 | 6.08 | 95.65 | 42.19 | 96.96 | 94.18 | 5.97 | 87.66 |
| MM-2d-2 | 5.98 | 95.62 | 42.34 | 97.16 | 94.55 | 5.86 | 87.97 |
| MM-2d-3 | 6.54 | 96.54 | 42.33 | 97.58 | 94.94 | 5.41 | 89.42 |
| MM-4d-1 | 5.87 | 96.32 | 42.33 | 96.60 | 93.88 | 5.72 | 89.04 |
| MM-4d-2 | 5.76 | 96.50 | 42.27 | 97.60 | 95.10 | 5.55 | 89.74 |
| MM-4d-3 | 6.05 | 96.17 | 42.32 | 97.03 | 94.51 | 5.89 | 89.19 |
| XH-0d-1 | 6.03 | 96.50 | 42.30 | 95.98 | 93.72 | 6.10 | 88.70 |
| XH-0d-2 | 6.10 | 96.10 | 42.29 | 95.87 | 93.65 | 6.18 | 88.21 |
| XH-0d-3 | 6.07 | 96.27 | 42.22 | 94.94 | 92.70 | 6.31 | 87.35 |
| XH-2d-1 | 6.21 | 95.78 | 42.36 | 95.29 | 93.21 | 6.39 | 87.50 |
| XH-2d-2 | 6.29 | 95.56 | 42.17 | 94.96 | 92.97 | 6.58 | 86.90 |
| XH-2d-3 | 5.98 | 95.89 | 42.26 | 95.30 | 92.81 | 6.16 | 88.18 |
| XH-4d-1 | 6.43 | 96.39 | 42.35 | 96.58 | 94.26 | 5.90 | 88.63 |
| XH-4d-2 | 6.42 | 96.04 | 41.95 | 95.90 | 93.85 | 6.14 | 87.90 |
| XH-4d-3 | 6.10 | 95.95 | 42.20 | 96.21 | 93.99 | 5.95 | 88.32 |
表1 转录组测序数据统计与质量评估
Table 1 Statistics and quality assessment of transcriptome sequencing data
| 样本Sample | 干净数据Clean data (Gb) | Q30 (%) | GC含量 GC content (%) | 总比对率Total mapped reads | 唯一映射的读段Unique mapped reads | 基因区间Intergenic (%) | 外显子Exon(%) |
|---|---|---|---|---|---|---|---|
| MM-0d-1 | 5.88 | 96.29 | 42.35 | 97.11 | 94.01 | 5.95 | 88.38 |
| MM-0d-2 | 6.16 | 95.50 | 42.19 | 97.14 | 94.26 | 6.21 | 87.17 |
| MM-0d-3 | 5.74 | 96.37 | 42.37 | 97.38 | 94.42 | 5.69 | 88.99 |
| MM-2d-1 | 6.08 | 95.65 | 42.19 | 96.96 | 94.18 | 5.97 | 87.66 |
| MM-2d-2 | 5.98 | 95.62 | 42.34 | 97.16 | 94.55 | 5.86 | 87.97 |
| MM-2d-3 | 6.54 | 96.54 | 42.33 | 97.58 | 94.94 | 5.41 | 89.42 |
| MM-4d-1 | 5.87 | 96.32 | 42.33 | 96.60 | 93.88 | 5.72 | 89.04 |
| MM-4d-2 | 5.76 | 96.50 | 42.27 | 97.60 | 95.10 | 5.55 | 89.74 |
| MM-4d-3 | 6.05 | 96.17 | 42.32 | 97.03 | 94.51 | 5.89 | 89.19 |
| XH-0d-1 | 6.03 | 96.50 | 42.30 | 95.98 | 93.72 | 6.10 | 88.70 |
| XH-0d-2 | 6.10 | 96.10 | 42.29 | 95.87 | 93.65 | 6.18 | 88.21 |
| XH-0d-3 | 6.07 | 96.27 | 42.22 | 94.94 | 92.70 | 6.31 | 87.35 |
| XH-2d-1 | 6.21 | 95.78 | 42.36 | 95.29 | 93.21 | 6.39 | 87.50 |
| XH-2d-2 | 6.29 | 95.56 | 42.17 | 94.96 | 92.97 | 6.58 | 86.90 |
| XH-2d-3 | 5.98 | 95.89 | 42.26 | 95.30 | 92.81 | 6.16 | 88.18 |
| XH-4d-1 | 6.43 | 96.39 | 42.35 | 96.58 | 94.26 | 5.90 | 88.63 |
| XH-4d-2 | 6.42 | 96.04 | 41.95 | 95.90 | 93.85 | 6.14 | 87.90 |
| XH-4d-3 | 6.10 | 95.95 | 42.20 | 96.21 | 93.99 | 5.95 | 88.32 |
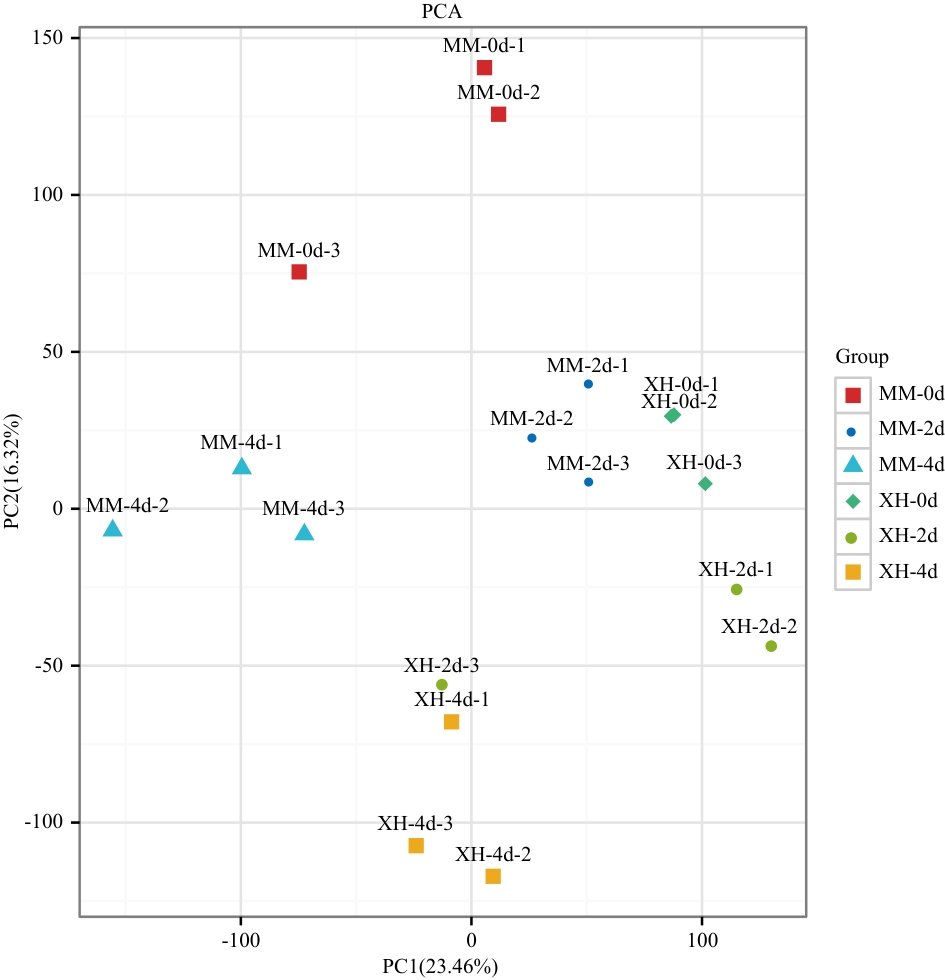
图2 供试番茄18个样本基于RNA-seq的PCA分布不同坐标表示不同主成分,百分比表示相应主成分对样品差异的贡献值,每个点表示1个样品,不同分组的样本用不同颜色和形状表示
Fig. 2 PCA distribution of 18 samples of the tested tomatoes based on RNA-seqThe different coordinates refer to different principal components, and the percentages refer to the contribution value of the corresponding principal component to the sample’s variance. Each point indicate a sample, and samples in different groups are marked by different colors and shapes
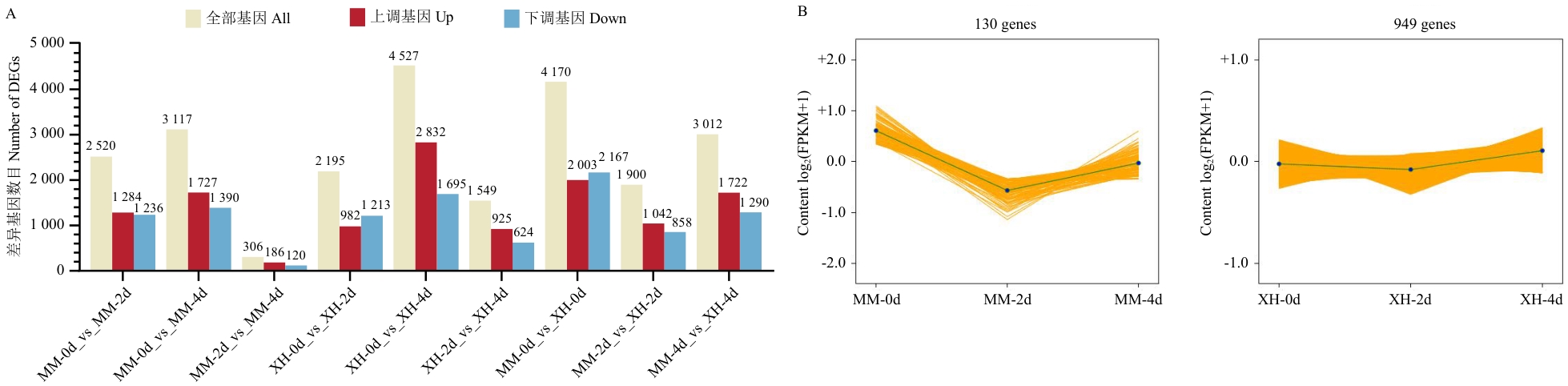
图3 不同比较策略下DEGs统计(A)及抗、感病组内共表达趋势分析(B)
Fig. 3 Statistical analysis of DEGs under different comparison strategies (A) and analysis of the co-expression trend within the resistant and susceptible groups (B)
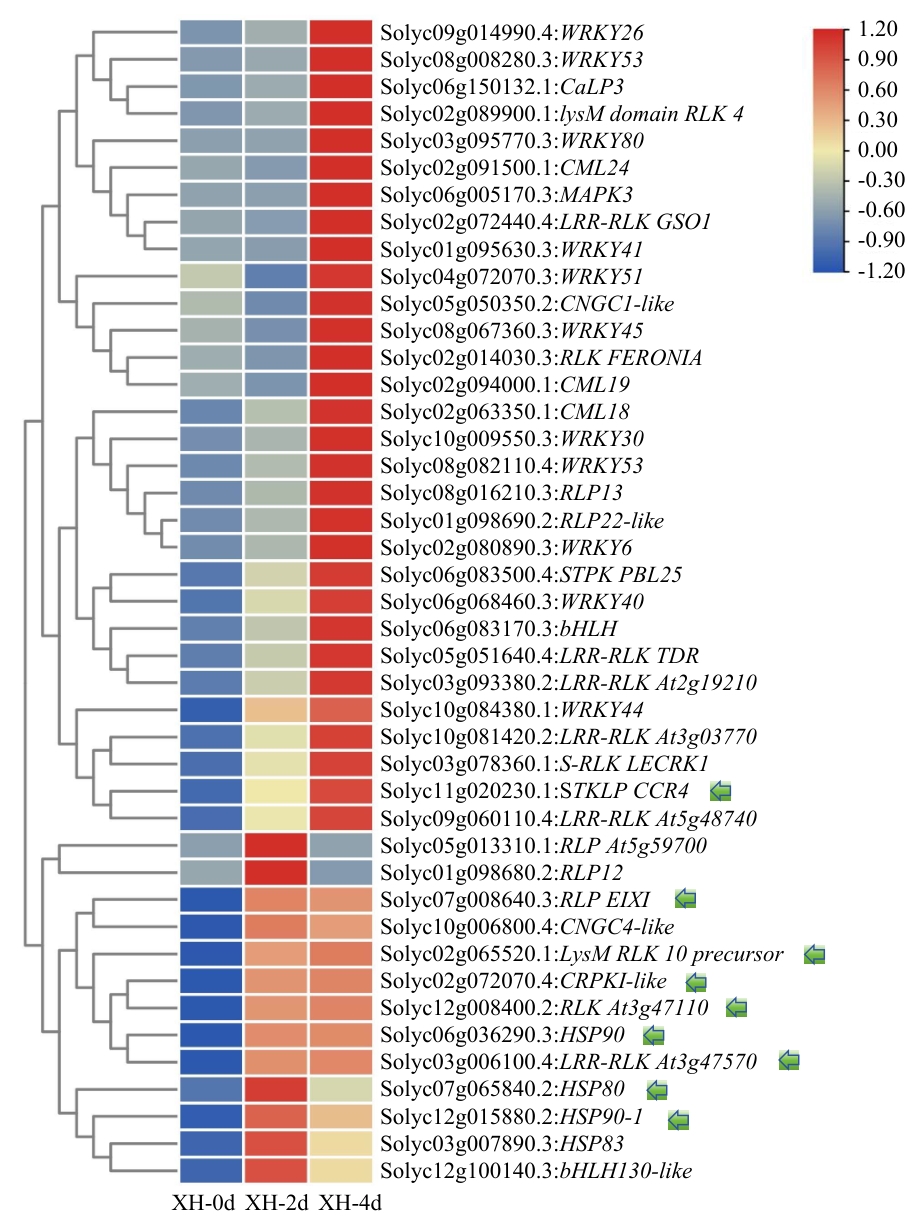
图5 XH组富集到植物‒病原菌互作途径中DEGs的FPKM聚类分析热图表示XH组不同处理阶段DEGs的FPKM值,模块颜色代表FPKM的大小,绿色箭头代表共同筛选的DEGs
Fig. 5 Cluster analysis of FPKM values of DEGs enriched in the plant-pathogen interaction pathway in the XH groupThe heatmap shows the FPKM values of DEGs in the XH group at different treatment stages, the colors of the modules indicate the size of FPKM, and the green arrows indicate the DEGs that were jointly selected

图6 组间比较MM_vs_XH中富集到植物‒病原菌互作途径DEGs的log2FC聚类分析聚类热图代表组间比较MM_vs_XH中不同接种阶段DEGs的log2FC统计,圆圈颜色和大小代表FPKM的数值高低,绿色箭头代表共同筛选的DEGs
Fig. 6 Cluster analysis of log2FC values for DEGs enriched in the plant-pathogen interaction pathway during inter-group comparisons (MM_vs_XH)The cluster heatmap indicates the statistical log2FC values of DEGs in the MM_vs_XH group comparison at different inoculation stages, the colors and sizes of the circles indicate the numerical values of FPKM, and the green arrows indicate the DEGs that are jointly selected
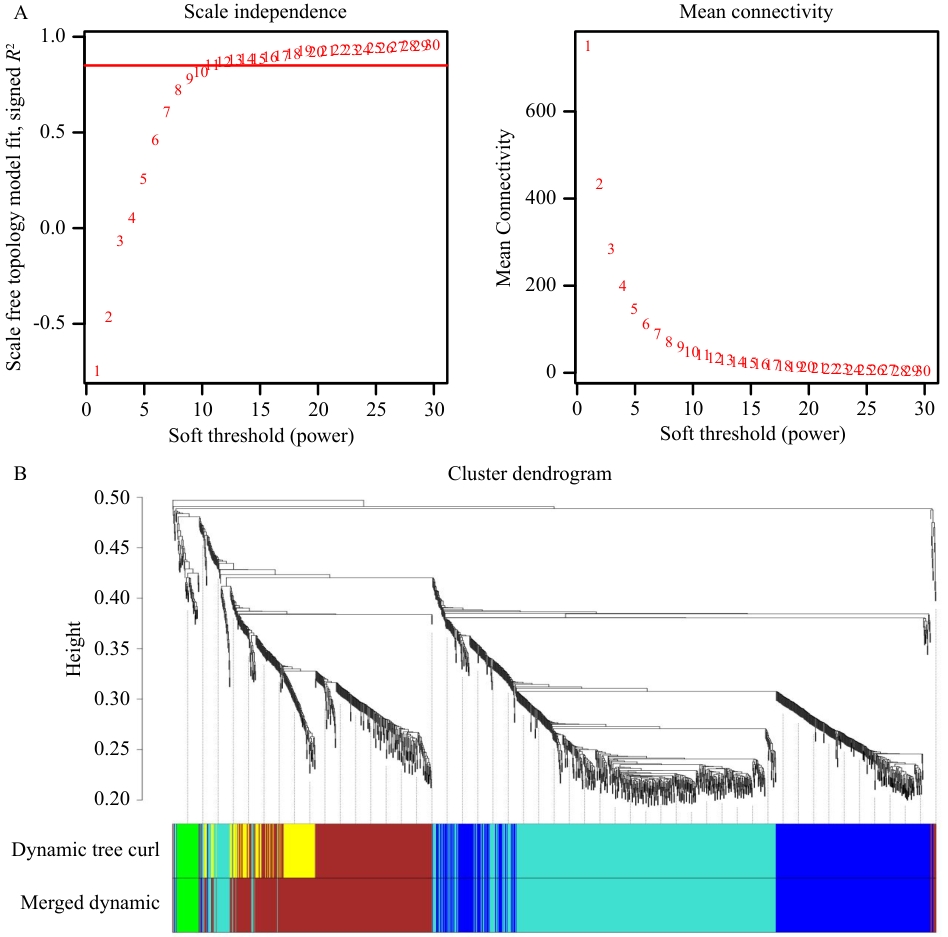
图7 层次聚类及模块划分A:最佳软阈值的选择;B:DEGs的系统聚类树和基因模块划分,不同颜色代表不同的模块
Fig. 7 Hierarchical clustering and module divisionA: Selection of the optimal soft threshold. B: The system clustering tree of DEGs and the division of gene modules, with different colors indicating different modules
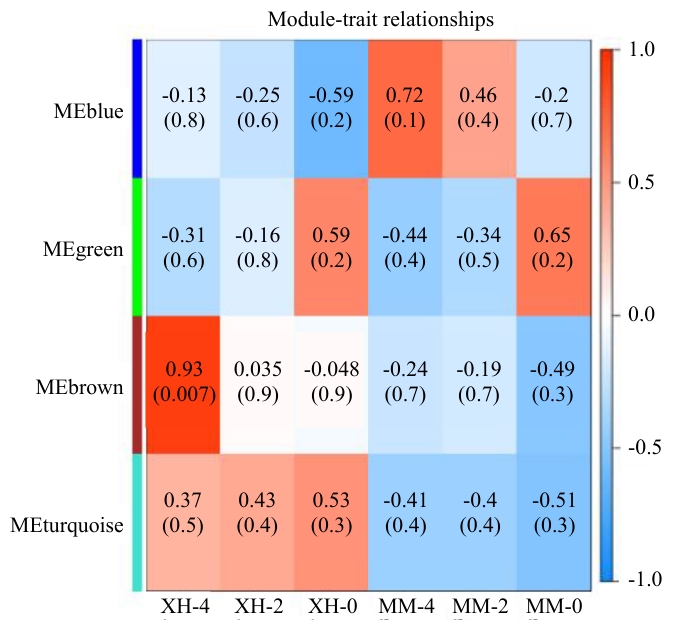
图8 模块与不同样本之间的相关性分析横坐标代表不同材料各接种时期,纵坐标代表模块名称,模块颜色越深代表相关性越强
Fig. 8 Correlation analysis between the module and different samplesThe horizontal axis indicates different materials and different inoculation periods, the vertical axis indicates the module name, and the darker the color of the module, the stronger the correlation
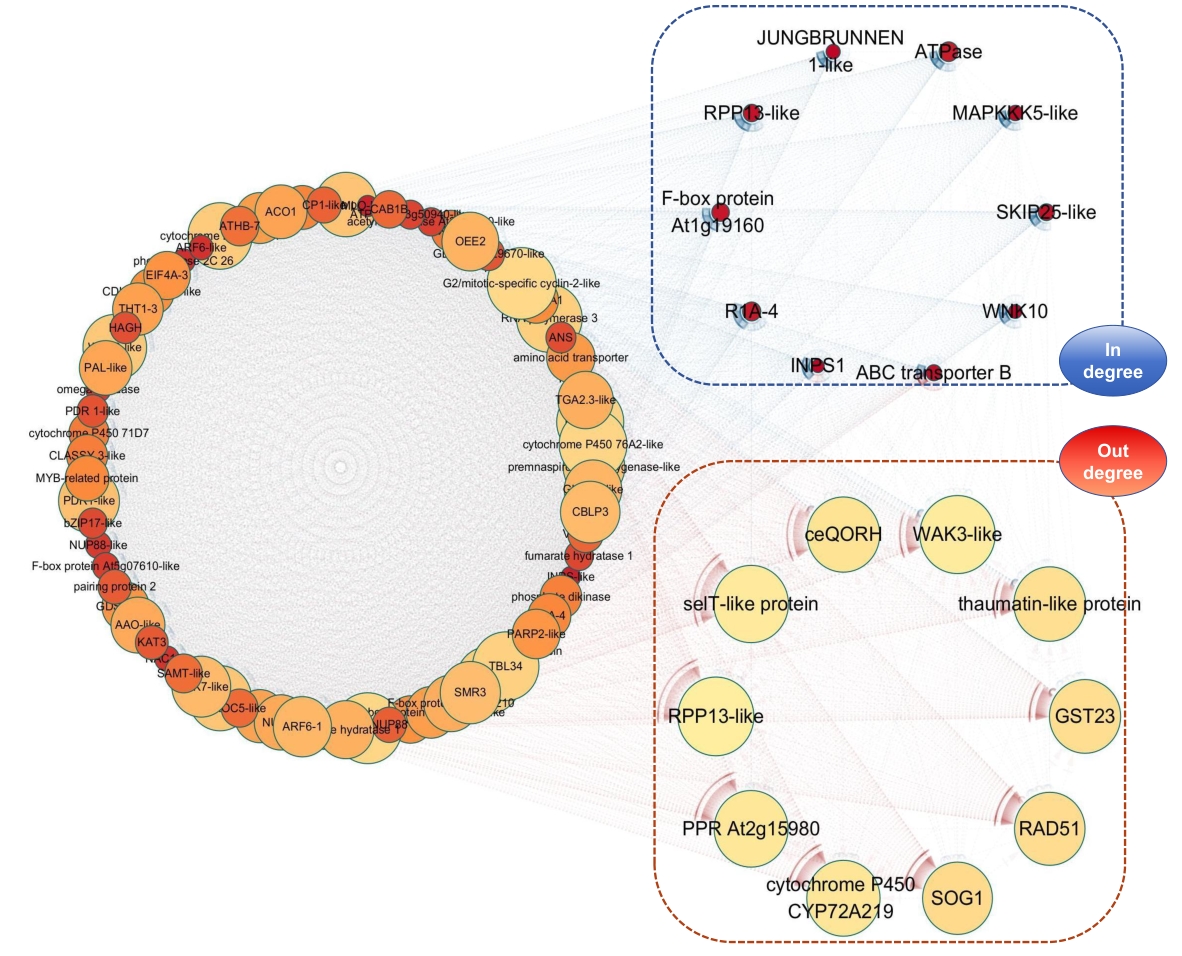
图9 共表达网络构建图结点的颜色越深代表in degree数量越多,圆圈越大代表out degree数量越多,虚线框内的nodes分别代表in degree和out degree数量排名前10的hub genes
Fig. 9 Construction diagram of co-expression networkThe darker the color of a node, the greater the number of in degree. The larger the circle, the greater the number of out degree. The nodes within the dotted box indicate the top ten hub genes in terms of in degree and out degree respectively
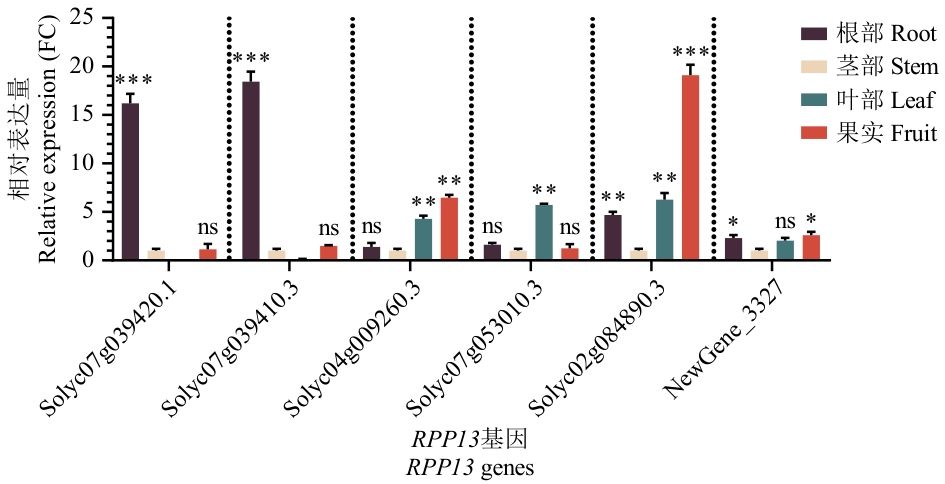
图10 RPP13基因的组织特异性表达分析ns:显著性差异(one-way ANOVA,P>0.05),* P<0.05,** P<0.01,*** P<0.001。下同
Fig. 10 Analysis of the tissue-specific expressions of the RPP13 genesns: Significant difference (one-way ANOVA, P>0.05), *:P<0.05, **P<0.01, *** P<0.001. The same below
| [1] | Gao J, Wang MZ, Chen LT, et al. First report of the root-knot nematode Meloidogyne incognita parasitizing Chenopodium album in China [J]. Plant Dis, 2024, 108(11): 3419. |
| [2] | Castañeda NEN, Alves GSC, Almeida RM, et al. Gene expression analysis in Musa acuminata during compatible interactions with Meloidogyne incognita [J]. Ann Bot, 2017, 119(5): 915-930. |
| [3] | Santini L, de Freitas Munhoz C, Bonfim MF Jr, et al. Host transcriptional profiling at early and later stages of the compatible interaction between Phaseolus vulgaris and Meloidogyne incognita [J]. Phytopathology, 2016, 106(3): 282-294. |
| [4] | 李琳, 刘薇, 张倩倩, 等. 设施蔬菜根结线虫的综合防治研究进展 [J]. 黑龙江农业科学, 2024(11): 114-120. |
| Li L, Liu W, Zhang QQ, et al. Research progress on comprehensive control of root-knot nematodes in facility vegetables [J]. Heilongjiang Agric Sci, 2024(11): 114-120. | |
| [5] | 许静, 项朝阳, 刘梅. 我国番茄产业成本收益分析及提质增效路径研究 [J]. 中国蔬菜, 2025(6): 1-8. |
| Xu J, Xiang ZY, Liu M. Study on cost-benefit analysis and quality-productivity improvement approaches in Chinese tomato industry [J]. China Veg, 2025(6): 1-8. | |
| [6] | 孙永珍, 贺靖, 魏芳, 等. “十三五”我国番茄产业发展及其国际竞争力评价 [J]. 中国瓜菜, 2023, 36(1): 112-116. |
| Sun YZ, He J, Wei F, et al. Evaluation on the development and international competitiveness of China’s tomato industry during the 13th Five-Year Plan period [J]. China Cucurbits Veg, 2023, 36(1): 112-116. | |
| [7] | Alekcevetch JC, de Lima Passianotto AL, Ferreira EGC, et al. Genome-wide association study for resistance to the Meloidogyne javanica causing root-knot nematode in soybean [J]. Theor Appl Genet, 2021, 134(3): 777-792. |
| [8] | 陈银霞, 王志泽, 冯成蒿, 等. 番茄抗根结线虫病Mi基因探索与WRKY转录因子参与抗病调控的研究进展 [J]. 江苏农业科学, 2025, 53(4): 16-22. |
| Chen YX, Wang ZZ, Feng CH, et al. Research progress of Mi gene against root-knot nematode disease in tomato and WRKY transcription factors involved in disease resistance regulation [J]. Jiangsu Agric Sci, 2025, 53(4): 16-22. | |
| [9] | El-Sappah AH, Islam MM, El-awady, HH, et al. Tomato natural resistance genes in controlling the root-knot nematode [J]. Genes, 2019, 10(11): 925. |
| [10] | Corbett BP, Jia LL, Sayler RJ, et al. The effects of root-knot nematode infection and mi-mediated nematode resistance in tomato on plant fitness [J]. J Nematol, 2011, 43(2): 82-89. |
| [11] | Yaghoobi J, Yates JL, Williamson VM. Fine mapping of the nematode resistance gene Mi-3 in Solanum peruvianum and construction of a S. lycopersicum DNA contig spanning the locus [J]. Mol Genet Genomics, 2005, 274(1): 60-69. |
| [12] | Wang YL, Yang WC, Zhang W, et al. Mapping of a heat-stable gene for resistance to southern root-knot nematode in Solanum lycopersicum [J]. Plant Mol Biol Report, 2013, 31(2): 352-362. |
| [13] | Dang YL, Wang ZZ, Nie WD, et al. SlWRKY30, a positive regulator of resistance to Meloidogyne incognita in Solanum lycopersicum [J]. Sci Hortic, 348: 114224. |
| [14] | Liu Y, Cao HY, Ling J, et al. Molecular cloning and functional analysis of the pepper resistance gene Me3 to root-knot nematode [J]. Hortic Plant J, 2023, 9(1): 133-144. |
| [15] | 李宁, 宫丽媛, 高升华, 等. 辣椒种质的抗性基因分子标记检测 [J]. 中国蔬菜, 2020(8): 19-32. |
| Li N, Gong LY, Gao SH, et al. Detection of resistance gene of Capsicum spp. germplasm by molecular markers [J]. China Veg, 2020(8): 19-32. | |
| [16] | Xu J, Narabu T, Mizukubo T, et al. A molecular marker correlated with selected virulence against the tomato resistance gene Mi in Meloidogyne incognita, M. javanica, and M. arenaria [J]. Phytopathology, 2001, 91(4): 377-382. |
| [17] | 叶德友, 钱春桃, 陈劲枫. 抗南方根结线虫黄瓜-酸黄瓜渐渗系的筛选及鉴定 [J]. 园艺学报, 2011, 38(12): 2281-2288. |
| Ye DY, Qian CT, Chen JF. Screening and identification of cucumber-sour cucumber introgression lines resistant to the root-knot nematode Meloidogyne incognita [J]. Acta Hortic Sin, 2011, 38(12): 2281-2288. | |
| [18] | Bhattarai KK, Atamian HS, Kaloshian I, et al. WRKY72-type transcription factors contribute to basal immunity in tomato and Arabidopsis as well as gene-for-gene resistance mediated by the tomato R gene Mi-1 [J]. Plant J, 2010, 63(2): 229-240. |
| [19] | Cap GB, Roberts PA, Thomason IJ. Inheritance of heat-stable resistance to Meloidogyne incognita in Lycopersicon peruvianum and its relationship to the Mi gene [J]. Theor Appl Genet, 1993, 85(6/7): 777-783. |
| [20] | Thor K, Jiang SS, Michard E, et al. The calcium-permeable channel OSCA1.3 regulates plant stomatal immunity [J]. Nature, 2020, 585(7826): 569-573. |
| [21] | Zhang DL, Yang XX, Wen ZY, et al. Proxitome profiling reveals a conserved SGT1-NSL1 signaling module that activates NLR-mediated immunity [J]. Mol Plant, 2024, 17(9): 1369-1391. |
| [22] | Moeder W, Phan V, Yoshioka K. Ca2+ to the rescue-Ca2+ channels and signaling in plant immunity [J]. Plant Sci, 2019, 279: 19-26. |
| [23] | Lin YL, Xu C, Li LL, et al. A conserved fungal effector disturbs Ca2+ sensing and ROS homeostasis to induce plant cell death [J]. Nat Commun, 2025, 16(1): 3523. |
| [24] | Wang DC, Wei LR, Liu T, et al. Suppression of ETI by PTI priming to balance plant growth and defense through an MPK3/MPK6-WRKYs-PP2Cs module [J]. Mol Plant, 2023, 16(5): 903-918. |
| [25] | Wen Q, Wang SY, Zhang XL, et al. Recent advances of NLR receptors in vegetable disease resistance [J]. Plant Sci, 2024, 348: 112224. |
| [26] | Joos HJ, Mauch-Mani B, Slusarenko AJ. Molecular mapping of the Arabidopsis locus RPP11 which conditions isolate-specific hypersensitive resistance against downy mildew in ecotype RLD [J]. Theor Appl Genet, 1996, 92(2): 281-284. |
| [27] | Serra H, Choi K, Zhao XH, et al. Interhomolog polymorphism shapes meiotic crossover within the Arabidopsis RAC1 and RPP13 disease resistance genes [J]. PLoS Genet, 2018, 14(12): e1007843. |
| [28] | Qiu DY, Xiao J, Xie WB, et al. Rice gene network inferred from expression profiling of plants overexpressing OsWRKY13, a positive regulator of disease resistance [J]. Mol Plant, 2008, 1(3): 538-551. |
| [29] | Mun JH, Yu HJ, Park S, et al. Genome-wide identification of NBS-encoding resistance genes in Brassica rapa [J]. Mol Genet Genomics, 2009, 282(6): 617-631. |
| [30] | Wang XT, Yang Q, Dai ZJ, et al. Identification of QTLs for resistance to maize rough dwarf disease using two connected RIL populations in maize [J]. PLoS One, 2019, 14(12): e0226700. |
| [31] | Prigozhin DM, Krasileva KV. Analysis of intraspecies diversity reveals a subset of highly variable plant immune receptors and predicts their binding sites [J]. Plant Cell, 2021, 33(4): 998-1015. |
| [32] | Guo SY, Zhang F, Du XY, et al. TaANK-TPR1 enhances wheat resistance against stripe rust via controlling gene expression and protein activity of NLR protein TaRPP13L1 [J]. Dev Cell, 2025, 60(12): 1702-1718.e6. |
| [33] | Chen YC, Wu W, Yang BH, et al. Grapevine VaRPP13 protein enhances oomycetes resistance by activating SA signal pathway [J]. Plant Cell Rep, 2022, 41(12): 2341-2350. |
| [34] | Yuan BQ, Li C, Wang QF, et al. Identification and functional characterization of the RPP13 gene family in potato (Solanum tuberosum L.) for disease resistance [J]. Front Plant Sci, 2024, 15: 1515060. |
| [35] | Chen L, Meng J, He XL, et al. Solanum lycopersicum microRNA1916 targets multiple target genes and negatively regulates the immune response in tomato [J]. Plant Cell Environ, 2019, 42(4): 1393-1407. |
| [36] | Zhang M, Zhang HY, Tan J, et al. Transcriptome analysis of eggplant root in response to root-knot nematode infection [J]. Pathogens, 2021, 10(4): 470. |
| [1] | 刘淼, 林涛, 贾乐松, 胡丰, 李涛, 李志万, 刘美芳, 郑方燕, 崔龙. 从野生到栽培:番茄果实色泽的演化与调控机制[J]. 生物技术通报, 2026, 42(3): 187-202. |
| [2] | 李亚妮, 韩鸿宇, 耿梦爽, 米若兰, 王韦琪, 于文静, 孟宪文, 李传友. ChiC基因调控番茄灰霉病抗性的机制研究[J]. 生物技术通报, 2026, 42(3): 255-262. |
| [3] | 程云霞, 张俊红, 叶杰. 番茄果实可溶性固形物积累的遗传调控研究进展[J]. 生物技术通报, 2026, 42(3): 145-155. |
| [4] | 颜晨琳, 李凡, 闫春婷, 程蛟文, 胡开林, 叶志彪, 宋建文. 番茄果实形态发育相关基因研究进展[J]. 生物技术通报, 2026, 42(3): 172-186. |
| [5] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [6] | 杜丹, 郭翔, 胡鑫, 潘宇. 质体发育调控果实成熟与品质的研究进展[J]. 生物技术通报, 2026, 42(3): 48-59. |
| [7] | 姜喆卉, 王小龙, 王守创, 周科. 番茄风味物质代谢途径解析与分子育种研究进展[J]. 生物技术通报, 2026, 42(3): 60-78. |
| [8] | 王潇奕, 李金焱, 邢醒, 朱鸿亮. 基于乙烯响应筛选调控番茄成熟且影响呼吸的基因及其功能分析[J]. 生物技术通报, 2026, 42(3): 275-282. |
| [9] | 李迎辉, 王杨博涵, 周浩博, 卢心如, 张珂欣, 于洋, 李传友, 孙传龙. 番茄VPE基因家族鉴定和抗逆功能分析[J]. 生物技术通报, 2026, 42(3): 263-274. |
| [10] | 高峥, 尹柳祎, 孟令品, 王晓慧, 孙佳欣, 宋扬, 温树波. 植物提取物抗牛病毒性腹泻病毒的研究进展[J]. 生物技术通报, 2026, 42(1): 31-41. |
| [11] | 苏秀敏, 韩文清, 王佼, 李鹏, 王秋兰, 李万星, 曹晋军. 哈茨木霉M408的分离鉴定、生物学特性及对番茄早疫病的生防效果[J]. 生物技术通报, 2025, 41(9): 277-288. |
| [12] | 朱丽娟, 张锴, 温晓蕾, 褚佳豪, 史凤玉, 王艳丽. 基于WGCNA挖掘野生大豆耐镉关键基因[J]. 生物技术通报, 2025, 41(8): 124-136. |
| [13] | 侯亚涛, 李迎辉, 邓磊, 李常保, 李传友, 孙传龙. 番茄果重基因功能型分子标记的开发及群体基因型分析[J]. 生物技术通报, 2025, 41(4): 98-105. |
| [14] | 王斌, 王玉昆, 肖艳辉. 丁香罗勒(Ocimum gratissimum)叶片响应镉胁迫的比较转录组学分析[J]. 生物技术通报, 2025, 41(3): 255-270. |
| [15] | 刘洁, 王飞, 陶婷, 张玉静, 陈浩婷, 张瑞星, 石玉, 张毅. 过表达SlWRKY41提高番茄幼苗抗旱性[J]. 生物技术通报, 2025, 41(2): 107-118. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||